Abstract
In somatic cells, recombination is a means of DNA damage repair. The most severe type of damage in nuclear DNA is double-strand breaks (DSBs) which may be repaired via either non-homologous end joining (NHEJ) or homologous recombination (HR). In this review, we will summarize the basic features, the mechanisms, and the key players of both repair modes in plants with a focus on the model plant Arabidopsis thaliana. NHEJ may result in insertion of sequences from elsewhere in the genome but is much more often associated with deletions. If more than one DSB is processed simultaneously via NHEJ, besides deletions, inversions or translocations may also arise. As the germ line is only set aside late in plant development, somatic changes may be transferred to the next generation. Thus, NHEJ might influence the evolution of plant genomes and indeed seems to be an important factor of genome shrinking. Deletions may also be due to DSB-induced recombination between tandem duplicated homologous sequences by single-strand annealing (SSA). Moreover, conservative HR using the synthesis-dependent strand annealing (SDSA) mechanism operates in somatic plant cells. The efficiency of SDSA is dependent on the genomic template used as matrix for the repair of the DSB. Besides DSBs, stalled replication forks may also be processed via HR. Several DNA processing enzymes are involved in the regulation of replication initiated HR, mostly in its suppression, and we summarize the current knowledge of these processes in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The control of genome stability is of upmost importance for all living organisms. There are various levels of genetic control to guarantee genome stability in somatic cells of eukaryotes. Most of the basic mechanisms are conserved between eukaryotic organisms, including plants, fungi, and animals. In humans and other mammals, loss of these controls may lead to the occurrence of various forms of cancer in the long run. In contrast to mammals, the germ line is only established late in development in plants. Therefore, changes in the nuclear genome that occur during somatic growth might be transferred to the next generation. Thus, control of genome stability in somatic cells is of significant importance for plant genome evolution. Transposable elements are an important cause of genome rearrangements (see reviews of Bureau und Kakutani in this issue). In the following, we summarize the current knowledge about the modes and regulation of DNA recombination in somatic cells of Arabidopsis thaliana, have a look at known similarities and differences in other plant species, and discuss their putative role in genome evolution. On the one hand, we will concentrate on the current knowledge of the molecular mechanisms of double-strand break (DSB) repair in plants and, on the other hand, on factors that suppress replication-associated hyperrecombination.
DSB repair
DSBs can be repaired without the involvement of identical or almost identical sequences via non-homologous end joining (NHEJ), which is also referred to as “illegitimate recombination.” Alternatively, DSBs can also be repaired by the use of homologies via homologous recombination (HR) [for recent reviews, see also (Lieberman-Lazarovich and Levy 2011; Waterworth et al. 2011)]. In meiosis, DSBs are induced in a controlled way and are repaired via HR using mostly the homologous chromosome as template (for details, see the review of Sanchez-Moran and Armstrong in this issue). As a rule, in somatic plant cells, the main mechanism of DSB repair is NHEJ, but depending on the state of the cell cycle and the template used for repair, a significant part of the DSBs might also be repaired by the use of HR. DSBs can arise in somatic cells not only due to internal processes such as replication and transposon excision but also due to external genotoxins like radiation and chemicals. Most of our knowledge on the repair of DSBs in plants comes from experiments in which DSBs were artificially induced by the use of sequence-specific endonucleases at defined sites in plant genomes [see below for examples and for review see (Puchta 2005)].
Recently, precise genome engineering in plants became much more feasible due to the development of engineered nucleases like zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), or the clustered regularly interspaced short palindromic repeats; CRISPR-associated (CRISPR/Cas) system. These nucleases can be used for NHEJ-mediated targeted mutagenesis approaches as well as HR-mediated genome editing techniques [for recent reviews, see (Puchta and Fauser 2013; Voytas 2013)].
