Abstract
The main aim of this study was to evaluate whether microRNA (miRNA) profiling could be a useful tool for in vitro developmental neurotoxicity (DNT) testing. Therefore, to identify the possible DNT biomarkers among miRNAs, we have studied the changes in miRNA expressions in a mixed neuronal/glial culture derived from carcinoma pluripotent stem cells (NT2 cell line) after exposure to methyl mercury chloride (MeHgCl) during the process of neuronal differentiation (2–36 days in vitro (DIV1)). The neuronal differentiation triggered by exposure to retinoic acid (RA) was characterized in the control culture by mRNA expression analysis of neuronal specific markers such as MAP2, NF-200, Tubulin βIII, MAPT-tau, synaptophysin as well as excitatory (NMDA, AMPA) and inhibitory (GABA) receptors. The results obtained from the miRNA expression analysis have identified the presence of a miRNA signature which is specific for neural differentiation in the control culture and another for the response to MeHgCl-induced toxicity. In differentiated neuronal control cultures, we observed the downregulation of the stemness phenotype-linked miR-302 cluster and the overexpression of several miRNAs specific for neuronal differentiation (e.g. let-7, miR-125b and miR-132). In the cultures exposed to MeHgCl (400 nM), we observed an overexpression of a signature composed of five miRNAs (miR-302b, miR-367, miR-372, miR-196b and miR-141) that are known to be involved in the regulation of developmental processes and cellular stress response mechanisms. Using gene ontology term and pathway enrichment analysis of the validated targets of the miRNAs deregulated by the toxic treatment, the possible effect of MeHgCl exposure on signalling pathways involved in axon guidance and learning and memory processes was revealed. The obtained data suggest that miRNA profiling could provide simplified functional evaluation of the toxicity pathways involved in developmental neurotoxicity in comparison with the transcriptomics studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are endogenous, small and noncoding RNAs that introduce an additional layer of post-translational regulatory control of gene expression. They regulate most cell processes from development and differentiation to cell death. It has been suggested that each miRNA could target up to a few hundred target genes resulting in the regulation of as much as one third of the protein expression codified in animals (Krek et al. 2005). Moreover, one third of miRNAs are highly conserved across different species and around 60 % of miRNAs conservation has been identified when comparing mouse with human (Roush and Slack 2008). They regulate gene expression at the post-transcriptional level through imperfect base-paring with specific sequences, located mostly in the 3′ UTRs (untranslated region) of the mRNA target. Often, the perfect complementarity between the miRNA and its target mRNA results in degradation of the mRNA whereas non-perfect complementarily induces translational inhibition.
Although a large percentage of known miRNAs are described in the brain, not much is known about their role during brain development. They are implicated in many developmental processes such as neurogenesis, neuronal differentiation, neurite outgrowth, synaptic plasticity and their expression is tightly regulated (Krichevsky et al. 2003; Miska et al. 2004).
Dicer ablation that results in the absence of all mature miRNAs has been used as a valuable tool to study the general role of miRNAs in regulatory pathways in the central nervous system (CNS). It has been shown that loss of Dicer reduces dendritic branching, affects spine morphology (Christensen and Schratt 2009), causes cell death of subpopulation of neurons: such as dopaminergic in the midbrain (Kim et al. 2007), Purkinje cells of the cerebellum (Schaefer et al. 2007) and neurons in the cortex (De Pietri Tonelli et al. 2008).
The most abundant brain-specific miRNA-124 has been shown to be also highly upregulated during the process of neuronal differentiation (Gao 2010). Indeed, many targets of miRNA-124 have been identified (Lim et al. 2005; Visvanathan et al. 2007; Makeyev et al. 2007), highlighting its primary function in supporting neuronal identity by downregulation of non-neuronal mRNAs (Cao et al. 2007). In addition, miRNA-124 is involved in the expression of neural specific genes through regulation of the RNA-binding protein PTBP1 (Makeyev et al. 2007) and it promotes expression of genes involved in neurite outgrowth, particularly those involved in controlling cytoskeleton reorganization (Yu et al. 2008). Furthermore, one of the possible mRNA targets of miRNA-124 is the cAMP response element bounding protein (CREB), a transcriptional activator required for long-term potentiation and enhancing synaptic facilitation (Barco et al. 2002). Additionally, CREB is also regulated by miRNA-132, affecting neurite outgrowth and branching, mainly through the Rho family GTPase activating protein p250GAP (Vo et al. 2005; Wayman et al. 2008). In mammals, another miRNA that plays an important role in neurite outgrowth is miRNA-388. During the developmental process, the AATK and miRNA-388 are co-expressed and both molecules are critical for regulating dendritic spine morphogenesis, synaptic plasticity and memory formation (Christensen and Schratt 2009). Several other miRNAs have been implicated in synaptic formation and plasticity such as miRNA-9, miRNA-284, miRNA-1, miRNA-138 (Christensen and Schratt 2009), miRNA-206, let-7 and miRNA-125a playing central roles in neuronal connectivity that is associated with learning and memory processes (Wang et al. 2012; Konopka et al. 2011). Recent studies also indicated an important role of let-7 and miRNA-125 at earlier stages of neurogenesis, such as neural tube closure and brain patterning process (Maller Schulman et al. 2008). Furthermore, miRNA-9 has been shown to promote neurogenesis, especially in mid- and hind brain (Leucht et al. 2008).
At the later stages of neuronal maturation, several studies claimed that there are miRNAs located in the dendritic shaft, such as miRNA-134, involved in the regulation of local protein synthesis at the mammalian synapses (Schratt et al. 2006). It is postulated that those newly synthesized proteins derived from local translation of mRNAs could be important for the establishment of certain forms of long-term memory (Sutton and Schuman 2006). Summing up, the above studies indicate that miRNAs are heavily involved in the regulation of neuronal developmental at different stages of cell differentiation and maturation.
The main aim of this study was to go one step further and evaluate whether miRNA profiling could be a useful tool for DNT testing using an in vitro approach. Therefore, to identify the possible DNT biomarkers among miRNAs we have studied the changes in miRNA expression during the process of neuronal differentiation in the control culture of carcinoma, pluripotent human stem cell-derived neurons and glial cells (NT2 cell line) and compared this with the profile of miRNAs expression after exposure to low concentration of methyl mercury chloride (MeHgCl). This is a well known DNT chemical that disturbs the process of neuronal differentiation, synapse formation and causes the damage to the process of learning and memory (Grandjean and Landrigan 2006; Hogberg et al. 2009, 2010). The exposure to MeHgCl was performed during the early stages of the neuronal differentiation process (1–36 DIV) since the previous in vitro studies from our laboratory (Hogberg et al. 2009; 2010) and epidemiological evidence (Claudio et al. 2000; Eriksson 1997; Tilson 2000; Rodier 1995) indicate that, compared to the adult CNS, the developing brain is more susceptible to toxicants. Moreover, the early exposure of neural progenitor cells to chemicals seems to be a more vulnerable time period than the later stages of neuronal differentiation (Stummann et al. 2009; Buzanska et al. 2009). Indeed, it has been proven that the human embryonic stem cell (hESC) at the stage of the neural precursor formation were already affected by non-cytotoxic concentrations of MeHg (Stummann et al. 2009). A similar conclusion was obtained based on the exposure of a human neural precursor cells derived from umbilical cord blood to various DNT compounds, including MeHgCl (Buzanska et al. 2009).
