Abstract
Boelen’s python (Morelia boeleni) is a montane New Guinea endemic found in highlands above 1000 m and below the tree line. The ecology, natural history, distribution, population size, and conservation status of this species are largely unknown. It has a protected status in Papua New Guinea but not in Indonesian Papua and several US and European zoos have active captive breeding programs that have been largely unsuccessful. To understand the degree of genetic diversity in wild and captive animals we undertook a genetic analysis of 90 M. boeleni for which we sequenced two mtDNA loci and one nuclear locus for a total of 1,418 bp of sequence data per individual. All 16 wild-caught M. boeleni from Indonesia and all captive M. boeleni are genetically uniform for all three loci. The single wild-caught animal from Papua New Guinea showed extremely low levels of genetic divergence and diversity from the Indonesian and captive samples. Data from two congeners, M. amethistina and M. viridis, suggests that M. boeleni have reduced genetic variation with a small effective population size possibly due to historical bottlenecks. These data demonstrate the need for further studies of genetic diversity of M. boeleni from across its range and raise particular concern for the limited genetic diversity of M. boeleni used captive breeding programs in zoological parks.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The tropical island of New Guinea has been identified as one of the world’s five High Biodiversity Wilderness Areas (Mittermeier et al. 2003). New Guinea occupies less than 0.6% of global land area yet harbors 5–7% of the world’s biodiversity (Beehler 1993; Dinerstein and Wikramanyake 1993; Myers et al. 2000). The herpetofauna of New Guinea currently accounts for approximately 5% of the world’s reptile and amphibian diversity yet surprisingly this is an underestimate of true diversity; species accumulation curves for both lizards and frogs continue to increase dramatically without hint of plateau (Austin et al. 2008). A defining feature of the island is the 2,500 km long central cordillera that forms the central highlands. The tallest peak exceeds 5,000 m and many other peaks exceed 3,000 m. The rugged highlands regions are geologically young (<5 mya) and the rugged montane topography encompasses a wide range of habitat types with a highly endemic fauna and flora (Hill and Gleadow 1989; Davies 1990; Audley-Charles 1991; Crowhurst et al. 1996).
A fairly recent discovery to science, Boelen’s Python (Morelia boeleni), is an uncommon snake found only in the New Guinea highlands above 1000 m and below the tree line (Brongersma 1953; O’Shea 1996). In Papua New Guinea, an independent country in the eastern half of the island (Figs. 1, 2), M. boeleni is a protected species and is afforded the same stringent legal status as birds of paradise (O’Shea 1996). In the western half of the island, the Indonesian province of Papua, M. boeleni has no country specific protection although both Indonesia and Papua New Guinea are CITES signatories and regulate trade of M. boeleni as a CITES Appendix II listed species. Virtually nothing is known about the natural history, ecology, or physiology of this montane endemic python. In addition, there is no data on population size, structure, or genetics of this species despite its conservation importance.
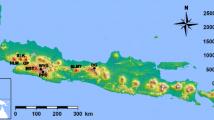
Map of the approximate distribution of M. boeleni along the central cordillera (in red from above 1000 m to the tree line) and disjunct distributions in the Huon Peninsula and Goodenough Island based on museum records and elevation (redrawn and modified with permission from the Bishop Museum Papuan database http://www.bishopmuseum.org/research/pbs/pngherps/index.html). The vertical line is the international border between Papua New Guinea to the east and the Indonesian province of Papua to the west. Samples of M. boeleni from Papua Indonesia came from the dashed box region (1) and the single M. boeleni from Papua New Guinea came from the dashed circle region (2). Although precise localities are known, we draw vague localities for protection of this species (Stuart et al. 2006). Samples of M. amethistina came from locality (3) and M. viridis came from localities (3) and (4)
The current distribution of montane forest in New Guinea is a large swath of habitat along the New Guinea central cordillera as well as several isolated mountain ranges that are separated from the central cordillera by lowland forests. Very little precise distributional data is available for M. boeleni but it is found throughout the central cordillera as well as in allopatric populations on the Huon Peninsula and Goodenough Island (Fig. 1). It is unknown if M. boeleni in the central cordillera represent one contiguous interbreeding population or a series of isolated mountain-top populations with limited, reduced, or no gene flow with one another, but limited locality data suggests the latter as there appear to be two disjunct populations in the central cordillera (Fig. 1). The two known allopatric populations on the Huon Peninsula and Goodenough Island are likely currently genetically isolated from the central cordillera population(s) because the intervening lowland forest is not suitable habitat. Morelia boeleni therefore consists of multiple allopatrically distributed populations that may be lineages of a single or multiple species.
