Abstract
The long-snouted seahorse Hippocampus guttulatus is one of the two European seahorse species. We describe the isolation of the first 12 microsatellite loci in this threatened species. These new markers were tested in non-invasive samples of 32 seahorses from NW Spain. The number of alleles ranged from 2 to 15 (mean: 6.3) and expected heterozygosity from 0.031 to 0.912 (mean: 0.500). All loci conformed to Hardy–Weinberg expectations and no genotypic disequilibrium was observed between any pair of loci. The theoretical exclusion probabilities for this set of loci, when no parental information exists or when one parent is known, were 0.973 and 0.998, respectively. This study indicates the usefulness of these novel loci for population analysis and kinship studies in Hippocampus guttulatus. Their potential application is extended to the other European seahorse species, since all loci were successfully cross-amplified in H. hippocampus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Seahorses (Syngnathidae, Gasterosteiformes) are emblematic and threatened fish with remarkable morphology and biology, including male pregnancy (Avise et al. 2002; CITES 2002). Only two of the 32 species in the world live in the Northeast Atlantic: Hippocampus guttulatus and H. hippocampus. No accurate biological and distribution data are available for these species, further research being needed to assess their conservation status (IUCN 2006). In order to address population genetic analysis of the long-snouted seahorse H. guttulatus, we have developed primers for 12 polymorphic microsatellite loci from enriched genomic libraries in this species. Microsatellites have proven to be powerful markers for population genetics and its application to conservation biology (Ellegren 2004). These loci are also useful for parentage analysis to support reproduction in captivity (Castro et al. 2004) and to study genetic mating systems (Avise et al. 2002).
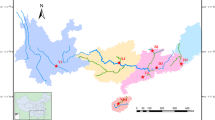
The genetic data presented here represent the first population analysis in this European seahorse. H. guttulatus was collected in Galician coasts (NW Spain), using non-invasive sampling of dorsal fin in live seahorses, and tissues stored in 100% ethanol. Genomic DNA was isolated by Chelex resin (Estoup et al. 1996) and standard phenol–chloroform methods.
Microsatellites were isolated using a partial enriched genomic library from muscle of a single dead seahorse according to the FIASCO protocol (Fast Isolation by AFLP of Sequences Containing repeats; Zane et al. 2002). Linked genomic fragments were enriched using a (AC)17 biotinylated probe as described by Pardo et al. (2006).
Polymorphism was preliminarily evaluated in four individuals using 2.5% agarose gels dyed with ethidium bromide using unlabelled primers. Allelic variation of putative polymorphic markers was confirmed in 32 wild seahorses using an ABI 3100 automated sequencer (Applied Biosystems). The forward primer of each pair was 5′ fluorescently labelled (Applied Biosystems). Genotyping was carried out using the genemapper 3.7 software (Applied Biosystems), with GenScan 500LIZ as internal size standard.
One hundred and fifty six clones out of 263 sequenced, revealed characteristic short tandem repeats of microsatellite loci. Primers pairs were designed for 52 sequences, where enough large flanking regions existed. Nineteen of these were successfully amplified and 12 of them result polymorphic (Table 1). Ten out of the variable loci contained dinucleotide repeats and the remaining ones presented tetranucleotide motifs. Eight microsatellites were perfect, three imperfect and one compound. The final yield of variable microsatellite markers rendered by this enrichment protocol was 4.6%, higher than in other fish studies using the same procedure (2.3%, Zane et al. 2002; 0.6%, Carreras-Carbonell et al. 2004).
