Abstract
Cannabidiol (CBD) is a non-psychoactive plant cannabinoid that is clinically used in a 1:1 mixture with the psychoactive cannabinoid Δ9-tetrahydrocannabinol (THC) for the treatment of neuropathic pain and spasticity in multiple sclerosis. Our group previously reported that CBD exerts anti-inflammatory effects on microglial cells. In addition, we found that CBD treatment increases the accumulation of the endocannabinoid N-arachidonoyl ethanolamine (AEA), thus enhancing endocannabinoid signaling. Here we proceeded to investigate the effects of CBD on the modulation of lipid-related genes in microglial cells. Cell viability was tested using FACS analysis, AEA levels were measured using LC/MS/MS, gene array analysis was validated with real-time qPCR, and cytokine release was measured using ELISA. We report that CBD significantly upregulated the mRNAs of the enzymes sterol-O-acyl transferase (Soat2), which synthesizes cholesteryl esters, and of sterol 27-hydroxylase (Cyp27a1). In addition, CBD increased the mRNA of the lipid droplet-associated protein, perilipin2 (Plin2). Moreover, we found that pretreatment of the cells with the cholesterol chelating agent, methyl-β-cyclodextrin (MBCD), reversed the CBD-induced increase in Soat2 mRNA but not in Plin2 mRNA. Incubation with AEA increased the level of Plin2, but not of Soat2 mRNA. Furthermore, MBCD treatment did not affect the reduction by CBD of the LPS-induced release of the proinflammatory cytokine IL-1β. CBD treatment modulates cholesterol homeostasis in microglial cells, and pretreatment with MBCD reverses this effect without interfering with CBD’s anti-inflammatory effects. The effects of the CBD-induced increase in AEA accumulation on lipid-gene expression are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The non-psychoactive plant cannabinoid, cannabidiol (CBD) in a 1:1 mixture with the psychoactive plant cannabinoid, Δ9-tetrahydrocannabinol (THC) has been clinically approved for the treatment of neuropathic pain and spasticity in multiple sclerosis (Russo and Guy 2006). In addition to their anti-nociceptive effects, several cannabinoids were shown to modulate immune cell signaling, function, and viability (Raz and Goldman 1976; Zimmerman et al. 1977; Jan et al. 2007; Lee et al. 2008; Liu et al. 2011; Ignatowska-Jankowska et al. 2009). CBD was shown to suppress proliferation and/or induce apoptosis in various immune cell types including mouse peritoneal macrophages, mouse thymocytes, splenocytes, and monocytes (Raz and Goldman 1976; Lee et al. 2008; Wu et al. 2008, 2010; Kaplan et al. 2008; Wu and Jan 2010). In addition, transformed immune cells were shown to be highly susceptible to CBD-induced apoptosis (Gallily et al. 2003; McKallip et al. 2002, 2006; Ligresti et al. 2006). Other immunosuppressive effects of CBD were reported in various culture and animal models (Jan et al. 2007; Weiss et al. 2006; Carrier et al. 2006; Malfait et al. 2000; Ruiz-Valdepenas et al. 2011). A study by Ignatowska-Jankowska et al. (2009) reported in vivo immunosuppressive effects of CBD on blood-derived lymphocyte subsets (including T, B, T helper, and T cytotoxic cells, but not NK or NKT) following repeated systemic administration in rats.
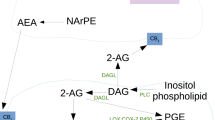
Our group has been characterizing the activity of CBD in microglial cells. Microglial cells are considered to be the resident macrophages of the central nervous system and constitute 5–20% of the CNS glial mass (Tambuyzer et al. 2009). We recently observed that in a mouse model of multiple sclerosis (experimental autoimmune encephalomyelitis), CBD administration during disease onset ameliorates clinical symptoms. This effect is accompanied by decreased inflammation and neuronal damage, as well as decreased microglial activation and T-cell recruitment into the spinal cord (Kozela et al. 2011). Moreover, we previously reported that CBD and THC inhibit lipopolysaccharide (LPS)-activated proinflammatory pathways in BV-2 microglial cells (Kozela et al. 2010). We found that CBD and THC exert their effects through different and partially overlapping pathways to inhibit LPS-induced proinflammatory cytokines. While both cannabinoids decrease the LPS-induced STAT1 transcription, only CBD reduces the activity of the NF-kB pathway and up-regulates STAT3 activation. We further characterized the differential transcription profiles and measured the endocannabinoid levels following incubation of BV-2 cells with CBD and THC. We found that CBD stimulates genes regulating the cellular stress response (Juknat et al. 2011). In addition, mass spectrometric analysis revealed CBD-induced increases in accumulation of the endocannabinoid, N-arachidonoyl ethanolamine (AEA, also known as anandamide), as well as additional N-acyl ethanolamine (NAEs) counterparts including the saturated N-palmitoyl ethanolamine and N-stearoyl ethanolamine that were specifically upregulated within lipid rafts Rimmerman et al. 2011).