From the regulatory point of view, new alternative methods are urgently needed to speed up the process of chemical testing to be able to identify those with DNT potential. So far, the information on the potential developmental toxicity induced by chemicals is limited since only a few compounds (arsenic, lead, methyl mercury, polychlorinated biphenyls and toluene) have been studied (Grandjean and Landrigan 2006). Currently, there are official DNT guidelines in Europe (OECD, TG 426, 2007) and the USA (U.S. EPA U.S. Environmental Protection Agency 1998) that are entirely based on animal studies but rarely used as they are complex, time consuming and not suitable for testing high number of chemicals. Therefore, referring to the document published by the National Academy of Sciences (NAS) in USA, Toxicity Testing of the 21st century (NRC 2007) we used the human NTERA-2/cl.D1 (NT2) cell line which can be differentiated from neural progenitor cells into mature neuronal- and glial-like cells in the presence of retinoic acid. The process of neuronal differentiation in vitro has been characterized in detail in our previous studies (Laurenza et al. 2013). NT2 cells can reach advanced stages of neuronal differentiation which is measured by high expression (both mRNA and protein) of critical neuronal markers such as NF-200, MAP2, MAPT-tau, synaptophysin, NMDA, AMPA and GABA receptors and glial marker (GFAP).
In this study, the mechanisms of toxicity induced by MeHgCl were determined based on the changes in miRNA expression. Those miRNAs which expression was up- or downregulated most significantly were evaluated in terms of whether they could serve as the potential DNT biomarkers for the cellular perturbations induced by MeHgCl. Therefore, the available databases of miRNAs/mRNA gene ontology and bioinformatics tools were used to link miRNA expression to their mRNA targets to be able to interpret the results in terms of cell function modification. Based on the obtained results, it is expected that miRNA profiling could provide simplified functional evaluation of the cellular pathways involved in developmental neurotoxicity mechanisms in comparison with the transcriptomics approach.
Materials and methods
NT2 cells maintenance and differentiation towards neuronal phenotype
NTERA-2/cl.D1 cells were differentiated into neurons as described previously in Pleasure et al. (1992). Undifferentiated NT2 cells (ATCC) were cultured in standard tissue culture flasks (Nunc) and maintained in Opti-MEM I (Gibco) supplemented with 5 % fetal bovine serum (FBS) (HyClone), 100 U/ml penicillin (P) and 100 μg/ml streptomycin (S) (Gibco). The differentiation process was performed by culturing the cells in Dulbecco's modified Eagle's medium—high glucose (DMEM-HG) (Gibco) supplemented with 10 % FBS, 1 % P/S, and 10 μM retinoic acid (RA) (Sigma) up to 36 DIV at 37 °C in a humidified atmosphere of 5 % of CO2. To obtain fully differentiated neurons, the cells were trypsinized at 36 DIV, and split in standard medium (DMEM-HG with 5 % FBS, P/S) supplemented with a mixture of mitosis inhibitors composed of 1 μM cytosine arabinoside (Sigma), 10 μM fluorodeoxyuridine (Sigma) and 10 μM uridine (Sigma). Cells were then seeded in plates coated with 10 μg/ml poly-d-lysine (Sigma) and 0.26 mg/ml Matrigel and cultured for the next 8 weeks (up to 96 DIV) at 37 °C in a humidified atmosphere of 5 % of CO2. The medium was changed twice a week.
mRNAs expression analysis of neuronal markers using Real-Time PCR
The total RNA extraction was performed according to the manufacturer's protocol of RNeasy Mini Kit (Qiagen). Any DNA contamination was removed by digestion process using an RNase-free DNase kit (Qiagen). RNA concentration and protein contamination were assessed spectrophotometrically (Biophotometer; Eppendorf) according to the 260/280 nm optical density ratio.
Reverse transcription was performed as follows: 500 ng RNA was incubated with 2.5 mM PCR Nucleotide Mix (Promega, Milan Andorra, Italy) and 12.5 μg/ml random primers (Promega) for 5 min at 65 °C using a Perkin-Elmer Geneamp PCR system 9600. Subsequently 2 U/μl RnaseOut inhibitor (Invitrogen), 10 U/μl M-MLV virus reverse transcriptase (Promega) and the samples were incubated for 10 min at 25 °C for annealing, 60 min at 37 °C for cDNA synthesis and 15 min at 70 °C for inactivation of enzymes. An AbiPrism 7000 sequence detector system in conjunction with TaqMan® Universal PCR master Mix and TaqMan® Real-Time PCR Assay-on-Demand (Applera) were used for investigating the expression of different genes and the housekeeping gene (beta actin) according to the manufacturer's protocol. TaqMan® primers were supplied by Applied Biosystems and designed to yield products spanning exon–intron boundaries to prevent possible genomic DNA contaminations from total RNA isolation. The primers used were as follows: microtubule-associated 2 (MAP2, Hs01103234_g1), neurofilament heavy polypeptide 200 kDa (NEFh, Hs00606024_m1), microtubule-associated protein tau (MAPT, Hs00902193_m1), nestin (Nes, Hs00707120_s1), synaptophysin (Syp, Hs00300531_m1), Tubulin, beta 3 (TUBB3, Hs00801390_s1), NMDAr (NMDA receptor regulated 2, NARG2, Hs00973298_g1), GABAr (gamma-aminobutyric acid (GABA) A receptor alpha 1, GABRA1, Hs00968132_m1), AMPAr (glutamate receptor ionotropic AMPA 1, GRIA1, Hs00181348_m1) and beta actin (ACTB, Hs99999903_m1). Relative RNA quantification was performed by normalizing the data to the control and to the beta actin content (housekeeping) on the specific day in culture, using the comparative Ct method (Livak and Schmittgen 2001).
Immunocytochemistry
NT2 cultures at 96 DIV of mixed neuronal/glial cell population were fixed for 20 min with 4 % paraformaldehyde in PBS at room temperature. After washing with PBS, the cells were permeabilized for 20 min with 0.1 % Triton X100 followed by a blocking step (1 % bovine serum albumin and 0.1 % Triton X100) for 1 h at room temperature.
Primary antibodies (all from Sigma) diluted in the blocking solution against neurofilament 200 (rabbit, 1:1,000, N4142, Sigma) were applied to the cells over night at 4 °C. Subsequently, the secondary antibody goat anti-rabbit IgG Alexa 488 (1:1000, A11034, Gibco, Invitrogen) was applied. Cell nuclei were stained by 4-6-diamidino-2-phenylindole (DAPI) diluted at 1:5000 purchased from Molecular Probes Europe. Controls for specificity of immunostaining were performed by omitting the primary antibody from the procedure. All stained cultures were examined by fluorescent microscope (Olympus 1X70).
Assessment of cell viability after exposure to methylmercury chloride (MeHgCl) using Alamar Blue Assay
The cell cultures were exposed to MeHgCl (Sigma-Aldrich) from 2–36 DIV during the cell differentiation process in the presence of RA. The medium with fresh portion of 400 nM MeHgCl was replaced twice a week using 350 μM of stock solutions prepared in water. Three biological replicates were performed per each experimental condition (control and toxicant exposure).
The 400 nM concentration of MeHgCl was chosen based on preliminary range-finding experiments, in which wide ranges of concentrations have been tested using the Alamar Blue (AB) (resazurin, Sigma) cell viability assay.
