Abstract
Recent studies have suggested that the initiation and progression of hepatocellular carcinoma (HCC) are closely associated with lipopolysaccharide (LPS) of intestinal bacteria. However, the role of LPS in immune regulation of HCC remains largely unknown. An orthotopic Hepa1-6 tumor model of HCC was constructed to analyze the effect of LPS on the expression of immune checkpoint molecules PD-1 and PD-L1. Then we verified the regulation of PD-L1 by LPS in HCC cells. Based on the previous finding that lncRNA MIR155HG regulates PD-L1 expression in HCC cells, we analyzed the relationship of LPS signaling pathway molecules with PD-L1 and MIR155HG by bioinformatics. The molecular mechanism of MIR155HG regulating PD-L1 expression induced by LPS was investigated by RNA pull-down followed by mass spectrometry, RNA immunoprecipitation, fluorescence in situ hybridization, and luciferase reporter assay. Finally, the HepG2 xenograft model was established to determine the role of MIR155HG on PD-L1 expression in vivo. We showed that LPS induced PD-1 and PD-L1 expression in mouse tumor tissues and induced PD-L1 expression in HCC cells. Mechanistically, upregulation of METTL14 by LPS promotes the m6A methylation of MIR155HG, which stabilizes MIR155HG relying on the “reader” protein ELAVL1 (also known as HuR)-dependent pathway. Moreover, MIR155HG functions as a competing endogenous RNA (ceRNA) to modulate the expression of PD-L1 by miR-223/STAT1 axis. Our results suggested that LPS plays a critical role in immune escape of HCC through METTL14/MIR155HG/PD-L1 axis. This study provides a new insight for understanding the complex immune microenvironment of HCC.
Graphical abstract
1. LPS plays a critical role in immune escape of HCC, especially HCC with cirrhosis.
2. Our study reveals that LPS regulates PD-L1 by m6A modification of lncRNA in HCC.
3. MIR155HG plays an important role in LPS induced PD-L1 expression.
4. LPS-MIR155HG-PD-L1 regulatory axis provides a new target for the treatment of HCC.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is the most common type of primary liver cancer with poor prognosis and invasiveness (Villanueva, 2019). The development of HCC is closely related to metabolites by intestinal microflora, known as pathogen-associated molecular patterns. Despite of no direct contact between the liver and microorganisms, lipopolysaccharide (LPS) can enter the liver through the gut-liver axis, thereby promoting the progress of HCC (Roderburg and Luedde, 2014).
LPS is a unique chemical component in the outer wall of gram-negative bacteria. It can enhance the invasive ability and chemotherapy resistance of liver cancer by maintaining the stemness of cancer stem cells, as well as induce the epithelial-mesenchymal transition by activating toll-like receptor 4 (TLR4) (Lai et al. 2016; Sekiya et al. 2016; Strain and Crosby, 2000). Recent studies have shown that LPS/TLR4 signaling can mediate the differentiation of hepatic progenitor cell into myofibroblasts; enhance the production of interleukin-6 and tumor necrosis factor-α, which in turn constitutes tumor microenvironment (TME); and promote proliferation and malignant transformation of hepatic progenitor cell (Liu et al. 2020a, b). These studies show the importance of exploring the relationship between LPS and TME in HCC.
Although immunotherapy has become a major focus of research in cancer, due to the complex TME, (Ringelhan et al. 2018), the curative effect of immunotherapy is poor in liver cancer compared with other types of cancers. It has been proved that LPS can stimulate target cells through TLR4 binding to the adaptor protein myeloid differentiation marker 88 (MyD88) (Nallasamy et al. 2018), mediating anti-tumor immunity (Han et al. 2017; Melssen et al. 2019) and promoting immune escape (Song et al. 2020). However, whether LPS plays a role in the regulation of TME in HCC remains unclear.
Long noncoding RNAs (lncRNAs) are a class of transcripts exceeding 200 bases without protein-coding potential, which is involved in the regulation of physiological and pathological processes (Yao et al. 2019). Recently, emerging studies found that lncRNAs can participate in the process of immune regulation including tumor immune resistance and immune escape (Hu et al. 2019; Huang et al. 2018). For example, lncRNA KCNQ1OT1 induced sorafenib resistance and programmed death-ligand 1 (PD-L1) mediating immune escape of HCC via sponging miR‑506 (Zhang et al. 2020a, b); lncRNA MALAT1 redirected the polarization of macrophages toward the M2 subset by adsorbing miR-140, leading to the transformation of HCC microenvironment to immunosuppression (Hou et al. 2020); lncRNA FENDRR was reported to be involved in the immune regulation, and knocking down FENDRR reduced the secretion of TGF-β, IL-10, and VEGF in HCC cells. Further research showed that FENDRR inhibited the immunosuppressive function of Treg cells through competitive binding to miR-423-5p (Yu et al. 2019). MIR155 host gene (MIR155HG), also known as B cell integration cluster, was considered to be a lncRNA (Tam et al. 1997). Most of the current reports on MIR155HG indicated that MIR155HG played a function in tumor development and immune regulation. Although the underlying mechanisms need to be further investigated, our previous study found that MIR155HG was closely related to immunity in HCC (Peng et al. 2019).
Herein, we found that LPS could promote the m6A methylation of MIR155HG by upregulating the expression of METTL14. The m6A modification of MIR155HG by METTL14 enhanced its stability relying on the “reader” protein ELAVL1. MIR155HG sponges miR-223-3p to upregulate STAT1, which regulates the expression of PD-L1 and mediates the immune escape of liver cancer.
Materials and methods
Human specimens
All of the tissue and blood samples in this scientific research are from Fujian Medical University Union Hospital. The ethics committee granted an informed consent form for ethics and morals. The clinical medical characteristics of the patients are shown in Table S1, S2, and S3. All HCC patients were diagnosed based on tissue samples, and pathology confirmed the HCC patients with cirrhosis. The staging of HCC patient was accurately measured according to the 8th edition of the AJCC.