Here we report that incubation of BV-2 microglial cells with CBD in serum-less medium increases AEA accumulation and decreases cell viability. Remarkably, the cholesterol chelating agent methyl-β-cyclodextrin (MBCD) rescues the cells from CBD-induced cell death. We further investigated the effects of CBD in 5% serum-containing growth medium (when cell viability was not affected). We show that CBD, but not THC or AEA, upregulated the gene transcript of the cholesterol metabolic enzyme sterol-O-acyltransferase-2 (Soat2; also known as Acat2) that produces intracellular cholesteryl esters from cholesterol and long-chain acyl CoA. Additionally, CBD and AEA, but not THC, significantly upregulated the gene transcript of the lipid droplet associated gene Perilipin2 (Plin2; previously known as adipose differentiation-related protein, adfp, adipophilin). When pretreated with MBCD, we found that Soat2 mRNA levels were reduced to normal levels, while Plin2 transcript levels remained upregulated. The triglyceride synthesis enzyme, diacylglycerol-O-acyl transferase-2 (Dgat2), was unchanged by all of these conditions. Importantly, MBCD treatment did not change the anti-inflammatory effects of CBD on LPS-treated microglial cells.
Methods
Reagents
LPS (Escherichia coli serotype 055:B5) was purchased from Sigma (St. Louis, MO, USA). THC and CBD were obtained from the National Institute on Drug Abuse (NIDA, Baltimore, MD, USA). Cannabinoid solutions were prepared before the experiments from their stocks in ethanol. Final concentration of ethanol in culture medium was 0.1%. MBCD was purchased from Sigma, its stock solution was prepared in H2O. [2H8]-N-arachidonoyl ethanolamine ([2H8]-AEA) and AEA were purchased from Cayman Chemical (Ann Arbor, MI). Other reagents were purchased as previously described (Bradshaw et al. 2006).
Cell Culture
The BV-2 murine microglial cell line, originally generated by E. Blasi (University of Perugia, Perugia, Italy; Blasi et al. 1990), was kindly provided by Prof. E.J. Choi from the Korea University (Seoul, Korea). Cells were grown under 5% CO2 at 37°C in Dulbecco’s modified Eagle’s medium containing high glucose (Gibco-BRL, Gaithersburg, MD, USA) supplemented with 5% heat-inactivated fetal bovine serum, streptomycin (100 μg/ml), and penicillin (100 U/ml) (Biological Industries Ltd., Kibbutz Beit Haemek, Israel).
Lipid Analysis of BV-2 Cells and Medium
BV-2 cells were plated (1 × 106 cells per 10 cm plate) and grown for 24 h in growth medium. Media was replaced with 5 ml fresh growth medium without serum 1 h before treatment. BV-2 cells were then treated with either vehicle (ethanol 0.1%) or CBD (10 μM). Cells together with media were scraped into 7 ml 100% HPLC grade methanol at three time points (10, 30, and 60; n = 2–3 per condition per experiment). The solution was then vortexed and spun down at 1000 rpm for 5 min. Supernatant samples were spiked with internal standard, and HPLC grade water was added to make a 75% aqueous solution. Lipids were then extracted as previously described (Rimmerman 2011). Mass spectrometric analysis proceeded as previously described (Rimmerman et al. 2008).
Cell Viability
Cell viability was measured by flow cytometry using propidium iodide (PI) staining according to Riccardi and Nicoletti (2006). Briefly, microglial cells (1 × 106 cells in 10-ml plates) were grown in growth medium containing 5% serum as above. Growth medium was replaced with serum-less growth medium. The cells were pretreated with MBCD (0.5 mM) or vehicle for 30 min, and then treated with CBD or vehicle. The cells were scraped 2 h after CBD treatment and spun down for 5 min at 2000 rpm. The pellets were washed twice with Dulbecco’s PBS without Ca++/Mg++ and fixed in 70% ethanol at −20°C overnight. Cells were then washed in PBS and incubated with RNase A (0.2 mg/ml) at 37°C, PBS rinsed and stained with PI (50 μg/ml) for 15 min on ice. The FACS analysis of 20,000 cells (for each sample) was measured using a flow cytometer (Becton–Dickinson FACS Calibur, San Jose, CA). The PI emission was detected in the FL2 channel using emission filter of 585 nm. The data were analyzed using CellQuestA software. Apoptotic cells were defined as cells in the sub-G1 phase with hypodiploid DNA content (Nicoletti et al. 1991).
Isolation of RNA and Microarray Transcript Analysis
RNA was extracted from BV-2 cells grown in 5% serum and treated for 6 h with either CBD or THC (unless otherwise described) using PerfectPure RNA extraction kit (5 PRIME, Darmstadt, Germany), and RNA samples (2 μg) were reverse transcribed using the QuantiTect Reverse Transcription Kit from Qiagen (Hilden, Germany) including DNase treatment of contaminating genomic DNA. Quality control of input RNA was performed with Agilent Bioanalyzer Nanochips (Agilent Technologies, Palo Alto, CA, USA) to assure reproducible and reliable microarray results. Total RNA (200 ng) was amplified, labeled, and hybridized on Illumina MouseRef-8 v1.1 Expression BeadChip (Illumina Inc, San Diego, CA), querying the expression of >24,000 RefSeq-curated gene targets and 822 random sequences used for the assessment of background noise. Slides were processed and scanned with Illumina BeadStation platform according to the manufacturer’s protocol. Raw data were analyzed using Bioconductor packages (www.bioconductor.org; Gentleman et al. 2004). Data were normalized using quantile normalization, and analysis of differential expression was performed using a linear model fitting (LIMMA package, Smyth 2005). Ingenuity analysis was performed as previously described (Juknat et al. 2011).
