Abstract
Acute exacerbation of chronic obstructive pulmonary disease (AECOPD) as a serious event has high mortality and medical costs. Systemic inflammation and immune response are the major factors influencing the outcome and quality of patient with AECOPD. On basis of identification and validation of AECOPD-specific inflammatory biomarkers, the present study aimed to identify AECOPD-specific immunomodulatory mediators by evaluating dynamic genomic and proteomic profiles of peripheral blood mononuclear cells (PBMCs) and plasma in patients with AECOPD on day 1, 3, and 10 after the hospital admission, to compare with healthy controls or patients with stable COPD. We found that genes and proteins of C1QC and C1RL were co-differentially up-expressed in patients with COPD or AECOPD, while haptoglobin (HP), ORM1, SERPING1, and C3 were identified as a panel of AECOPD-specific immunomodulatory mediators. We also found that inflammatory stimuli could up-regulate osteopontin (OPN)-associated HP expression through the PI3K signal pathway in A549 cells. Block of autocrine production of OPN by gene inhibition could reduce HP production from inflammation-induced lung epithelial cells. The complex network of AECOPD- or COPD-specific immunomodulatory mediators will benefit the development of precision or personalized medicine strategies for prevention and treatment of AECOPD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chronic obstructive pulmonary disease (COPD) is a syndrome characterized with progressive pulmonary dysfunction inflammation and airway obstruction (Guirguis-Blake et al. 2016). COPD is a major and growing public health burden, ranking the fourth leading cause of death in China (Fang et al. 2011). Acute exacerbations of COPD (AECOPD) are often induced by the infection, with a sudden, severe, and progressive lung function and inflammation (Celli and Barnes 2007; Rubinsztajn et al. 2016). AECOPD has high economic and social burden, and lacks early and specific diagnoses and therapies, since the disease often becomes significant or irreversible (Murtagh et al. 2005; Vestbo et al. 2013). Due to the complex of inflammatory mediators and the difficulty of new technique for COPD, there is a great lack of understanding molecular mechanisms and disease-specific biomarkers as defined recently (Fang et al. 2013b; Liu et al. 2014; Niu et al. 2016; Wang 2011; Wang and Ward 2012; Wu et al. 2014a).
AECOPD can be also worsened by local and systemic inflammations (He et al. 2010; Prins et al. 2016). Our previous studies found the over-production of cytokines and chemokines in the circulation of patients with AECOPD, of which some showed disease-specific and severity-associated (Chen et al. 2012a; Chen et al. 2012b; Wu et al. 2014b), attracted inflammatory cells from the circulation to the lung, and amplified the inflammatory process of COPD (Barnes 2014; Hogg et al. 2004). CD8+ lymphocytes increased as one of COPD characters (Barnes et al. 2003) and monocytes became more active in the development of AECOPD (Bhattacharya et al. 2011; Melguizo et al. 2013). By integrating proteomics with clinical informatics, we also found that osteopontin (OPN) played an important role in the occurrence of AECOPD and was associated with the disease severities in patients with AECOPD (Chen et al. 2012a). OPN is a secreted glycophosphoprotein and contributes to diverse physiological and pathological processes, including bone formation, tumor growth and metastasis, inflammation, and immune responses (Denhardt et al. 2001; Wai and Kuo 2004). During inflammation response, it is widely accepted that OPN acts as a well-characterized cytokine with both pro-inflammatory and anti-inflammatory functions (Zhao et al. 2010). In the immune system, OPN is expressed by various immune cells (e.g., macrophages, dendritic cells, and T lymphocytes) and modulates both innate and adaptive immune responses (Ashkar et al. 2000; Chabas et al. 2001; Wang and Denhardt 2008). However, the concrete mechanisms among them remains unclear.
To make an extension of one of our previous studies and furthermore explore the potential association and interaction between OPN and specific immunomodulatory mediators in AECOPD. The present study aimed to screen AECOPD-specific immunomodulatory mediators of peripheral blood mononuclear cells (PBMCs) and circulating plasma using the concept of clinical bioinformatics integrating genomics, proteomics, bioinformatics, and clinical informatics (Chen and Wang 2011; Wang 2011). Finally, we found that haptoglobin (HP), ORM1, C3, and SERPING1 were identified as a panel of AECOPD-specific biomarkers. Of those genes, it revealed a slight, but a significantly positive correlation between OPN and HP protein levels and then we focused on the role of OPN in the regulation of HP expression and production in human alveolar epithelial cells.
Materials and methods
Patient population
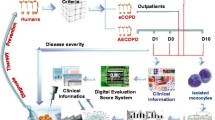
Two hundred twenty patients were recruited in the study, including 60 healthy controls, 80 AECOPD patients, and 80 stable COPD patients in Zhongshan Hospital, from 2011 October to 2012 March. Inclusion criteria for patients with COPD were as follows: FEV1 <80% of predicted value adjusted for age, weight, and height, and an improvement in FEV1 following bronchodilator inhalation <12% of baseline FEV1. Asthmatic patients showing a persistent airflow obstruction were excluded. Stable COPD was defined by American Thoracic Society/European Respiratory Society consensus criteria as no requirement for increased treatment above maintenance therapy, other than bronchodilators, for 30 days (Celli and Mac Nee 2004). AECOPD was the reason for hospital admission and characterized by a worsening of the patient respiratory symptoms that was beyond normal day-to-day variations and leaded to a change in medication (Celli and Barnes 2007; Rodriguez-Roisin 2000). Healthy controls were enrolled from blood donors in Zhongshan Hospital. Subjects with respiratory diseases, or any family history of lung disease, were excluded. PBMCs and plasma were harvested once from healthy controls and patients with stable COPD as well as patients with AECOPD on the admission day and 3 and 10 days after the admission. Each group is matched according to age, gender, and smoking status previously, and we randomly selected six patients in each groups representative of the whole population. All subjects were given informed consent and the protocol was approved by the Ethical Evaluation Committee of Zhongshan Hospital. The aliquots of plasma were collected in potassium-EDTA tubes, centrifuged at 2000 rpm for 20 min, and then stored at −80 °C until analyses.