Cell lines
The human HCC cell line HepG2, Huh7, and mouse hepatoma cell line Hepa1-6 were all cultured in DMEM media, with penicillin and streptomycin, which was filled with high glucose supplemented containing 12% (v/v) fetal bovine serum (FBS, Gibco), 37 °C, containing 5% carbon dioxide. HepG2 and Huh7 cells were STR-authenticated. All cells used in this research were routinely screened and found to be free of mycoplasma.
Mouse studies
Male C57BL/6 mice were injected with 1 × 107 Hepa1-6 cells to create orthotopic model of hepatoma by surgery treatment. After 1 week, the mice were randomly divided into 2 groups, each injected with LPS (10 mg/kg) or PBS, once every 3 days, for a total of 4 injections. Three days after the last injection, the mice were executed with anesthetic, and the indicators were tested by immunohistochemistry (IHC). In order to better test the efficacy of MIR155HG, HepG2 cells (5 × 106) transfected with sh-MIR155HG or sh-NC were injected subcutaneously into male BALB/c nude mice approximately 6 weeks old. The mice were killed under anesthesia after 8 weeks, and the tumor tissues were removed to carry out subsequent experiments. The animal experiments were undertaken in accordance with the guidelines of National Institute of Health Guide, with the approval of the Animal Ethics Committee of Fujian Medical University.
RNA pull-down and mass spectrometry analysis
RNA pull-down assays were processed according to the instructions of Pierce™ Magnetic RNA–Protein Pull-Down Kit manufactured by Thermo Scientific (Waltham, MA). MIR155HG was transcribed using MEGAscript ®Kit (Thermo Scientific) with a DNA template containing a T7 promoter in vitro. Biotin-labeled DNAs were captured by magnetic beads coated with streptavidin and then mixed with proteins obtained from HepG2 at 4 °C for overnight stay. Proteins were eluted from RNA protein complex and then analyzed with mass spectrometry. One-shot mass spectrometry analyses were used for checking the purity of proteins. LC–MS/MS was carried out by Beijing Protein Innovation Co. Ltd. (Beijing, China).
Analysis of RNA immunoprecipitation
According to the instructions of RNA immunoprecipitation (RIP) kit (Millipore, Darmstadt, Germany), HepG2 cells were washed with PBS and collected for RNA immunoprecipitation analysis. ELAVL1, AGO2 antibody, or IgG (4 μg) were added, and cell lysates were incubated at 4 °C overnight. Subsequently, the compounds were treated with proteinase K, and immunoprecipitation RNA was separated and obtained with Trizol, reverse transcribed to cDNA, and then analyzed by qRT-PCR. The primers used for this analysis are described in Table S4.
Luciferase reporter assay
The sequence of MIR155HG or 3′-UTR of STAT1 and binding sites mutated sequences were, respectively, subcloned into pmirGLO luciferase reporter vector. Co-transfect the pmirGLO vector with MIR155HG or mutation sequence and miR-223-3p mimic into HCC cells. Co-transfect MIR155HG- wild-type/mutant (WT/Mut) or STAT1-WT/Mut with miR-223-3p mimic or NC into HCC cells. In order to examine the effect of METTL14 on MIR155HG RNA methylation, the pmirGLO-MIR155HG or MIR155HG m6A site mutation vectors were co-transfected with METTL14. Accurately measure the luciferase activity of HCC cells according to the dual luciferase reporter gene monitoring system (Promega).
T cell–mediated tumor cell killing assay
In this scientific research, peripheral blood mononuclear cells (PBMCs) are derived from individuals who are physically and mentally healthy. PBMCs were cultured in T cell medium supplemented with IL-2 (10 ng/mL, PeproTech) and human CD3/CD28/CD2 T cell activator (25ul/mL) purchased from STEMCELL for 1 week. Tumor cells treated accordingly were cocultured with activated cells in 96-well plates 12 h. The ratios of cancer cells to activated cells is 1:3. The living cancer cells were then stained with crystal violet.
Flow cytometry sorting
After digestion, the cells were centrifuged and re-suspended with PBS buffer (containing 2%FBS) to prepare single cell suspension and count. The amount of antibody was determined according to the counting results, and the antibody was incubated in ice for 30 min. After centrifugation, the cells were resuspended with 500μL PBS buffer (containing 2%FBS) centrifugation. Cell concentration was adjusted with 100uL PBS buffer. BD FACSAria III was used for flow cytometry sorting.
Statistical analysis
Statistical analysis was conducted by GraphPad Prism 7 software (San Diego, CA). All data are presented as mean ± SD. For multiple groups, significance was measured by one-way ANOVA. Student’s t test was used to compare two groups of independent samples. The level of significance was set at 0.05.
For other experimental methods (see Supplemental Material).
Results
The expression of PD-L1 and the level of LPS were upregulated in HCC patients with cirrhosis
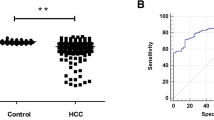
To explore the level of PD-L1 expression in HCC patients with or without cirrhosis, we first queried the published clinical datasets TCGA and found that the mRNA expression of PD-L1 in HCC patients with cirrhosis was significantly higher than that without cirrhosis (Fig. 1A). Subsequently, we validated the bioinformatics data in a clinical sample cohort of HCC tissues by qRT-PCR and found the mRNA level of PD-L1 was significantly higher in HCC patients with cirrhosis than without cirrhosis (Fig. 1B). Furthermore, we tested the expression of PD-L1 in clinical HCC tissue by Immunofluorescence (IF) histochemistry. The results showed that the expression level of PD-L1 was significantly increased in HCC patients with cirrhotic (Fig. 1C). Next, Limulus lysate test indicated that patients with cirrhotic liver cancer had a significantly higher plasma LPS than that of those without liver cirrhosis (Fig. 1D). Therefore, we speculate that the increased PD-L1 expression in HCC patients with cirrhotic might be associated with high LPS.