NHEJ—the mechanisms
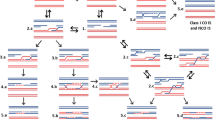
NHEJ is the main mode of DSB repair in somatic plant cells. This is documented by the fact that foreign DNA is frequently integrated randomly, without use of homology, into plant genomes. Recent analyses indicate that there are at least two different mechanisms of NHEJ that can be discriminated by the key players involved as well as by the patterns of the formed junctions (Fig. 1). Long known is the “canonical” or “classical” NHEJ (cNHEJ) pathway. More recently, an “alternative NHEJ” (aNHEJ) pathway was discovered (Mladenov and Iliakis 2011). Both pathways are characterized by the distinct patterns of repair junctions obtained. In cNHEJ, the DSB is repaired without much sequence loss and without microhomologies at the junction. This is due to the fact that in cNHEJ, the two double-stranded ends are protected from degradation by binding of a heterodimer of KU70 and KU80 before ligation takes place with little or even without any end processing. The enzyme ligase 4 is specifically involved in this last ligation step. The aNHEJ pathway is less well characterized and to a certain extent reminiscent of the single-strand annealing (SSA) pathway of HR (see below), since a particular amount of 3′-resection of the broken ends occurs by different kinds of exonucleolytic enzyme complexes. Thereby, single strands are produced that in case of few complementary nucleotides can anneal with one another. After trimming of the ends, religation occurs. aNHEJ thus results in a small number of identical base pairs at the junction site (microhomologies) combined with the deletion of some nucleotides. For Arabidopsis, it was demonstrated that PARP1 and XRCC1 are involved in this pathway (Charbonnel et al. 2011; Jia et al. 2013). Involvement of the same proteins has been shown for mammals as well. It seems that proteins from both pathways, e.g., KU80 and PARP1, compete for DSB ends, since in a ku80 mutant of Arabidopsis, an increased error-prone rejoining frequency with increased end degradation was found (Osakabe et al. 2010). Possibly, a third NHEJ pathway operates in Arabidopsis as some DNA ends are found to be joined in the absence of factors of cNHEJ and aNHEJ (Charbonnel et al. 2011).
The two main pathways of non-homologous end joining (NHEJ) in plant cells. Both the classical or canonical NHEJ (cNHEJ) pathway and the alternative NHEJ (aNHEJ) pathway operate efficiently in plant cells. Main actors in cNHEJ are the KU70/KU80 heterodimer and ligase 4. Due to their protection, only minimal processing of the broken ends occurs before ligation and thus at maximum a few base pairs are lost after rejoining. In contrast, broken ends are significantly processed during aNHEJ until short stretches of nucleotides that are complementary in both strands anneal with one another and overhangs are removed before ligation occurs. Therefore, microhomologies and larger deletions are often found after aNHEJ-mediated DSB repair
NHEJ—an important source of genome rearrangements
The outcome of NHEJ will often result in genomic change. Even small deletions occurring during cNHEJ as well as larger deletions associated with aNHEJ might result in a change of important information within genes or regulatory elements. However, NHEJ might also be associated with insertions. Sequences from elsewhere in the genome can be inserted into a break (Salomon and Puchta 1998). In most of these cases, microhomologies were found at the junction sites so that either an aNHEJ mechanism or a mode of copying similar to the synthesis-dependent strand annealing (SDSA) model described below for homologous DSB repair could be postulated to explain the phenomenon. As the inserted genomic sequences are still found at their original sites after DSB repair, it is highly likely that a copying mechanism operates in a majority of cases (Salomon and Puchta 1998). The suggested SDSA-like mechanism for the generation of insertions at DSBs is depicted in Fig. 2.
The potential of larger genomic rearrangements is given if more than one break is induced simultaneously. Multiple breaks are often induced not only after exposure to genotoxins but also during replication or following the activation of several copies of transposons. Although in the vast majority of cases the originally linked DSB ends are rejoined, sometimes erroneous joining occurs between ends that were not linked previously. Thus, new combinations of genetic information will be generated (Fig. 3). If two breaks arise within the same chromosome, a deletion (Siebert and Puchta 2002) or an inversion can take place. If the two breaks are ectopic on different chromosomes, translocations of chromosome arms via NHEJ can occur (Pacher et al. 2007). In case of several DSBs, even the reshuffling of sequence blocks between chromosomes might occur. Most often, these chromosomal rearrangements do not lead to viable progeny. Even if in some cases rearranged chromosomes might be propagated during somatic growth, it is by no means guaranteed that they can pass through meiosis. Nevertheless, even if only a small portion of rearranged chromosomes is transmitted to the next generation, somatic genomic rearrangements might have an influence on speciation. Inversions, for example, can be found in cultivars within the same species. In inverted regions, meiotic recombination is suppressed (Drouaud et al. 2006; Fransz et al. 2000).
NHEJ-mediated genome rearrangements. In case more than one DSB occurs at a given time point in a genome, different kinds of rearrangements might occur due to joining of previously unlinked broken ends. If two breaks occur within one chromosome, the intermediate sequences could be deleted (a), or if the fragment is rejoined in opposite orientation into the chromosome, an inversion will occur (b). If two breaks occur simultaneously in the arms of two different chromosomes, translocation can occur (c). If multiple breaks are present in the genome, even multiply rearranged chromosomes might be produced (d)
NHEJ—a major cause for genome shrinking?