Reverse Transcription and Real-Time Quantitative PCR
Primers for mouse Soat2 (NM_146064.1), mouse Plin2 (NM_007408.2), mouse Dgat2 (NM_026384.2), and mouse β-microglobulin (BMG; NM_009735.3) were designed using PrimerQuest IDT (Integrated DNA Technologies, Inc, San Diego, CA, USA). The following primers were used: Soat2, forward TCATCTTCTTCGCCTTCCTGCACT, reverse AGAGCCACAGCCCATCTTGATACA; Plin2, forward AGCTTCTTAGAACCAAGGGCCTCA, reverse AAGACTAACACAGGCCACTCACCA; Dgat2, forward AGCTCACTTCTAGTGTGTCCTGCT, reverse ACATGGCCTCTGCCTCTCAAGAAT; BMG, forward ATGGGAAGCCGAACATACTG, reverse CAGTCTCAGTGGGGGTGAAT. Expression of mRNAs was determined by qPCR, using BMG as a normalizing gene, as previously described (Butovsky et al. 2006). Normal and mock reversed transcribed samples (in the absence of reverse transcriptase), as well as no template controls (total mix without cDNA), were run for each of the examined mRNAs. qPCR reactions were subjected to an initial step of 15 min at 95°C to activate the HotStar Taq DNA polymerase, followed by 40 cycles consisting of 15 s at 94°C, 30 s at 60°C, and 30 s at 72°C. Fluorescence was measured at the end of each elongation step. Data were analyzed using the Rotor-Gene software (Corbett Life Sciences, Australia), and a threshold cycle value C t was calculated from the exponential phase of each PCR sample. Expression levels of mRNAs were calculated and expressed in relative units of SYBR Green fluorescence.
ELISA of Released IL1β
Microglial cells were pretreated with cannabinoids for 2 h and stimulated with LPS (100 ng/ml) in the presence of 5% serum. Supernatants of cultured microglia were collected 4 h after LPS stimulation, spun down (5 min at 2000 rpm) to remove remaining cells, and the levels of IL-1β were measured by ELISA using specific monoclonal antibodies (R&D Systems, Minneapolis, MN, USA). The serum in the media did not interfere with the assay.
Statistical Analysis
SPSS was used for statistical analysis. Data are expressed as the mean ± SEM and analyzed for statistical significance using one-way analysis of variance (ANOVA), followed by Bonferroni or LSD post hoc tests. P < 0.05 was considered significant. Gene array analysis data were converted to % change using the formula 100 − (100/2signal log ratio). Signal log 2 ratio data with significance, SEM, and probe sequence are presented in Supplementary Table 1.
Results
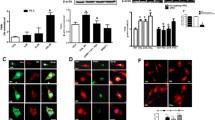
BV-2 cells were incubated with CBD (10 μM, 2 h) in serum-less growth medium. This resulted in the transfer of ~50% of the cell counts to the SubG1 cell cycle phase, indicating cell cycle arrest and cell death (Fig. 1). No effect was observed with vehicle in the absence of CBD. Pretreatment with MBCD (the cholesterol chelating/lipid raft disruptor agent) prevented the CBD-induced cell death. THC, the psychoactive cannabinoid present in marijuana, had no effect on cell viability (10 μM; n = 5; SubG1 = 1.50 ± 0.32% of cell counts). This effect of CBD on cell viability was observed only in serum-free medium. In the presence of 5% serum, BV-2 cell viability was not significantly affected up to 12 h after CBD incubation (Kozela et al. 2010).
CBD induction of cell death under serum-less conditions is reversed by pretreatment with MBCD. BV-2 cells in medium without serum were pretreated with vehicle (water) or MBCD (0.5 mM) for 30 min, followed by a 2-h incubation with CBD (10 μM). CBD incubation resulted in increased cell death as revealed by PI incorporation. # P < 0.0001, for SubG1 phase, VEH + EtOH versus VEH + CBD. Pretreatment with MBCD reversed this effect in the SubG1 phase VEH + CBD versus MBCD + CBD, # P < 0.0001; one-way ANOVA with Bonferroni post hoc
Since CBD was previously shown to enhance AEA signaling by inhibiting its enzymatic degradation or uptake (Watanabe et al. 1996; Rakhshan et al. 2000; Bisogno et al. 2001; Rimmerman 2011), we assayed the effect of CBD treatment on AEA levels in BV-2 cells with serum-less growth medium (Fig. 2). We found that indeed, AEA accumulation was time-dependently increased (measured up to 1 h following incubation with CBD).