Gene microarray analysis
PBMCs were isolated by using CPT tubes (Becton Dickinson, Franklin lakes, NT, USA) according to manufacturer’s instructions. Following centrifugation, cells were lysed and DNAse-free total RNA preparation was performed by using Trizol reagent (Invitrogen life technologies, Carlsbad, CA) and the Rneasy kit (QIAGEN, Valencia, CA) according to the manufacturer’s recommendations. RNA concentrations were determined by Nanodrop ND-1000 (Nanodrop Technologies, Wilmington, DE, USA). RNA quality was assessed on an Agilent 2100 Bioanalyzer; samples with a RNA Integrity Score of >6.0 were used in this study. The Human 12 x135K Gene Expression Array (NimbleGen Systems, Inc., Madison, WI, USA) with about 45,000 human genes and transcripts represented with public domain annotations was applied for this study. Sample labeling and array hybridization were performed according to the protocol of One-Color Microarray-Based Gene Expression Analysis (NimbleGen Systems) described previously (Wu et al. 2014b).
Protein microarray analysis
An antibody-based membrane array for measuring inflammatory and immunomodulatory mediators (A Custom Raybio® Human Inflammation Antibody Array kit) was purchased from Raybiotech (Norcross, Ga, USA) with selected 507 factors. Each antibody was spotted in duplicate onto one membrane. The antibody microarray consists of numerous affinity reagents arrayed on a solid surface, and proteins that bind specific target proteins to unique locations on the array are subsequently detected (Mac Beath 2002). Briefly, membranes immobilized with capture antibodies were blocked with 5% bovine serum albumin/TBS for 1 h, and then incubated with 1 ml samples in ten-fold dilution with 5% bovine serum albumin/TBS for 2 h at room temperature. Membranes were then incubated individually or collectively with biotin-conjugated antibodies, after extensive washes with TBS/0.1% Tween 20 and TBS thrice to remove unbound proteins. Membranes were then incubated with horseradish peroxidase-conjugated streptavidin at 2.5 pg/ml for 1 h at room temperature. Unbound materials were washed out with TBS/0.1% Tween 20 and TBS. Buffers C and D were then mixed and loaded onto the membranes to cover the entire surface for 5 min. Finally, spots were detected by exposing to Kodak X-Omat radiographic film for 1 min for image. Each film was scanned with Scanalyze software, and spots were digitized into densities. The data were exported into Microsoft Excel, and for each spot the net density was determined by subtracting the background density from the total raw density. The relative spot density in each membrane equals to [the average of inflammatory mediator spot density − blank density] / [the average of positive control density − blank density] × 100%. According to the protocol from the manufacture, the positive signals are used to identify the orientation and compare the relative expression levels among the different membranes. Horseradish peroxidase-conjugated antibody served as a positive control as 12 spots and was also used to identify the orientation. The positive control of density in the microarray was the known protein in the controlled concentration to control the quality of sample load, the density of spots, and the operation.
Digital evaluation score system
Digital evaluation score system (DESS) is a score index to translate clinical descriptions and information into clinical informatics, which took into account patient symptoms, signs, doctor examination, biochemical analyses, and clinical imaging, as described previously (Chen et al. 2012a). Variables in the DESS included symptoms in Supplement Table 1, signs in Supplement Table 2, and clinical biochemical analysis in Supplement Table 3. For the assessment of severity, each component was then assigned with 0, 1, 2, and 4 as shown in Supplement Tables 1, 2, and 3. The score of 4 as the maximal value indicates far more above normal range or much severer condition, while 0 as the minimal value indicates that it is within physiological range. Several variables were 0 or 4, e.g., orthopnea at night, chill, three depression sign, or barrel chest, etc. The value of 3 was missed in the scoring system for exponential values in order to better define the severity stages. After compiling patients’ data, the points of each variable were added, so that the DESS scores ranged from 0 to 264 points, with higher scores indicating a severer condition. Patients were scored on the day when sample were collected.
Cell lines and reagents
Human lung cancer cell line A549 cells were obtained from Shanghai Institute for Biological Science. Cells were cultured in RPMI 1640 supplemented with 100 U/ml penicillin, 100 mg/ml streptomycin, and 10% heat-inactivated fetal bovine serum (FBS). All cells were maintained at 37 °C in a humidified incubator with 5% carbon dioxide. Human recombinant OPN was purchased from R&D Systems China Co. Ltd. (Shanghai, China). Lipopolysaccharide (LPS) was purchased from Sigma (Escherichia coli, 055: B5, St. Louis, MO). LY294002, a specific inhibitor of PI3K, PD98059, a specific inhibitor of Erk1/2, were purchased from Biovision Company (California, USA). HP antibody for Western blot was purchased from Abcam (HK, China). All reagents were prepared and used as recommended by their suppliers.
