Abstract
Nucleosome depletion in the promoters has been indicated in yeasts, suggesting that nucleosome depletion in promoter might be a fundamental feature of eukaryotic transcriptional regulation. We compared the relationship between histone H3 acetylation at lysine 9 (K9) in promoter, gene expression level, and nucleosome density in the vicinity of the transcription start site (TSS), in HepG2 cells (human hepatocellular liver carcinoma cells). We found that the density of nucleosome is relatively low in the close vicinity of TSS flanked by H3 K9 significantly acetylated promoter, compared with that for genes without marked H3 K9 acetylation in promoter, regardless of their transcriptional activation status. Our results imply that the relative nucleosome depletion in the vicinity of TSS is not necessarily associated with active transcription, but with histone H3 K9 acetylation in promoter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Eukaryotic genomic DNA is packaged with histone proteins to form chromatin. The most fundamental repeating unit of chromatin is nucleosome. Chromatin structure is dependent on the modification of nucleosome core histones (Strahl & Allis 2000, Jenuwein & Allis 2001, Rice & Allis 2001). Chromatin structure, particularly in the promoter region, has been reported to be associated with gene expression. However, a recent study indicated that chromatin structure is not associated with gene expression, but with gene density (Gilbert et al. 2004).
Nucleosome depletion in the yeast promoters has been indicated independently (Bernstein et al. 2004, Lee et al. 2004, Liu et al. 2005, Sekinger et al. 2005). Their results suggested that nucleosome depletion in promoters might be a fundamental feature of eukaryotic transcriptional regulation. However, Bernstein et al. (2005) suggested that the marked nucleosome depletion evident in active promoters is not a characteristic of mammalian chromatin, based on analysis using immunoprecipitated chromatin from human cells with a modification-insensitive antibody against histone H3.
Recently, high-density genomic oligonucleotide microarrays have been developed as a tool to analyze chromatin structure (Cawley et al. 2004, Bernstein et al. 2005). In this study we carried out three lines of experiments using 25-base oligonucleotides arrays (35 bp resolution) that tile nonrepetitive portions of human chromosomes 21 and 22. First we sought to identify histone H3 K9 significantly acetylated sites using immunoprecipitated chromatin from HepG2 cells with an antibody against histone H3 acetylated at K9. It has been reported that modification of histone H3 at K9 regulates chromatin structure and function (Rice & Allis 2001). H3 K9 acetylation is enriched in transcriptionally active genes (Li et al. 2004, Liang et al. 2004, Bernstein et al. 2005, Roh et al. 2005). Secondly we measured the density of nucleosome using immunoprecipitated chromatin with a modification-insensitive antibody against histone H3. Lastly we measured the density of nuclesome using mapping of nucleosomal DNAs digested by MNase. We analyzed a variation of the density of nucleosome in the vicinity of TSS in relation to the histone H3 K9 acetylation status of promoter and gene expression, using the results.
Materials and methods
Chromatin immunoprecipitation
Approximately 2.5 × 107 HepG2 cells (JCRB1054, Japan Health Sciences Foundation) grown to 80% confluency in MEM with 10% FBS and 1% sodium pyruvate, were fixed with 1% formaldehyde (at room temperature for 10 min with rotation), incubated for 5 min after addition of glycine to a final concentration of 0.125 M, centrifuged, washed with ice-cold PBS, washed with TE (pH 8), and resuspended in TE buffer (1 ml). Samples were sonicated on ice with a Bronson 450 Sonifier for ten 1-min cycles (60% duty; output 4) and centrifuged at 13 000 × g for 10 min. The supernatant was transferred to a 15-ml tube and adjusted to a volume or 2.5 ml with addition of NaCl (final concentration, 100 mM), NP-40 (final concentration, 0.4%) and 250 μl BSA–blocked protein G–Sepharose 4 Fast Flow (Amersham Biosciences). The solution was incubated at 4°C for 1 h with rotation. Protein G–Sepharose beads were removed by column filtration. The pre-cleared solution was divided into aliquots of 1 ml (for binding with antibody to histone H3 acetylated at K9 or modification-insensitive antibody to histone H3), 1 ml (for no-antibody control), and 500 μl (input DNA control). The samples were incubated at 4°C overnight with diluted antibody (10-fold in IP buffer) against acetyl H3 K9 (5 μg; lot no. 28741, catalog no. 07-352, Upstate) or modification-insensitive antibody against histone H3 (5 μg; lot no. 69098, catalog no. ab1791, Abcam), supplemented with 50 μl Dynabeads Protein G (Dynal Biothech), and incubated at 4°C for 1 h with rotation. The supernatant was removed with Magical Trapper (Toyobo). The beads were washed 10 times with IP buffer, incubated at 65°C for 3.5 h after addition of 500 μl TE and 20 μl 5 M NaCl, incubated at 65°C for 0.5 h after addition of 1 μl RNase (Nippon Gene), and then incubated at 45°C for 1 h after addition of 10 μl 0.5 M EDTA, 20 μl 1 M Tris · HCl (pH 6.5), and 2 μl proteinase K (Nippon Gene). DNA was purified by extracting twice with phenol–chloroform (1:1) and precipitating in isopropanol.