PD-L1 and LPS were upregulated in HCC with cirrhosis. A The expression of PD-L1 of HCC patients with cirrhosis or not in TCGA. B The mRNA levels of PD-L1 in HCC tissues were determined by qRT-PCR. C IF histochemistry analysis of PD-L1 expression in HCC tissues. D The levels of LPS in plasma of HCC patients. *P < 0.05, **P < 0.01
LPS upregulated PD-L1 expression and mediated T cell suppression in HCC
Cancer cells can skillfully evade immune surveillance by manipulating immune checkpoint molecules such as PD-L1. LPS is known to be involved in the formation of liver cancer microenvironment, but whether it participates in immune escape is not completely understood. We used Hepa1-6 cells to construct an orthotopic liver cancer model to explore the effect of LPS on PD-L1 expression in C57BL/6 mice. The experimental flowchart was shown in Fig. S1A. We observed that LPS intraperitoneally injection enhanced tumor growth in orthotopic liver cancer model (Fig. S1B). Next, by using the immunohistochemical serial section staining, we found that the expression of PD-1 (Fig. S1C) and PD-L1 (Fig. 2A) in hepatoma tissue of mice injected with LPS was higher than those PBS group. Subsequently, we validated the expression of PD-L1 in hepatoma cell lines (HepG2, Huh7, and Hepa1-6) by qRT-PCR and found that the mRNA of PD-L1 was upregulated after LPS treatment (Fig. 2B, C). Western blot and immunofluorescence results also showed that HCC cells treated with LPS significantly upregulated PD-L1 protein levels (Fig. 2D, E, S2, S11A and 11B).
LPS upregulated the expression of PD-L1 in HCC cells. A The expression of PD-L1 of mice tumor tissues in LPS (10 mg/kg) and PBS group (n = 5 per group). B–E The HCC cells were treated with LPS (0, 1, 10, 100, 200 ug/mL) for 0, 3, 6, 12, and 24 h, and the expression of PD-L1 was assessed by qRT-PCR (B, C) (mean ± SD, n = 3) and western blot (D, E). F T cell–mediated tumor cell killing assay of HCC cells treated with LPS (100 ug/mL) for 12 h or not. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001
PD-L1 on tumor cells and macrophages binds to PD-1 on activated cytotoxic T lymphocytes (CTLs), inhibits T cell activation, and protects tumor cells from undergoing T cell–mediated killing. To examine the activity and function of PD-L1 upregulated by LPS, we performed a T cell–mediated cancer cell killing assay. The result revealed that HepG2 and Huh7 cells treated by LPS were more resistant to human T cell–mediated cytolysis than control (Fig. 2F and S10C).
LPS promoted PD-L1 expression via m6A methylation modification of MIR155HG by upregulating METTL14
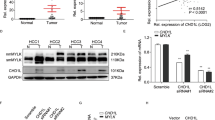
Based on our previous study that PD-L1 mRNA was positively correlated with MIR155HG level in HCC tissues (Peng et al. 2019), we speculated that MIR155HG might be involved in the process of LPS induced PD-L1. We measured the expression of MIR155HG in HCC cells (HepG2 and Huh7) after LPS treatment. The level of MIR155HG was increased after treatment with LPS (Fig. 3A, B). To determine the effects of MIR155HG on regulation of PD-L1, we overexpressed MIR155HG by transfecting the full-length sequence carried by pcDNA3.1 (Fig. S3A). The qRT-PCR and western blot results showed that the mRNA (Fig. S3B) and protein (Fig. 3C) level of PD-L1 were increased after MIR155HG overexpression in HCC cells. Immunofluorescent assays further showed that MIR155HG enhanced PD-L1 expression (Fig. S4A and S11C). Conversely, knockdown of MIR155HG using siRNA or short hairpin RNAs (shRNAs) decreased PD-L1 levels in HCC cells (Fig. 3D, S3C–S3G, S4B and S11D). Furthermore, downregulation of MIR155HG significantly decreased the expression of PD-L1 induced by LPS (Fig. 3E and S11E). Interestingly, the MIR155HG overexpressed cells were more resistant to T cell–mediated cytolysis than control (Fig. S4C and S10D).
LPS promoted PD-L1 expression via regulated m6A methylation modification of MIR155HG through METTL14. A, B The expression of MIR155HG in HCC cells after LPS stimulated (0, 1, 10, 100, 200 ug/mL) for 0, 3, 6, 12, and 24 h. C, D The expression of PD-L1 was detected by western blot after MIR155HG overexpression (C) or knockdown (D). E The protein of PD-L1 was measured by western blot in HCC cells transfected with sh-MIR155HG after LPS stimulation. F The levels of METTL3 and METTL14 in Huh7 cell stimulated by LPS (100 ug/mL) for 12 h. G The expression of METTL14 and PD-L1 in Huh7 cell overexpressed METTL14 was assayed by western blot. (H) The expression of METTL14 and PD-L1 protein in HCC cells knocking down METTL14. I Silver staining of proteins in HepG2 cells that were bound to MIR155HG. The highlighted regions were identified as ELAVL1 by mass spectrometry. J RNA pull-down was performed to determine the interaction of ELAVL1 with full-length or truncations of MIR155HG. K RIP analysis to determine the bound of MIR155HG with ELAVL1 using anti-ELAVL1 antibody. L Luciferase reporter assays identify MIR155HG methylation modification sites by METTL14. The data are representative of three independent experiments (mean ± SD). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001
The RNA modifications have been found in mammalian cells, which regulate the function of RNA and play a critical role in the epigenetic modulation of RNA metabolism. N6-methyladenosine (m6A) is one of the most prevalent modifications in mRNA and non-coding RNA (Meyer and Jaffrey, 2014). Studies have shown that LPS promotes the development of liver cancer through m6A modification-mediated epigenetic regulation (Ding et al. 2020). We speculated whether LPS regulation of MIR155HG dependent on m6A modification. Firstly, we detected the expression of RNA methyltransferase METTL14 and METTL3 after treated with LPS and found that both METTL3 and METTL1 were increased in Huh7 cell after LPS stimulation (Fig. 3F,S11F). In order to evaluate the biological function of METTL3 and METTL14, we overexpressed METTL3 and METTL14 in HCC cells respectively (Fig. S11G). The results showed that overexpression of METTL14 significantly increased the level of PD-L1 in Huh7 cell (Fig. 3G,S11G), while METTL3 failed to elicit the alterations (Fig. S5A). Additionally, we found that the RNA levels of MIR155HG and PD-L1 were increased after overexpression of METTL14 in HCC cells (Fig. S5B, S5C). Knockdown of METTL14 decreased the MIR155HG and PD-L1 level in HepG2 and Huh7 cells (Fig. S5D, Fig. 3H,S11H). Further experiments showed that knocking down METTL14 could restore the change of MIR155HG caused by LPS (Fig. S5E). Collectively, these results revealed the critical role of METTL14 in LPS regulation of MIR155HG.