Historically, live specimens M. boeleni have been imported into the USA since the mid 1970s. International Species Information System (ISIS) records indicate that 127 legal specimens have entered zoological institutions and currently 44 (21 male; 23 female) of those specimens are alive. There are 102 (38 male; 64 female) specimens accountable in private ownership; this population includes animals in USA, UK, Spain, Japan, and Germany. Most of these specimens are thought to have originated from one population found in the Western Highlands of Papua, Indonesia. This species has gained the reputation of being difficult to maintain and reproduce in captivity. However, this reputation has diminished due to current natural history studies (Spataro and Baldogo: http://www.boelenspythons.com/home.html) and exporters offering captive hatched specimens. Throughout their short captive history, there has been a concentrated effort at breeding this species yet efforts have been successful only four times, with none of these occurring in zoological institutions. Current wild populations seem to be non-threatened, receiving little pressure from its two known predators: humans and the New Guinea Harpy Eagle (Harpyopsis novaeguineae). Their status may change, however, as the human population increases or more natural resources are discovered on the island resulting in more habitat destruction. For these reasons this species should be considered a target species in need of a well-structured wild and captive management plan.
Upon discovering this species, Brongersma (1953) described it as a member of the genus Liasis (Gray 1842), where it remained until Kluge (2003) placed it within Morelia (Gray 1842), based on morphological and genetic analysis. Morelia boeleni are referred to by the following common names: Boelen’s Python, Black Python, and Sanca bulan. Genetic analysis has shown that this species’ closest extant relative is Morelia amethistina (Harvey et al. 2000; Rawlings et al. 2008), which also occurs on the island of New Guinea.
The objectives of this study are to elucidate general patterns of genetic structure for M. boeleni and to understand the levels of genetic variation in captive animals in zoological parks that are used for captive breeding purposes. We use DNA sequence data from two mitochondrial (mtDNA) genes and one nuclear (nDNA) gene to examine the amount of genetic variation from wild and captive animals. The mtDNA cytochrome b (cytb) gene has been widely used to detect population and specific level variation in boas and pythons (Austin 2000; Keogh et al. 2001; Vences et al. 2001; Auliya et al. 2002; Rawlings and Donnellan 2003; Lawson et al. 2004; Rawlings et al. 2008; Tzika et al. 2008) as well as other snakes. The second mtDNA gene region sequenced, cytochrome oxidase I (COI), has also been used extensively in snakes, reptiles, and other animals (Daniels et al. 2002; Melville et al. 2004; Utiger and Schatti 2004; Plaisance et al. 2008). The nDNA gene region sequenced, melanocortin-1 receptor (MC1R), has been shown to be variable in snakes, lizards, and other vertebrates (Theron et al. 2001; Rosenblum et al. 2004, 2007; Fajardo et al. 2008). This is the first phylogeographic study for any montane New Guinea reptile. As such these genetic data will provide a better understanding of estimates of the timing of population divergence and inferences of the historical population-level processes that generate and sustain the current patterns of genetic variation. Results of this study have implications for conservation of this species in the wild as well as providing important genetic data for captive breeding programs.