Cell viability was determined every week during the 5 weeks of treatment with MeHgCl, using the AB (resazurin) assay (O'Brien et al. 2000). The blue coloured indicator dye resazurin is reduced into fluorescent resorufin by red-ox reactions in viable cells. Resazurin (10 μl of 100 μM stock) in Hank's Buffered Salt Solution was added directly to the 96-well plates, without removing the medium (100 μl). The plate was incubated for 6 h at 37 °C, 5 % CO2. After the incubation period, the fluorescence of the resazurin metabolite (resorufin) was measured at 530 nm/590 nm (excitation/emission) in a multi-well fluorometric reader (Tecan i-control). Three independent experiments were performed in six replicates; the results were expressed as a percentage of the mean value for the untreated cultures.
miRNAs expression profiling in the control and MeHgCl treated cultures
RNA was extracted using the MIRVANA kit (AMBION) according to manufacturer's instructions. This protocol allows the isolation of total RNA enriched with miRNAs. RNA concentration and quality were determined by Nanodrop and 2100 Agilent Bioanalyzer. The RIN was always above 8.5.
Total RNA was reverse transcribed with Taqman MicroRNA Reverse Transcription Kit using Megaplex™ RT Primers (Applied Biosystems). Real-time PCR reactions were carried out on pre-configured microfluidic cards (Taqman Array MicroRNA Cards, set A,V2.2 , Applied Biosystems) allowing the detection of about specific 380 unique assays and four candidate endogenous control assays. The microfluidics cards were evaluated with Applied Biosystems 7900HT Sequence Detection system. Three biological replicates were performed per every experimental condition.
Stemness markers expression profiling in the control and treated cultures
The expression of a set of genes known as stemness markers of human embryonic stem cells was evaluated using TaqMan® Human Stem Cell Pluripotency microfluidic cards (Applied Biosystems). The expression of 90 well-defined genes validated as markers for pluripotency plus 6 endogenous controls was evaluated after retro-transcription of total RNA with High Capacity cDNA reverse Transcription Kit (Applied Biosystems) by real-time PCR using customized microfluidic cards (Applied Biosystems). The microfluidic cards were analysed with Applied Biosystems 7900HT Sequence Detection system. Each measurement was performed in two independent experiments.
Analysis of microRNA and mRNA expression
Experimental data were then analysed by SDS 2.3 software (Applied Biosystems). The Ct values were exported from SDS 2.3 software (Applied Biosystems) and used as raw data for analysis of qRTPCR data. The R software (Gentleman et al. 2004) and the packages HTqPCR (Dvinge and Bertone 2009) and linear models in micro-array analysis (Limma) (Smyth 2005) were used for the manipulation and analysis of the Ct values.
Measurements with a threshold cycle greater than 35 were discarded. Data normalization was calculated using U6 as an endogenous control for miRNAs and the geometric means of ACTB, CTNNB1, EEF1A1, GAPD, and RAF1.
Statistical significance was assessed using Limma in HTqPCR. Statistical comparisons were generated for differentiated neuronal cultures samples between the treated and control samples. Detectors that showed a fold change greater than 2 or less than 0.5 with a p value smaller than 0.01 were considered as differentially expressed (the p value is not corrected for multiple testing). The miRror algorithm (Friedman et al. 2010) was used to identify the predicted targets for regulated microRNA that were further analysed for identification of pathways enrichment. The analysis was conducted online using Database for Annotation, Visualization and Integrated Discovery (DAVID) (Huang et al. 2009); the indicated p value represents the EASE Score, a modified Fisher Exact p value, for gene-enrichment analysis.
Statistical analysis
GraphPad Prism 4.0 (GraphPad software, San Diego, CA) was used for statistical analyses. The data are the means of four independent experiments performed in six replicates (cell viability assay) and duplicates (real-time PCR analysis) (mean ± SD). One-way ANOVA was performed to assess differences between treated and control culture in the AB assay. For the statistical analysis of the real-time PCR experiments, differences between 43 against 78 DIV studied were assessed by two-way ANOVA. Statistical significance was indicated as follows *p < 0.05, **p < 0.01 and ***p < 0.001.
Results
Characterization of NT2 cells differentiation
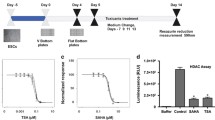
Initially, the single cells showed a granular appearance (Fig. 1a) and after 35 DIV in the presence of RA, they formed a very dense multilayered culture with some aggregates on the top, occasionally connected by processes, resembling neuronal culture (Fig. 1b). At this time point, the high cell density did not allow a good identification of neuronal and glial cell morphology. After RA treatment, the neuronal differentiation protocol was continued up to 96 DIV (Fig. 1c) to verify if neurons reached morphological maturation as expected (Laurenza et al. 2013). The differentiated NT2 culture at 96 DIV showed large cell aggregates, linked to each other by thick bundles of neurites creating a dense neuronal network, which positively stained for Neurofilament 200 (NF-200) (Fig. 1c). Recently, the neuronal and glial differentiation of NT2 neural precursors has already been characterized in detail by our group (Laurenza et al. 2013). Here, we performed gene expression studies by semi-quantitative RT-PCR at 43 and 78 DIV to make sure that the process of neuronal differentiation initially triggered by RA was progressing as expected. Indeed, the increasing expression of neuronal specific markers such as MAP2 (dendrite), NF-200 (cytoskeleton), Tubulin βIII (cytoskeleton), MAPT-tau (axon) and synaptothysin (protein of synaptic vesicles) was observed (Fig. 1d). The neuronal maturation was also confirmed by the expression of both excitatory (NMDA, AMPA) and inhibitory (GABA) receptors (Fig. 1e). The glial cells were present as well since GFAP mRNA expression was confirmed by RT-PCR analysis and immunostaining (data not shown) suggesting that a mixed neuronal/glial culture was obtained, confirming the results obtained in our previous studies (Laurenza et al. 2013).
Representative pictures of NT2 cell culture morphology during the neuronal differentiation process. Cells after 24 h of seeding (a) were exposed to 10 μM retinoic acid up to 36 DIV (b) to induce neural differentiation. The process of neuronal differentiation was followed up to 96 DIV. At this time, groups of neurons were linked by the dense network of neurites, positively stained against NF-200 (green) with cell nuclei co-stained by DAPI (c). White bars correspond to 500 μm. Evaluation of the mRNA expression of neuronal markers (d) and receptors (AMPA, GABA and NMDA) (e) at 43 and 78 DIV of cell differentiation by real-time RT-PCR. Gene expression levels in (d) and (e) were normalized to the housekeeping gene (actin) and the mRNA expression at 43 DIV. Data are presented as mean ± S.E.M. of three independent experiments performed in duplicate
Based on these and our previous results (Laurenza et al. 2013), we can conclude that human carcinoma, pluripotent NT2 stem cell-derived model is a suitable test system for studying developmental neurotoxicity as the key developmental processes can be followed up including the neural precursor cells commitment towards advanced stages of neuronal and glial differentiation and maturation.
