Abstract
Genetics is considered as an important risk factor in the pathological changes of Parkinson’s disease (PD). Substantia nigra (SN) is thought to be the most vulnerable area in this process. In recent decades, however, few related long non-coding RNAs (lncRNAs) in the SN of PD patients had been identified and the functions of those lncRNAs had been studied even less. In this study, we sought to investigate the lncRNA expression profiles and their potential functions in the SN of PD patients. We screened lncRNA expression profiles in the SN of PD patients using the lncRNA mining approach from the ArrayExpress database, which included GSE20295. The samples were from 11 of PD and 14 of normal tissue samples. We identified 87 lncRNAs that were altered significantly in the SN during the occurrence of PD. Among these lncRNAs, lncRNA AL049437 and lncRNA AK021630 varied most dramatically. AL049437 was up-regulated in the PD samples, while AK021630 was down-regulated. Based on the results, we focused on the potential roles of the two lncRNAs in the pathogenesis of PD by the knockdown of the expression of AL049437 or AK021630 in human neuroblastoma SH-SY5Y cell line. Results indicated that the reduction in AL049437 level increased cell viability, mitochondrial transmembrane potential (Δψm), mitochondrial mass, and tyrosine hydroxylase (TyrH) secretion. By contrast, the knockdown of AK021630 resulted in the opposite effect. Based on these results, we speculated that lncRNA AL049437 likely contributed to the risk of PD, while lncRNA AK021630 likely inhibited the occurrence of PD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Parkinson’s disease (PD) is the second most common neurodegenerative disease in the world, affecting approximately 1 % of individuals older than 60 years (Samii et al. 2004). Neuronal loss in the substantia nigra pars compacta (SNc) and the subsequent loss of striatal dopamine content are considered responsible for the classical motor features of PD (Dexter and Jenner 2013; Samii et al. 2004; Sherer et al. 2001). Various epidemiological and experimental investigations have shown that old age, environmental risk factors, and genetic factors are related to the pathogenesis of PD (Bonifati 2014; Campdelacreu 2014; Collier et al. 2011; Wirdefeldt et al. 2011).
Only 2.94 % of the genome has the GENCODE-annotated exons of protein-coding genes, while other genes are transcribed as non-coding RNAs (ncRNAs) (An integrated encyclopedia of DNA elements in the human genome 2012). ncRNAs are divided into housekeeping ncRNAs and regulatory ncRNAs. Regulatory ncRNAs are further classified into two categories according to nucleotide length. Those regulatory ncRNAs with less than 200 nucleotides are often known as short/small ncRNAs, including microRNAs (miRNAs), small interfering RNAs, and Piwi-associated RNAs. Another subset of regulatory ncRNAs, with length greater than 200 nucleotides, is known as long non-coding RNAs (lncRNAs) (Nagano and Fraser 2011).
lncRNAs could regulate gene expression at post-transcriptional level, such as translation (Wei et al. 2015), RNA maturation (Li et al. 2015; Savic et al. 2014), and transport (Lau 2014; Mathieu et al. 2014). They also exert their effects on transcriptional gene silencing through the regulation of chromatin structure (Guttman and Rinn 2012). Recent research shows that lncRNA may play roles in central nervous system (CNS) development, neuron function and maintenance, and neurodegenerative diseases including PD, Alzheimer’s disease (AD), Huntington’s disease (HD), and amyotrophic lateral sclerosis (ALS) (Wu et al. 2013). Mutations in the lncRNA of PTEN (phosphatase and tensin homolog)-induced kinase 1 (PINK1) could cause early-onset PD (Hatano et al. 2004). The modulations of lncRNA and alternative splicing were identified in Parkinson’s leukocytes by RNA sequencing (Soreq et al. 2014). However, the lncRNA expressions in the SN of PD patients have not been systematically demonstrated. In addition, the potential function of various lncRNAs in PD pathogenesis has not been fully investigated.
The pathological changes of PD occur mainly in the SN. In this study, we revealed a set of lncRNAs which were differentially expressed in the SN of PD by analyzing publicly available data sets from the ArrayExpress. We then investigated the possible roles of lncRNA AL049437 and AK021630 in the pathogenesis of PD.
Materials and Methods
Microarray Data of PD
Affymetrix Human Genome U133A Array for the SN of PD and normal tissue were downloaded from the GEO: GSE20295. GSE20295 consisted of 11 Parkinson’s disease and 14 normal tissues.