Real-time PCR
Quantitative RT-PCR was carried out using real-time PCR with the SYBR Green reporter. Cell cultures were washed in PBS and RNA was isolated using a guanidinium isothiocyanate/chloroform-based technique (Trizol, Life Technologies). OD 260 nm was used to determine RNA yield. RNA was subsequently reverse transcribed to cDNA with the SuperScript First-strand Synthesis System (Invitrogen, USA). Quantitative RT-PCR was carried out afterward. Primer (Invitrogen) concentrations (10 nM) were optimized before use. SYBR Green PCR master kit was used with the appropriate concentrations (10 nM) of forward and reverse primers in a total volume of 20 μl. Quantitative RT-PCR was carried out using an ABI 7000 PCR instrument (Eppendorf, Hamburg, Germany) with the two-stage program parameters provided by the manufacturer, as follows: 1 min at 95 °C, followed by 40 cycles of 5 s at 95 °C and 30 s at 60 °C. The data shown was normalized to GAPDH expression and averaged between three repeated experiments. For data analysis, the raw threshold cycle (CT) value of each sample was first normalized to the house keeping gene, giving us the normalized CT value. The normalized CT was then calibrated to control cell samples, giving the calibrated CT value. Primer sequences were shown in Supplement Table 4.
Western blot analysis
To measure the expression of HP and the signal pathway between OPN and HP, A549 cells were cultured in 6 well plate (1 × 105 cells/well) for 24 h and treated with LY294002 and PD98059 at 10, 20, and 30 μM for another 2 h. Then cells were stimulated with or without OPN at 1 μg/ml for 12 h. Intracellular protein was extracted by RIPA lysis immediately. Protein samples (40 μg) were mixed with an equal volume of 5× SDS sample buffer, boiled for 5 min, and then separated through 10% SDS-PAGE gels. After electrophoresis, proteins were transferred to PVDF membranes by electrophoretic transfer. Membranes were blocked in 5% dry milk (2 h), rinsed, and incubated with primary antibodies (diluted at their instructions) in TBS thrice (TBST) at 4 °C overnight. Primary antibody was then removed by washing in TBST, and labeled by incubating with 0.1 mg/ml peroxidase-labeled secondary antibodies (against mouse and rabbit) for 2 h. Following three washes in TBST, bands were visualized by ECL and exposed to X-ray film. All results were calculated by Phoretix 1D software.
RNA interference and transfection
Three different sequences targeted to three different sites in OPN messenger RNA (mRNA) were designed and provided by Ribobio (Guangzhou, Guangdong, China). The sense and antisense strands of RNA interference (siRNAs) are shown in the Supplement Table 4. Transfection was performed in cells using Lipofectamine 2000 (Invitrogen, USA) according to manufacturer’s protocol. Each experiment included controls containing the transfection reagent with a siRNA control. Transfected cells were detected by real-time PCR and Western blot analysis. SiRNA-OPN-3 was validated for the most efficient interference of OPN and selected for our further study (Data not shown).
Statistical analysis
Statistical analysis were performed by SPSS software (SPSS 18.0; SPSS Inc.; Chicago, IL). Signal densities of microarrays among the three groups were analyzed with one-way ANOVA, followed by an unpaired Student’s t test to compare the difference between two groups, when the ANOVA test indicated significant. The subset of mediators with significance among groups was then selected. Correlation analysis between total DESS and selected mediator intensities was performed by the nonparametric spearman correlation test. All data were expressed as mean ± SEM, and p value of <0.05 was considered statistically significant.
Results
Baseline characteristics of patients with stable COPD or AECOPD
Clinical phenotypes of subjects, such as age, gender, smoking status, lung function test, and emphysema score, were described in Table 1. The baseline FEV1/FVC% and FEV1/pred % of AECOPD patients were similar to those of stable patients, since lung function tests were not performed at the onset of AECOPD due to the severity of disease. There was no significant difference of emphysema extent between stable and AECOPD patients (p = 0.47). DESS scores of subjects from each group were shown in Supplement Table 5. DESS values of patients with stable COPD or AECOPD were significantly higher than in control subjects, as shown in Table 2 (p < 0.01). DESS scores represented the severity of AECOPD and declined as the condition improved. Mean values of DESS in AECOPD patients were 85.7, 70.5, or 36.7 on hospital admission day 1 (AE-1), day 3 (AE-3), or day 10 (AE-10), respectively, and DESS values on AE-1 were significantly higher than AE-3 and AE-10 (p < 0.05 or less, Table 2).
Co-differentially expressed genes of PBMCs in patients with stable COPD or AECOPD
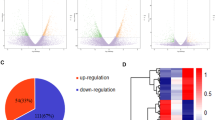
Co-differentially expressed genes of PBMCs in patients with stable COPD or AECOPD from the healthy were selected as COPD specific genes, of which 75 up-expressed (fold changes >5), while 23 down-expressed (fold changes <5), as shown in Fig. 1a. Of those, 16 were specific immunomodulatory genes (14 up-expressed or 2 down-expressed), including DEFA4, LTF, FN1, ELF3, ADCY4, TPSB2, ORM2, RAB17, S100A14, COL3A1, P8, DEFA1, C1QC, C1RL, UBASH3, and IL24 (Fig. 1a). Co-differentially expressed genes of PBMCs in patients with AECOPD on days 1, 3, and 10 from either stable COPD or healthy controls (Fig. 1b) were considered as AECOPD-specific genes. About 82 genes up-expressed and 48 down-expressed in patients with AECOPD, more than twofold as compared with stable COPD or healthy. Of those, 24 were specific immunomodulatory genes (15 up-expressed and 9 down-expressed), including PVRL2, MMP2, CXCL2, ELF3, JAG1, A2M, HP, C1QB, SERPING1, C1RL, ORM1, ECM1, C1QC, C3, FN1, SLC11A1, UBASH3A, LGALS3, DDX3X, TNFAIP3, IFNGR1, VSIG4, CD55, or CD8B (Fig. 1b).