CBD time dependently increases AEA accumulation in BV-2 cells under serum-less conditions. BV-2 cells were incubated in medium without serum with CBD (10 μM) for 10, 30, or 60 min. AEA was extracted and measured using LC/MS/MS. Data are representative of two independent experiments with similar results. CBD incubation significantly increased AEA accumulation at 30 min: # P < 0.002, n = 2, one-way ANOVA with LSD post hoc; and at 60 min: *P < 0.001, n = 3, one-way ANOVA with LSD post hoc
Due to the effect of CBD on AEA levels in the presence and in the absence of serum in the medium, as well as its affect on NAE distribution in lipid rafts (Rimmerman 2011), it was of interest to study whether CBD is modulating lipid-related pathways in these cells. As a first step, we analyzed the effects of CBD and THC on lipid-related genes using the gene array illumina platform; the results of the assay are described in Table 1, as well as Supplementary Table 1. We discovered that the expression of Soat2 mRNA encoding the cholesteryl ester synthetic enzyme was significantly upregulated by CBD (700%), but only 72% by THC. Additionally, the levels of the gene transcript for sterol 27-hydroxylase (Cyp27a1), a cholesterol metabolizing enzyme, were significantly increased by CBD (83%) and less so by THC (29%). Another lipid-related gene that was affected by CBD was Plin2, which was significantly upregulated following CBD by 135%, and was not affected by THC (−9%). Diacylglycerol-O-acyl transferase 2 (Dgat2), which encodes the enzyme for triglyceride synthesis, was slightly reduced by CBD (−37%), with almost no effect by THC. The treatments by CBD or THC did not affect the expression of the AEA metabolizing enzyme fatty acid amide hydrolase (FAAH). However, the expression of the mRNA for the protein tyrosine phosphatase, non-receptor, 22 (Ptpn22), shown to be involved in AEA synthesis in a macrophage cell line (Liu et al. 2006), was increased by 92% following incubation with CBD, and only by 31% following THC.
The results of the gene array analysis for Soat2, Plin2, and Dgat2 were validated by real-time qPCR (Fig. 3). This technique was used to test the effects of CBD, THC, MBCD, MBCD followed by CBD, and exogenously applied AEA (10 μM), on the expression of these gene products. The results obtained by qPCR for the effects of CBD and THC were in agreement with those obtained by the gene array method. Soat2 mRNA levels were significantly increased by CBD (190%) and non-significantly by THC (Fig. 3a). Plin2 mRNA levels were significantly increased by CBD (73%) and non-significantly by THC (Fig. 3b). Dgat2 mRNA levels were non-significantly affected by these cannabinoids (Fig. 3c). MBCD by itself had no significant effects on the expression of any of these genes; however, it eliminated the induction by CBD of the expression of Soat2 mRNA (reduction from 190% by CBD to 35% by CBD in the presence MBCD). Conversely, MBCD did not affect the induction of Plin2 mRNA by CBD (from 73% increase by CBD to 87% increase by CBD in the presence of MBCD).
CBD increases the mRNA levels of the lipid related genes Soat2 and Plin2. Cells in medium containing serum were preincubated for 30 min with vehicle or MBCD and then incubated with either vehicle, CBD (10 μM), THC (10 μM), or AEA (10 μM) for 6 h. RNA was extracted for real-time qPCR analysis. The bar graphs show the changes in gene expression compared to vehicle (presented as 1.0) for the following genes: a Soat2; b Plin2; c Dgat2. a CBD significantly upregulated Soat2 gene expression, # P < 0.001, n = 4, one-way ANOVA with LSD post hoc. b CBD and AEA significantly upregulated Plin2 gene expression, # P < 0.05, n = 4, one-way ANOVA with LSD post hoc. CBD + MBCD significantly upregulated Plin2 gene expression, *P < 0.005, n = 4, one-way ANOVA with LSD post hoc. c There were no significant differences in the expression of Dgat2 by the various treatments
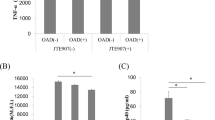
MBCD is known to affect membrane cholesterol and the structure of lipid rafts; however, one cannot rule out a direct chelating effect on CBD, thus modulating its activity in the cells. Therefore, we tested whether the anti-inflammatory effects, previously described for CBD (Kozela et al. 2010), persist following pretreatment with MBCD. The release of the inflammatory cytokine IL-1β is markedly upregulated in BV-2 cells following LPS treatment (Kozela et al. 2010). We found that this increase by LPS is significantly inhibited by CBD as well as by CBD in the presence of MBCD (Fig. 4). Interestingly, MBCD by itself significantly enhanced the effect of LPS. This explains why the amount of released IL-1β is somewhat higher in the presence of LPS + MBCD + CBD than in the presence of LPS + CBD; however, the level of inhibition by CBD remained the same (~70%) in both cases.
CBD inhibits the LPS-induced release of IL-1β from BV-2 cells. Cells in medium containing serum were preincubated for 30 min with MBCD or vehicle followed by incubation with CBD (10 μM, 2 h) or vehicle and then stimulated with 100 ng/ml LPS for 4 h. Cell-free media was collected, and the release of IL-1β was measured by ELISA. Data are representative of three independent experiments with similar results. All data are expressed as % change compared to LPS-induced IL-1β release. CBD significantly inhibited the LPS-induced release of IL-1β, # P < 0.005, n = 3, one-way ANOVA with LSD post hoc. MBCD + CBD significantly inhibited LPS-induced release of IL-1β, *P < 0.05, n = 3, one-way ANOVA with LSD post hoc. MBCD alone significantly enhanced the LPS-induced release of IL-1β, ^ P < 0.001, n = 3, one-way ANOVA with LSD post hoc. LPS by itself increased the release compared with control by about 100-fold (Kozela et al. 2010)
Discussion
We recently reported that in BV-2 microglial cells, CBD significantly increased the production of the endocannabinoid AEA (Rimmerman 2011). This occurred within 60 min in 5% serum-containing growth medium. Since the production of other N-acyl ethanolamines was also affected by CBD, we hypothesized that the contributing mechanisms may involve inhibition of AEA uptake and/or hydrolysis, as previously suggested (Watanabe et al. 1996; Rakhshan et al. 2000; Bisogno et al. 2001). Here we show that the level of AEA is significantly increased by CBD even in growth medium lacking serum. However, under these conditions, many of the BV-2 cells are susceptible to cell cycle arrest and cell death. A similar susceptibility to cell death in no serum-containing growth medium was previously reported by Raz and Goldman (1976), following incubation of primary peritoneal macrophages with CBD.