Microarray Analysis
The raw CEL files were downloaded from GEO database and background was adjusted using Robust Multichip Average. GATExplorer was used to process microarrays on a local computer for analyses of gene expressions of lncRNAs (Risueno et al. 2010), which has been used by Hu et al. (2014) and Chen et al. (2014). This GATExplorer provides a series of R packages, designed to be used with Bioconductor tools, which allowed simple probe mapping of the data included in GATExplorer. A type of files called lncRNA Mapper was also obtained from GATExplorer, which included the probes that were not mapped to any coding region but were mapped to a database for non-coding RNA of human derived from RNAdb (Pang et al. 2005). A customized R script was used to perform a microarray expression calculation according to the re-mapping data (file ncrnamapperhgu133acdf_3.0). Each lncRNA should include at least a minimum of 3 probe mappings in the corresponding lncRNA entity.
Cell Culture and Treatment
The human neuroblastoma SH-SY5Y cell line (CRL-2266) was purchased from American Type Culture Collections (ATCC) (Rockville, MD, USA). Human neuroblastoma SH-SY5Y cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10 % (v/v) fetal bovine serum (FBS) plus 1 % penicillin/streptomycin at 37 °C under 5 % CO2 and 95 % oxygen in humidified air, and the medium was changed every 2 days. When SH-SY5Y cells in 60-mm dishes were 70 % confluent, they were used for further treatment.
Rotenone (Sigma-Aldrich) was dissolved in dimethyl sulfoxide (DMSO). SH-SY5Y cells were incubated for 24 h with rotenone at different concentrations (10–500 nM).
siRNAs Interference
The siRNA oligos were synthesized by Shanghai GenePharma Co., Ltd. The sequence of siRNA duplexes were as follows: non-specific siRNA (negative oligo): sense 5′-UUCUCCGAACGUGUCACGUTT-3′, antisense 5′-ACGUGACACGUUCGGAGAATT-3′; siAL049437: sense 5′-GAGCUUGAAUAGUGAUCUUTT-3′, antisense 5′-AAGAUCACUAUUCAAGCUCTT-3′; and siAK021630: sense 5′-GCAUUGUUCAUUUCCGCUATT -3′, antisense 5′-UAGCGGAAAUGAACAAUGCTT-3′. Cells were transfected with 100 nM of siRNA oligos using lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. The knockdown efficiency was determined by qRT-PCR at 24 h post transfection. The transfected cells were used for subsequent experiments.
The experiment was divided into four groups: (1) a control group, namely SH-SY5Y cells with lipofectamine 2000 plus reagent in OptiMEM; (2) non-specific siRNA group in which SH-SY5Y cells transfected with negative oligo were incubated with rotenone for 24 h; (3) siAL049437 group in which SH-SY5Y cells were transfected with siAL049437 oligo and then incubated with rotenone for 24 h; and (4) siAK021630 group in which SH-SY5Y cells were transfected with siAK021630 oligo and then incubated with rotenone for 24 h.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
RNA was isolated using RNeasy Mini Kit (Qiagen). Reverse transcriptase (Invitrogen) was employed for random-primed first-strand complementary DNA (cDNA) synthesis. qRT-PCR was carried out on ABI Prism 7300 detection system using SYBR Green PCR master mix. The ΔΔCt method was used to quantify comparatively the amount of mRNA level. Primer sequences were listed below: AL049437: 5′-GAAAGGGAGGGGACGTTTCA-3′ (forward) and 5′-GGCTCCCATGACCCTCTTTT-3′ (reverse); AK021630, 5′-AACACCAGCCAAGAGGTAGC-3′ (forward) and 5′-CCACTCAAGGCCAGTTCAGT-3′ (reverse).
Cell Viability Assay
The viability of SH-SY5Y cells under various treatment conditions was assayed using CCK-8 (Wako, Richmond, VA, USA) in accordance with the manufacturer’s protocol. CCK-8 solution was added to each well at a final concentration of 10 % (v/v) and incubated for 2.5 h at 37 °C. The absorbance was measured at 490/630 nm by the microplate reader (Bio-Rad). The experiments were repeated, with at least six replications of each treatment.