A number of studies on genome size evolution in angiosperms indicated that the ancestral genome was most likely relatively small, with a tendency towards an increase in DNA content during evolution due to polyploidization and propagation of retrotransposons. So the question arose whether plants have indeed a “one-way ticket to obesity” (Bennetzen and Kellogg 1997). However, it soon became clear that organisms can also lose DNA and genomes can shrink. As NHEJ is often associated with deletions, it is now generally accepted that NHEJ contributes to genome shrinking over evolutionary time periods. DNA removal from the rice and Arabidopsis genome could be documented for sequence variation within several retrotransposon families. The identified nucleotide sequence changes in the two termini of individual retrotransposons were mainly small deletions associated with microhomologies indicative for NHEJ events (Devos et al. 2002; Bennetzen et al. 2005).
One can indeed speculate that genomes might evolve differently due to subtle variations in the availability of particular DSB repair factors; canonical and alternative NHEJ might work with slightly higher or lower efficiencies in different plant species. The greater the efficiency of aNHEJ, which is accompanied by larger deletions in comparison to cNHEJ, the more DNA might be lost over time. Alternatively, also faster resection of DNA ends by more active exonucleases or a less efficient DSB end protection due to the lack of sufficient numbers of KU70/KU80 heterodimers might lead to the loss of more DNA over time during genome evolution. The ratio of NHEJ-mediated insertions to deletions might also differ in a species-specific manner. In an early experimental study, differences in the processing of DSBs could indeed be documented between Arabidopsis and tobacco. Focusing on larger deletions and insertions, it could be demonstrated that after site-specific induction of a DSB within a negative selectable marker gene in tobacco, the deletion size was smaller while the number of insertion events was larger than in a comparable experiment in Arabidopsis. Thus, a direct correlation between genome size and NHEJ repair patterns was postulated (Kirik et al. 2000). Although such a correlation could not be seen in a recent study concentrating on small deletions at the break site without selection (Lloyd et al. 2012), bioinformatics data confirm that differences in genome size can be correlated with species-specific differences in DSB repair: In a recent study, it could be documented that A. thaliana has lost six times more introns than Arabidopsis lyrata since the divergence of the two species but gained very few introns. The pattern of the intron gains and losses revealed microhomologies between the splice sites of several gained and lost introns, suggesting that aNHEJ might not only be an important pathway for intron gain and loss but indeed might also be an important cause for species-specific differences in genome size (Fawcett et al. 2012).
HR mechanisms
Single-strand annealing
Single-strand annealing (SSA) is a repair mechanism that can be used efficiently if two homologous sequences are arranged in close proximity in a tandem orientation. Such a situation occurs quite regularly in plant genomes as in the case of local gene duplication or also in the clustered ribosomal genes. The mechanism is not conservative, since all sequence information between the respective repeats is lost. Thus, besides aNHEJ, SSA seems to be another mechanism by which DNA might be lost during genome evolution. Indeed, the presence of multiple single long terminal repeats (LTRs) as a result of LTR retrotransposon loss in the genomes of various plant species indicates that such events might also be transferred to the germ line.
The mechanism of SSA itself is quite simple to visualize (Fig. 4): Following DSB induction, single-stranded overhangs are produced via exonuclease-catalyzed resection. Both ends now carry single-stranded regions that are at least partially complementary to each other. These molecules can then directly anneal in this region and a chimeric DNA structure is formed. If this structure contains 3′-overhangs, the respective parts of the sequence will be trimmed; otherwise, single-stranded gaps will be filled in via DNA synthesis and finally nicks in the backbone will be sealed by a DNA ligase.
The two main pathways of HR in somatic plant cells. Repair can either occur via single-strand annealing (SSA) or synthesis-dependent strand annealing (SDSA). After the formation of a DSB (I), in both pathways, resection and production of 3′ overhanging single strands occur (II). In the case of SSA, direct annealing of both complementary single strands is the next step (III). Thus, all sequence information between the repeats is lost in the rejoined molecule (IV). In the case of SDSA, one 3′ single strand invades a homologous double strand and initiates a copying process that elongates the invading strand (III). Later on, the strand is set free and reannealed (IV) and the respective double strand is repaired (V). Thus, via SDSA, a restored dsDNA molecule without loss of genetic information is obtained
SSA can proceed in somatic plant cells almost as efficient as NHEJ: A study performed in tobacco analyzing an endonuclease-induced break indicated that up to one out of three DSBs is repaired via SSA, whereas the other are repaired by either cNHEJ or aNHEJ (Siebert and Puchta 2002). SSA can, in principle, also occur between two DNA molecules that were not linked before. If several DSBs are induced, chromosomal translocation by a chromosome arm exchange can occur via regions of ectopic homology. This could already be demonstrated by the use of a site-specific endonuclease in tobacco (Pacher et al. 2007). No proteins exclusively for SSA have been characterized in plants.