Materials and methods
We examined 108 specimens representing three python species from New Guinea. Two congeners of M. boeleni, M. amethistina and M. viridis, were incorporated to compare molecular divergence and diversity within the genus. DNA was extracted from 90 M. boeleni shed skins or tissue, and from liver from six M. amethistina, and 12 M. viridis, using a guanidine thiocyanate or ammonium acetate salt extraction protocol (http://socrates.berkeley.edu/~fujita/protocols/DNA_Extraction.pdf). Sheds of M. boeleni were donated from various zoological parks and private collectors (see “Appendix”). One nuclear (MC1R) and two mitochondrial (cytb and COI) gene fragments were amplified via PCR for each individual; each reaction contained the following: 25–50 ng template, 5 pmoles each primer (see Table 1 for primer sequences), 1.25 nmoles each dNTP, 1× PCR Buffer (New England Biolabs, Ipswich, MA), 0.5 units Taq polymerase (New England Biolabs, Ipswich, MA), and nuclease-free H2 0–25 μl. Primers for MC1R were designed from squamte sequences in GenBank. Amplicons were purified using Exo-SAP IT (USB Corp., Cleveland, OH). Cleaned products were cycle sequenced with Big Dye 3.1 (Applied Biosystems, Foster City, CA) following manufacturer’s protocols. Excess dye terminator was removed via sephadex column filtration, and products were electrophoresed on an ABI-3100 Genetic Analyzer (Applied Biosystems, Foster City, CA). Sequence output was aligned and edited with Sequencher 4.7 (Gene Codes Corp., Ann Arbor, MI).
Of the 90 individuals of M. boeleni, 17 total sheds were classified as ‘wild-caught’, 16 from Indonesian Papua, and one museum specimen (BPBM 11611) was collected ~1200 km away in Morobe Province in eastern Papua New Guinea (PNG) (Fig. 1). The remaining 73 individual M. boeleni are of unknown origin but are presumed to have come from Indonesia as this has been the source of all known exports. All M. amethistina were from one locality (Milne Bay Province, PNG) and the M. viridis were collected from two localities (Sandaun and Milne Bay Provinces, PNG) ~1600 km distant (Fig. 1; “Appendix”).
To compliment the empirical data, we performed a power analysis in order to assess the ability to detect presence of rare haplotypes with a sample of this size (16). Under the conservative assumption that these individuals were collected from the same locality, we calculated the probability of detecting more than one haplotype (P >1) as 1 − ρ 16, while varying the proportion (ρ) of the most common haplotype in the population between 0.5 and 1.
Results
All wild-caught specimens from Indonesia and all captive individuals of M. boeleni were genetically uniform across all three loci. No single nucleotide polymorphisms or insertion/deletion mutations were detected from the 1,418 bp of total aligned sequence data (cytb: 370 bp; COI: 553 bp; MC1R: 495 bp). For the nuclear locus no heterozygotes were detected as inferred from lack of double peaks in the electropherograms (Brumfield et al. 2003). The single haplotype seen in the cytb fragment sequenced from our Indonesian samples was identical to that obtained by Harvey et al. (2000). The single individual from eastern New Guinea (Morobe Province, PNG) ~1200 km distant from the Indonesian locality differed from the western samples by ≤1.1% sequence divergence for both mitochondrial genes, but the MC1R sequence was identical to the Indonesian M. boeleni. In contrast, sequences from the six M. amethistina and 12 M. viridis both yielded multiple haplotypes for each gene with mean intrapopulation divergence ranging from 0.24 to 0.41% and mean interpopulation divergence ranging from 6.8 to 7.5% for the mitochondrial genes and mean intrapopulation divergence ranging from 0.20 to 0.38% for the nuclear locus (Table 2). Results from the power analysis are shown in Fig. 3 and all sequences are deposited in GenBank (FJ864817-FJ865133).