Measurements of Mitochondrial Membrane Potential and Mitochondrial Mass
Mitochondrial membrane potential (Δψm) was measured using the tetramethylrhodamine methyl ester (TMRM; Life Technologies). TMRM is a cell-permeable cationic fluorescent dye that is readily sequestered by active mitochondria. Cells with different treatments were incubated with 50 nM TMRM at 37 °C for 30 min, washed with PBS twice, and fixed. Fluorescence images were acquired in a fluorescence confocal microscope (Leica Wetzlar, Germany) and the red photo multiplier channel was used for emission. The wavelengths of excitation and emission for TMRM are 530 and 590 nm, respectively. Images were taken from six random fields per coverslip. The fluorescence intensity was analyzed at a wavelength of 530 nm excitation and 590 nm emission by microplate reader (BioTek).
Mitochondrial mass was estimated by fluorescence of MitoTracker Green FM (Life Technologies), independent of the mitochondrial membrane potential. The cells with different treatments were incubated with 100 nM MitoTracker Green FM at 37 °C for 30 min, washed with PBS twice, and fixed. The fluorescence intensity was analyzed at a wavelength of 490 nm excitation and 516 nm emission by microplate reader (BioTek).
Western Blot Assays
SH-SY5Y cells were harvested at 24 h after transfection and lysed in Triton X-100 lysis buffer for 30 min at 4 °C. The primary antibodies included rabbit polyclonal anti-tyrosine hydroxylase (TyrH) antibody (Abcam, 1:500 dilutions) and anti-GAPDH antibody (Abcam, 1:1000 dilution). The secondary antibody included goat anti-rabbit IgG conjugated with HRP (horseradish peroxidase) at a dilution of 1:1000. GAPDH was used as an internal control. Representative images from one of three independent experiments were shown. The band intensity was quantified using ImageJ software.
Statistical Analysis
All data were expressed as mean ± S.D. Statistical analyses were accomplished using SPSS 16.0 for windows (SPSS, Chicago, IL). The statistical evaluations of data were performed using one-way analysis of variance (ANOVA) and t test. P < 0.05 was considered statistically significant.
Results
LncRNA Expression Profile in PD Patients
Affymetrix Human Genome U133A Array consisted of ~698 lncRNAs collected from the Parkinson’s disease samples. To identify differentially expressed genes, BRB-ArrayTools was used to compare the samples of SN from PD with the normal tissue samples. The raw P value was corrected by the Benjamin and Hochberg method to circumvent the multi-test bias (Chen et al. 2014). The fold change value larger than 4 or less than 0.25 and false discovery rate (FDR) <0.01 were selected as cutoff criteria for DEGs.
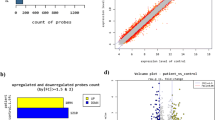
In total, the expressions of 87 lncRNAs in the SN were significantly altered during the pathogenesis of PD (Table 1). In the 87 lncRNAs of the SN from PD samples, the levels of 42 lncRNAs were elevated and those of 45 lncRNAs declined. It was worth noting that LncRNA AL049437 was most significantly up-regulated in the PD samples, while AK021630 was most significantly down-regulated (Fig. 1a, b). Thus, lncRNAs of AL049437 and AL021630 were further investigated in the pathogenesis of PD.
Cluster heat map of 87 differentially expressed lncRNAs from 11 of PD and 15 of normal tissue samples. Each column represents one sample and the row represents lncRNAs. Gene expression levels are shown as follows: blue high expression; green low expression (a). A volcano plot representation of differential expression test results indicated the magnitude (log2-transformed fold change, x-axis) and significance (P value, inverted y-axis) of all lncRNAs (b)
Changes of Cell Viabilities in the Knockdown of Two lncRNAs (AL049437 and AK021630)
To explore the roles of lncRNA AL049437 and AL021630 in neuronal activities in PD, we first detected the expression of lncRNAs (AL049437 and AK021630) in SH-SY5Y cells. Results showed that the levels of lncRNA AL049437 were significantly down-regulated, while those of lncRNA AK021630 were up-regulated (Fig. 2a, b). To further explore its roles in neuronal apoptosis in PD, RNAi specific to lncRNA AL049437 and AK021630 was employed, which significantly reduced lncRNA AL049437 and AK021630 levels in SH-SY5Y cells (supplementary Fig. 1 A, B). To establish a PD model in vitro, we exposed SH-SY5Y cells to rotenone at different concentrations (10–500 nM) for 24 h and chose 500 nM as the most appropriate concentration for the PD cell model (cell viability reduced to 53.5 ± 5.86 % (P < 0.05) vs. control) (Fig. 2a), which was also consistent with a previous report (Condello et al. 2011). We then interfered the expression of lncRNA AL049437 and AK021630 in this PD cell model, and found that knockdown of AL049437 increased SH-SY5Y cell viability, while knockdown of AK021630 reduced SH-SY5Y cell viability, comparing with non-specific siRNA group (Fig. 2b).