Changes in the miRNAs expression during the process of neuronal differentiation
According to the published data, miRNAs control spatial and temporal gene expression to allow proper patterning and neuronal differentiation (Cochella and Hobert 2012). In order to determine whether our in vitro test system could mimic the expected changes in miRNAs expression, we particularly analysed those miRNAs that are known markers of pluripotency (stemness) and neuronal differentiation. The variations in the expression of 381 miRNAs were measured in the undifferentiated culture and after the exposure to RA (2–36 DIV) by using microfluidic cards “Taqman Array MicroRNA Cards” (Applied Biosystems). A heat map shows 243 miRNAs differentially expressed with a log2 FC (fold of change) value higher than 1 or lower than −1 (Fig. 2a) with p value <0.01. In the undifferentiated cells, we have identified statistically significant overexpression of several miRNAs belonging to the miRNA-302 cluster (p < 0.001), known to be implicated in the pluripotency maintenance (Barroso-del Jesus et al. 2008). The expression levels of these miRNAs was 8.5- (miRNA-302a), 11.0- (miRNA-302b), 8.7- (miRNA-302c) and 11.7- (miRNA-367) fold higher in undifferentiated cells when compared with the differentiated cells (p < 0.001) (Fig. 2b).
Heat map representing changes in miRNA expression in undifferentiated (1 DIV) and differentiated NT2 cells triggered by 10 μM retinoic acid (36 DIV) (a). Each row represents a miRNA in undifferentiated or differentiated culture (three independent experiments for each culture) and the changes in the levels are represented by red colour for higher expression and green for low expression level. Quantification of the expression levels of some of the most significantly regulated (up or down) miRNAs during the process of neural differentiation in NT2 cells (b). Data are expressed in log2 FC (fold change), comparing differentiated (36 DIV) to undifferentiated cell samples (1 DIV). miRNAs specific for stemness were downregulated during the differentiation process (negative fold change) but miRNAs specific for neuronal differentiation showed an increase of their expression in the differentiated cells (positive fold change). Representation of miRNA expression relevant to neuronal differentiation at 36 DIV was measured by Ct value (c) as their expression levels in undifferentiated cells were too low to be quantified. ***p < 0.001
In the presence of RA, the process of neuronal differentiation was triggered and several miRNAs known to be involved in neuronal development showed a significant increase in expression. In particular, we observed higher expression (on average 8.2-fold change) of some members of let-7 family (let-7e, let7-g) (Fig. 2b) which is known to be involved in the processes of neural lineage differentiation (p < 0.001) (Kawahara et al. 2012). Similarly, the expression of miR-125b (a member of miRNA-125 family) that is involved in mammalian neuronal development (Sempere et al. 2004) was 5.1-fold higher in the differentiated cells and miR-132, involved in dendrite outgrowth (Vo et al. 2005), increased 3.1-fold (p < 0.001) (Fig. 2b). The expression of miR-10a and b and of other let-7 family members (let-7a, b, c, d, and f) increased upon differentiation, in agreement with previous studies (Christensen and Schratt 2009; Parsons 2012), but it was very low (or undetectable) in the undifferentiated cells (Fig 2c).
Evaluation of miRNAs expression after exposure to MeHgCl
MicroRNA expression is implicated not only in maintaining normal neuronal function and homeostasis but is also highly deregulated when neurotoxicity is triggered (Kaur et al. 2012). The NT2 cell culture was exposed to 400 nM MeHgCl (2–36 DIV) that was non-cytotoxic during the first 2 weeks of treatment, however it led to slight cytotoxicity with the prolonged treatment, producing a level of toxicity of 28 % (±8 % SD) after 5 weeks of exposure (Fig. 3a).
Evaluation of cytotoxicity induced by different concentrations of MeHgCl during the process of neuronal differentiation (2–36 DIV) measured each week using Alamar Blue assay (a). Data are presented as mean ± S.D. of three independent experiments performed in six replicates compared to the control sample (untreated). Heat map representing changes in miRNA expression (red and green colour for high and low expression levels, respectively) after treatment with 400 nM MeHgCl (tox) and compared to the control (diff) at 36 DIV (b) followed by their quantification (c) that is expressed as log2 FC (fold change). The analysis is based on three independent experiments for each condition. *p < 0.05, **p < 0.01, and ***p < 0.001
At 36 DIV, MeHgCl affected the expression of five miRNAs that displayed consistent (p < 0.01) and significant (log2 fold change >1 or < −1) up- or down- regulation (Fig. 3b). Three of these, miR-302b, miR-367 and miR-372, are known to be involved in maintaining the pluripotent phenotype of stem cells (Suh et al. 2004). The miR-302b and miR-367 that belong to the same miRNA cluster, were overexpressed 5.3- and 6.1-fold change, respectively (p < 0.005 and p < 0.007) and miR-372 was increased by 9.5-fold (p < 0.007) (Fig. 3c). The expression of the miR-141 and the miR-196b was 2.4- and 3.2-fold higher in the treated cells, respectively, compared to the control cultures (p < 0.01 and p < 0.001) (Fig. 3c). The miRNA-141 belongs to the miR-200 family, a group of miRNAs known to be involved in oxidative stress response (Magenta et al. 2011; Mateescu et al. 2011) and cell differentiation (Braceen et al. 2008). The changes observed in miRNA-141 expression could be linked to the well known mechanism of MeHgCl-induced toxicity (Ceccatelli et al. 2010). miR-196b is known to regulate the antero-posterior axis formation during development (Amiel et al. 2012). Based on in vivo experiments miR-196b inhibits translation of HOX target genes and indirectly regulates the RA signalling pathway involved in HOX gene expression (He et al. 2011).
Furthermore, we studied the involvement of the miRNAs that we found deregulated during neural differentiation or after MeHgCl exposure by term enrichment analysis to identify their possible targets using the Kyoto Encyclopedia of Genes and Genomes (KEGG) annotation (www.genome.jp/kegg/pathway.html) which provides functional evaluation of deregulated genes. KEGG (Kyoto Encyclopedia of Genes and Genomes) PATHWAY database is a collection of manually drawn pathway maps representing molecular interaction and reaction networks (Kanehisa et al. 2010). Targets of the miRNAs involved in neuronal differentiation process of NT2 cells (Fig. 2b) presented a significant enrichment in the mTOR signalling pathway (enrichment value 2.17 %, p value < 0.05) (Fig. 4). Based on this analysis, we identified the targets that belong to this pathway such as phosphoinositide-3-kinase (PIK3R3), ribosomal protein S6 kinase (RPS6KA3) and tuberous sclerosis 1 (TSC1).
Representation of mTOR signalling pathway showing the genes (green boxes) identified as possible targets of miRNAs which expression changed (up- or downregulated) during the differentiation process of the control culture (KEGG database: http://www.genome.jp/kegg/kegg2.html) (p value <0.05)
In the human NT2 cells exposed to MeHgCl the pathway enrichment tool indicated that four relevant molecules (Netrin, Eph, CXCL12, Cofilin) acting in the axon guidance pathway (enrichment value 2.98 %, p value < 0.02) could be possible targets of the miRNAs which were overexpressed after exposure to MeHgCl (Fig. 5) suggesting that the processes of synaptogenesis and neuronal networking could be affected.
Representation of the axon guidance pathway showing the genes downregulated (yellow boxes) since they were identified as possible targets of miRNAs which expression was up regulated in the culture exposed to 400 nM MeHgCl during 2–36 DIV (KEGG database: http://www.genome.jp/kegg/kegg2.html) (p value <0.02)
Evaluation of mRNA expression of stemness and neuronal differentiation genes after MeHgCl treatment
After exposure to MeHgCl (400 nM) during neuronal differentiation, mRNA expression of 91 different genes was analysed using the “TaqMan Human Stem Cell pluripotency microfluidic card” that contained a well-defined set of validated gene expression markers used in order to characterize human embryonic stem (ES) cell identity.