A power curve representing the probability of detecting more than one haplotype (P >1) in a sample of 16, given the known proportion of the most common haplotype (ρ). The probability to the right of A (ρ = 0.83) is less than 0.95, and in a population where the ρ > 0.996 (B), a sample of this size will fail to detect diversity in an average of 95% of cases
Discussion
The lack of genetic variation for all 89 M. boeleni from wild-caught and captive sheds from Indonesia was unexpected. Although only 16 of the 89 Indonesian M. boeleni were recorded as wild-caught we would still expect to detect some degree of haplotype diversity with this sample size, given the results from the congeners and molecular results from other pythons (Table 2; Austin 2000; Harvey et al. 2000; Rawlings and Donnellan 2003; Rawlings et al. 2008). These results may be attributed to differences in lineage histories, however, results of the power analysis suggest that a sample of this size is likely to reveal diversity even when a single haplotype comprises the majority of a population. While an analysis that incorporated the 2 N nuclear data would have more power to detect diversity (>95% detectable on average when a single allele is 91% of the population and <5% when a single allele is 99.8% of the population). The single M. boeleni from eastern New Guinea (Morobe Province, PNG) showed limited mtDNA divergence [COI (0.82%); cytb (1.1%)] despite being 1200 km distant. In contrast, M. viridis was sampled from two localities 1600 km apart and showed an almost ten-fold increase in mtDNA divergence and increased haplotype diversity [COI (6.8%), 4 haplotypes; cytb (7.5%), 5 haplotypes]. For the nuclear locus MC1R, M. amethistina from a single population showed intrapopulation level divergence (0.88%) for two haplotypes and M. viridis from two populations also showed intrapopulation level divergence (0.20%). These data from the two congeners closely mirror data collected for other pythons and suggests that M. boeleni shows reduced genetic diversity and divergence at both mtDNA and nuclear genomes (Austin 2000; Harvey et al. 2000; Rawlings and Donnellan 2003; Rawlings et al. 2008).
Low levels of genetic variability have been observed in many other vertebrates (O’Brian et al. 1985; Gray 1995; Kretzmann et al. 1996; Rivera et al. 2006; Vargas-Ramirez et al. 2007). Reduced levels of genetic variability are typically explained by historical bottleneck(s) in population size from either natural or human mediated processes. Inbreeding depression results from an increase in homozygosity and this loss of genetic variability has a variety of negative consequences including decreased fitness from reduced fecundity and lower survival. In addition, small or declining populations may be susceptible to mutational meltdown where deleterious mutations do not have the opportunity to be eliminated by natural selection (Amos and Balmford 2001).
The genetic uniformity of all 89 M. boeleni from Indonesia can best be explained by all our samples from captive and wild-caught animals coming from a single highly genetically uniform population. We only have precise locality data for four sheds collected from the wild by MS. The other sheds from wild-caught animals and the one shed from the USF&W confiscation may or may not come from our one known Indonesian locality but this would be the most parsimonious assumption based on the genetic data. Even if this were the case, the lack of haplotype diversity is surprising as most wild reptile populations have moderate amounts of haplotype diversity that would be detected with this sample size and is evident from our much smaller congener samples (Austin 2000; Harvey et al. 2000; Rawlings and Donnellan 2003; Rawlings et al. 2008). Given the broad, allopatric, and topographically varied distribution of M. boeleni our a priori expectation was that the species would show at least some degree of genetic variation even if most samples were from a single locality. One possible explanation for the genetic uniformity of a single population of M. boeleni would be a recent bottleneck associated with paleoclimatological oscillations. Montane reptiles may be particularly sensitive to rapid climate change and alterations or fluctuations in environmental conditions that shift populations up or down an elevational gradient may lead to genetic bottlenecks as one or a few individuals are able to colonize new climate suitable elevations. In particular, multiple rapid oscillations of climate may cause severe bottlenecks. Our results are still surprising given that examination of genetic diversity of populations that are resulted from postglacial recolonization still show moderate haplotype diversity (Clark et al. 2003; Fuerst and Austin 2004; Ursenbacher et al. 2006).
Future research will be needed to obtain genetic material from M. boeleni from across its range throughout the central cordillera and in particular the two known allopatric populations on the Huon Peninsula and Goodenough Island (Fig. 1). In addition, use of more variable markers such as microsatellites may provide further refinement of our estimates of genetic structure in M. boeleni (Jordan et al. 2002). Further natural history studies are needed to increase our knowledge on this species. This data will hopefully help unlock the secrets as to why this is such a difficult species to reproduce in captivity and in return larger captive populations will be established. Institutions maintaining this species are encouraged to put forth more effort to assure their success with this species based on genetic and natural history information. Current CITES status should remain in effect, with better regulations of exportation on specimens originating from Indonesia.