The increases that we observed in AEA accumulation were not accompanied by changes in the mRNA for FAAH (the main gene product known to participate in AEA metabolism (Maccarrone et al. 2010; Ueda et al. 2010), or in the AEA transport-associated gene product, fatty acid binding protein 5 (FABP5; Kaczocha et al. 2009). Conversely, the mRNA level of Ptpn22, an enzyme that catalyzes AEA production from phospho-AEA (Liu et al. 2006), was markedly increased by CBD (~90%) and much less so by THC (~30%). It is therefore possible that this enzyme contributes to the observed increase in AEA levels. This possible link needs to be further investigated by manipulating phospho-AEA metabolism (e.g., using specific phosphatase inhibitors to reduce AEA biosynthesis via this pathway). We further observed that the CBD-induced microglial cell death (in no serum-containing growth medium) could be prevented by pre-incubation with MBCD. MBCD was reported to inhibit lipid-mediated cell death, including endocannabinoid-induced cell death in several cellular models (Gajate and Mollinedo 2001; Sarker and Maruyama 2003; Sarker et al. 2003; Biswas et al. 2003; Yang et al. 2010). In this regard, Sarker and Maruyama (2003) reported that AEA-induced cell death was inhibited by MBCD in various cell models including, C6 glioma, HL-60, Neuro-2A, HEK-293, CHO, SMC, Jurkat, and PC12. They suggested that cholesterol-rich lipid rafts play an important role in this AEA-induced apoptosis. Biswas et al. (2003) further showed that AEA-induced cell death in hepatoma cells (as well as primary hepathocytes) was inhibited by MBCD and by mevastatin (an HMG-CoA reductase inhibitor). Both treatments lead to a reduction in membrane cholesterol levels. Additionally, they provided evidence for the presence of a molecular interaction between AEA and cholesterol and suggested that membrane cholesterol determines the fate of hepatocytes exposed to AEA. In our microglial cell system, CBD, but not THC or AEA, affected the expression of Soat2, and CBD’s effect was reversed by pretreatment with MBCD. The fact that AEA by itself did not increase Soat2 suggests that the effects on cholesterol homeostasis are not due to the mere increase in AEA accumulation. Moreover, these results suggest that the effects on cholesterol homeostasis are cannabinoid receptor-independent, since CBD by itself does not efficiently interact with the classical cannabinoid receptors (CB1 and CB2), while the molecules that do interact with these receptors (THC and AEA) did not induce these effects.
The effects of CBD on cholesterol homeostasis were previously reported by Cornicelli et al. (1981). They showed that CBD inhibited cholesteryl ester formation in human fibroblasts and in aortic medial cells (e.g., following treatment with 30 μM CBD, cholesteryl ester formation in human fibroblasts was reduced by ~80%). They found that the inhibition of cholesteryl ester formation occurred without inhibiting HMG-CoA reductase activity (the enzyme targeted by statins to lower cholesterol levels). In agreement with their results, we did not observe CBD-induced changes in HMG-CoA reductase mRNA levels in the gene array (Table 1 and Supplementary Table 1). However, we detected an 83% increase in the expression of the gene transcript Cyp27a1. Cyp27a1 encodes sterol 27-hydroxylase, an enzyme with a main role in the metabolism and removal of cholesterol. It has been shown that human deficiencies in Cyp27a1 lead to abnormal cholesterol metabolism resulting in cerebrotendinous xanthomatosis (CTX) disease. Clinical symptoms of CTX include cataract, premature atherosclerosis, and osteoporosis (Pikuleva 2006).
An additional gene product that was significantly affected by CBD, but also by AEA, was Plin2. Plin2 is a member of the PAT-family of proteins which share a conserved N-terminal sequence and localize to intracellular lipid storage droplets (Heid et al. 1998; Fujimoto et al. 2008; Kimmel et al. 2010). We cannot rule out the possibility that the effect of CBD on Plin2 gene transcription arises via the effect of CBD on the induction of AEA accumulation, since AEA itself increases Plin2 gene transcription. Furthermore, there is evidence suggesting that lipid droplets serve as AEA accumulation and metabolic sites (Oddi et al. 2008; Maccarrone et al. 2010; Kaczocha et al. 2010). This raises the possibility that these sites could allow interaction between Plin2 and AEA. This idea needs to be furthered investigated in the future.