Next, we explored the molecular mechanism of METTL14 affecting MIR155HG in HCC cells. In order to identify the potential proteins interacting with MIR155HG, we performed RNA pull-down experiment using biotinylated MIR155HG, followed by mass spectrometry. ELAVL1 (a protein that can stabilizes the RNA bound to it) (Visvanathan et al. 2018) was identified as an interacting protein of MIR155HG (Fig. 3I, S6, S7A–S7D), which was validated by western blot (Fig. 3J). To determine the nucleotide sequence of MIR155HG that binds ELAVL1, we prepared a series of MIR155HG deletion mutants. As shown in Fig. 3J, mutants including nt 1 to 750 of MIR155HG exhibited binding to ELAVL1, while the antisense and mutants containing 1–375 fragments did not bind to ELAVL1. These results illustrated that the 375–1474 (full-length) nucleotide sequence of MIR155HG is critical for the interaction of MIR155HG and ELAVL1. The interaction of MIR155HG with ELAVL1 was further confirmed by RIP (Fig. 3K). To examine the role of ELAVL1, we detected the expression of MIR155HG after knockdown ELAVL1 in HCC cells. As is shown in S7E, knockdown of ELAVL1 by siRNA decreased the level of MIR155HG. Knocking down ELAVL1 could restore the change of MIR155HG caused by LPS (Fig. S7F). The online prediction tool SRAMP revealed that three m6A sites are distributed in the MIR155HG, of which 751 sites had the highest confidence (Fig. S8A). We then constructed the plasmids with MIR155HG 751 sites mutant for the luciferase reporter assay to determine the effect of m6A modification on the expression of MIR155HG (Fig. S8B). As shown in Fig. 3L, METTL14 increased wild-type fusion MIR155HG luciferase activity but not mutant MIR155HG. These results suggested that the m6A modification regulating MIR155HG level is determined by METTL14.
MIR155HG regulated PD-L1 expression via miR-223-STAT1 axis
LncRNAs can exert their molecular biological effects through a variety of mechanisms. The subcellular localization of lncRNA determines its different effects (Salmena et al. 2011). Fluorescence in situ hybridization (FISH) confirmed that MIR155HG is located in the cytoplasm (Fig. 4A). Then we tried to scientifically study whether MIR155HG interacted with miRNA to regulate the expression of PD-L1. The predictive analysis results of Starbase3.0 showed that miR-223-3p is likely to interacted with MIR155HG (Fig. 4B). To further validate the interaction of MIR155HG with miR-223-3p, we established a wild-type and mutant reporter vectors of MIR155HG (Fig. 4B). The luciferase reporting gene examination indicated that transfection of miR-223-3p mimics reduced the luciferase activity of the wild-type MIR155HG reporter plasmid but not reduce the luciferase activity of MIR155HG mutant plasmid in HepG2 and Huh7 cells (Fig. 4C). Next, we used qRT-PCR to detect the level of miR-223-3p and found that after overexpression of MIR155HG in HCC cells, the level of endogenous miR-223-3p was significantly decreased (Fig. 4D).
MIR155HG regulated PD-L1 expression through the miR-223-3p-STAT1 axis. A Cytoplasmic localization of MIR155HG by RNA fluorescence in situ hybridization in HepG2 and Huh7 cells. B Interacting sequences on MIR155HG for miR-223-3p were obtained from starBase 3.0. The putative binding sites of miR-223-3p was mutated in MIR155HG. MIR155HG and mutated sequences were constructed into luciferase reporter vector, respectively. C HCC cells co-transfected with miR-223-3p mimics and WT or Mut MIR155HG vector were measured for luciferase activity. D The expression of miR-223-3p in HCC cells overexpressed MIR155HG. E Complementary sequences of miR-223-3p to 3′UTR of STAT1. 3′UTR sequences of STAT1 as well as mutant sequences were constructed onto the reporter vector. F The expression of STAT1 mRNA in HCC cells transfected with miR-223-3p mimics. G STAT1 and PD-L1 protein expression in HCC cells transfected with miR-223-3p mimics was determined by western blot. H Luciferase reporter assays were performed in HepG2 and Huh7 cells after co-transfection with miR-223-3p mimics and WT or Mut 3′UTR vector of STAT1. I RIP assay was conducted using AGO2 antibody to confirm the interaction between miR-223-3p, MIR155HG, and STAT1 in HepG2 cells. J MIR155HG and miR-223-3p mimics or control was co-transfected in HCC cells, and the expression of STAT1 and PD-L1 was detected. The data are representative of three independent experiments (mean ± SD). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001
Based on the above results, we further considered how the MIR155HG/miR-223-3p axis regulates the expression of PD-L1 in HCC cells? Interestingly, we found that STAT1 may possibly be a target gene of miR-223-3p by starBase 3.0 (Fig. 4E). Multiple articles have reported that STAT1 upregulates the expression of PD-L1 to mediate immune escape (Sasidharan Nair et al. 2018; Zhang et al. 2020a, b). Therefore, we speculated that MIR155HG/miR-223-3p axis is more likely to regulate PD-L1 expression by affecting STAT1. We subsequently examined the expression level of STAT1 after overexpression of miR-223-3p in HCC cells. The results showed that the mRNA and protein level of STAT1 decreased after overexpression of miR-223-3p (Fig. 4F, G,S11I) and the level of PD-L1 protein also decreased (Fig. 4G,S11I). Figure 4E indicated two possible interaction sites between miR-223-3p and STAT1. In order to determine the binding domain between miR-223-3p and STAT1, we constructed luciferase reporter vectors containing WT or Mut 3’UTR of STAT1, respectively. As shown in Fig. 4H, the overexpression of miR-223-3p reduces the luciferase activity of WT and Mut1 instead of Mut2 vectors, which indicates that miR-223-3p modulated STAT1 mainly through the Mut2 site. Since AGO2 is a component of RNA-induced silencing complex (RISC), it participates in the key regulatory function of miRNA (Ye et al. 2015), we performed RIP experiments with AGO2 antibody in HepG2 cells. The STAT1, MIR155HG, and miR-223-3p were immunoprecipitated by AGO2 (Fig. 4I). Furthermore, overexpression of MIR155HG elevated the expression level of STAT1 and PD-L1, which counteracted by transfection of miR-223-3p mimics as shown in Fig. 4J and S11J.