Genealogical relationships and the harbored genetic diversity are important data for the use of developing a comprehensive captive breeding plan for any threatened or endangered species (Miller 1995). This study highlights the important need for further genetic sampling of M. boeleni from precise localities from across its range in order to assess if this species is actually highly genetically uniform and thus susceptible to inbreeding depression and in need of increased conservation protection and management.
References
Amos W, Balmford A (2001) When does conservation genetics matter? Heredity 87:257–265. doi:10.1046/j.1365-2540.2001.00940.x
Audley-Charles MG (1991) Tectonics of the New Guinea area. Annu Rev Earth Planet Sci 19:17–41. doi:10.1146/annurev.ea.19.050191.000313
Auliya M, Mausfeld P, Bohme W (2002) Review of the reticulated python (Python reticulatus Schneider, 1801) with the description of new subspecies from Indonesia. Naturwissenschaften 89:201–213. doi:10.1007/s00114-002-0320-4
Austin CC (2000) Molecular phylogeny and historical biogeography of Pacific island boas (Candoia). Copeia 2000:341–352. doi:10.1643/0045-8511(2000)000[0341:MPAHBO]2.0.CO;2
Austin CC, Hayden CJ, Bigilale I, Dahl C, Anaminiato J (2008) Checklist and comments on the terrestrial amphibian and reptile fauna from Utai, Northwestern Papua New Guinea. Herpetol Rev 39:40–46
Beehler BM (1993) Papua New Guinea conservation needs assessment. In: Beehler BM (ed) Mapping PNG’s biodiversity. Washington, DC, pp 193–209
Brongersma LD (1953) Notes on New Guinean reptiles and amphibians. II. Proc K Ned Akad Wet C 53:317–325
Brumfield RT, Beerli P, Nickerson DA, Edwards SV (2003) The utility of single nucleotide polymorphisms in inferences of population history. Trends Ecol Evol 18:249–256. doi:10.1016/S0169-5347(03)00018-1
Clark AM, Moler PE, Possardt EE, Savitzky AH, Brown WS, Bowen BW (2003) Phylogeography of the timber rattlesnake (Crotalus horridus) based on mtDNA sequences. J Herpetol 37:145–154. doi:10.1670/0022-1511(2003)037[0145:POTTRC]2.0.CO;2
Crowhurst PV, Hill KC, Foster DA, Bennett AF (1996) Thermochronological and geochemical constraints on the tectonic evolution of northern Papua New Guinea. In: Hall R, Blundell D (eds) Tectonic Evolution of Southeast Asia. Geological Society Special Publication No. 106
Daniels S, Heideman RN, Hendricks M, Wilson B (2002) A molecular phylogeny for the South African limbless lizard taxa of the subfamily Acontinae (Sauria: Scincidae) with special emphasis on relationships within Acontias. Mol Phylogenet Evol 24:315–323. doi:10.1016/S1055-7903(02)00217-8
Davies HL (1990) Structure and evolution of the border region of New Guinea. In: Carmen GJ, Carmen Z (eds) Petroleum exploration in Papua New Guinea: proceedings of the first PNG petroleum convention, Port Moresby
Dinerstein E, Wikramanyake ED (1993) Beyond “hotspots:” how to prioritize investments to conserve biodiversity in the Indo-Pacific region. Conserv Biol 7:53–65
Fajardo V, Gonzalez I, Martin I, Rojas M, Hernandez PE, Garcia T, Martin R (2008) Differentiation of European wild boar (Sus scrofa scrofa) and domestic swine (Sus scrofa domestica) meats by PCR analysis targeting the mitochondrial D-loop and the nuclear melanocortin receptor 1 (MC1R) genes. Meat Sci 78:314–322. doi:10.1016/j.meatsci.2007.06.018
Fuerst GS, Austin CC (2004) Population genetic structure of the Prairie Skink (Eumeces septentrionalis): nested Clade analysis of post Pleiostocence populations. J Herpetol 38:257–268. doi:10.1670/190-03A