Numerous molecular targets have been reported for cannabinoids (De Petrocellis and Di Marzo 2010; De Petrocellis et al. 2011), including CB1, CB2, GPR55, GPR18, the transient receptor potential vanilloid type-1 and type-2 (TRPV1; TRPV2) channels, peroxisome proliferator-activating receptor γ (PPARγ), and FAAH. We and others found that BV-2 microglial cells express CB1, CB2, GPR55, GPR18, and TRPV2 (Pietr et al. 2009; Stella 2010; McHugh et al. 2010; Rimmerman 2011). Using gene array analysis, we observed that CBD did not change the relative levels of CB1, CB2, GPR18, and FAAH transcripts. In contrast, PPARγ transcript was significantly (37%) up regulated. Further experiments are needed in order to reveal the molecular targets of CBD in microglia, which modulate the observed changes in lipid-related gene transcription.
In conclusion, we report here several effects of CBD on lipid metabolism. CBD affects genes that control sterol metabolism; these include increases in Soat2 and Cyp27a1. CBD also increases accumulation of AEA, as well as the transcription of Ptpn22 which was shown to enhance AEA production. In addition, CBD and AEA increase the transcription of Plin2, which is present in lipid droplets, a suggested site for AEA metabolism.
References
Bisogno T, Hanus L, De Petrocellis L, Tchilibon S, Ponde DE, Brandi I, Moriello AS, Davis JB, Mechoulam R, Di Marzo V (2001) Molecular targets for cannabidiol and its synthetic analogues: effect on vanilloid VR1 receptors and on the cellular uptake and enzymatic hydrolysis of anandamide. Br J Pharmacol 134:845–852
Biswas KK, Sarker KP, Abeyama K, Kawahara K, Iino S, Otsubo Y, Saigo K, Izumi H, Hashiguchi T, Yamakuchi M, Yamaji K, Endo R, Suzuki K, Imaizumi H, Maruyama I (2003) Membrane cholesterol but not putative receptors mediates anandamide-induced hepatocyte apoptosis. Hepatology 38:1167–1177
Blasi E, Barluzzi R, Bocchini V, Mazzolla R, Bistoni F (1990) Immortalization of murine microglial cells by a v-raf/v-myc carrying retrovirus. J Neuroimmunol 27:229–237
Bradshaw HB, Rimmerman N, Krey JF, Walker JM (2006) Sex and hormonal cycle differences in rat brain levels of pain-related cannabimimetic lipid mediators. Am J Physiol Regul Integr Comp Physiol 291:R349–R358
Butovsky E, Juknat A, Elbaz J, Shabat-Simon M, Eilam R, Zangen A, Altstein M, Vogel Z (2006) Chronic exposure to Delta9-tetrahydrocannabinol downregulates oxytocin and oxytocin-associated neurophysin in specific brain areas. Mol Cell Neurosci 31:795–804
Carrier EJ, Auchampach JA, Hillard CJ (2006) Inhibition of an equilibrative nucleoside transporter by cannabidiol: a mechanism of cannabinoid immunosuppression. Proc Natl Acad Sci USA 103:7895–7900
Cornicelli JA, Gilman SR, Krom BA, Kottke BA (1981) Cannabinoids impair the formation of cholesteryl ester in cultured human cells. Arteriosclerosis 1:449–454
De Petrocellis L, Di Marzo V (2010) Non-CB1, non-CB2 receptors for endocannabinoids, plant cannabinoids and synthetic cannabimimetics: focus on G-protein coupled receptors and transient receptor potential channels. J Neuroimmune Pharmacol 5:103–121
De Petrocellis L, Ligresti A, Moriello AS, Allara M, Bisogno T, Petrosino S et al (2011) Effects of cannabinoids and cannabinoid-enriched Cannabis extracts on TRP channels and endocannabinoid metabolic enzymes. Br J Pharmacol. doi:10.1111/j.1476-5381.2010.01166.x
Fujimoto T, Ohsaki Y, Cheng J, Suzuki M, Shinohara Y (2008) Lipid droplets: a classic organelle with new outfits. Histochem Cell Biol 130:263–279
Gajate C, Mollinedo F (2001) The antitumor ether lipid ET-18-OCH(3) induces apoptosis through translocation and capping of Fas/CD95 into membrane rafts in human leukemic cells. Blood 98:3860–3863
Gallily R, Even-Chena T, Katzavian G, Lehmann D, Dagan A, Mechoulam R (2003) Gamma-irradiation enhances apoptosis induced by cannabidiol, a non-psychotropic cannabinoid, in cultured HL-60 myeloblastic leukemia cells. Leuk Lymphoma 44:1767–1773
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JY, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Heid HW, Moll R, Schwetlick I, Rackwitz HR, Keenan TW (1998) Adipophilin is a specific marker of lipid accumulation in diverse cell types and diseases. Cell Tissue Res 294:309–321
Ignatowska-Jankowska B, Jankowski M, Glac W, Swiergel AH (2009) Cannabidiol-induced lymphopenia does not involve NKT and NK cells. J Physiol Pharmacol 60(Suppl 3):99–103