LM-PCR and array hybridization
Oligonucleotides (5′-accgcgcgtaatacgactcactataggg-3′ and 5′-ccctatagtgagtcgtattaca-3′; 5′-terminus is phosphorylated) were annealed while the temperature was decreased gradually from 99°C to 15°C over 90 min. The blunted DNA sample was ligated with the annealed oligonucleotides by using T4 DNA ligase. The cassette DNA fragments were amplified by using Blend Taq Plus (Toyobo) with the oligonucleotide 5′-accgcgcgtaatacgactcactataggg-3′. PCR amplification was done under the following conditions: denaturation at 95°C for 1 min; 25 cycles of 95°C for 30 s, 55°C for 30 s, 72°C for 2 min; and a final extension at 72°C for 7 min. Amplified DNA was purified, fragmented with DNase I (Epicentre), and end-labeled with biotin-ddATP by using terminal deoxytransferase (Roche) (Kapranov et al. 2002, Kampa et al. 2004). Arrays were hybridized for 18 h at 45°C, washed, and scanned using the Affymetrix GeneChip System. Each sample was hybridized in triplicate.
Nucleosome binding DNA and array hybridization
Approximately 5 × 106 HepG2 cells grown to 80% confluence in MEM with 10% FBS and 1% sodium pyruvate were centrifuged, washed with ice-cold PBS, resuspended in the 10 ml cell lysis buffer (250 mM sucrose, 10 mM Tris-HCl [pH 7.4], 4 mM MgCl2, 0.1% PMSF, 0.1% Triton X-100), and lysed with 10 strokes of the Dounce homogenizer. The nuclei were pelleted by centrifugation at 2000 × g for 10 min at 4°C, and resuspended in the 2 ml nuclei wash buffer (250 mM sucrose, 10 mM Tris-HCl [pH 7.4], 4 mM MgCl2, 0.1 mM PMSF). The suspension was overlaid onto a 4 ml 30% sucrose cushion in nuclei wash buffer, and centrifuged at 2400 × g for 5 min at 4°C. The nuclei were washed twice in the nuclei digest buffer (10 mM NaCl, 10 mM Tris-HCl [pH 7.5], 3 mM MgCl2, 1 mM CaCl2, 0.1 mM PMSF), and resuspended in the 1 ml nuclei digest buffer. After addition of 20 U MNase (Takara), the suspension was incubated at 37°C for 10 min. The digestion was terminated by adding 20 μl EDTA. The digested nuclei were incubated at 45°C for 0.5 h after the addition of 2 μl RNase, and then incubated at 45°C for 1 h after the addition of 20 μl 0.5 M EDTA, 40 μl 1 M Tris-HCl [pH 6.5], and 4 μl proteinase K. DNA was purified by extracting twice with phenol–chloroform (1:1) and precipitating in isopropanol. The extracted DNA was dephosphorylated with alkaline phosphatase (calf intestine) (Takara) by the standard protocol, fragmented with DNase I (Epicentre), and end-labeled with biotin-ddATP by using terminal deoxytransferase (Roche). Arrays were hybridized for 18 h at 45°C, washed, and scanned using the Affymetrix GeneChip System. The sample was hybridized in triplicate.