Gray JE (1842) Description of some hitherto unrecorded species of Australian Reptiles and Batrachians. Zool Misc, London 5:1–57
Gray EM (1995) DNA fingerprinting reveals a lack of genetic variation in northern populations of the western pond turtle (Clemmys marmorata). Conserv Biol 9:1244–1255. doi:10.1046/j.1523-1739.1995.9051244.x
Harvey MB, Barker DG, Ammerman LK, Chippindale PT (2000) Systematics of pythons of the Morelia amethistina complex (Serpentes: Boidae) with the description of three new species. Herpetological Monogr 14:139–185. doi:10.2307/1467047
Hill KC, Gleadow AJ (1989) Uplift and thermal history of the Papuan fold belt, Papua New Guinea: apatite fission track analysis. Aust J Earth Sci 36:515–539. doi:10.1080/08120098908729507
Jordan PW, Goodman AE, Donnellan S (2002) Microsatellite primers for Australian and New Guinean pythons isolated with an efficient marker development method for related species. Mol Ecol Notes 2:78–82. doi:10.1046/j.1471-8286.2002.00135.x
Keogh JS, Barker DG, Shine R (2001) Heavily exploited but poorly known: systematics and biogeography of commercially harvested pythons (Python curtus group) in Southeast Asia. Biol J Linn Soc Lond 73:113–129. doi:10.1111/j.1095-8312.2001.tb01350.x
Kluge AG (2003) Aspidites and the phylogeny of pythonine snakes. Records of the Australian Museum, Supplement 19
Kretzmann MB, Gilmartin WG, Meyer A, Zegers GP, Fain SR, Taylor BF, Costa DP (1996) Low genetic variability in the Hawaiian Monk Seal. Conserv Biol 11:482–490. doi:10.1046/j.1523-1739.1997.96031.x
Lawson R, Slowinski JB, Brubrink FT (2004) A molecular approach to discerning the phylogenetic placement of the enigmatic snake Xenophidion schaeferi among the Alethinophidia. J Zool 263:285–294. doi:10.1017/S0952836904005278
Melville J, Schulte JA, Larson A (2004) A molecular study of phylogenetic relationships and evolution of antipredator strategies in Australian Diplodactylus geckos, subgenus Strophurus. Biol J Linn Soc Lond 82:123–138. doi:10.1111/j.1095-8312.2004.00324.x
Miller PS (1995) Selective breeding programs for rare alleles: examples form the Przewalski’s Horse and California Condor pedigrees. Conserv Biol 9:1262–1273. doi:10.1046/j.1523-1739.1995.9051262.x
Mittermeier RA, Mittermeier CG, Brooks TM, Pilgrim JD, Konstant WR, Da Fonseca GAB, Kormos C (2003) Wilderness and biodiversity conservation. Proc Natl Acad Sci USA 100:10309–10313. doi:10.1073/pnas.1732458100
Myers N, Mittermeier RA, Mittermeier CG, Da Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858
O’Brian SJ, Roelke ME, Marker L, Newman A, Winkler CA, Meltzer D, Colly L, Evermann JF, Bush M, Wildt DE (1985) Genetic basis for species vulnerability in the Cheetah. Science 227:1428–1434. doi:10.1126/science.2983425
O’Shea M (1996) A guide to the snakes of Papua New Guinea. Independent Publishing, Independent Group Pty Ltd, Port Moresby, PNG
Plaisance L, Rousset V, Morand S, Littlewood DTJ (2008) Colonization of Pacific islands by parasites of low dispersal ability: phylogeography of two monogenean species parasitizing butterflyfishes in the South Pacific Ocean. J Biogeogr 35:76–87
Rawlings LH, Donnellan SC (2003) Phylogeographic analysis of the green python, Morelia viridis, reveals a cryptic species. Mol Phylogenet Evol 27:36–44. doi:10.1016/S1055-7903(02)00396-2
Rawlings LH, Rabosky DL, Donnellan SC, Hutchinson MN (2008) Python phylogenetics: inference from morphology and mitochondrial DNA. Biol J Linn Soc Lond 93:603–619. doi:10.1111/j.1095-8312.2007.00904.x