Synthesis-dependent strand annealing
In contrast to SSA, no sequence information is lost in classical HR reactions, but the DSB is repaired by the use of a homologous donor molecule. If the sequence of the template is identical to the broken site, the information content of the broken double strand can be restored completely. Thus, this kind of conservative reaction is in principle not mutagenic. There are two general mechanisms for the conservative repair of DSBs that differ in their outcome. In case of the so-called DSB repair (DSBR) mechanism (Szostak et al. 1983), double Holliday junctions are formed between the participating DNA molecules as recombination intermediates which have to be resolved. This resolution results in either crossovers (COs) or gene conversions. DSBR is an important mechanism during meiotic recombination (see review by Armstrong et al., this issue), where COs are the cause for the mixing of the parental genomes. In somatic plant cells, SDSA is the main mechanism of conservative HR repair.
In SDSA, after resection of the DSB 3′-end, invasion of a single strand into a homologous template occurs (Fig. 4). Thus a so-called D-loop is formed and DNA synthesis is initiated using the newly paired strand as a template. At a certain time point, synthesis stops and the extended strand is set free from the D-loop and hybridizes with complementary sequences of a single strand resulting from the resection of the other end of the break. After repair of the single-strand gaps, the genetic information of the broken double-stranded DNA molecule is restored. Thus, a gene conversion is achieved. That SDSA and not the DSBR mechanism is mainly used for the repair of DSBs in somatic cells could be shown by the use of a site-specific endonuclease for DSB induction: A main difference between DSBR and SDSA is that they predict different outcomes if only homology to one end of the DSB is available. To discriminate between the two pathways, Agrobacterium-mediated transformation experiments were performed in tobacco using a T-DNA with homology to only one end of the break compared with the results of experiments in which a T-DNA harboring homologous sequences to both ends of the break in the target locus was used. Whereas the DSBR model predicts that homology to both ends of the break is essential for recombination, SDSA-like events should only require a homologous region at one end. The recombination frequencies obtained with the T-DNA showing homology to both ends of the DSB were only one-third higher than those obtained using the one-ended construct (Puchta 1998). Thus, obviously, homology to only one end of the DSB is sufficient for an efficient HR reaction to occur in somatic plant cells and the SDSA model is most appropriate for its description. The small difference between the two experiments could be easily explained by one-sided invasion of the homologous end of the respective T-DNA via SDSA into the genomic locus.
That SDSA is the prevalent mode of conservative HR in somatic plant cells seems also meaningful in the light of genome stability. If DSBR operated efficiently in somatic plant cells, crossovers would occur regularly. Since multiple repeated sequences are found at ectopic positions in plant chromosomes, COs between such repeated ectopic sequences would occur. However, such COs would result in dicentric and acentric chromosomes and thus endanger genome stability in somatic cells.
A number of proteins have been characterized that have important functions in SDSA. In contrast, no proteins have been characterized in plants that are exclusively required for SSA, yet. The main difference between SDSA and SSA is that only in SDSA a strand exchange reaction is required. It has been demonstrated that in this reaction, homologues of the recombinase RecA, AtRAD51, and AtXRCC3, as well as the SWI2/SNF2 family ATPase AtRAD54, are involved (Fig. 4, III right). Neither of these proteins is involved in SSA (Roth et al. 2012). Other proteins such as the DNA helicases AtRECQ4A and AtFANCM as well as nucleases like AtMUS81 play some role in SDSA and at the same time a minor role in SSA (Mannuss et al. 2010; Roth et al. 2012). AtRECQ4A might be involved in strand resection in SSA and SDSA (Fig. 4, II). There are strong indications that in SSA, the AtRAD1/AtRAD10 heterodimer, a structure-specific flap endonuclease-like enzyme, is involved in trimming of the complementary strand before ligation (Fig. 4, III left) (Dubest et al. 2002). Surprisingly, a AtRAD51-independent role of the AtRAD51 paralogues AtXRCC2, AtRAD51B, and AtRAD51D in SSA was reported recently (Da Ines et al. 2013).