Analysis of Affymetrix tiling array data
The genomic regions enriched for chromatin modification were detected by the standard protocol described previously (Kampa et al. 2004, Bernstein et al. 2005). The hybridization intensities of >1 M pairs of 25-mer probe sequences corresponding to genomic regions spaced at 35 bp on the genome were measured with the treatment and control samples in three replicates. A shift of the intensities in treatment relative to control data in a 400 bp window centered at each probe was evaluated by a Wilcoxon rank sum test, which assigned a p-value to the probe. A p-value cutoff, 10−4, was applied to define the genomic intervals enriched for chromatin modification. We used the Affymetrix software, GTRANS and GTAS (www.affymetrix.com/support/developer/downloads/TilingArrayTools) for the p-value calculation and the selection of the enrichment intervals, using 200 bp and 10−4 for the bandwidth and the p-value cutoff, respectively. Although the probe sequences were extracted from nonrepetitive regions of the two chromosomes, some of them still show various levels of duplication on the entire genome. We aligned all the 25-mer probe sequences to the whole human genome (NCBI version h33) to examine their level of duplication, and found that about 12% of them match perfectly to two or more sites. Since highly duplicated probes (particularly those repeated within a small region) may not measure accurately the hybridization intensity of a unique genomic site, we did not consider regions that contain probes matching to more than five sites on the entire genome. We determined the genomic intervals covered by the probes and therefore potentially detectable as fragments enriched for chromatin modification (blanket intervals), assuming that all the probes detected p-values below the cutoff. The sum of the blanket intervals amounts to 34.1 Mb.
Determination of transcription start sites on chromosomes 21 and 22
Transcription start sites were identified by aligning mRNA sequences from the RefSeq database (ftp.ncbi.nih.gov/refseq) by using Blat (Kent 2002) and Sim4 (Florea et al. 1999). We selected sequences aligned at more than 97% identity over at least 90% of the length including the initial 100 bp. In addition, we examined the coverage of the promoter region (± 1 kb around each TSS) by the probes, and discarded those lacking sufficient coverage (more than 100-bp overlap with the blanket intervals). We considered 293 and 535 TSS (separated at least 10 bp) determined for 260 and 491 loci on chromosomes 21 and 22, respectively. We assigned a representative transcript to each of the loci having multiple transcripts; for a locus with at least one H3 K9 significantly acetylated region detected, we used a transcript with TSS flanked by the maximum number of enriched regions, whereas one with the largest total region aligned to genome was used as a representative if a locus has no TSS flanked by a fragment enriched for H3 K9 acetylation.
Estimation of the expression level of genes in HepG2
The Affymetrix five-base density array data (Cheng et al. 2005) for cytosolic polyA+ RNA from the HepG2 cell line were used to estimate the expression level of genes in the two chromosomes. We used the average of signal intensities of the probes located in exons of each gene as an indicator of the expression level. Only genes mapped by 50 or more probes were used in this analysis.
Results and discussion
Identification of H3 K9 significantly acetylated sites
We used agarose gel electrophoresis to compare the immunoprecipitated DNA fragments with those generated in the absence of antibody (data not shown). We then used the ligation-mediated PCR (LM-PCR) method (Mueller & Wold 1989, Ren et al. 2000) to amplify the precipitated DNA fragments and unenriched control DNA (whole cell extract), and hybridized the bulk products from each reaction to tiling arrays of human chromosome 21 and 22. As done in a previous work (Bernstein et al. 2005), we used a significance threshold of p-value (p) < 10−4 to define genomic regions enriched for chromatin modification. Of sites identified as p-value (p) < 10−4, 93% were confirmed by quantitative PCR to have enrichment ratios of at least 4-fold (Bernstein et al. 2005). We identified 284 H3 K9 significantly acetylated sites, 228 (80.3%) of which overlapped the histone H3 K9/K14 significantly acetylated sites reported in the previous study (Bernstein et al. 2005).
To analyze the positional distribution of the significantly acetylated sites in relation to TSS, we considered a genomic interval spanning 1 kb upstream and downstream from TSS as promoter region. The oligonucleotide tiling array that we used covers 34.1 Mb of chromosomes 21 and 22, in which 828 TSSs are located over 751 loci (please see Materials and methods section). A representative TSS in each of the 751 loci was used for the subsequent study. Although a total of 1.07 Mb (3.1%) of the promoter regions is covered by the tiling array probes, 172 (60.6%) of the 284 H3 K9 significantly acetylated sites were located in promoter regions. This result is consistent with previous works (Bernstein et al. 2005, Roh et al. 2005), indicating that distribution of H3 K9 significant acetylation is biased toward promoter regions.