MIR155HG promoted the expression of PD-L1 in vivo
In order to further verify the molecular biological efficacy of MIR155HG in vivo, xenograft tumor model was constructed. HepG2 cells were transfected with sh-MIR155HG and sh-NC. The transfected HepG2 cells were injected into nude mice. We observed that the expression of MIR155HG in sh-MIR155HG group was significantly decreased (Fig. 5A). On the contrary, the expression of miR-223-3p was higher in sh-MIR155HG group (Fig. 5B), while the levels of STAT1 and PD-L1 mRNA in knockdown MIR155HG group were lower (Fig. 5C, D). Next, we further evaluated the protein expression of STAT1 and PD-L1 in tumor using IHC and found that the levels of STAT1 and PD-L1 in sh-MIR155HG group were lower (Fig. 5E, F).
The regulation of PD-L1 by MIR155HG was examined in vivo. A–D HepG2 cells that were stably transfected with sh-MIR155HG or control shRNA were subcutaneously injected into nude mice. Xenograft tissues were subjected to qRT-PCR for MIR155HG (A), miR-223-3p (B), STAT1 (C), and PD-L1 (D). E, F Immunohistochemical comparison of the expression of STAT1 (E) and PD-L1 (F) in sh-NC versus sh-MIR155HG group (n = 5 per group). *P < 0.05, **P < 0.01
To assess the biological role of MIR155HG in HCC, we overexpressed MIR155HG in HepG2 and Huh7 cells. CCK-8 assays indicated that MIR155HG overexpression had no influence on the proliferation of HepG2 and Huh7 cells (Fig. S9A). Transwell assay also showed that MIR155HG overexpression did not significantly affected the migration of HepG2 and Huh7 cells (Fig. S9B). In order to explore the effect of MIR155HG in the regulation of tumor growth of HCC in vivo, MIR155HG knockdown of HepG2 cells were subcutaneously injected. The results suggested that decreased MIR155HG expression had no obviously effect on tumor growth in nude mice (Fig. S9C). We also analyzed the association of MIR155HG with prognosis of HCC patient and found that MIR155HG was not associated with overall survival (Fig. S9D) and disease-free survival (Fig. S9E).
The METTL14-MIR155HG-STAT1 axis was upregulated in HCC patients with cirrhosis
It was well documented that the level of LPS was increased in the plasma and liver tissue of patients with cirrhosis and LPS in turn further promoted hepatocarcinogenesis (Roderburg and Luedde, 2014). In this study, we found that LPS was upregulated in HCC patients with cirrhosis and LPS regulated the expression of PD-L1 by METTL14-MIR155HG-STAT1 axis. To validate the molecular activation of the PD-L1 pathway regulated by LPS in cirrhotic HCC patients, we investigated the TCGA data and found that the expression of METTL14 mRNA and MIR155HG in HCC patients with cirrhosis was significantly higher than that without cirrhosis (Fig. 6A, B). We further detected the expression of MIR155HG in clinical samples by qRT-PCR; the result was consistent with that of TCGA (Fig. 6C). Moreover, the TCGA data also showed that the level of STAT1 mRNA is higher in HCC patients with cirrhosis (Fig. 6D). Subsequently, we examined the expression of METTL14 and STAT1 protein in clinical HCC tissue by IHC. The results also suggested that METTL14 and STAT1 were significantly increased in HCC patients with cirrhotic (Fig. 6E, F).
METTL14-MIR155HG-STAT1 regulatory axis was activated in HCC patients with cirrhosis. A, B The expression of METTL14 (A) and MIR155HG (B) in HCC patients with or without cirrhosis of TCGA. C The levels of MIR155HG in clinical tissues of HCC. D The expression of STAT1 mRNA in TCGA. E, F The expression of METTL14 (E) and STAT1 (F) in HCC patients detected by immunohistochemistry. G LPS regulated PD-L1 expression through m6A modification of MIR155HG. *P < 0.05, **P < 0.01
Discussion
Although immune checkpoint blockade therapy has raised hope for HCC patients, the therapeutic efficacy is limited (Finn et al. 2020). Considering the complexity of the immune landscape of HCC, an intense investigation into the mechanisms of immune microenvironment is urgently required. Intestinal microbial products enter the liver through the gut-liver axis and contribute to the progression of liver cancer (Schwabe and Greten, 2020). Despite a number of studies which have confirmed the involvement of LPS in the formation and development of liver cancer, there are few articles revealing the relationship between LPS and tumor immunity. Herein, we found that LPS can regulate immune escape by inducing the expression of PD-L1 in HCC. Our results provided a new perspective for understanding the immune status of liver cancer.