Dose-dependent effect of rotenone on cell viability determined by the CCK-8 method. The levels of lncRNAs (AL049437 and AK021630) in SH-SY5Y cells were detected by qRT-PCR. It showed that the levels of lncRNA AL049437 were significantly down-regulated, while those of lncRNA AK021630 were up-regulated (a, b). With the concentration of rotenone increasing, the cell viability was reduced. Compared with control, the dose of 500 nM of rotenone reduced cell viability to 53.5 ± 5.86 %. # P < 0.01 versus control (c). The effect of the knockdown of lncRNAs (AL049437 or AK021630) on the cell viabilities was demonstrated by CCK-8 (d). Compared with non-specific siRNA group, knockdown of AL049437 increased the cell viability, while knockdown of AK021630 reduced the cell viability. # P < 0.05 vs. the sample with non-specific siRNA (d)
Effects of Knockdown of Two lncRNAs (AL049437 and AK021630) on the Δψm and The Mitochondrial Mass
Mitochondria play an important role in the pathogenesis of PD (Chaturvedi and Beal 2013). For analyses of the effect of the two lncRNAs (AL049437 and AK021630) on the Δψm and the mitochondrial mass, AL049437 lncRNA or AK021630 lncRNA was knocked down. The Δψm was tested by TMRM and the mitochondrial mass was detected by MitoTracker Green FM. Compared with non-specific siRNA group, knockdown of AL049437 increased the quantity and density of TMRM fluorescent signal, while the knockdown of AK021630 led to decreases in the quantity and density of TMRM fluorescent signal examined by fluorescence confocal microscope and microplate reader (Fig. 3a–e). The results suggested that knockdown of AL049437 increased the Δψm expression, while knockdown of AK021630 reduced the Δψm expression.
Effects of the knockdown of the lncRNA (AL049437 or AK021630) on the Δψm and the mitochondrial mass. The Δψm was tested by TMRM and the mitochondrial mass was detected by MitoTracker Green FM (a–d). The density and quantity of TMRM fluorescent signal expressed in fluorescence confocal microscope. Compared with non-specific siRNA group, knockdown of AL049437 increased the density and quantity of TMRM fluorescent signal, while the knockdown of AK021630 decreased the density and quantity of TMRM fluorescent signal (e). The density of TMRM and MitoTracker Green FM fluorescent signal was examined by a microplate reader. Compared with non-specific siRNA group, the knockdown of AL049437 increased the density of MitoTracker Green FM fluorescent signal, while the knockdown of AK021630 reduced the density of MitoTracker Green FM fluorescent signal (f). # P < 0.05 compared with non-specific siRNA
In comparison with non-specific siRNA group, knockdown of AL049437 increased the fluorescent density of MitoTracker Green FM and knockdown of AK021630 reduced the fluorescent density of MitoTracker Green FM (Fig. 3f). The result indicated that the knockdown of AL049437 increased the mitochondrial mass and the knockdown of AK021630 caused a decrease in the mitochondrial mass.
Effects of Knockdown of lncRNAs (AL049437 and AK021630) on the TyrH Expression
Tyrosine hydroxylase (TyrH) is one of the major agents in determining dopamine levels (Daubner et al. 2011). For analyses of the effect of the two lncRNAs (AL049437 and AK021630) on the TyrH expression, AL049437 lncRNA or AK021630 lncRNA was knocked down and TyrH expressions were detected by Western blot analysis.
Results indicated that, compared with non-specific siRNA group, knockdown of AL049437 up-regulated the expression of TyrH, while knockdown of AK021630 down-regulated the expression of TyrH (Fig. 4). Thus, we speculated that lncRNA AL049437 increased the risk of PD, while lncRNA AK021630 inhibited the occurrence of PD.