The results showed a significant upregulation of 6 stemness markers (Fig. 6b) and 2 genes correlated to the neural differentiation process, in the cells treated with MeHgCl compared to the control cells (Fig. 6a). The transcription factor Nanog that is critically involved in self-renewal processes of undifferentiated embryonic stem cells (Pan and Thomson 2007) was upregulated by 16.4-fold (Fig. 6b). Additionally, it seems to be linked to the transcriptional induction of the embryonic stem cell-specific miR-302–367 cluster. In fact, the miRNA-302 cluster promoter presents a sequence of potential binding sites for Nanog (Barroso-delJesus et al. 2008).
Quantification of the mRNA expression levels of cell differentiation markers (a) and stemness markers (b) in NT2 cells exposed to MeHgCl (400 nM) during 2–36 DIV in comparison to control culture (untreated). Each measurement was performed in two independent experiments (p value ≤ 0.05) and the results are expressed as log2 FC (fold change)
Additionally, Nodal, known as transforming factor β, also showed an overexpression of 3.6-fold in the treated cells (Fig. 6b). This factor is involved in brain development, inhibition of neuroepithelial cell differentiation, stem cell maintenance and in the determination of lateral mesoderm left/right symmetry (http://www.ebi.ac.uk/QuickGO). Its signalling is important for cell fate determination during very early stages of development (Dougan et al. 2003).
Another gene overexpressed after the exposure to MeHgCl (4.4-fold change) was the transcription factor CP2. It is present in many gene ontology (GO) terms associated with development and it plays an important role in the epithelial–mesenchymal transition (EMT) (Rodda et al. 2001). The germ cell nuclear factor (NR6A1), the liver receptor homolog-1 (NR5A2) and the interferon-induced transmembrane protein 1 (IFITM1) were all overexpressed in the treated cells (2.4-, 2.1- and 2.2-fold, respectively). These three genes are also involved in embryo development (http://www.ebi.ac.uk/QuickGO) and NR6A1 is involved in neurogenesis and germ cell development (Lei et al. 1997). At the same time, after the treatment, we observed the upregulation of only two markers specific for neuronal/glial differentiation: eomesodermin (EOMES; 2.4 folds) and the glial cell missing1 protein (GCM1, 2.4-fold of change). Both of them are DNA-binding proteins involved in glial and neuronal development (http://www.ebi.ac.uk/QuickGO; Chotard et al. 2005).
Gene ontology term enrichment of miRNAs molecular targets regulated by RA-triggered neuronal differentiation or by exposure to MeHgCl
To identify potential roles of miRNAs in the cell differentiation process and in response to the toxic compound exposure, we focused on the most significantly regulated miRNAs identified previously (Figs. 2a and 3b). Since the changes in the expression of these miRNAs could affect the regulation of their target genes, we used a combinatorial approach to identify the potential target genes of this set of miRNAs. Using the miRror algorithm which integrates data from a dozen miRNA target prediction programs, several genes were predicted as the possible targets of the regulated miRNAs. As an outcome, 108 total targets of neuronal differentiation process-related miRNAs and 133 targets of MetHgCl exposure-related miRNAs were identified.
The predicted gene targets were loaded in the DAVID database for GO term enrichment analysis to detect functional pathways alterations. We focused on the most relevant GO terms associated with the list of deregulated miRNAs predicted gene targets and correlated with the biological functions such as apoptosis, cell cycle regulation and specific neuronal functions. Many of these GO terms were enriched following the analysis, highlighting a specific functional profile.
The neuronal differentiation triggered by RA (Table 1) seemed to affect the expression of miRNAs implicated in the control of RNA biosynthesis processes, protein complex biosynthesis, regulation of transcription from RNA, metabolic pathways (GO:0051254 ∼ positive regulation of RNA metabolic process, p value 0.00; GO:0051173 ∼ positive regulation of nitrogen compound metabolic process, p value 0.01; GO:0010604 ∼ positive regulation of macromolecule metabolic process p value 0.02), early neural specific differentiation processes, such as cell migration (GO:0016477, p value 0.02), cell location (GO:0016477, p value 0.03), myelination (GO:0042552, p value 0.04), axon ensheathment (GO:0042552, p value 0.04) and also later neuronal system functions relevant to cerebral cortex (GO:0021987, p value 0.03) and diencephalon development (GO:0021536, p value 0.03) (Table 1).
On the other hand, the treatment with MeHgCl induced the deregulation of miRNAs mostly implicated in signalling pathways involved in cell maturation (GO:0048469, p value 0.04), phosphorylation (GO:0016310, p value 0.005); programmed cell death pathways: induction of apoptosis (e.g. GO:0006917, p value 0.01), regulation of apoptosis (GO:0042981, p value 0.03) and some neuronal specific processes such as synaptic vesicles transport (GO:0048489, p value 0.02), neuronal development (GO:0048666, p value 0.05), cell migration (GO:0016477, p value 0.05), sodium ion transport (GO: 0006814, p value 0.01), vesicle-mediated transport (GO:0016192, p value 0.02) and processes linked to learning and memory activity (GO:0007611, p value 0.02) (Table 2). Interestingly, ubiquitin-dependent proteasome pathway could be one of the MeHgCl targets (GO:0050801, p value 0.01) involved in toxicity since it was significantly affected.
Discussion
MicroRNAs affect many steps required for the development of the nervous system, from neuronal differentiation and patterning to plasticity (Kosik 2006; Feng and Feng 2011). In this study, we characterized miRNA variations in human carcinoma pluripotent stem cell-derived neuronal cell culture (NT2 cell line) to determine whether they are specific for neuronal differentiation. NT2 cell line is a convenient and robust model for studying DNT, because key processes such as the commitment of human neural stem cells to the neuronal lineage and their subsequent differentiation into neuronal and glial-like cells are taking place. Indeed, upon exposure to RA (36 DIV) several miRNAs known to be involved in the process of neuronal differentiation such as miRNA-132 (Vo et al. 2005; Wanet et al. 2012), miRNA-125 family (Sempere et al. 2004) and the let-7 family of miRNAs (Abbott et al. 2005; Pasquinelli and Ruvkun 2002), were upregulated. miRNA-132 is induced by synaptic activity and involved in dendritic branching and spine density in in vitro models (Vo et al. 2005) suggesting that cells reached an advanced stage of differentiation. Let-7 family cluster is known to be involved in the processes of neural lineage differentiation and stem cell commitment, both in embryonal stem cells and embryocarcinoma cells, resembling the expression of the other brain-enriched miRNAs (Rybak et al. 2008; Wulczyn et al. 2007). These results suggest that neural precursors of NT2 cell line responded to RA-triggered neuronal differentiation, confirmed further by the expression of neuronal specific markers studied at the mRNA levels (Fig. 1d and e) and cell morphology immunostaining (Fig. 1c). On the other hand, as expected, members of the miR-302 cluster which expression is important for pluripotency maintenance were downregulated in differentiated neurons. Similar results were obtained in the studies of our laboratory where the miRNAs expression was determined in H9 cell line derived from human embryonic stem cell and differentiated towards neuronal phenotype (Nerini-Molteni et al. 2012). Further bioinformatics analysis using DAVID database for GO term enrichment detected important functional pathways alterations taking place during the process of neuronal differentiation. miRNAs involved in the control of RNA and protein biosynthesis and metabolic processes as well as in neuronal specific developmental processes such as myelination, escheatment of axons, cell proliferation cell migration (Table 1) were identified. The deregulated miRNAs presented a particular enrichment in the mTOR signalling pathway which includes targets such as phosphoinositide-3-kinase PIK3R3, ribosomal protein S6 kinase RPS6KA3 and tuberous sclerosis 1 (TSC1). These gene transcripts are targets of many miRNAs that are regulated during cell differentiation and maturation. Interestingly all of them are predicted targets of the miRNA-302 cluster, which is downregulated (Fig. 2b) during the differentiation process. Moreover, mTOR signalling is a key pathway for initiating neuronal differentiation as it facilitates coordination of cell cycle and differentiation program (Fishwick et al. 2010). These results suggest that miRNA expression could serve as reliable descriptor of the neuronal differentiation process using human NT2 cell line since, as expected, in the presence of RA the overexpression of miRNAs specific for different stages of neuronal development was observed. A similar approach, based on profiling of miRNA expression for characterization of the RA-induced neuronal differentiation has been also applied for neuroblastoma-derived neuronal models (Stallings et al. 2011).