a Co-differentially expressed genes of PBMCs from patients with stable COPD or AECOPD, as compared to the healthy controls. b Co-differentially expressed genes of PBMCs from patients with AECOPD, as compared to both stable COPD and healthy controls. c Co-differentially expressed proteins of plasma from patients with stable COPD or AECOPD, as compared to the healthy controls. d Co-differentially expressed proteins of plasma from patients with AECOPD, as compared to both stable COPD and healthy controls. C9 complement component C9, C7 complement component C7, FGB fibrinogen beta chain, KRT1 keratin type II cytoskeletal 1, APOA1 apolipoprotein A-I, F2 prothrombin, SERPINC1 antithrombin-III, HRG histidine-rich glycoprotein, PGLYRP2 N-acetylmuramoyl-l-alanine amidase, VTN vitronectin, SERPINF2 alpha-2-antiplasmin, KLKB1 plasma kallikrein, PPBP platelet basic protein, ATRN attractin, NCAM1 neural cell adhesion molecule 1, HPX hemopexin, IGHA1 Ig alpha-1 chain C region, APOE apolipoprotein E, AZGP1 zinc-alpha-2-glycoprotein, AGT angiotensinogen, C4BPA C4b-binding protein alpha chain, IGJ immunoglobulin J chain, F12 coagulation factor XII, SAA1 serum amyloid A protein, PROS1 vitamin K-dependent protein S, C4BPB C4b-binding protein beta chain, CRP C-reactive protein pentraxin-related, APCS serum amyloid P-component, CFH complement factor H, IGHG3 Ig gamma-3 chain C region, AMBP protein AMBP, CFI complement factor I isoform CRA_b, C1R complement C1r subcomponent, C1QA complement C1q subcomponent subunit A, CFHR5 complement factor H-related protein 5, NCAM1 neural cell adhesion molecule 1
Co-differentially expressed protein of plasma in patients with stable COPD or AECOPD
In the protein level, we found 22 COPD-specific proteins in the circulation up-expressed and 29 down-expressed in patients with stable COPD and AECOPD, more than 1.2-fold as compared to the healthy controls (Fig. 1c). Of them, 20 were specific immunomodulatory mediators (6 up-expressed and 14 down-expressed), including complement C1q subcomponent subunit C, complement C1r subcomponent-like protein, complement component C9, complement component C7, fibrinogen beta chain, and others (Fig. 1c). To seek for AECOPD-specific proteins, co-differentially expressed proteins of plasma from patients with AECOPD on days 1, 3, and 10 were compared to those from either stable COPD or healthy controls. As compared with both stable COPD and healthy controls, about 84 AECOPD-specific protein were selected, of which 50 up-expressed and 34 down-expressed more than 1.2-fold (Fig. 1d). Of those, 38 were specific immunomodulatory mediators (23 up-expressed and 15 down-expressed), including apolipoprotein A-I and II, prothrombin, antithrombin-III, vitronectin, alpha-2-antiplasmin, plasma kallikrein, platelet basic protein, attractin, neural cell adhesion molecule 1, and others, as shown in Fig. 1d.
Both genes and proteins were co-differentially up-expressed in patients with COPD or AECOPD
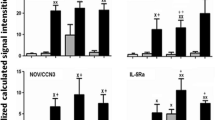
Figure 2 demonstrated that gene and protein expressions of C1QC or C1RL in patients with stable COPD or AECOPD were significantly higher than those in healthy controls (p < 0.01). Gene and protein expressions of HP, ORM1, C3, and SERPING1 in patients with AECOPD on days 1, 3, and 10 were significantly higher than those in either stable COPD or healthy controls (p < 0.01). There was no significant difference of C1QC, C1RL, ORM1, C3, and SERPING1 expression between different groups stimulated by LPS (Fig. 3). The details of these specific genes are listed in Table 3.
Gene and protein levels of C1QC (a), C1RL (b), SERPING1 (c), HP (d), ORM1 (e), and C3 (f) in healthy controls, patients with stable COPD or AECOPD on days 1, 3, and 10. Double asterisks stand for p values less than 0.01, as compared with healthy controls. Double plus signs stand for p values less than 0.01, as compared with stable COPD
The expression of immunomodulatory mediators stimulated by LPS in human lung epithelial cells A549. A549 cells grown in complete medium were left untreated (control) or treated with LPS (0.01, 0.1, and 1 μg/ml) for 6 h. The mRNA expression of C1QC (a), C1RL (b), C3 (c), ORM1 (d), SERPING1 (e), and HP (f) from A549 in response to LPS. Data were normalized to control, each data point represents mean ± SEM of three experiments. Asterisk and double asterisks stand for p values less than 0.05 and 0.01, respectively, as compared with control
Effects of PI3K and Erk1/2 inhibitors on OPN-induced HP production
HP mRNA or protein in LPS-stimulated A549 cells up-expressed significantly from 3 or 12 h and on with a peak at 6 or 24 h, respectively (p < 0.05 or 0.01, respectively, Fig. 4a or 4c). Figure 4b, d demonstrated that the transfection of OPN siRNA blocked LPS-induced HP production (p < 0.05 or 0.01, respectively). To characterize the individual contribution of OPN on HP production, we further stimulated the A549 cells with human recombinant OPN. Treatment with OPN at the concentration of 1000 ng/ml could significantly increase HP mRNA (Fig. 5a) and protein (Fig. 5b) expression in lung epithelial cells. Treatments with Erk1/2 inhibitor PD98059 did not influence the expression of HP in A549 cells (Fig. 5c, d), while PI3K inhibitor LY294002 at 20 or 30 μM significantly inhibited OPN-induced HP production (p < 0.05 or 0.01, respectively, Fig. 5e, f).