Effects of the knockdown of lncRNAs (AL049437 or AK021630) on the TyrH expression. TyrH expressions were detected by Western blot analysis. Compared with non-specific siRNA group, the knockdown of AL049437 up-regulated the expression of TyrH, while knockdown of AK021630 down-regulated the expression of TyrH. # P < 0.05 vs. the sample with non-specific siRNA
Discussion
In this study, we found 87 lncRNAs in the SN, which were differentially expressed between the PD and normal tissue samples. Among these lncRNAs, AL049437 and AL021630 were changed most dramatically and thus were further investigated in the pathogenesis of PD. Our results indicated that AL049437 may induce the occurrence of PD, while AK021630 may inhibit the development of PD.
Genetics is an important contributing factor in the pathogenesis of PD. Approximately 15 % PD patients have a first-degree relative suffering from PD (Samii et al. 2004). 5 % PD patients have a mutation of one of several specific nuclear genes including α-synuclein (SNCA), Parkin, PTEN (phosphatase and tensin homolog)-induced kinase 1 (PINK1), DJ-1 or leucine-rich repeat kinase 2 (LRRK2) gene, and HTR2A (Bonifati 2014; Klein and Schlossmacher 2006; Samii et al. 2004). PD can also result from mitochondrial gene variations and mutations (Klein and Schlossmacher 2006). In addition, miRNA has been illuminated to be related to the PD pathogenesis (Srivastava et al. 2011).
Unlike miRNAs, lncRNAs have only recently emerged as a major class of eukaryotic transcripts and have been shown to be involved in the regulation of gene expression through diverse mechanisms that are not yet fully understood. For example, they are known to be able to serve as mediators of mRNA decay, structural scaffolds for nuclear substructures, host genes for miRNAs, and regulators of chromatin remodeling (Wapinski and Chang 2011). lncRNAs participate in a wide range of biological processes and diseases, including PD pathogenesis. For example, lncRNA PINK1 mutation has been demonstrated to be an important cause of PD (Morais et al. 2009). By RNA sequencing, lncRNA and alternative splicing modulations have been validated in blood leukocytes of PD patients (Soreq et al. 2014). In their study, RNA sequencing was used to unbiasedly and comprehensively investigate the differentially expressed transcripts. At this stage, RNA sequencing is a rather costly and time-consuming approach. For the pathogenic research of PD, the RNA from blood is no good from the SN. In our study, a large group of lncRNA-specific probes was fortuitously represented on the commonly used microarray platform (Affymetrix HG-U133 plus 2.0), so we initially mined previously published gene expression microarray data from the ArrayExpress, and conducted lncRNA profiling on the samples from the SN of PD patients. Although only a fraction of human lncRNAs (~698) were included in the analysis, it might better confirm that lncRNAs would participate in the pathogenesis of PD.
In this research, results indicated that 87 lncRNAs apparently changed in the SN of PD samples by probing and integrating currently available microarray data. 42 lncRNAs were elevated in PD samples and 45 lncRNAs declined.
The suppression of complex I which transports electrons in the inner mitochondrial membrane can induce parkinsonism. Several PD-associated genes interfere with the pathways regulating mitochondrial function, morphology, and dynamics (Andalib et al. 2014; Sai et al. 2012; Winklhofer and Haass 2010). So the level of mitochondrial integrity is very important in the pathogenesis of PD. In this research, we further investigated the functions of two lncRNAs of AL049437 and AK021630 in PD pathogenesis. In the patients with PD, AL049437 was found to be up-regulated most significantly, whereas AK021630 was found to be down-regulated most significantly. The cell model of rotenone for parkinsonism was applied. The rotenone models for parkinsonism in vitro and in vivo were widely used in experiments (Chiu et al. 2015; Johnson and Bobrovskaya 2015). Knockdown of AL049437 increased the cell viability, Δψm, mitochondrial mass, and TyrH in cell model of parkinsonism, while AK021630 had the opposite effect. These results suggested that lncRNA AL049437 might contribute to the risk of PD, while lncRNA AK021630 might inhibit the occurrence of PD. Nevertheless, given that both AL049437 and AK021630 intersect protein-coding genes, it is possible that down-regulation of lncRNAs could interfere with mRNA levels of the overlapping protein-coding genes. Thus, the observed phenotypes may be due to a direct effect of lncRNAs and/or an indirect effect through regulation of overlapping protein-coding genes. Further experiments will clearly address the exact mechanisms.