Furthermore, we have evaluated whether miRNAs could function as biomarkers of response to cellular perturbations as they are supposed to regulate numerous targets of cell stress response induced by exposure to a toxicant. Such an approach for toxicity evaluation is in line with the recently proposed large-scale shift in toxicity testing (NRC 2007) that should be focused on the mechanistic in vitro assays using human models. In this context, we used miRNA profiling as a tool to evaluate whether it could provide information on cellular pathways activation triggered by the repeated exposure to MeHgCl. Significant variations were found in five miRNAs (miRNA-141, 196b, 367, 302b and 372), although less significant changes occurred in a larger number of genes. By using term enrichment analysis, we investigated possible clustering of the target genes of these miRNAs under relevant functions. Taking into consideration the experimentally validated targets present in miRror database, we identified 108 “hits” as possible target genes of the deregulated miRNAs. Further analysis based on GO term enrichment and DAVID functional clustering indicated that MeHgCl exposure deregulated miRNAs targets implicated in apoptosis and neuronal developmental processes such as synaptic vesicle transport, cell migration and differentiation as well as biological processes linked to learning and memory (Table 2). These biological functions, identified as possible targets of MeHgCl, are also described in in vivo studies (Fox et al. 2012). Indeed, the relatively low concentrations of methyl mercury could result in cognitive and motor dysfunctions (Dolbec and Mergler 2000; Grandjean et al. 1997; Lebel et al. 1998). In fact, the threshold of methyl mercury concentrations in the brain resulting in the clinical effects has been suggested to be as low as 0.3 ppm (1.5 μM) (Burbacher et al 1990).
It is worth emphasizing that MeHgCl effects on learning and memory processes, well known in epidemiological studies (Yokoo et al. 2003), were also identified in this in vitro study. Moreover, this suggestion is further backed up by the observed overexpression of miRNAs involved in the regulation of the molecules such as netrin, Eph, CXCL12 and cofilin that are involved in axon guidance pathway suggesting that methyl mercury could inhibit the axonal growth cone, resulting in the decreased neuronal networking, possibly linked to damaged process of learning and memory (Fox et al 2012). This observation is in agreement with the well-established view that methyl mercury inhibits neurite outgrowth based on both in vivo (Rand 2010) and in vitro studies using various models such as human neural precursor-based model (Stiegler et al. 2011; Krug et al. 2013), primary cultures (Harrill et al. 2011) or cell lines (Radio and Mundy 2008). The obtained data suggest that in vitro miRNA profiling might be a useful tool for possible DNT biomarker identification, however, the interpretation of the obtained results has to be confirmed by further studies where target genes of deregulated miRNAs will be defined by studying their mRNA expression.
Furthermore, miRNAs that regulate the process of ubiquitin-dependent protein degradation were also strongly upregulated (p < 0.01), indicating that MeHgCl could cause the deficiency in clearance of cellular by-products. The same pathway of toxicity, induced by MeHgCl was also identified in similar studies of our laboratory using H9 cell line (Nerini-Molteni et al. 2012).
miRNA expression profiling in human pluripotent stem cell-derived neuronal models could be a promising approach towards system-biology-based predictive human toxicology. The identification of miRNAs and their targets could be useful as first screening tool to identify affected cellular pathways that might be further studied by different endpoints. Although further studies are needed with larger amount of tested chemicals using different cell models, our results suggest that miRNA expression analysis could be a powerful tool, providing a different level of information on pathway activation or perturbation, leading to the improved predictive toxicology.
Conclusions
In conclusion, the obtained data suggest that the applied in vitro neuronal model derived from human carcinoma stem cells (NT2 cell line) and miRNA profiling is a relevant approach for in vitro neurodevelopmental toxicity testing. Profiling of miRNAs and identification of their functional targets after exposure to a toxicant allow to apply a more mechanistic approach for toxicity evaluation, leading to the identification of the potential biomarkers of neurotoxicity. Although further investigations are needed, our results suggest that miRNA profiling is a reliable tool in pathway toxicity analysis and could improve predictive human toxicology, especially when based on human in vitro models.
Abbreviations
- AATK:
-
Apoptosis-associated tyrosine kinase
- AB assay:
-
Alamar blue assay
- CNS:
-
Central nervous system
- DAVID:
-
Database for Annotation Visualization and Integrated Discovery
- DIV:
-
Days in vitro
- DNT:
-
Developmental neurotoxicity
- EMT:
-
Epithelial–mesenchymal transition
- ES:
-
Cell embryonic stem cell
- GO term:
-
Gene ontology term
- hESC:
-
Human embryonic stem cell
- MeHgCl:
-
Methyl mercury chloride
- miRNA:
-
MicroRNA
- NT2 cell line:
-
NTERA-2 cell line
- RA:
-
Retinoic acid
- UTRs:
-
Untranslated regions
References
Abbott AL, Alvarez-Saavedra E, Miska EA, Lau NC, Bartel DP, Horvitz HR, et al. The let-7 microRNA family members mir-48, mir-84, and mir-241 function together to regulate developmental timing in Caenorhabditis elegans. Dev Cell. 2005;9:403–14.
Amiel J, de Pontual L, Henrion-Caude A. miRNA, development and disease. Adv Genet. 2012;80:1–36.
Barco A, Alarcon JM, Kandel ER. Expression of constitutively active CREB protein facilitates the late phase of long-term potentiation by enhancing synaptic capture. Cell. 2002;108(5):689–703.
Barroso-del Jesus A, Romero-López C, Lucena-Aguilar G, Melen GJ, Sanchez L, Ligero G, et al. Embryonic stem cell-specific miR302-367 cluster: human gene structure and functional characterization of its core promoter. Mol Cell Biol. 2008;28(21):6609–19.
Braceen CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF, et al. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res. 2008;68(19):7846–54.
Burbacher TM, Rodier PM, Weiss B. Methylmercury developmental neurotoxicity: a comparison of effects in humans and animals. Neurotoxicol. Teratol. 1990;12:191–202.
Buzanska L, Sypecka J, Nerini-Molteni S, Compagnoni A, Hogberg HT, del Torchio R, et al. A human stem cell-based model for identifying adverse effects of organic and inorganic chemicals on the developing nervous system. Stem Cells. 2009;27(10):2591–601.
Cao X, Pfaff SL, Gage FH. A functional study of miR-124 in the developing neural tube. Genes Dev. 2007;21(5):531–6.