LPS induces increased production of HP in human lung epithelial cells A549. A549 cells were stimulated with LPS (1 μg/ml) for indicated times. Real-time PCR shows LPS-induced expression of HP mRNA with a peak at 6 h of stimulation and continues to increase at 24 h (a). LPS induced HP protein expression for indicated times. Western blot shows LPS-induced expression of HP protein from 12 h of stimulation to peak at 24 h (c). A549 cells were transfected with OPN siRNA (A549-SHR) and nonspecific control siRNA (A549-NC), respectively. Subsequently, cells were stimulated with LPS (1 μg/ml). Total RNA was extracted and subjected to reverse transcription followed by real-time-PCR to detect HP mRNA at 6 h (b). The level of HP protein was measured by Western blot at 24 h (d). Each data point represents mean ± SEM of three experiments. Asterisks and double asterisks stand for p values less than 0.05 and 0.01, in comparison with control, plus sign and double plus signs stand for p values less than 0.05 and 0.01, as compared to LPS and control siRNA, respectively
Effects of PI3K and ERK inhibitors on OPN-induced HP production. A549 cells were stimulated with OPN (10, 100, and 1000 ng/ml) for 6 h to detect HP mRNA (a) or 24 h to detect its HP protein (b). The mRNA and protein of HP from cells was measured after the culture with DMSO alone (control), OPN at 1000 ng/ml plus DMSO, ERK-specific inhibitor PD98089 (c, d), or PI3K-specific inhibitor LY294002 (e, f), at doses of 10, 20, or 30 μM. Asterisks and double asterisks stand for p values less than 0.05 and 0.01. Plus sign and double plus signs stand for p values less than 0.05 and 0.01, as compared to OPN and DMSO, respectively
Discussion
Systemic inflammation and immune response are key processes in the pathogenesis of COPD and AECOPD, especially in the lung characterized by increased recruitment of inflammatory cells and over-production of inflammatory mediators in the lung tissues, sputum, and bronchoalveolar lavage fluid from COPD patients (Chen et al. 2010). It is responsible for the initiation and acceleration of the secondary inflammatory reactions (Wang et al. 2007). Systemic cytokine patterns were found to vary between different stages and severities of COPD and AECOPD (Valipour et al. 2008). We developed a new protocol of biomarker evaluation by comparing systemic profiles of inflammatory mediators among different study groups and disease stages, integrating clinical informatics and bioinformatics, and understanding the biological function and signal networks of COPD (Chen et al. 2012a; Chen et al. 2012b). Systemic cytokine profiles in COPD patients were associated with airway and parenchymal phenotypes (Bon et al. 2009), reduced lung function, and other clinical variables (de Torres et al. 2011; Thorleifsson et al. 2009). The present study furthermore integrated genomics of PBMCs, and proteomics of circulating proteins, and clinical phenotypes, to select and validate COPD-specific, AECOPD-specific, or severity-specific immunomodulatory mediators from systemic inflammatory mediators.
PBMCs play a critical and important role in the occurrence of AECOPD due to less capacity for balancing the pro-inflammatory immune response caused by infection and for secreting adequate amounts of anti-inflammatory cytokines (Rupp et al. 2003). The present study specially focused on the AECOPD-specific immunomodulatory mediators by evaluating the dynamic gene and protein profiles of PBMCs or plasma from patients with AECOPD on days 1, 3, and 10 after the hospital admission, to compare with healthy controls or patients with stable COPD. We found 36 COPD-specific immunomodulatory mediator genes more than 1.2-fold, as well as 62 AECOPD-specific immunomodulatory genes more than 1.2-fold. We selected COPD or AECOPD-specific immunomodulatory mediator profiles in the co-existent expression of gene and protein and the consistence of alterations on basis of the disease, stages, and severity. To our knowledge, it is for the first time to select and validate COPD-specific and AECOPD-specific immunomodulatory biomarker by integrating genomics, proteomics, with clinical informatics. We found two profile patterns of systemic immunomodulatory mediators, e.g., C1QC and C1RL as COPD-specific biomarkers; HP, ORM1, C3, and SERPING1 as AECOPD-specific biomarkers.
We found that the expression of HP was up-regulated in response to inflammatory stimuli. HP protein as a major positive acute-phase reactant elevated in the circulation of various diseases (Beckman et al. 1986; Zhao et al. 2007). HP can transport hemoglobin to the liver and facilitate hemoglobin catabolism to prevent tissue injury, through antioxidant activity (Gutteridge 1987), angiogenesis (Cid et al. 1993), prostaglandin synthesis (Langlois and Delanghe 1996), and reverse cholesterol transport (Salvatore et al. 2007). HP was recently found to contribute the host defense response to infection and inflammation by receptor-ligand activation of the immune system (Huntoon et al. 2013). There is increasing evidence that HP has biological function in the lung (Landi et al. 2011; Okamoto et al. 2012; Wen et al. 2012). HP levels decreased in patients with asthma and/or rhinitis (Khazaei et al. 2012; Larsen et al. 2006). We showed the further evidence that the inflammatory agent could induce gene and protein expression of HP in lung epithelial cells. HP could be synthesized and over-expressed in the human lung alveolar epithelial cells during inflammation, probably involved in lung injury and repair. The HP level increased at the acute-phase and more severe state of COPD.