In conclusion, this study first provided the altered expressions of lncRNAs in the SN of PD. Our results also suggested that lncRNAs AL049437 and AK021630 are involved in the pathogenesis of PD. Thus, these two lncRNAs might serve as therapeutic targets for PD.
References
An integrated encyclopedia of DNA elements in the human genome (2012) Nature 489:57–74. doi:10.1038/nature11247
Andalib S, Vafaee MS, Gjedde A (2014) Parkinson’s disease and mitochondrial gene variations: a review. J Neurol Sci 346:11–19. doi:10.1016/j.jns.2014.07.067
Bonifati V (2014) Genetics of Parkinson’s disease–state of the art, 2013. Parkinsonism Relat Disord 20(Suppl 1):S23–S28. doi:10.1016/S1353-8020(13)70009-9
Campdelacreu J (2014) Parkinson disease and Alzheimer disease: environmental risk factors. Neurologia 29:541–549. doi:10.1016/j.nrl.2012.04.001
Chaturvedi RK, Beal MF (2013) Mitochondrial diseases of the brain. Free Radic Biol Med 63:1–29. doi:10.1016/j.freeradbiomed.2013.03.018
Chen H, Xu J, Hong J, Tang R, Zhang X, Fang JY (2014) Long noncoding RNA profiles identify five distinct molecular subtypes of colorectal cancer with clinical relevance. Mol Oncol 8:1393–1403. doi:10.1016/j.molonc.2014.05.010
Chiu CC et al (2015) Neuroprotective effects of aldehyde dehydrogenase 2 activation in rotenone-induced cellular and animal models of parkinsonism. Exp Neurol 263:244–253. doi:10.1016/j.expneurol.2014.09.016
Collier TJ, Kanaan NM, Kordower JH (2011) Ageing as a primary risk factor for Parkinson’s disease: evidence from studies of non-human primates. Nat Rev Neurosci 12:359–366. doi:10.1038/nrn3039
Condello S, Curro M, Ferlazzo N, Caccamo D, Satriano J, Ientile R (2011) Agmatine effects on mitochondrial membrane potential and NF-kappaB activation protect against rotenone-induced cell damage in human neuronal-like SH-SY5Y cells. J Neurochem 116:67–75. doi:10.1111/j.1471-4159.2010.07085.x
Daubner SC, Le T, Wang S (2011) Tyrosine hydroxylase and regulation of dopamine synthesis. Arch Biochem Biophys 508:1–12. doi:10.1016/j.abb.2010.12.017
Dexter DT, Jenner P (2013) Parkinson disease: from pathology to molecular disease mechanisms. Free Radic Biol Med 62:132–144. doi:10.1016/j.freeradbiomed.2013.01.018
Guttman M, Rinn JL (2012) Modular regulatory principles of large non-coding RNAs. Nature 482:339–346. doi:10.1038/nature10887
Hatano Y et al (2004) Novel PINK1 mutations in early-onset parkinsonism. Ann Neurol 56:424–427. doi:10.1002/ana.20251
Hu Y et al (2014) A long non-coding RNA signature to improve prognosis prediction of colorectal cancer. Oncotarget 5:2230–2242
Johnson ME, Bobrovskaya L (2015) An update on the rotenone models of Parkinson’s disease: their ability to reproduce the features of clinical disease and model gene-environment interactions. Neurotoxicology 46:101–116. doi:10.1016/j.neuro.2014.12.002
Klein C, Schlossmacher MG (2006) The genetics of Parkinson disease: implications for neurological care. Nat Clin Pract Neurol 2:136–146. doi:10.1038/ncpneuro0126
Lau E (2014) Non-coding RNA: zooming in on lncRNA functions. Nat Rev Genet 15:574–575. doi:10.1038/nrg3795
Li J et al (2015) Increased new lncRNA-mRNA gene pair levels in human cumulus cells correlate with oocyte maturation and embryo development. Reprod Sci 22:1008–1014. doi:10.1177/1933719115570911
Mathieu EL, Belhocine M, Dao LT, Puthier D, Spicuglia S (2014) Functions of lncRNA in development and diseases. Med Sci (Paris) 30:790–796. doi:10.1051/medsci/20143008018