In this study, we identified that the PD-L1 may be associated with LPS in HCC (Fig. 1). Subsequently, we further investigated its function and mechanism of LPS in regulating PD-L1 expression. We found that the expression of PD-L1 in HCC cells increased significantly under LPS stimulation and that MIR155HG has an important impact on this process. It has been reported that the expression of MIR155HG in macrophages and dendritic cells is increased significantly after TLR activation (Ceppi et al. 2009; Tam et al. 1997). A study found that MIR155HG was increased significantly in mouse infected with influenza A virus (Maarouf et al. 2019). Other articles also found that there was a close relationship between MIR155HG and immunity in multiple types of tumors (Peng et al. 2019; Wang et al. 2018a, b; Xu et al. 2020). These results indicated that MIR155HG played a great significance on immune regulation. FISH suggested that MIR155HG localized primarily to the cytoplasm of HCC cells. Some studies suggested that lncRNA located in the cytoplasm and containing microRNA response element (MRE) can function through ceRNA mechanism (Tay et al. 2011). Our results confirmed that MIR155HG was competitively binding miR-223-3p to exert regulatory effects by act as a ceRNA. This was different from what other researchers have reported. For example, Wu et al. found that MIR155HG influenced mesenchymal transition in glioma through derivative miR-155 (Wu et al. 2017); Niu et al. suggested that MIR155HG-encoded micropeptide interacted with the heat-shock cognate protein 70 (HSC70), disrupting the HSC70-HSP90 interaction to regulate antigen presentation and T cell priming (Niu et al. 2020). Our study further revealed that MIR155HG upregulated the expression of PD-L1 by competitively binding to miR-223-3p with the 3′UTR of STAT1 (Fig. 6G). This provided new evidence for STAT1 activation which upregulates PD-L1 expression on tumor cells (Liu et al. 2020a, b; Sasidharan Nair et al. 2018) and also illustrated that STAT1 determined the direction of immune response in tumor microenvironment to some extent (Yu et al. 2009).
The m6A modification has been reported to participate in various cellular processes such as mRNA maturation, protein translation, and molecular structure change (Zhao et al. 2017). As one of the most prevalent RNA modifications, RNA m6A methylation regulators exist in almost all types of RNAs, including mRNA metabolism (Haussmann et al. 2016), miRNA biogenesis, and lncRNA function (Dai et al. 2020). Epigenetic regulation mediated by m6A modification has profound influence on the progression of malignancies, including HCC (Wang et al. 2018a, b). A recent study identified that LPS mediates epigenetic regulation through m6A modification promotes growth and invasion of HCC cell (Ding et al. 2020). This conclusion is further imprinted in the current study from a novel perspective that lncRNA modified by m6A participates in the regulation of immune microenvironment in HCC. We suggested that LPS upregulated RNA methyltransferase METTL14 promotes m6A methylation modification of MIR155HG. Meanwhile, MIR155HG was found to interact with ELAVL1 (also known as HuR) by RNA pull-down assay. Visvanathan et al. previously reported that ELAVL1 bound m6A modified RNA and mediated its stabilization compared to unmethylated RNA (Visvanathan et al. 2018). Therefore, we considered that MIR155HG with m6A methylation binds to ELAVL1 and stabilizes MIR155HG (Fig. 6G). Our observation provided a convincing argument in favor of a role for m6A modification of lncRNAs in cellular physiological processes regulation (Ni et al. 2019; Zheng et al. 2019). The observation further suggested that intestinal microbiota modulates host gene expression through m6A modification to influence epigenetics.
Multiple evidences indicated that duration of exposure to microbiota or microbiota-derived factors resulted in liver fibrosis and provoked hepatic malignant potential. For example, Seki et al. found that after performing bile duct ligation, antibiotic cocktail efficiently suppressed the increase of LPS in plasma compared with intact intestinal flora mouse. They further confirmed that intestinal flora was a major source of LPS, and gut-derived LPS drives fibrogenesis (Seki et al. 2007). Roderburg also pointed out that the gut microbiota played a critical role in the pathogenesis of HCC by creating an LPS-dependent proinflammatory microenvironment in the liver (Roderburg and Luedde, 2014). Currently, the relationship between liver cirrhosis and the immunosuppressive factor PD-L1 remains inconclusive. Some reports suggested that cirrhosis was associated with the expression of PD-L1 (Pei et al. 2019), while others did not (Chen et al. 2020). In this study, we suggested that the LPS in serum and the PD-L1 expression in tumor were higher in HCC patients with cirrhosis compared with those of patients without cirrhosis. This result supports the conclusion that there is a correlation between cirrhosis and PD-L1 expression in HCC, highlighting that gut microbial products, particularly LPS, are important contributors to this correlation. Our study gives the following hints: (1) the immune microenvironment of HCC patients with cirrhosis is more complex relatively, and LPS can promote immune escape and accelerate the transformation of cirrhosis to cancer; (2) it is necessary to consider the immunosuppressive of HCC patients with cirrhosis during immunotherapy in clinical.
The present study revealed that LPS caused immune escape of HCC by affecting the level of m6A modification of MIR155HG to upregulate PD-L1. The current results not only provided novel insights into the molecular mechanisms underlying HCC induced by LPS but also paved the way for the development of more effective immunotherapeutic strategies for HCC.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
References
Ceppi M, Pereira PM, Dunand-Sauthier I, Barras E, Reith W, Santos MA, Pierre P. MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells. Proc Natl Acad Sci U S A. 2009;106:2735–40.
Chen L, Huang X, Zhang W, Liu Y, Chen B, Xiang Y, Zhang R, Zhang M, Feng J, Liu S, Duan T, Chen X, Wang W, Pan T, Yan L, Jin T, Li G, Li Y, Xie T, Sui X. Correlation of PD-L1 and SOCS3 co-expression with the prognosis of hepatocellular carcinoma patients. J Cancer. 2020;11:5440–8.
Dai F, Wu Y, Lu Y, An C, Zheng X, Dai L, Guo Y, Zhang L, Li H, Xu W, Gao W. Crosstalk between RNA m(6)A Modification and non-coding rna contributes to cancer growth and progression. Mol Ther Nucleic Acids. 2020;22:62–71.
Ding H, Zhang X, Su Y, Jia C, Dai C. GNAS promotes inflammation-related hepatocellular carcinoma progression by promoting STAT3 activation. Cell Mol Biol Lett. 2020;25:8.