HP could decrease the reactivity of lymphocytes and neutrophils as a natural antagonist for receptor-ligand activation of the immune system (Oh et al. 1990) or directly affects T cells and suppresses T helper cells through down-regulation of the cytokine production. The evidence from our previous studies and others suggested that OPN might play a critical role in the physiological and pathological processes of lung diseases by modulating both innate and adaptive immune responses (Chen et al. 2012a; Wai and Kuo 2004). We also found that the level of OPN is significantly positive correlated with HP expression. To explore the correlation between OPN and HP, we measured the expression of HP after transfection of OPN siRNA or human recombinant OPN stimulation and the results demonstrated that both endogenous and exogenous OPN could induce over-production of HP.
OPN resembles a matrix protein and binds to integrin and CD44 family receptors, which in turn activates focal adhesion kinase (FAK), NF-kappaB pathways, PI3K/Akt, and MAPK/Erk1/2 signaling pathways that induce distinct patterns of cytokine/chemokine expression and specific immune responses (Lin and Yang-Yen 2001; Philip and Kundu 2003; Xie et al. 2014). There is evidence suggesting that PI3K plays an important role in the acute and chronic lung inflammation (Chen et al. 2011; Fang et al. 2013a). Our data furthermore indicate that the potential mechanism by which OPN is regulated to produce HP over-produced through the signal pathway of PI3K/Akt axis. he over-production of HP by OPN was fully prevented by PI3K inhibitor, rather than Erk1/2 inhibitor. It implies that the OPN-PI3K/Akt-HP chain can be the potential of new therapeutic target in lung cancer or chronic lung diseases.
However, this study is limited by several factors, we randomly selected six patients in each groups representative of the whole population due to the high cost of testing. The present manuscript is a preliminary study, it needs to be verified in a large number of patients. Additionally, the alveolar epithelial cell A549 we employed is a transformed, immortalized lung cancer cell line. Although A549 is a well-established alveolar epithelial cell model, there are still some differences between tumor cells and normal alveolar epithelial cells. Responses of the normal bronchial epithelial cells or epithelial cell harvested from COPD patients need to be furthermore evaluated in future.
In conclusion, we explored the feasibility and reliability of a new protocol of disease-specific biomarker evaluation by integrating genomics and proteomic profiles of immunomodulatory mediators in different AECOPD stages, with clinical informatics. We found four AECOPD-specific immunomodulatory mediator candidates in patients different from both healthy and stable COPD patients. Of them, the expression of HP is up-regulated through OPN-dependent signal in response to stimuli. Both endogenous and exogenous OPN could increase the over-production HP through the PI3K/Akt pathway. The complex network of AECOPD- or COPD-specific immunomodulatory mediators will benefit the development of precision or personalized medicine strategies.
References
Ashkar S, Weber GF, Panoutsakopoulou V, Sanchirico ME, Jansson M, Zawaideh S, et al. Eta-1 (osteopontin): an early component of type-1 (cell-mediated) immunity. Science. 2000;287(5454):860–4.
Barnes PJ. Cellular and molecular mechanisms of chronic obstructive pulmonary disease. Clin Chest Med. 2014;35(1):71–86.
Barnes PJ, Shapiro SD, Pauwels RA. Chronic obstructive pulmonary disease: molecular and cellular mechanisms. The European respiratory journal : official journal of the European Society for Clinical Respiratory. Physiology. 2003;22(4):672–88.
Beckman G, Eklund A, Frohlander N, Stjernberg N. Haptoglobin groups and lung cancer. Hum Hered. 1986;36(4):258–60.
Bhattacharya S, Tyagi S, Srisuma S, Demeo DL, Shapiro SD, Bueno R, et al. Peripheral blood gene expression profiles in COPD subjects. Journal of clinical bioinformatics. 2011;1(1):12.
Bon JM, Leader JK, Weissfeld JL, Coxson HO, Zheng B, Branch RA, et al. The influence of radiographic phenotype and smoking status on peripheral blood biomarker patterns in chronic obstructive pulmonary disease. PLoS One. 2009;4(8):e6865.
Celli BR, Barnes PJ. Exacerbations of chronic obstructive pulmonary disease. The European respiratory journal: official journal of the European Society for Clinical Respiratory. Physiology. 2007;29(6):1224–38.
Celli BR, Mac Nee W. Standards for the diagnosis and treatment of patients with COPD: a summary of the ATS/ERS position paper. The European respiratory journal: official journal of the European Society for Clinical Respiratory. Physiology. 2004;23(6):932–46.
Chabas D, Baranzini SE, Mitchell D, Bernard CC, Rittling SR, Denhardt DT, et al. The influence of the proinflammatory cytokine, osteopontin, on autoimmune demyelinating disease. Science. 2001;294(5547):1731–5.
Chen C, Fang X, Wang Y, Li Y, Wang D, Zhao X, et al. Preventive and therapeutic effects of phosphoinositide 3-kinase inhibitors on acute lung injury. Chest. 2011;140(2):391–400.
Chen H, Song Z, Qian M, Bai C, Wang X. Selection of disease-specific biomarkers by integrating inflammatory mediators with clinical informatics in AECOPD patients: a preliminary study. J Cell Mol Med. 2012a;16(6):1286–97.
Chen H, Wang D, Bai C, Wang X. Proteomics-based biomarkers in chronic obstructive pulmonary disease. J Proteome Res. 2010;9(6):2798–808.
Chen H, Wang X. Significance of bioinformatics in research of chronic obstructive pulmonary disease. Journal of clinical bioinformatics. 2011;1:35.