Various templates can be used by SDSA
Several different kinds of homologies are present for most sequences in plant genomes that could be used for SDSA-mediated HR (Fig. 5). In the S and G2 phases of the cell cycle, homologies from the sister chromatid are available. Current knowledge indicates that this is the kind of template that is most efficiently used. Unfortunately, due to the fact that a HR reaction utilizing the sister chromatid does not lead to any sequence change, the frequencies of such events could not be determined for plants. Besides being present on the sister chromatid, the same sequence is also present on the homologous chromosome in all diploid cells. Using the homing endonuclease I-SceI, an early study revealed that a DSB in a transgene was repaired by a mutated allelic template in about 1 out of 10,000 cases (Gisler et al. 2002). Thus, in contrast to meiotic recombination, the allelic sequence on the homolog is not a frequently used template for repair in somatic cells. A further alternative would be the usage of a homology on a different chromosome as a repair template (“ectopic” recombination). Indeed, such a situation was also tested using I-SceI and two homologous transgenes, one carrying an I-SceI site in an ectopic position in the tobacco genome. Interestingly, it turned out that about 1 out of 10,000 DSBs is repaired by the use of an ectopic sequence. The detailed molecular analysis of the recombinants indicated that HR did not occur at both ends of the DSB in all cases and that a combination of HR and NHEJ took place (Puchta 1999), which is in accordance with the mechanism of SDSA: After homology-dependent elongation of one break end and subsequent ejection of the elongated strand from the heteroduplex, this end was ligated with the second free end without use of homology (NHEJ). Notably, in an independent study based on DSB induction by a transposon, a similar efficiency of DSB-induced ectopic HR was reported in Arabidopsis (Shalev and Levy 1997). These results demonstrate that neither allelic sequences on the homologous chromosome nor ectopic sequences on heterologous chromosomes are efficient templates for DSB repair and breaks are in the overwhelming majority of cases repaired by NHEJ.
Templates for homologous recombination (HR). For SDSA, a homologous template is required for the copying process. Here, a number of different kinds of templates are available in the nucleus. a Homologous sequences on the sister chromatid in G2 and S phases of the cell cycle. b Intrachromosomal homologies. c Allelic sequences on the homologous chromosome. d Ectopic sequences on a different chromosome
Replication-associated HR
Various lines of experimental evidence indicate that besides DSBs, also the processing of replication-associated intermediates like stalled replication forks can be channeled into HR. In contrast to DSBs, recombination-inducing replication intermediates cannot be repaired by pathways that require two free DNA ends, notably NHEJ and SSA. It is therefore expected that the function of HR is more important during S phase than during the gap phases of the cell cycle, in which at least in mammals the NHEJ and the SSA pathways are predominantly active (Shrivastav et al. 2008). Replication forks can stall due to a number of different reasons, e.g., the presence of modified bases that are not suitable as a template for the replicative DNA polymerases, cross-linked DNA strands inhibiting their separation, or the increased tension in dsDNA due to supercoiling through the inactivation of topoisomerases.
Typically, lesions stalling replicative DNA polymerase are not repaired at the replication fork but bypassed in order to finish replication and repaired at a later time point. A common mechanism uses so-called translesion polymerases that can synthesize past modified DNA bases. Besides lesion bypass, several models have been put forward in which HR-like processes can be used by the cell to elongate daughter strands past DNA lesions. When a DNA strand cannot be elongated due to damage in one parental strand, the second daughter strand can still be elongated by DNA polymerase in a process called “overshoot synthesis.” In a process very similar to the SDSA mechanism, the shorter daughter strand can invade the sister duplex, form a D-loop, and, by copying sequence information from the intact sister strand, avoid using the damaged strand as a template. After rejection of the invaded strand and reannealing to its own parental strand, the lesion has been bypassed and replication can proceed. In a similar manner, following overshoot synthesis, the replication fork can be regressed so that both newly synthesized daughter strands can anneal to each other. By this way, a four-branched DNA structure is formed that is also called a “chicken foot.” Here, the shorter daughter strand can be elongated by a DNA polymerase with the sequence information of its longer sister. Following DNA synthesis, the replication fork can be formed again, with both daughter strands past the lesion and ready for replication to proceed (Fig. 6).