Finn RS, Ryoo BY, Merle P, Kudo M, Bouattour M, Lim HY, Breder V, Edeline J, Chao Y, Ogasawara S, Yau T, Garrido M, Chan SL, Knox J, Daniele B, Ebbinghaus SW, Chen E, Siegel AB, Zhu AX, Cheng AL. Pembrolizumab as second-line therapy in patients with advanced hepatocellular carcinoma in KEYNOTE-240: a randomized, double-blind, phase III trial. J Clin Oncol. 2020;38:193–202.
Han S, Wang C, Qin X, Xia J, Wu A. LPS alters the immuno-phenotype of glioma and glioma stem-like cells and induces in vivo antitumor immunity via TLR4. J Exp Clin Cancer Res. 2017;36:83.
Haussmann IU, Bodi Z, Sanchez-Moran E, Mongan NP, Archer N, Fray RG, Soller M. m(6)A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature. 2016;540:301–4.
Hou ZH, Xu XW, Fu XY, Zhou LD, Liu SP, Tan DM. Long non-coding RNA MALAT1 promotes angiogenesis and immunosuppressive properties of HCC cells by sponging miR-140. Am J Physiol Cell Physiol. 2020;318:C649–63.
Hu Q, Ye Y, Chan LC, Li Y, Liang K, Lin A, Egranov SD, Zhang Y, Xia W, Gong J, Pan Y, Chatterjee SS, Yao J, Evans KW, Nguyen TK, Park PK, Liu J, Coarfa C, Donepudi SR, Putluri V, Putluri N, Sreekumar A, Ambati CR, Hawke DH, Marks JR, Gunaratne PH, Caudle AS, Sahin AA, Hortobagyi GN, Meric-Bernstam F, Chen L, Yu D, Hung MC, Curran MA, Han L, Lin C, Yang L. Oncogenic lncRNA downregulates cancer cell antigen presentation and intrinsic tumor suppression. Nat Immunol. 2019;20:835–51.
Huang D, Chen J, Yang L, Ouyang Q, Li J, Lao L, Zhao J, Liu J, Lu Y, Xing Y, Chen F, Su F, Yao H, Liu Q, Su S, Song E. NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat Immunol. 2018;19:1112–25.
Lai FB, Liu WT, Jing YY, Yu GF, Han ZP, Yang X, Zeng JX, Zhang HJ, Shi RY, Li XY, Pan XR, Li R, Zhao QD, Wu MC, Zhang P, Liu JF, Wei LX. Lipopolysaccharide supports maintaining the stemness of CD133(+) hepatoma cells through activation of the NF-κB/HIF-1α pathway. Cancer Lett. 2016;378:131–41.
Liu F, Liu J, Zhang J, Shi J, Gui L, Xu G. Expression of STAT1 is positively correlated with PD-L1 in human ovarian cancer. Cancer Biol Ther. 2020a;21:963–71.
Liu WT, Jing YY, Gao L, Li R, Yang X, Pan XR, Yang Y, Meng Y, Hou XJ, Zhao QD, Han ZP, Wei LX. Lipopolysaccharide induces the differentiation of hepatic progenitor cells into myofibroblasts constitutes the hepatocarcinogenesis-associated microenvironment. Cell Death Differ. 2020b;27:85–101.
Maarouf M, Chen B, Chen Y, Wang X, Rai KR, Zhao Z, Liu S, Li Y, Xiao M, Chen JL. Identification of lncRNA-155 encoded by MIR155HG as a novel regulator of innate immunity against influenza A virus infection. Cell Microbiol. 2019;21:e13036.
Melssen MM, Petroni GR, Chianese-Bullock KA, Wages NA, Grosh WW, Varhegyi N, Smolkin ME, Smith KT, Galeassi NV, Deacon DH, Gaughan EM, Slingluff CL Jr. A multipeptide vaccine plus toll-like receptor agonists LPS or polyICLC in combination with incomplete Freund’s adjuvant in melanoma patients. J Immunother Cancer. 2019;7:163.
Meyer KD, Jaffrey SR. The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat Rev Mol Cell Biol. 2014;15:313–26.
Nallasamy P, Chava S, Verma SS, Mishra S, Gorantla S, Coulter DW, Byrareddy SN, Batra SK, Gupta SC, Challagundla KB. PD-L1, inflammation, non-coding RNAs, and neuroblastoma: Immuno-oncology perspective. Semin Cancer Biol. 2018;52:53–65.
Ni W, Yao S, Zhou Y, Liu Y, Huang P, Zhou A, Liu J, Che L, Li J. Long noncoding RNA GAS5 inhibits progression of colorectal cancer by interacting with and triggering YAP phosphorylation and degradation and is negatively regulated by the m(6)A reader YTHDF3. Mol Cancer. 2019;18:143.
Niu L, Lou F, Sun Y, Sun L, Cai X, Liu Z, Zhou H, Wang H, Wang Z, Bai J, Yin Q, Zhang J, Chen L, Peng D, Xu Z, Gao Y, Tang S, Fan L, Wang H. A micropeptide encoded by lncRNA MIR155HG suppresses autoimmune inflammation via modulating antigen presentation. Sci Adv. 2020;6:eaaz2059.
Pei R, Zhang W, Wang S, Huang X, Zou Y. Prognostic value of PD-L1 in patients with hepatocellular carcinoma. Clin Lab 2019;65.
Peng L, Chen Z, Chen Y, Wang X, Tang N. MIR155HG is a prognostic biomarker and associated with immune infiltration and immune checkpoint molecules expression in multiple cancers. Cancer Med. 2019;8:7161–73.
Ringelhan M, Pfister D, O’Connor T, Pikarsky E, Heikenwalder M. The immunology of hepatocellular carcinoma. Nat Immunol. 2018;19:222–32.
Roderburg C, Luedde T. The role of the gut microbiome in the development and progression of liver cirrhosis and hepatocellular carcinoma. Gut Microbes. 2014;5:441–5.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146:353–8.
Sasidharan Nair V, Toor SM, Ali BR, Elkord E. Dual inhibition of STAT1 and STAT3 activation downregulates expression of PD-L1 in human breast cancer cells. Expert Opin Ther Targets. 2018;22:547–57.