Comparison between H3 K9 significantly acetylated promoters and gene expression level
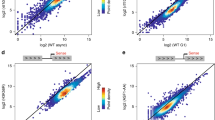
We compared the expression levels between genes with and without H3 K9 significantly acetylated sites in their promoters. We estimated the gene expression level using the Affymetrix five-base density array data of HepG2 (Cheng et al. 2005) (please see Materials and methods section). Figure 1 indicates that histone H3 K9 significantly acetylated promoters are associated with high levels of gene expression (expression level > 50), whereas H3 K9 non-significantly acetylated promoters are associated with both a high level (level > 50) and a low level of gene expression (level ≤ 10). Although genes with histone H3 K9 significantly acetylated promoters can have high levels of expression, H3 K9 significant acetylation is not essential to transcriptional activity (Figure 1). Promoters with a low level of gene expression are not significantly acetylated (Figure 1). Thus, H3 K9 deacetylation is associated with gene silencing.
Relationship between the expression level and H3 K9 acetylation in the promoter region. Genes with a high level of H3 K9 acetylation detected in the promoter region show a notably higher level of expression than those without significant acetylation detected in the promoter region. The average signal intensity of the Affymetrix five-base density array data for cytosolic polyA+ RNA from the HepG2 cell line (Cheng et al. 2005) of the probes located in exons of each gene is compared between genes with H3 K9 acetylation enrichment detected in the promoter region and those without the acetylation enrichment. We considered only a representative transcript for each locus.
Location of H3 K9 significantly acetylated sites in the promoters
It is believed that the K9 residue of histone H3 in the transcriptionally active promoter is often acetylated. However, the positional relationship between acetylated site and TSS has not been rigorously studied. Although Bernstein et al. (2005) showed the relationship between promoter, H3 K9 acetylated site, and H3 K4 methylated site, the positional relationship between acetylated site and TSS within the promoter region was not analyzed. We investigated the precise positional distribution of the H3 K9 significantly acetylated sites in the promoter region, in relation to TSS. We analyzed the location of the nearest significantly acetylated site to TSS within the promoter region. As a result only 16 of the 172 H3 K9 significantly acetylated sites in promoters coincide with the TSS (Figure 2). On the other hand, 68 and 88 of them are located upstream and downstream of TSS, respectively (Figure 2). Therefore, histone H3 K9 significantly acetylated promoters have a low level of H3 K9 acetylation in the close vicinity of TSS.
Location of the nearest significant H3 K9 acetylation site to transcription start site within promoter region. As an example, the nearest significant H3 K9 acetylation site to TSS of SMTN on chromosome 22 on the Integrated Genome Browser (Affymetrix). Blue bars show p-values. Green indicates the 5′ region of SMTN from Refseq database. Dot line indicates p = 10−4 threshold line.
Nucleosome density in the vicinity of TSS based on signal intensities of ChIP fragment using modification-insensitive antibody against histone H3
To elucidate whether or not the low level of H3 K9 acetylation in the vicinity of TSS is related to the absence of nucleosome in the region, the density of nucleosome at or nearest to TSS was compared between three sets of genes showing distinct combinations of the H3 K9 acetylation in promoter and level of gene expression. We compared the hybridization intensities of ChIP fragment using a modification-insensitive antibody against histone H3 at TSS of: (a) genes with H3 K9 significantly acetylated promoter, (b) genes without H3 K9 significantly acetylated promoter and showing a low expression (level ≤ 10), and (c) genes without H3 K9 significantly acetylated promoter and showing a high expression (level > 50).
Comparison between signal intensity distributions of set (a), (b), and (c) shows that the distribution of set (a) is shifted to the lower side (Figure 3A), indicating that histone H3 depletion (nucleosome depletion) prevails in the vicinity of TSS in histone H3 K9 significantly acetylated promoters. On the other hand, the distributions of set (b) and set (c) are so similar (Figure 3A), indicating that nucleosome is formed in the vicinity of TSS in histone H3 K9 non-significantly acetylated promoters regardless of the level of gene expression. The comparison of the signal intensity distribution between each of the three sets and all the promoters yielded similar results (Panel A of Supplementary Figure S1). We also compared the signal intensity distribution between two groups of significantly acetylated promoters with distinct levels of acetylation and found that the group of higher level of acetylation shows lower intensity distribution (Supplementary Figure S2).