Ceccatelli S, Daré E, Moors M. Methylmercury-induced neurotoxicity and apoptosis. Chem Biol Interact. 2010;188(2):301–8.
Chotard C, Leung W, Salecker I. Glial cells missing and gcm2 cell autonomously regulate both glial and neuronal development in the visual system of Drosophila. Neuron. 2005;48(2):237–51.
Christensen M, Schratt GM. microRNA involvement in developmental and functional aspects of the nervous system and in neurological diseases. Neurosci Lett. 2009;466(2):55–62.
Claudio L, Kwa WC, Russell AL, Wallinga D. Testing methods for developmental neurotoxicity of environmental chemicals. Toxicol Appl Pharmacol. 2000;164:1–14.
Cochella L, Hobert O. Embryonic priming of a miRNA locus predetermines postmitotic neuronal left/right asymmetry in C. elegans. Cell. 2012;151(6):1229–42.
De Pietri Tonelli D, Pulvers JN, Haffner C, Murchison EP, Hannon GJ, Huttner WB. miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development. 2008;135(23):3911–21.
Dolbec J, Mergler D, Sousa Passos CJ, Sousa dM, Lebel J. Methylmercury exposure affects motor performance of a riverine population of the Tapajos River, Brazilian Amazon. Int Arch Occup Environ Health. 2000;73:195–203.
Dougan ST, Warga RM, Kane DA, Schier AF, Talbot WS. The role of the zebrafish nodal-related genes squint and cyclops in patterning of mesendoderm. Development. 2003;130(9):1837–51.
Dvinge H, Bertone P. HTqPCR: high-throughput analysis and visualization of quantitative real-time PCR data in R. Bioinformatics. 2009;25:3325–6.
Eriksson P. Developmental neurotoxicity of environmental agents in the neonate. Neurotoxicology. 1997;18:719–26.
Feng W, Feng Y. MicroRNAs in neural cell development and brain diseases. Sci China Life Sci. 2011;54(12):1103–12.
Fishwick KJ, Li RA, Halley P, Deng P, Storey KG. Initiation of neuronal differentiation requires PI3-kinase/TOR signalling in the vertebrate neural tube. Dev Biol. 2010;338(2):215–25.
Fox DA, Grandjean P, de Groot D, Paule MG. Developmental origins of adult diseases and neurotoxicity: epidemiological and experimental studies. Neurotoxicology. 2012;33(4):810–6.
Friedman Y, Naamati G, Linial M. MiRror: a combinatorial analysis web tool for ensembles of microRNAs and their targets. Bioinformatics. 2010;26(15):1920–1.
Gao FB. Context-dependent functions of specific microRNAs in neuronal development. Neural Dev. 2010;5:25. doi:10.1186/1749-8104-5-25.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5(10):R80.
Grandjean P, Landrigan PJ. Developmental neurotoxicity of industrial chemicals. Lancet. 2006;368(9553):2167–78.
Grandjean P, Weihe P, White RF, Debes F, Araki S, Yokoyama K, et al. Cognitive deficit in 7-year-old children with prenatal exposure to methylmercury. Neurotoxicol Teratol. 1997;19:417–28.
Harrill JA, Freudenrich TM, Robinette BL, Mundy WR. Comparative sensitivity of human and rat neural cultures to chemical-induced inhibition of neurite outgrowth. Toxicol Appl Pharmacol. 2011;256(3):268–80.
He X, Yan YL, Eberhart JK, Herpin A, Wagner TU, Schartl M, et al. miR-196 regulates axial patterning and pectoral appendage initiation. Dev Biol. 2011;357(2):463–77.
Hogberg HT, Kinsner-Ovaskainen A, Hartung T, Coecke S, Bal-Price AK. Gene expression as a sensitive endpoint to evaluate cell differentiation and maturation of the developing central nervous system in primary cultures of rat cerebellar granule cells (CGCs) exposed to pesticides. Toxicol Appl Pharmacol. 2009;235:268–86.
Hogberg HT, Kinsner-Ovaskainen A, Coecke S, Hartung T, Bal-Price AK. mRNA expression is a relevant tool to identify developmental neurotoxicants using an in vitro approach. Toxicol Sci. 2010;113:95–115.
Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Kanehisa M, Goto S, Furumichi M, Tanabe M, Hirakawa M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 2010;38(Database issue):D355–60.
Kaur P, Armugam A, Jeyaseelan K. MicroRNAs in Neurotoxicity. J Toxicol. 2012; 870150
Kawahara H, Imai T, Okano H. MicroRNAs in neural stem cells and neurogenesis. Front Neurosci. 2012;6:30.
Kim J, Inoue K, Ishii J, Vanti WB, Voronov SV, Murchison E, et al. A microRNA feedback circuit in midbrain dopamine neurons. Science. 2007;317(5842):1220–4.
Konopka W, Schütz G, Kaczmarek L. The microRNA contribution to learning and memory. Neuroscientist. 2011;17(5):468–74.
Kosik KS. Neuroscience gears up for duel on the issue of brain versus deity. Nature. 2006;439(7073):138.
Krek A, Grün D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, et al. Combinatorial microRNA target predictions. Nat Genet. 2005;37(5):495–500.
Krichevsky AM, King KS, Donahue CP, Khrapko K, Kosik KS. A microRNA array reveals extensive regulation of microRNAs during brain development. RNA. 2003;9(10):1274–81.
Krug AK, Balmer NV, Matt F, Schönenberger F, Merhof D, Leist M. Evaluation of a human neurite growth assay as specific screen for developmental neurotoxicants. Arch Toxicol. 2013; May 14.
Laurenza I, Pallocca G, Mennecozzi M, Scelfo B, Pamies D, Bal-Price A. A human pluripotent carcinoma stem cell-based model for in vitro developmental neurotoxicity testing: effects of methylmercury, lead and aluminum evaluated by gene expression studies. Int J Dev Neurosci. 2013;S0736–5748(13):00038–5.
Lebel J, Mergler D, Branches F, Lucotte M, Amorim M, Larribe F, et al. Neurotoxic effects of low-level methylmercury contamination in the Amazonian Basin. Environ Res. 1998;79:20–32.
Lei W, Hirose T, Zhang LX, Adachi H, Spinella MJ, Dmitrovsky E, et al. Cloning of the human orphan receptor germ cell nuclear factor/retinoid receptor-related testis-associated receptor and its differential regulation during embryonal carcinoma cell differentiation. J Mol Endocrinol. 1997;18(2):167–76.
Leucht C, Stigloher C, Wizenmann A, Klafke R, Folchert A, Bally-Cuif L. MicroRNA-9 directs late organizer activity of the midbrain–hindbrain boundary. Nat Neurosci. 2008;11(6):641–8.
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature. 2005;433(7027):769–73.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Magenta A, Cencioni C, Fasanaro P, Zaccagnini G, Greco S, Sarra-Ferraris G, et al. miR-200c is upregulated by oxidative stress and induces endothelial cell apoptosis and senescence via ZEB1 inhibition. Cell Death Differ. 2011;18(10):1628–39.
Makeyev EV, Zhang J, Carrasco MA, Maniatis T. The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol Cell. 2007;27(3):435–48.
Maller Schulman BR, Liang X, Stahlhut C, DelConte C, Stefani G, Slack FJ. The let-7 microRNA target gene, Mlin41/Trim71 is required for mouse embryonic survival and neural tube closure. Cell Cycle. 2008;7(24):3935–42.