Jan TR, Su ST, Wu HY, Liao MH (2007) Suppressive effects of cannabidiol on antigen-specific antibody production and functional activity of splenocytes in ovalbumin-sensitized BALB/c mice. Int Immunopharmacol 7:773–780
Juknat A, Pietr M, Kozela E, Rimmerman N, Levy R, Coppola G, Geschwind D, Vogel Z (2011) Differential transcriptional profiles mediated by exposure to the cannabinoids cannabidiol and Δ9-tetrahydrocannabinol in BV-2 microglial cells. Br J Pharmacol (accepted)
Kaczocha M, Glaser ST, Deutsch DG (2009) Identification of intracellular carriers for the endocannabinoid anandamide. Proc Natl Acad Sci USA 106:6375–6380
Kaczocha M, Glaser ST, Chae J, Brown DA, Deutsch DG (2010) Lipid droplets are novel sites of N-acylethanolamine inactivation by fatty acid amide hydrolase-2. J Biol Chem 285:2796–2806
Kaplan BL, Springs AE, Kaminski NE (2008) The profile of immune modulation by cannabidiol (CBD) involves deregulation of nuclear factor of activated T cells (NFAT). Biochem Pharmacol 76:726–737
Kimmel AR, Brasaemle DL, McAndrews-Hill M, Sztalryd C, Londos C (2010) Adoption of PERILIPIN as a unifying nomenclature for the mammalian PAT-family of intracellular lipid storage droplet proteins. J Lipid Res 51:468–471
Kozela E, Pietr M, Juknat A, Rimmerman N, Levy R, Vogel Z (2010) Cannabinoids Delta(9)-tetrahydrocannabinol and cannabidiol differentially inhibit the lipopolysaccharide-activated NF-kappaB and interferon-beta/STAT proinflammatory pathways in BV-2 microglial cells. J Biol Chem 285:1616–1626
Kozela E, Lev N, Kaushansky N, Eilam R, Rimmerman N, Levy R, Ben-Nun A, Juknat A Vogel Z (2011) Cannabidiol inhibits pathogenic T-cells, decreases spinal microglial activation and ameliorates multiple sclerosis-like disease in C57BL/6 mice. Br J Pharmacol. doi:10.1111/j.1476-5381.2011.01379.x
Lee CY, Wey SP, Liao MH, Hsu WL, Wu HY, Jan TR (2008) A comparative study on cannabidiol-induced apoptosis in murine thymocytes and EL-4 thymoma cells. Int Immunopharmacol 8:732–740
Ligresti A, Cascio MG, Pryce G, Kulasegram S, Beletskaya I, De Petrocellis L, Saha B, Mahadevan A, Visintin C, Wiley JL, Baker D, Martin BR, Razdan RK, Di Marzo V (2006) New potent and selective inhibitors of anandamide reuptake with antispastic activity in a mouse model of multiple sclerosis. Br J Pharmacol 147:83–91
Liu J, Wang L, Harvey-White J, Osei-Hyiaman D, Razdan R, Gong Q, Chan AC, Zhou Z, Huang BX, Kim HY, Kunos G (2006) A biosynthetic pathway for anandamide. Proc Natl Acad Sci USA 103:13345–13350
Liu DZ, Hu CM, Huang CH, Wey SP, Jan TR (2011) Cannabidiol attenuates delayed-type hypersensitivity reactions via suppressing T-cell and macrophage reactivity. Acta Pharmacol Sin 31:1611–1617
Maccarrone M, Dainese E, Oddi S (2010) Intracellular trafficking of anandamide: new concepts for signaling. Trends Biochem Sci 35:601–608
Malfait AM, Gallily R, Sumariwalla PF, Malik AS, Andreakos E, Mechoulam R, Feldmann M (2000) The nonpsychoactive cannabis constituent cannabidiol is an oral anti-arthritic therapeutic in murine collagen-induced arthritis. Proc Natl Acad Sci USA 97:9561–9566
McHugh D, Hu SS, Rimmerman N, Juknat A, Vogel Z, Walker JM et al (2010) N-arachidonoyl glycine, an abundant endogenous lipid, potently drives directed cellular migration through GPR18, the putative abnormal cannabidiol receptor. BMC Neurosci 11:44
McKallip RJ, Lombard C, Fisher M, Martin BR, Ryu S, Grant S, Nagarkatti PS, Nagarkatti M (2002) Targeting CB2 cannabinoid receptors as a novel therapy to treat malignant lymphoblastic disease. Blood 100:627–634
McKallip RJ, Jia W, Schlomer J, Warren JW, Nagarkatti PS, Nagarkatti M (2006) Cannabidiol-induced apoptosis in human leukemia cells: a novel role of cannabidiol in the regulation of p22phox and Nox4 expression. Mol Pharmacol 70:897–908
Nicoletti I, Migliorati G, Pagliacci MC, Grignani F, Riccardi C (1991) A rapid and simple method for measuring thymocyte apoptosis by propidium iodide staining and flow cytometry. J Immunol Methods 139:271–279
Oddi S, Fezza F, Pasquariello N, De Simone C, Rapino C, Dainese E, Finazzi-Agro A, Maccarrone M (2008) Evidence for the intracellular accumulation of anandamide in adiposomes. Cell Mol Life Sci 65:840–850
Pietr M, Kozela E, Levy R, Rimmerman N, Lin YH, Stella N et al (2009) Differential changes in GPR55 during microglial activation. FEBS Lett 583:2071–2076