Population genetic parameters were estimated using the program cervus 3.0 (Marshall et al. 1998) in 32 seahorses from NW Spain (Table 2). Number of alleles ranged from 2 (Hgu-USC10 and Hgu-USC11) to 15 (Hgu-USC7). Expected heterozygosity and PIC ranged from 0.031 to 0.912 and from 0.030 to 0.889, respectively, at loci Hgu-USC11 and Hgu-USC7. All loci conformed to Hardy–Weinberg and genotype linkage equilibrium expectations (genepop 3.1 program; Raymond and Rousset 1995), after sequential Bonferroni correction. These data are compatible with random mating, and suggest no close linkage between loci. However, null alleles might be present at two loci (Hgu-USC1 and Hgu-USC7; Table 2) as revealed by micro-checker 2.2.3 program (van Oosterhout et al. 2004), suggesting caution in their application for kinship analysis. The probability of exclusion of a false parent for this set of loci when the other parent is unknown or known was 0.973 and 0.998, respectively, using cervus program (Table 2). All these data support the usefulness of this set of loci for population genetic studies and parentage analysis. They will permit to evaluate wild and cultured genetic resources, to address genetic mating system in this species and to support the development of its reproduction in captivity.
We also evaluated the cross-amplification of these loci in two individuals of the short-snouted seahorse H. hippocampus, using a range of MgCl2 (1.5–2 mM) and temperature (48–60°C) PCR conditions. All loci were successfully amplified, some of them showing interspecific size differences (Table 1). This set of markers are potentially useful to obtain appropriate population data for conservation and management of genetic resources of the two threatened European seahorses.
References
Avise JC, Jones AG, Walker D et al. (2002) Genetic mating systems and reproductive natural histories of fishes: lessons for ecology and evolution. Annu Rev Genet 36:19–45
Carreras-Carbonell J, Macpherson E, Pascual M (2004) Isolation and characterization of microsatellite loci in Tripterygion delaisi. Mol Ecol Notes 4:438–439
Castro J, Bouza C, Presa P, Pino-Querido A et al. (2004) Potential sources of error in parentage assessment of turbot (Scophthalmus maximus) using microsatellite loci. Aquaculture 242:119–135
CITES (2002) Convention on International trade in endangered species of wild fauna and flora. Twelth Meeting of the Conference of the Parties, Santiago de Chile, Chile, 3–15 November
Ellegren H (2004) Microsatellites: simple sequences with complex evolution. Nat Rev Genet 5:435–445
Estoup A, Largiadèr CR, Perrot E et al. (1996) Rapid one-tube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Mol Mar Biol Biotechnol 5:295–298
IUCN (2006) 2006 IUCN red list of threatened species.<www.iucnredlist.org>. Cited 26 September 2006
Marshall TC, Slate J, Kruuk LEB et al. (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655
Pardo BG, Hermida M, Fernández C, Bouza C, Perez M, Llavona A, Sanchez L, Martinez P (2006) A set of highly polymorphic microsatellites useful for kinship and population analysis in turbot (Scopthalmus maximus L). Aquac Res. DOI 10.1111/j.1365-2109.2006.01600.x
Raymond M, Rousset F (1995) GENEPOP version 1.2.: population genetics software for exact tests and ecumenicism. J Hered 86:248–249
van Oosterhout C, Hutchinson WF, Wills DPM et al. (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Zane L, Bargelloni L, Patarnello T (2002) Strategies for microsatellite isolation: a review. Mol Ecol 11:1–16
Acknowledgements
BG Pardo and A López contributed equally to this work. This work was supported by a grant from the Spanish Ministerio de Ciencia y Tecnología (CGL2005–05927-C03-03), as part of a coordinated research (Hippocampus project, 2005/PC091). We wish to thank Miquel Planas (Instituto Investigaciones Marinas, CSIC, Vigo), Antonio Vilar (Aquarium Finisterrae) and Lucía Molina (University of Las Palmas de Gran Canarias) for supplying seahorse samples. We also thank Susana Sánchez, María López, Sonia Gómez, Lucía Insua and María Portela for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pardo, B.G., López, A., Martínez, P. et al. Novel microsatellite loci in the threatened European long-snouted seahorse (Hippocampus guttulatus) for genetic diversity and parentage analysis. Conserv Genet 8, 1243–1245 (2007). https://doi.org/10.1007/s10592-006-9241-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-006-9241-7