Recombination processes at the replication fork. a Following a lesion in the leading strand, replication cannot proceed and the replication fork stalls. To bypass the damage and proceed with replication, one daughter strand can invade into the sister chromatid. There, formation of a D-loop in a SDSA-like mechanism allows the elongation of the strand. Alternatively, replication fork reversal leads to the formation of a chicken foot structure that also facilitates strand elongation using the sister chromatid. In both cases, the replication fork can be reformed with the lesion site bypassed. b Unrepaired nicks in one strand lead to the formation of one-sided DSBs when they are encountered by the replication fork. Since no second free end is present, NHEJ and SSA mechanisms cannot be used. Therefore, HR mechanisms such as strand invasion of the sister chromatid need to be utilized for repair
In addition to lesion bypass pathways that directly involve the sister chromatid at the replication fork as explained above, there is also the possibility of the formation of a DNA break during replication. Such a break might happen due to the encounter of a nick in one strand by the replication fork or the action of an endonuclease following stalling of the fork. Even though a DSB is formed, in this case, there is no second end present with which it could be ligated by NHEJ. Therefore, such a so-called one-sided DSB must be repaired using homologous sequences, most probably from the sister chromatid due to its proximity by cohesion.
Using reporter lines in Arabidopsis that measure the relative stability of repeated sequences in the genome [for review, see (Puchta and Hohn 2012)], a number of factors have been investigated on their possible roles in HR in vivo. Quantification of HR events in wild type and mutant plants in the same recombination reporter background gives insight into the respective roles of these proteins and, by creating double mutants, into whether or not the factors contribute to different pathways. By measuring HR frequencies without the induction of DSBs by genotoxins, it is expected to find HR events that occur due to repair of naturally occurring endogenous DNA lesions. In contrast, treatment of such plant lines with DSB-forming agents, e.g., bleomycin, should mostly give classic HR events for DSB repair. Recent studies showed that a number of mutant lines display differences in HR frequencies depending on the presence or absence of genotoxin-induced DSBs. Mutants of RECQ4A, the Arabidopsis homolog of the RecQ family helicase Sgs1 in yeast and BLM in humans, show no change in HR frequency after treatment with bleomycin, but a strong increase of spontaneous HR events compared with wild type (Hartung et al. 2008). Thus, RECQ4A is involved in the suppression of HR events that are not initiated by a DSB but by endogenous DNA lesions. Even more striking, mutants of AtFANCM, which is a homolog of the human Fanconi anemia gene FANCM and yeast Mph1, display more spontaneous but fewer bleomycin-induced HR events compared with wild type (Knoll et al. 2012). AtFANCM therefore either has a function in regulating somatic HR outcomes depending on whether the initiating lesion was a DSB or not, or it has two different functions depending on factors such as cell cycle, DNA lesion, or HR pathway used. RECQ4A and FANCM are involved in different pathways of HR suppression as the double mutant showed a higher HR frequency of both single mutants. Interestingly, treatment with genotoxins that do not induce DSBs but are expected to affect replication, such as the cross-linking agent cisplatin, has shown a clear function of these factors in HR following such lesions. Following cisplatin induction, the overall frequency of HR events is higher than in untreated plants. However, in contrast to the results after bleomycin treatment, recq4A and fancm mutants here display increased HR frequencies compared with wild type plants. Similar results have been obtained after treatment with the methylating agent methyl methanesulfonate (MMS) (our unpublished results). This indicates that the spontaneous HR events witnessed in untreated plants might mostly be due to HR at stalled replication forks. It seems that for the correct processing of HR at stalled replication forks, a number of different DNA helicases are required. Most probably, they are needed to channel the DNA strands into the different possible pathways by unwinding intermediates. Moreover, some proteins containing helicase domains have also been shown to promote the annealing of DNA strands during recombination or to facilitate the transformation of DNA intermediates not by an unwinding but by a translocase action. The yeast Swi2/Snf2 family protein Rad5, for example, has been shown to promote the formation of chicken foot intermediates through replication fork regression in vitro. The Rad5 homolog of Arabidopsis, AtRAD5A, is also involved in the repair of replication fork-stalling DNA lesions and may be required for such a function (Mannuss et al. 2010).
The importance of HR for genome stability in somatic plant cells has mostly been researched in respect to classic DSBs. As discussed above, however, there seem to be also other instances when HR is active to protect cells even though no DSB is even formed. For future research, it will be most interesting to define how HR proteins interplay with the replisome in more detail and with other damage repair and bypass pathways which are interconnected with HR in plant cells.