Relationship between nucleosome occupancy in the vicinity of the transcription start site, acetylation enrichment, and level of gene expression. The level of nucleosome occupancy around TSS has been measured by hybridizing ChIP DNA fragments obtained with a modification-insensitive antibody against histone H3 (A) and by hybridizing nucleosomal DNA fragments digested by MNase (B). The normalized distribution of signal intensities detected within±200 bp from TSS was compared between three classes of TSS: (1) TSS flanked by significantly acetylated sites in the promoter region, (2) TSS without a significantly acetylated site in the promoter region, showing low a level of transcription and (3) TSS without a significantly acetylated site in the promoter region, showing a high level of transcription. The signal intensity at each probe position was computed using the Affymetrix software, GTRANS www.affymetrix.com/support/developer/downloads/TilingArrayTools). A Wilcoxon signed-rank test was applied to the hybridization intensities (background-subtracted intensity, PM − MM where PM and MM indicate intensities detected by a 25-mer perfectly matching and another one-base-mismatching to genome, respectively) measured at probes located within ±200 bp of every probe location, and the pseudomedian generated by the Wilcoxon test was assigned as an estimate of the signal intensity to the probe position. The signal intensity is given in log2 scale. A Mann–Whitney test yielded p < 10−4 in all the four pairwise comparisons of the intensity distributions between TSS flanked by significantly acetylated sites (1) and TSS without a significantly acetylated site in the promoter, showing either a high (2) or low (3) level of gene expression (top and middle panels). We could not find a significant difference in nucleosome occupancy between the two groups of TSS flanked by a non-significantly acetylated promoter with high and low gene expression, (2) and (3), with the Mann–Whitney test yielding p > 0.05 (bottom panel). In these plots we did not include probes that detected intensity 0 (zero intensity). The zero intensity is generated by probes in which MM detected a higher hybridization intensity than PM, thus indicating extremely weak signal intensities. The proportions of probes that detected zero intensity are 44% (369/835), 24% (637/2649), and 28% (195/693) in data (A), and 37% (306/835), 26% (693/2649), and 27% (185/693) in data (B), for (1), (2) and (3), respectively. The TSS flanked by significantly acetylated promoters (1) show a significantly higher ratio of zero-intensity probes than others, whereas the two groups of TSS with non-significantly acetylated promoters (2) and (3) show similar ratios of zero-intensity probes. Therefore the results of the Mann–Whitney test remain invariant with inclusion of the zero-intensity probes.
These results indicate that comparing signal intensities is more useful than p-value threshold method (Bernstein et al. 2005) in analysis of hybridization data of the ChIP fragment using a modification-insensitive antibody against histone H3.
Nucleosome density in the vicinity of TSS based on nucleosomal DNA mapping data
Next, we compared the hybridization intensities of nucleosomal DNA fragments digested by MNase in the vicinity of TSS. This experiment is the simplest and most direct method for the measurement of nucleosome density, because it does not need PCR (please see Materials and methods section). The result (Figure 3B) is consistent with Figure 3A. The comparison of the signal intensity distribution between each of the three sets and all the promoters yielded similar results (Panel B of Supplementary Figure S1). Therefore, results from the two different experiments indicate that nucleosome depletion in the vicinity of TSS is not necessarily associated with active transcription, but with histone H3 K9 acetylation in the promoter.
Nucleosome depletion in the vicinity of TSS is a fundamental feature of eukaryotic transcriptional regulation
The relation between histone H3 K9 significantly acetylated promoters and the densities of nucleosomes in the vicinity of the TSS is largely uninvestigated, especially in human cells. Our results strongly support the theory that nucleosome depletion in the vicinity of TSS is a fundamental feature of eukaryotic transcriptional regulation. Figure 4 illustrates a model for typical transcriptional silencing through regulating the nucleosome formation from gene expression with an H3 K9 significantly acetylated promoter. If a protein or a protein complex binds in the vicinity of TSS, recruiting histone remodeling complex with a histone deacetylase, the histones H3 around TSS become deacetylated at K9. This event is probably the first step to gene silencing through regulating the formation of nucleosomes in the vicinity of TSS. On the other hand, some transcriptionally active promoters are not significantly acetylated at K9 of histone H3 (Figure 1). This strongly suggests that gene silencing needs other chromatin structural changes after the nucleosome formation in the vicinity of TSS.