Rivera PC, Gardenal CN, Chiaraviglio M (2006) Sex-biased dispersal and high levels of gene flow among local populations in the argentine boa constrictor, Boa constrictor occidentalis. Austral Ecol 31:948–955. doi:10.1111/j.1442-9993.2006.01661.x
Rosenblum EB, Hoekstra HE, Nachman MW (2004) Adaptive reptile color variation and the evolution of the MC1R gene. Evolution Int J Org Evolution 58:1794–1808
Rosenblum EB, Hickerson M, Moritz C (2007) A multilocus perspective on colonization accompanied by selection and gene flow. Evolution Int J Org Evolution 61:12971–12985. doi:10.1111/j.1558-5646.2007.00251.x
Stuart BL, Rhodin AGJ, Grismer LL, Hansel T (2006) Scientific description can imperil species. Science 312:1137. doi:10.1126/science.312.5777.1137b
Theron E, Hawkins K, Bermingham E, Ricklefs RE, Mundy NI (2001) The molecular basis of an avian plumage polymorphism in the wild: A melanocortin-1-receptor point mutation is perfectly associated with the melanic plumage morph of the bananaquit, Coereba flaveola. Curr Biol 11:550–557. doi:10.1016/S0960-9822(01)00158-0
Tzika AC, Koenig S, Miller R, Garcia G, Remy C, Milinkovitch MC (2008) Population structure of an endemic vulnerable species, the Jamaican boa (Epicrates subflavus). Mol Ecol 17:533–544. doi:10.1111/j.1365-294X.2008.03967.x
Ursenbacher S, Carlsson M, Helfer V, Tegelstrom H, Fumagalli L (2006) Phylogeography and Pleistocene refugia of the adder (Vipera berus) as inferred from mitochondrial DNA sequence data. Mol Ecol 15:3425–3437. doi:10.1111/j.1365-294X.2006.03031.x
Utiger U, Schatti B (2004) Morphology and phylogenetic relationships of the Cyprus racer, Hierophis cypriensis, and the systematic status of Coluber gemonensis gyarosensis Mertens (Reptilia : Squamata : Colubrinae). Rev Suisse Zool 11:225–238
Vargas-Ramirez M, Chiari Y, Castano-Mora OV, Menken SBJ (2007) Low genetic variability in the endangered Colombian endemic freshwater turtle Podocnemis lewyana (Testudines, Podocnemididae). Contrib Zool 76:1–7
Vences M, Glaw F, Kosuch J, Bohme W, Veith M (2001) Phylogeny of South American and Malagasy boine snakes: molecular evidence for the validity of Sanzinia and Acrantophis and biogeographic implications. Copeia 4:1151–1154. doi:10.1643/0045-8511(2001)001[1151:POSAAM]2.0.CO;2
Acknowledgments
We thank the following museums, zoological parks, and individuals for donating genetic material for the genetic work: The Denver, Fort Worth, Houston, Milwaukee, Oklahoma City, Riverside, St. Louis, and San Diego Zoos as well as the Bishop Museum; A. Allison, J. Baylin, R. Beard, D. Bellis, N. Bottini, N. Hoover, M. Jodney, T. Koegen, Y. Kuto, J. Leware, R. Maugg, F. Memmo, J. Rosenstarch, O. Robert, G. Schiavino, B. Simpson, M. Smith, J. Sola, D. Taylor, K. Tepedelen, S. Wari and G. Womer. We thank B. Roy, V. Kula, and B. Wilmot from the PNG Department of Environment and Conservation, and J. Robins from the PNG National Research Institute who have provided research assistance in Papua New Guinea. This manuscript was improved from comments from the Austin lab group. This research was funded by National Science Foundation grants DEB 0445213 and DBI 0400797 to CCA.
Author information
Authors and Affiliations
Corresponding author
Appendix
Rights and permissions
About this article
Cite this article
Austin, C.C., Spataro, M., Peterson, S. et al. Conservation genetics of Boelen’s python (Morelia boeleni) from New Guinea: reduced genetic diversity and divergence of captive and wild animals. Conserv Genet 11, 889–896 (2010). https://doi.org/10.1007/s10592-009-9931-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-009-9931-z