Mateescu B, Batista L, Cardon M, Gruosso T, de Feraudy Y, Mariani O, et al. miR-141 and miR-200a act on ovarian tumorigenesis by controlling oxidative stress response. Nat Med. 2011;17(12):1627–35.
Miska EA, Alvarez-Saavedra E, Townsend M, Yoshii A, Sestan N, Rakic P, et al. Microarray analysis of microRNA expression in the developing mammalian brain. Genome Biol. 2004;5(9):R68.
Nerini-Molteni S, Mennecozzi M, Fabbri M, Sacco MG, Vojnits K, Compagnoni A, et al. MicroRNA profiling as a tool for pathway analysis in a human in vitro model for neural development. Curr Med Chem. 2012;19(36):6214–23.
NRC. Toxicity testing in the 21st century: a vision and a strategy. Washington, D.C.: The National Academies Press; 2007.
O'Brien J, Wilson I, Orton T, Pognan F. Investigation of the Alamar Blue (resazurin) fluorescent dye for the assessment of mammalian cell cytotoxicity. Eur J Biochem. 2000;267:5421–6.
OECD (2007). Test Guideline 426. OECD Guideline for Testing of Chemicals. Developmental Neurotoxicity Study. Available: http://www.oecd.org/document/55/0,3343,en_2649_34377_2349687_1_1_1_1,00.html
Pan G, Thomson JA. Nanog and transcriptional networks in embryonic stem cell pluripotency. Cell Res. 2007;17(1):42–9.
Parsons XH. MicroRNA profiling reveals distinct mechanisms governing cardiac and neural lineage-specification of pluripotent human embryonic stem cells. J Stem Cell Res Ther. 2012;13:2(3).
Pasquinelli AE, Ruvkun G. Control of developmental timing by microRNAs and their targets. Annu Rev Cell Dev Biol. 2002;18:495–513.
Pleasure SJ, Page C, Lee VM. Pure, postmitotic, polarized human neurons derived from NTera 2 cells provide a system for expressing exogenous proteins in terminally differentiated neurons. J Neurosci. 1992;12:1802–15.
Radio NM, Mundy WR. Developmental neurotoxicity testing in vitro: models for assessing chemical effects on neurite outgrowth. Neurotoxicology. 2008;29(3):361–76.
Rand MD. Drosophotoxicology: the growing potential for Drosophila in neurotoxicology. Neurotoxicol Teratol. 2010;32(1):74–83.
Rodda S, Sharma S, Scherer M, Chapman G, Rathjen P. CRTR-1, a developmentally regulated transcriptional repressor related to the CP2 family of transcription factors. J Biol Chem. 2001;276(5):3324–32.
Rodier PM. Developing brain as a target of toxicity. Environ Health Perspect. 1995;103:73–6.
Roush S, Slack FJ. The let-7 family of microRNAs. Trends Cell Biol. 2008;18(10):505–16.
Rybak A, Fuchs H, Smirnova L, Brandt C, Pohl EE, Nitsch R, et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nat Cell Biol. 2008;10(8):987–93.
Schaefer A, O'Carroll D, Tan CL, Hillman D, Sugimori M, Llinas R, et al. Cerebellar neurodegeneration in the absence of microRNAs. J Exp Med. 2007;204(7):1553–8.
Schratt GM, Tuebing F, Nigh EA, Kane CG, Sabatini ME, Kiebler M, et al. A brain-specific microRNA regulates dendritic spine development. Nature. 2006;439(7074):283–9.
Sempere LF, Freemantle S, Pitha-Rowe I, Moss E, Dmitrovsky E, Ambros V. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 2004;5(3):R13.
Smyth GK. Limma: linear models for microarray data. In: Bioinformatics and computational biology solutions using R and bioconductor. Springer: New York; 2005. p. 397–420.
Stallings RL, Foley NH, Bray IM, Das S, Buckley PG. MicroRNA and DNA methylation alterations mediating retinoic acid induced neuroblastoma cell differentiation. Semin Cancer Biol. 2011;21(4):283–90.
Stiegler NV, Krug AK, Matt F, Leist M. Assessment of chemical-induced impairment of human neurite outgrowth by multiparametric live cell imaging in high-density cultures. Toxicol Sci. 2011;121(1):73–87.
Stummann TC, Hareng L, Bremer S. Hazard assessment of methylmercury toxicity to neuronal induction in embryogenesis using human embryonic stem cells. Toxicology. 2009;257(3):117–26.
Suh MR, Lee Y, Kim JY, Kim SK, Moon SH, Lee JY, et al. Human embryonic stem cells express a unique set of microRNAs. Dev Biol. 2004;270(2):488–98.
Sutton MA, Schuman EM. Dendritic protein synthesis, synaptic plasticity, and memory. Cell. 2006;127(1):49–58.
Tilson HA. Neurotoxicology risk assessment guidelines: developmental neurotoxicology. Neurotoxicology. 2000;21:189–94.
U.S. EPA (U.S. Environmental Protection Agency), 1998. U.S. Environmental Protection Agency Health Effects Test Guidelines. OPPTS 870.6300. Developmental Neurotoxicity Study. U.S. EPA 712-C-98-239. Available:http://www.epa.gov/opptsfrs/publications/OPPTS_Harmonized/870_Health_Effects_Test_Guidelines/Series/870-6300.pdf
Visvanathan J, Lee S, Lee B, Lee JW, Lee SK. The microRNA miR-124 antagonizes the antineural REST/SCP1 pathway during embryonic CNS development. Genes Dev. 2007;21(7):744–9.
Vo N, Klein ME, Varlamova O, Keller DM, Yamamoto T, Goodman RH, et al. A cAMP-response element binding protein-induced microRNA regulates neuronal morphogenesis. Proc Natl Acad Sci U S A. 2005;102(45):16426–31.
Wanet A, Tacheny A, Arnould T, Renard P. miR-212/132 expression and functions: within and beyond the neuronal compartment. Nucleic Acids Res. 2012;40(11):4742–53.
Wang W, Kwon EJ, Tsai LH. MicroRNAs in learning, memory, and neurological diseases. Learn Mem. 2012;19(9):359–68.
Wayman GA, Davare M, Ando H, Fortin D, Varlamova O, Cheng HY, et al. An activity-regulated microRNA controls dendritic plasticity by down-regulating p250GAP. Proc Natl Acad Sci U S A. 2008;105(26):9093–8.
Wulczyn FG, Smirnova L, Rybak A, Brandt C, Kwidzinski E, Ninnemann O, et al. Post-transcriptional regulation of the let-7 microRNA during neural cell specification. FASEB J. 2007;21(2):415–26.
Yokoo EM, Valente JG, Grattan L, Schmidt SL, Platt I, Silbergeld EK. Low level methylmercury exposure affects neuropsychological function in adults. Environ Health. 2003;2(1):8.
Yu JY, Chung KH, Deo M, Thompson RC, Turner DL. MicroRNA miR-124 regulates neurite outgrowth during neuronal differentiation. Exp Cell Res. 2008;314(14):2618–33.
Acknowledgments
M. Fabbri is a student of the PhD program in Biotechnology, School of Biological and Medical Sciences, University of Insubria (Italy).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pallocca, G., Fabbri, M., Sacco, M.G. et al. miRNA expression profiling in a human stem cell-based model as a tool for developmental neurotoxicity testing. Cell Biol Toxicol 29, 239–257 (2013). https://doi.org/10.1007/s10565-013-9250-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10565-013-9250-5