References
Bernstein BE, Kamal M, Lindblad-Toh K et al. (2005) Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120: 169–181.
Bernstein BE, Liu CL, Humphrey EL et al. (2004) Global nucleosome occupancy in yeast. Genome Biol 5: R62.
Cawley S, Bekiranov S, Ng HH et al. (2004) Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116: 499–509.
Cheng J, Kapranov P, Drenkow J et al. (2005) Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 308: 1149–1154.
Florea L, Hartzell G, Zhang Z et al. (1999) A computer program for aligning a cDNA sequence with a genomic DNA sequence. Genome Res 8: 967–974.
Gilbert N, Boyle S, Fiegler H et al. (2004) Chromatin architecture of the human genome: gene-rich domains are enriched in open chromatin fibers. Cell 118: 555–566.
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293: 1074–1080.
Kampa D, Cheng J, Kapranov P et al. (2004) Novel RNAs identified an in-depth analysis of the transcriptome of human chromosomes 21 and 22. Genome Res 14: 331–342.
Kapranov P, Cawley SE, Drenkow J et al. (2002) Large-scale transcriptional activity in chromosomes 21 and 22. Science 296: 916–919.
Kent W (2002) BLAT: the BLAST-like alignment tool. Genome Res 12: 656–664.
Lee C-K, Shibata Y, Rao B et al. (2004) Evidence for nucleosome depletion at active regulatory regions genome-wide. Nat Genet 36: 900–905.
Li T, Vu TH, Ulaner GA et al. (2004) Activating and silencing histone modifications form independent allelic switch regions in the imprinted Gnas gene. Hum Mol Genet 13: 741–750.
Liang G, Lin JCY, Wei V et al. (2004) Distinct localization of histone H3 acetylation and H3-K4 methylation to the transcription start sites in the human genome. Proc Natl Acad Sci USA 101: 7357–7362.
Liu CL, Kaplan T, Kim M et al. (2005) Single-nucleosome mapping of histone modifications in S. cerevisiae. PLoS Biol 3: e328.
Mueller PR, Wold B (1989) In vivo footprinting of a muscle specific enhancer by ligation-mediated PCR. Science 246: 780–786.
Ren B, Robert F, Wyrick JJ et al. (2000) Genome-wide location and function of DNA binding proteins. Science 290: 2306–2309.
Rice JC, Allis CD (2001) Histone methylation versus histone acetylation: new insights into epigenetic regulation. Curr Opin Cell Biol 13: 263–273.
Roh T-Y, Cuddapah S, Zhao K (2005) Active chromatin domains are defined by acetylation islands revealed by genome-wide mapping. Genes Dev 19: 542–552.
Sekinger EA, Moqtaderi Z, Struhl K (2005) Intrinsic histone-DNA interaction and low nucleosome density are important for preferential accessibility of promoter regions in yeast. Mol Cell 18: 735–748.
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403: 41–45.
Acknowledgements
We thank Tom Gingeras and his research group in Affymetrix, Affymetrix Japan, Harukazu Suzuki, Jun Kawai, and Haruhiko Koseki for helpful advice and useful suggestions. This work was supported in part by a Research Grant for the Genome Network Project and a Research Grant for the RIKEN Genome Exploration Research Project from the Ministry of Education, Culture, Sports, Science and Technology of the Japanese Government, a Research Grant for Advanced and Innovational Research Program in Life Science, and the Strategic Programs for R&D of Riken to Y.H.
Author information
Authors and Affiliations
Corresponding author
Additional information
†These authors contributed equally to this work.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Nishida, H., Suzuki, T., Kondo, S. et al. Histone H3 acetylated at lysine 9 in promoter is associated with low nucleosome density in the vicinity of transcription start site in human cell. Chromosome Res 14, 203–211 (2006). https://doi.org/10.1007/s10577-006-1036-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-006-1036-7