Morais VA et al (2009) Parkinson’s disease mutations in PINK1 result in decreased Complex I activity and deficient synaptic function. EMBO Mol Med 1:99–111. doi:10.1002/emmm.200900006
Nagano T, Fraser P (2011) No-nonsense functions for long noncoding RNAs. Cell 145:178–181. doi:10.1016/j.cell.2011.03.014
Pang KC et al (2005) RNAdb–a comprehensive mammalian noncoding RNA database. Nucleic Acids Res 33:D125–D130. doi:10.1093/nar/gki089
Risueno A, Fontanillo C, Dinger ME, De Las Rivas J (2010) GATExplorer: genomic and transcriptomic explorer; mapping expression probes to gene loci, transcripts, exons and ncRNAs. BMC Bioinform 11:221. doi:10.1186/1471-2105-11-221
Sai Y, Zou Z, Peng K, Dong Z (2012) The Parkinson’s disease-related genes act in mitochondrial homeostasis. Neurosci Biobehav Rev 36:2034–2043. doi:10.1016/j.neubiorev.2012.06.007
Samii A, Nutt JG, Ransom BR (2004) Parkinson’s disease. Lancet 363:1783–1793. doi:10.1016/S0140-6736(04)16305-8
Savic N et al (2014) lncRNA maturation to initiate heterochromatin formation in the nucleolus is required for exit from pluripotency in ESCs. Cell Stem Cell 15:720–734. doi:10.1016/j.stem.2014.10.005
Sherer TB, Betarbet R, Greenamyre JT (2001) Pathogenesis of Parkinson’s disease. Curr Opin Investig Drugs 2:657–662
Soreq L, Guffanti A, Salomonis N, Simchovitz A, Israel Z, Bergman H, Soreq H (2014) Long non-coding RNA and alternative splicing modulations in Parkinson’s leukocytes identified by RNA sequencing. PLoS Comput Biol 10:e1003517. doi:10.1371/journal.pcbi.1003517
Srivastava G, Dixit A, Prakash O, Singh MP (2011) Tiny non-coding RNAs in Parkinson’s disease: implications, expectations and hypes. Neurochem Int 59:759–769. doi:10.1016/j.neuint.2011.07.009
Wapinski O, Chang HY (2011) Long noncoding RNAs and human disease. Trends Cell Biol 21:354–361. doi:10.1016/j.tcb.2011.04.001
Wei N et al (2015) PU.1 antisense lncRNA against its mRNA translation promotes adipogenesis in porcine preadipocytes. Anim Genet 46:133–140. doi:10.1111/age.12275
Winklhofer KF, Haass C (2010) Mitochondrial dysfunction in Parkinson’s disease. Biochim Biophys Acta 1802:29–44. doi:10.1016/j.bbadis.2009.08.013
Wirdefeldt K, Adami HO, Cole P, Trichopoulos D, Mandel J (2011) Epidemiology and etiology of Parkinson’s disease: a review of the evidence. Eur J Epidemiol 26(Suppl 1):S1–S58. doi:10.1007/s10654-011-9581-6
Wu P, Zuo X, Deng H, Liu X, Liu L, Ji A (2013) Roles of long noncoding RNAs in brain development, functional diversification and neurodegenerative diseases. Brain Res Bull 97:69–80. doi:10.1016/j.brainresbull.2013.06.001
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Yaohui Ni and Hua Huang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10571_2016_373_MOESM1_ESM.tif
RNAi-specific to lncRNA AL049437 and AK021630 reduced lncRNA AL049437 and AK021630 levels in SH-SY5Y cells. (A) siRNA at different concentration was employed to knock down the levels of lncRNA AL049437 after 12 and 24 h treatment. (B) siRNA at different concentration was employed to knock down the levels of lncRNA AK021630 after 12 and 24 h treatment. P < 0.05 vs. the sample with non-specific siRNA (TIFF 6756 kb)
Rights and permissions
About this article
Cite this article
Ni, Y., Huang, H., Chen, Y. et al. Investigation of Long Non-coding RNA Expression Profiles in the Substantia Nigra of Parkinson’s Disease. Cell Mol Neurobiol 37, 329–338 (2017). https://doi.org/10.1007/s10571-016-0373-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-016-0373-0