Schwabe RF, Greten TF. Gut microbiome in HCC - Mechanisms, diagnosis and therapy. J Hepatol. 2020;72:230–8.
Seki E, De Minicis S, Osterreicher CH, Kluwe J, Osawa Y, Brenner DA, Schwabe RF. TLR4 enhances TGF-beta signaling and hepatic fibrosis. Nat Med. 2007;13:1324–32.
Sekiya S, Miura S, Matsuda-Ito K, Suzuki A. Myofibroblasts derived from hepatic progenitor cells create the tumor microenvironment. Stem Cell Reports. 2016;7:1130–9.
Song H, Liu Y, Li X, Chen S, Xie R, Chen D, Gao H, Wang G, Cai B, Yang XJC, Medicine, T. Long noncoding RNA CASC11 promotes hepatocarcinogenesis and HCC progression through EIF4A3-mediated E2F1 activation. 2020;10:e220.
Strain AJ, Crosby HA. Hepatic Stem Cells. Gut. 2000;46:743–5.
Tam W, Ben-Yehuda D, Hayward WS. bic, a novel gene activated by proviral insertions in avian leukosis virus-induced lymphomas, is likely to function through its noncoding RNA. Mol Cell Biol. 1997;17:1490–502.
Tay Y, Kats L, Salmena L, Weiss D, Tan SM, Ala U, Karreth F, Poliseno L, Provero P, Di Cunto F, Lieberman J, Rigoutsos I, Pandolfi PP. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell. 2011;147:344–57.
Villanueva A. Hepatocellular Carcinoma. N Engl J Med. 2019;380:1450–62.
Visvanathan A, Patil V, Arora A, Hegde AS, Arivazhagan A, Santosh V, Somasundaram K. Essential role of METTL3-mediated m(6)A modification in glioma stem-like cells maintenance and radioresistance. Oncogene. 2018;37:522–33.
Wang S, Chai P, Jia R, Jia R. Novel insights on m(6)A RNA methylation in tumorigenesis: a double-edged sword. Mol Cancer. 2018a;17:101.
Wang W, Zhao Z, Yang F, Wang H, Wu F, Liang T, Yan X, Li J, Lan Q, Wang J, Zhao J. An immune-related lncRNA signature for patients with anaplastic gliomas. J Neurooncol. 2018b;136:263–71.
Wu X, Wang Y, Yu T, Nie E, Hu Q, Wu W, Zhi T, Jiang K, Wang X, Lu X, Li H, Liu N, Zhang J, You Y. Blocking MIR155HG/miR-155 axis inhibits mesenchymal transition in glioma. Neuro Oncol. 2017;19:1195–205.
Xu J, Liu F, Li Y, Shen L. A 1p/19q codeletion-associated immune signature for predicting lower grade glioma prognosis. Cell Mol Neurobiol 2020.
Yao RW, Wang Y, Chen LL. Cellular functions of long noncoding RNAs. Nat Cell Biol. 2019;21:542–51.
Ye Z, Jin H, Qian Q. Argonaute 2: a novel rising star in cancer research. J Cancer. 2015;6:877–82.
Yu H, Pardoll D, Jove R. STATs in cancer inflammation and immunity: a leading role for STAT3. Nat Rev Cancer. 2009;9:798–809.
Yu Z, Zhao H, Feng X, Li H, Qiu C, Yi X, Tang H, Zhang J. Long non-coding RNA FENDRR acts as a miR-423-5p sponge to suppress the Treg-mediated immune escape of hepatocellular carcinoma cells. Mol Ther Nucleic Acids. 2019;17:516–29.
Zhang H, Zhu C, He Z, Chen S, Li L, Sun C. LncRNA PSMB8-AS1 contributes to pancreatic cancer progression via modulating miR-382-3p/STAT1/PD-L1 axis. J Exp Clin Cancer Res. 2020a;39:179.
Zhang J, Zhao X, Ma X, Yuan Z, Hu M. KCNQ1OT1 contributes to sorafenib resistance and programmed death-ligand-1-mediated immune escape via sponging miR-506 in hepatocellular carcinoma cells. Int J Mol Med. 2020b;46:1794–804.
Zhao BS, Roundtree IA, He C. Post-transcriptional gene regulation by mRNA modifications. Nat Rev Mol Cell Biol. 2017;18:31–42.
Zheng ZQ, Li ZX, Zhou GQ, Lin L, Zhang LL, Lv JW, Huang XD, Liu RQ, Chen F, He XJ, Kou J, Zhang J, Wen X, Li YQ, Ma J, Liu N, Sun Y. Long noncoding RNA FAM225A promotes nasopharyngeal carcinoma tumorigenesis and metastasis by acting as ceRNA to sponge miR-590-3p/miR-1275 and upregulate ITGB3. Cancer Res. 2019;79:4612–26.
Funding
This work was supported by the National Natural Science Foundation of China (82173167); the Startup Fund for Scientific Research of Fujian Medical University (2019XQ2020); the Joint Funds for the Innovation of Science and Technology, Fujian province (2017Y9100); and the Natural Science Foundation of Fujian Province (2018J01296).
Author information
Authors and Affiliations
Contributions
Conception and design: LP, BP, and NT. Development of methodology: BP and XZ. Data curation: ZW, JQ, and XW. Writing, review and/or revision of the manuscript: LP, BP, ZW, and NT. Study supervision: NT. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
Ethical consent was granted from the ethics committee of Fujian Medical University Union Hospital. The animal experiments were approved by the Animal Ethics Committee of Fujian Medical University.
Consent to participate
All the authors listed have consented to participate the study.
Consent for publication
All the listed authors agree with the publication of the paper.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peng, L., Pan, B., Zhang, X. et al. Lipopolysaccharide facilitates immune escape of hepatocellular carcinoma cells via m6A modification of lncRNA MIR155HG to upregulate PD-L1 expression. Cell Biol Toxicol 38, 1159–1173 (2022). https://doi.org/10.1007/s10565-022-09718-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10565-022-09718-0