Abbreviations
- aNHEJ:
-
Alternative NHEJ
- cNHEJ:
-
Classical NHEJ
- CO:
-
Crossover
- CRISPR/Cas:
-
Clustered regularly interspaced short palindromic repeats/CRISPR-associated
- DSB:
-
Double-strand break
- DSBR:
-
Double-strand break repair
- HR:
-
Homologous recombination
- LTR:
-
Long terminal repeat
- MMS:
-
Methyl methanesulfonate
- NHEJ:
-
Non-homologous end joining
- SDSA:
-
Synthesis-dependent strand annealing
- SSA:
-
Single-strand annealing
- TALEN:
-
Transcription activator-like effector nuclease
- ZFN:
-
Zinc finger nuclease
References
Bennetzen JL, Kellogg EA (1997) Do plants have a one-way ticket to genomic obesity? Plant Cell 9(9):1509–1514. doi:10.1105/tpc.9.9.1509
Bennetzen JL, Ma J, Devos KM (2005) Mechanisms of recent genome size variation in flowering plants. Ann Bot 95(1):127–132. doi:10.1093/aob/mci008
Charbonnel C, Allain E, Gallego ME, White CI (2011) Kinetic analysis of DNA double-strand break repair pathways in Arabidopsis. DNA Repair (Amst) 10(6):611–619. doi:10.1016/j.dnarep.2011.04.002
Da Ines O, Degroote F, Amiard S, Goubely C, Gallego ME, White CI (2013) Effects of XRCC2 and RAD51B mutations on somatic and meiotic recombination in Arabidopsis thaliana. Plant J 74(6):959–970. doi:10.1111/tpj.12182
Devos KM, Brown JK, Bennetzen JL (2002) Genome size reduction through illegitimate recombination counteracts genome expansion in Arabidopsis. Genome Res 12(7):1075–1079. doi:10.1101/gr.132102
Drouaud J, Camilleri C, Bourguignon P, Canaguier A, Bérard A, Vezon D, Giancola S, Brunel D, Colot V, Prum B, Quesneville H, Mézard C (2006) Variation in crossing-over rates across chromosome 4 of Arabidopsis thaliana reveals the presence of meiotic recombination “hot spots”. Genome Res 16(1):106–114. doi:10.1101/gr.4319006
Dubest S, Gallego ME, White CI (2002) Role of the AtRad1p endonuclease in homologous recombination in plants. EMBO Rep 3(11):1049–1054. doi:10.1093/embo-reports/kvf211
Fawcett JA, Rouze P, Van de Peer Y (2012) Higher intron loss rate in Arabidopsis thaliana than A. lyrata is consistent with stronger selection for a smaller genome. Mol Biol Evol 29(2):849–859. doi:10.1093/molbev/msr254
Fransz PF, Armstrong S, de Jong JH, Parnell LD, van Drunen C, Dean C, Zabel P, Bisseling T, Jones GH (2000) Integrated cytogenetic map of chromosome arm 4S of A. thaliana: structural organization of heterochromatic knob and centromere region. Cell 100(3):367–376
Gisler B, Salomon S, Puchta H (2002) The role of double-strand break-induced allelic homologous recombination in somatic plant cells. Plant J 32(3):277–284
Hartung F, Suer S, Knoll A, Wurz-Wildersinn R, Puchta H (2008) Topoisomerase 3alpha and RMI1 suppress somatic crossovers and are essential for resolution of meiotic recombination intermediates in Arabidopsis thaliana. PLoS Genet 4(12):e1000285. doi:10.1371/journal.pgen.1000285
Jia Q, den Dulk-Ras A, Shen H, Hooykaas PJ, de Pater S (2013) Poly(ADP-ribose)polymerases are involved in microhomology mediated back-up non-homologous end joining in Arabidopsis thaliana. Plant Mol Biol 82(4–5):339–351. doi:10.1007/s11103-013-0065-9
Kirik A, Salomon S, Puchta H (2000) Species-specific double-strand break repair and genome evolution in plants. EMBO J 19(20):5562–5566. doi:10.1093/emboj/19.20.5562
Knoll A, Higgins JD, Seeliger K, Reha SJ, Dangel NJ, Bauknecht M, Schropfer S, Franklin FC, Puchta H (2012) The Fanconi anemia ortholog FANCM ensures ordered homologous recombination in both somatic and meiotic cells in Arabidopsis. Plant Cell 24(4):1448–1464. doi:10.1105/tpc.112.096644
Lieberman-Lazarovich M, Levy AA (2011) Homologous recombination in plants: an antireview. Methods Mol Biol 701:51–65. doi:10.1007/978-1-61737-957-4_3
Lloyd AH, Wang D, Timmis JN (2012) Single molecule PCR reveals similar patterns of non-homologous DSB repair in tobacco and Arabidopsis. PLoS One 7(2):e32255. doi:10.1371/journal.pone.0032255