Pikuleva IA (2006) Cholesterol-metabolizing cytochromes P450. Drug Metab Dispos 34:513–520
Rakhshan F, Day TA, Blakely RD, Barker EL (2000) Carrier-mediated uptake of the endogenous cannabinoid anandamide in RBL-2H3 cells. J Pharmacol Exp Ther 292:960–967
Raz A, Goldman R (1976) Effect of hashish compounds on mouse peritoneal macrophages. Lab Invest 34:69–76
Riccardi C, Nicoletti I (2006) Analysis of apoptosis by propidium iodide staining and flow cytometry. Nat Protoc 1:1458–1461
Rimmerman N, Hughes HV, Bradshaw HB, Pazos MX, Mackie K, Prieto AL, Walker JM (2008) Compartmentalization of endocannabinoids into lipid rafts in a dorsal root ganglion cell line. Br J Pharmacol 153:380–389
Rimmerman N, Bradshaw HB, Kozela E, Levy R, Juknat A, Vogel Z (2011) Compartmentalization of endocannabinoids into lipid rafts in a microglial cell line which is devoid of caveolin-1. Br J Pharmacol. doi:10.1111/j.14765381.2011.01380.x
Ruiz-Valdepenas L, Martinez-Orgado JA, Benito C, Millan A, Tolon RM, Romero J (2011) Cannabidiol reduces lipopolysaccharide-induced vascular changes and inflammation in the mouse brain: an intravital microscopy study. J Neuroinflammation 8:5
Russo E, Guy GW (2006) A tale of two cannabinoids: the therapeutic rationale for combining tetrahydrocannabinol and cannabidiol. Med Hypotheses 66:234–246
Sarker KP, Maruyama I (2003) Anandamide induces cell death independently of cannabinoid receptors or vanilloid receptor 1: possible involvement of lipid rafts. Cell Mol Life Sci 60:1200–1208
Sarker KP, Biswas KK, Yamakuchi M, Lee KY, Hahiguchi T, Kracht M, Kitajima I, Maruyama I (2003) ASK1-p38 MAPK/JNK signaling cascade mediates anandamide-induced PC12 cell death. J Neurochem 85:50–61
Smyth GK (2005) Limma: linear models for microarray data. In: Gentleman R, Carey V, Dudoit S, Irizarry R, Huber W (eds) Bioinformatics and computational biology solutions using R and bioconductor. Springer, New York, pp 397–420
Stella N (2010) Cannabinoid and cannabinoid-like receptors in microglia, astrocytes and astrocytomas. Glia 58:1017–1030
Tambuyzer BR, Ponsaerts P, Nouwen EJ (2009) Microglia: gatekeepers of central nervous system immunology. J Leukoc Biol 85:352–370
Ueda N, Tsuboi K, Uyama T (2010) Enzymological studies on the biosynthesis of N-acylethanolamines. Biochim Biophys Acta 1801:1274–1285
Watanabe K, Kayano Y, Matsunaga T, Yamamoto I, Yoshimura H (1996) Inhibition of anandamide amidase activity in mouse brain microsomes by cannabinoids. Biol Pharm Bull 19:1109–1111
Weiss L, Zeira M, Reich S, Har-Noy M, Mechoulam R, Slavin S, Gallily R (2006) Cannabidiol lowers incidence of diabetes in non-obese diabetic mice. Autoimmunity 39:143–151
Wu HY, Jan TR (2010) Cannabidiol hydroxyquinone-induced apoptosis of splenocytes is mediated predominantly by thiol depletion. Toxicol Lett 195:68–74
Wu HY, Chu RM, Wang CC, Lee CY, Lin SH, Jan TR (2008) Cannabidiol-induced apoptosis in primary lymphocytes is associated with oxidative stress-dependent activation of caspase-8. Toxicol Appl Pharmacol 226:260–270
Wu HY, Chang AC, Wang CC, Kuo FH, Lee CY, Liu DZ, Jan TR (2010) Cannabidiol induced a contrasting pro-apoptotic effect between freshly isolated and precultured human monocytes. Toxicol Appl Pharmacol. doi:10.1016/j.physletb.2003.10.071
Yang Q, Liu HY, Zhang YW, Wu WJ, Tang WX (2010) Anandamide induces cell death through lipid rafts in hepatic stellate cells. J Gastroenterol Hepatol 25:991–1001
Zimmerman S, Zimmerman AM, Cameron IL, Laurence HL (1977) Delta1-tetrahydrocannabinol, cannabidiol and cannabinol effects on the immune response of mice. Pharmacology 15:10–23
Acknowledgments
We would like to thank Prof. Daniel Geschwind and Dr. Giovanni Coppola, from the Program in Neurogenetics, Department of Neurology, University of California, Los Angeles, CA, for running the microarray analysis. NR and AJ were supported by the Israeli Ministry of Immigrant Absorption, The Center for Absorption in Science. This work was supported by The Dr. Miriam and Sheldon G. Adelson Center for the Biology of Addictive Disease.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rimmerman, N., Juknat, A., Kozela, E. et al. The Non-Psychoactive Plant Cannabinoid, Cannabidiol Affects Cholesterol Metabolism-Related Genes in Microglial Cells. Cell Mol Neurobiol 31, 921–930 (2011). https://doi.org/10.1007/s10571-011-9692-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-011-9692-3