Chen H, Wang Y, Bai C, Wang X. Alterations of plasma inflammatory biomarkers in the healthy and chronic obstructive pulmonary disease patients with or without acute exacerbation. J Proteome. 2012b;75(10):2835–43.
Cid MC, Grant DS, Hoffman GS, Auerbach R, Fauci AS, Kleinman HK. Identification of haptoglobin as an angiogenic factor in sera from patients with systemic vasculitis. J Clin Invest. 1993;91(3):977–85.
de Torres JP, Casanova C, Pinto-Plata V, Varo N, Restituto P, Cordoba-Lanus E, et al. Gender differences in plasma biomarker levels in a cohort of COPD patients: a pilot study. PLoS One. 2011;6(1):e16021.
Denhardt DT, Noda M, O'Regan AW, Pavlin D, Berman JS. Osteopontin as a means to cope with environmental insults: regulation of inflammation, tissue remodeling, and cell survival. J Clin Invest. 2001;107(9):1055–61.
Fang X, Li K, Tao X, Chen C, Wang X, Wang L, et al. Effects of phosphoinositide 3-kinase on protease-induced acute and chronic lung inflammation, remodeling, and emphysema in rats. Chest. 2013a;143(4):1025–35.
Fang X, Netzer M, Baumgartner C, Bai C, Wang X. Genetic network and gene set enrichment analysis to identify biomarkers related to cigarette smoking and lung cancer. Cancer Treat Rev. 2013b;39(1):77–88.
Fang X, Wang X, Bai C. COPD in China: the burden and importance of proper management. Chest. 2011;139(4):920–9.
Guirguis-Blake JM, Senger CA, Webber EM, Mularski RA, Whitlock EP. Screening for chronic obstructive pulmonary disease: evidence report and systematic review for the US preventive services task force. JAMA. 2016;315(13):1378–93.
Gutteridge JM. The antioxidant activity of haptoglobin towards haemoglobin-stimulated lipid peroxidation. Biochim Biophys Acta. 1987;917(2):219–23.
He Z, Chen Y, Chen P, Wu G, Cai S. Local inflammation occurs before systemic inflammation in patients with COPD. Respirology. 2010;15(3):478–84.
Hogg JC, Chu F, Utokaparch S, Woods R, Elliott WM, Buzatu L, et al. The nature of small-airway obstruction in chronic obstructive pulmonary disease. N Engl J Med. 2004;350(26):2645–53.
Huntoon KM, Russell L, Tracy E, Barbour KW, Li Q, Shrikant PA, et al. A unique form of haptoglobin produced by murine hematopoietic cells supports B-cell survival, differentiation and immune response. Mol Immunol. 2013;55(3–4):345–54.
Khazaei HA, Nakhaei A, Dashti GA, Mohammadi M, Hejazenia F, Mehrangeez N, et al. Association of haptoglobin phenotypes with serum levels of IgE and IgA in allergic rhinitis patients. Iranian journal of immunology: IJI. 2012;9(4):254–60.
Landi C, Bargagli E, Magi B, Prasse A, Muller-Quernheim J, Bini L, et al. Proteome analysis of bronchoalveolar lavage in pulmonary langerhans cell histiocytosis. Journal of clinical bioinformatics. 2011;1:31.
Langlois MR, Delanghe JR. Biological and clinical significance of haptoglobin polymorphism in humans. Clin Chem. 1996;42(10):1589–600.
Larsen K, Macleod D, Nihlberg K, Gurcan E, Bjermer L, Marko-Varga G, et al. Specific haptoglobin expression in bronchoalveolar lavage during differentiation of circulating fibroblast progenitor cells in mild asthma. J Proteome Res. 2006;5(6):1479–83.
Lin YH, Yang-Yen HF. The osteopontin-CD44 survival signal involves activation of the phosphatidylinositol 3-kinase/Akt signaling pathway. J Biol Chem. 2001;276(49):46024–30.
Liu R, Wang X, Aihara K, Chen L. Early diagnosis of complex diseases by molecular biomarkers, network biomarkers, and dynamical network biomarkers. Med Res Rev. 2014;34(3):455–78.
Mac Beath G. Protein microarrays and proteomics. Nat Genet. 2002;32(Suppl):526–32.
Melguizo C, Prados J, Luque R, Ortiz R, Rama AR, Caba O, et al. Modulation of multidrug resistance gene expression in peripheral blood mononuclear cells of lung cancer patients and evaluation of their clinical significance. Cancer Chemother Pharmacol. 2013;71(2):537–41.
Murtagh E, Heaney L, Gingles J, Shepherd R, Kee F, Patterson C, et al. Prevalence of obstructive lung disease in a general population sample: the NICECOPD study. Eur J Epidemiol. 2005;20(5):443–53.
Niu F, Wang DC, Lu J, Wu W, Wang X. Potentials of single-cell biology in identification and validation of disease biomarkers. J Cell Mol Med. 2016; doi:10.1111/jcmm.12868.
Oh SK, Pavlotsky N, Tauber AI. Specific binding of haptoglobin to human neutrophils and its functional consequences. J Leukoc Biol. 1990;47(2):142–8.
Okamoto T, Miyazaki Y, Shirahama R, Tamaoka M, Inase N. Proteome analysis of bronchoalveolar lavage fluid in chronic hypersensitivity pneumonitis. Allergology international: official journal of the Japanese Society of Allergology. 2012;61(1):83–92.
Philip S, Kundu GC. Osteopontin induces nuclear factor kappa B-mediated promatrix metalloproteinase-2 activation through I kappa B alpha/IKK signaling pathways, and curcumin (diferulolylmethane) down-regulates these pathways. J Biol Chem. 2003;278(16):14487–97.