Mannuss A, Dukowic-Schulze S, Suer S, Hartung F, Pacher M, Puchta H (2010) RAD5A, RECQ4A, and MUS81 have specific functions in homologous recombination and define different pathways of DNA repair in Arabidopsis thaliana. Plant Cell 22(10):3318–3330. doi:10.1105/tpc.110.078568
Mladenov E, Iliakis G (2011) Induction and repair of DNA double strand breaks: the increasing spectrum of non-homologous end joining pathways. Mutat Res 711(1–2):61–72. doi:10.1016/j.mrfmmm.2011.02.005
Osakabe K, Osakabe Y, Toki S (2010) Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. Proc Natl Acad Sci U S A 107(26):12034–12039. doi:10.1073/pnas.1000234107
Pacher M, Schmidt-Puchta W, Puchta H (2007) Two unlinked double-strand breaks can induce reciprocal exchanges in plant genomes via homologous recombination and nonhomologous end joining. Genetics 175(1):21–29. doi:10.1534/genetics.106.065185
Puchta H (1998) Repair of genomic double-strand breaks in somatic plant cells by one-sided invasion of homologous sequences. Plant J 13(3):331–339. doi:10.1046/j.1365-313X.1998.00035.x
Puchta H (1999) Double-strand break-induced recombination between ectopic homologous sequences in somatic plant cells. Genetics 152(3):1173–1181
Puchta H (2005) The repair of double-strand breaks in plants: mechanisms and consequences for genome evolution. J Exp Bot 56(409):1–14. doi:10.1093/jxb/eri025
Puchta H, Fauser F (2013) Synthetic nucleases for genome engineering in plants: prospects for a bright future. Plant J. doi:10.1111/tpj.12338
Puchta H, Hohn B (2012) In planta somatic homologous recombination assay revisited: a successful and versatile, but delicate tool. Plant Cell 24(11):4324–4331. doi:10.1105/tpc.112.101824
Roth N, Klimesch J, Dukowic-Schulze S, Pacher M, Mannuss A, Puchta H (2012) The requirement for recombination factors differs considerably between different pathways of homologous double-strand break repair in somatic plant cells. Plant J 72(5):781–790. doi:10.1111/j.1365-313X.2012.05119.x
Salomon S, Puchta H (1998) Capture of genomic and T-DNA sequences during double-strand break repair in somatic plant cells. EMBO J 17(20):6086–6095. doi:10.1093/emboj/17.20.6086
Shalev G, Levy AA (1997) The maize transposable element Ac induces recombination between the donor site and an homologous ectopic sequence. Genetics 146(3):1143–1151
Shrivastav M, De Haro LP, Nickoloff JA (2008) Regulation of DNA double-strand break repair pathway choice. Cell Res 18(1):134–147
Siebert R, Puchta H (2002) Efficient repair of genomic double-strand breaks by homologous recombination between directly repeated sequences in the plant genome. Plant Cell 14(5):1121–1131
Szostak JW, Orr-Weaver TL, Rothstein RJ, Stahl FW (1983) The double-strand-break repair model for recombination. Cell 33(1):25–35
Voytas DF (2013) Plant genome engineering with sequence-specific nucleases. Annu Rev Plant Biol 64:327–350. doi:10.1146/annurev-arplant-042811-105552
Waterworth WM, Drury GE, Bray CM, West CE (2011) Repairing breaks in the plant genome: the importance of keeping it together. New Phytol 192(4):805–822. doi:10.1111/j.1469-8137.2011.03926.x
Acknowledgements
We regret that the majority of publications dealing with DNA recombination in plants could not be cited in this short review due to space limitations. We would like to thank Manfred Focke for his critical reading of the manuscript. Work on DNA recombination in our group has been funded over the years by the European Union, the Deutsche Forschungsgemeinschaft (DFG), and recently by the European Research Council (ERC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editors: Martin Lysak and Paul Fransz
Rights and permissions
About this article
Cite this article
Knoll, A., Fauser, F. & Puchta, H. DNA recombination in somatic plant cells: mechanisms and evolutionary consequences. Chromosome Res 22, 191–201 (2014). https://doi.org/10.1007/s10577-014-9415-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-014-9415-y