Prins HJ, Daniels JM, Lindeman JH, Lutter R, Boersma WG. Effects of doxycycline on local and systemic inflammation in stable COPD patients, a randomized clinical trial. Respir Med. 2016;110:46–52.
Rodriguez-Roisin R. Toward a consensus definition for COPD exacerbations. Chest. 2000;117(5 Suppl 2):398S–401S.
Rubinsztajn R, Przybylowski T, Maskey-Warzechowska M, Karwat K, Chazan R. Exacerbations of chronic obstructive pulmonary disease and quality of life of patients. Adv Exp Med Biol. 2016;884:69–74.
Rupp J, Kothe H, Mueller A, Maass M, Dalhoff K. Imbalanced secretion of IL-1beta and IL-1RA in chlamydia pneumoniae-infected mononuclear cells from COPD patients. The European respiratory journal: official journal of the European Society for Clinical Respiratory. Physiology. 2003;22(2):274–9.
Salvatore A, Cigliano L, Bucci EM, Corpillo D, Velasco S, Carlucci A, et al. Haptoglobin binding to apolipoprotein A-I prevents damage from hydroxyl radicals on its stimulatory activity of the enzyme lecithin-cholesterol acyl-transferase. Biochemistry. 2007;46(39):11158–68.
Thorleifsson SJ, Margretardottir OB, Gudmundsson G, Olafsson I, Benediktsdottir B, Janson C, et al. Chronic airflow obstruction and markers of systemic inflammation: results from the BOLD study in Iceland. Respir Med. 2009;103(10):1548–53.
Valipour A, Schreder M, Wolzt M, Saliba S, Kapiotis S, Eickhoff P, et al. Circulating vascular endothelial growth factor and systemic inflammatory markers in patients with stable and exacerbated chronic obstructive pulmonary disease. Clin Sci (Lond). 2008;115(7):225–32.
Vestbo J, Hurd SS, Agusti AG, Jones PW, Vogelmeier C, Anzueto A, et al. Global strategy for the diagnosis, management, and prevention of chronic obstructive pulmonary disease: GOLD executive summary. Am J Respir Crit Care Med. 2013;187(4):347–65.
Wai PY, Kuo PC. The role of Osteopontin in tumor metastasis. J Surg Res. 2004;121(2):228–41.
Wang KX, Denhardt DT. Osteopontin: role in immune regulation and stress responses. Cytokine Growth Factor Rev. 2008;19(5–6):333–45.
Wang X. Role of clinical bioinformatics in the development of network-based biomarkers. Journal of clinical bioinformatics. 2011;1(1):28.
Wang X, Adler KB, Erjefalt J, Bai C. Airway epithelial dysfunction in the development of acute lung injury and acute respiratory distress syndrome. Expert review of respiratory medicine. 2007;1(1):149–55.
Wang X, Ward PA. Opportunities and challenges of disease biomarkers: a new section in the journal of translational medicine. J Transl Med. 2012;10:240.
Wen CL, Chen KY, Chen CT, Chuang JG, Yang PC, Chow LP. Development of an alpha LISA assay to quantify serum core-fucosylated E-cadherin as a metastatic lung adenocarcinoma biomarker. J Proteome. 2012;75(13):3963–76.
Wu X, Chen L, Wang X. Network biomarkers, interaction networks and dynamical network biomarkers in respiratory diseases. Clinical and translational medicine. 2014a;3:16.
Wu X, Sun X, Chen C, Bai C, Wang X. Dynamic gene expressions of peripheral blood mononuclear cells in patients with acute exacerbation of chronic obstructive pulmonary disease: a preliminary study. Crit Care. 2014b;18(6):508.
Xie Y, Li Y, Kong Y. OPN induces fox M1 expression and localization through ERK 1/2, AKT, and p38 signaling pathway in HEC-1A cells. Int J Mol Sci. 2014;15(12):23345–58.
Zhao C, Annamalai L, Guo C, Kothandaraman N, Koh SC, Zhang H, et al. Circulating haptoglobin is an independent prognostic factor in the sera of patients with epithelial ovarian cancer. Neoplasia. 2007;9(1):1–7.
Zhao W, Wang L, Zhang L, Yuan C, Kuo PC, Gao C. Differential expression of intracellular and secreted osteopontin isoforms by murine macrophages in response to toll-like receptor agonists. J Biol Chem. 2010;285(27):20452–61.
Acknowledgements
The work was supported by Shanghai Leading Academic Discipline Project (B115), Zhongshan Distinguished Professor Grant (XDW), the National Nature Science Foundation of China (91230204, 81270099, 81320108001, 81270131, 81400035, 81570075, 81500058, 81500025), the Shanghai Committee of Science and Technology (12JC1402200, 12431900207, 11410708600), Zhejiang Provincial Natural Science Foundation (LZ15H010001), and Zhejiang Provincial Science Technology Department Foundation (WKJ-ZJ-1526).
Author information
Authors and Affiliations
Contributions
Conceived and designed the study: Xiangdong Wang; performed the biological experiments: Lin Shi; statistical analysis: Bijun Zhu and Menglin Xu. Wrote the paper: Lin Shi. All authors read and proofed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Shi, L., Zhu, B., Xu, M. et al. Selection of AECOPD-specific immunomodulatory biomarkers by integrating genomics and proteomics with clinical informatics. Cell Biol Toxicol 34, 109–123 (2018). https://doi.org/10.1007/s10565-017-9405-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10565-017-9405-x