Abstract
As the price of next-generation sequencing keeps decreasing, cost is becoming a less important discriminator for diagnostic laboratories in choosing the preferred type of approach to genetic testing. Genome-wide sequencing strategies will plausibly become the standard first-tier tools for genetic testing, with the potential for deeper understanding of the genetic architecture of cardiomyopathies and discovery of the underlying aetiology in the many patients in whom the genetic cause remains elusive. Routine usage of extended sequencing assays will also enable “genetic-first diagnostics”, particularly for those patients affected with syndromic conditions of unclear genetic origin, often resulting in costly and distressing diagnostic odysseys before reaching a diagnosis. However, access to genome-wide data for all patients will need to be managed with rigour and caution by (cardiovascular) genetic professionals to avoid erroneous variant pathogenicity assertions and over-reporting uncertain findings, both damaging scenarios to patients and their family members. Researchers will also be required to adopt robust methods to demonstrate novel genetic associations with disease, given the high “narrative potential” of such large datasets and the dangers of generating further false positive associations (that have previously blighted the field of cardiac genetics). Here, we discuss advantages and dangers associated with the routine adoption of whole-exome (and whole-genome) sequencing in diagnostic facilities and in the research setting in the context of cardiomyopathies but relevant to several other conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The price gap between targeted panel sequencing and genome-wide approaches such as whole-exome and whole-genome sequencing (WES and WGS, respectively) is narrowing as a consequence of the decreasing costs of next-generation sequencing (NGS) [1, 2]. Although the exact cost of each approach depends on several variables—including the desired sequencing depth and target coverage, the platform used and, for panels, the number of targeted genes—the per-sample cost of WES is increasingly comparable with that of targeted panels, and the cost figure for WGS is steadily below USD 2000 since 2017 (Fig. 1).
Indicative price per sample (excluding consumables and service costs) for whole-genome sequencing (green line), whole-exome sequencing (purple) and targeted gene panels (red) over the last decade, alongside price fold difference between the three approaches (inset). The price of targeted gene panels is assumed to have remained constant given the higher price per sequenced base in the early years was balanced by the smaller number of genes targeted by panels. Price figures have been obtained from the web, scientific literature and personal communications [3, 4]
So far, the most prevalent approach to genetic testing in cardiac conditions has been that of targeted gene panels comprising most or all the genes implicated in the disease in question, with this choice driven by the convenient combination of relatively low prices and excellent variant detection sensitivity [3]. Extended panels targeting genes implicated in multiple conditions, such as the TruSight Cardio panel by Illumina [4], provide the additional advantage of a standardized approach on all patients affected by an inherited cardiovascular disease. As cost is becoming a less important discriminator in choosing the sequencing strategy in diagnostic laboratories, the routine usage of WES (and WGS) for diagnostic purposes on all patients, irrespective of the disease in question, is a realistic scenario for the near future. In addition, the cost of such approaches would be one-off, without the need of new panel-based genetic tests in case new genes are associated with the disease later on or a different disorder is suspected.
Such adoption of WES as a “universal panel” would enable laboratories to benefit from increased standardization, to re-analyse genotype-negative cases over time as new gene-disease associations emerge and to implement gene-discovery approaches de facto constituting research activities but directly translating also into potential diagnoses that would often be missed with targeted gene panels. In addition, it would spare those (often paediatric) patients affected with complex disorders of unclear genetic origin the necessity to undergo multiple tests before reaching a diagnosis.
First-tier adoption of WGS has also been reported to improve the diagnostic pathway compared with targeted panels through higher diagnostic yield and reduced time-to-diagnosis for similar reasons, though with the additional advantage of detecting also copy number variants (CNV), gross chromosomal abnormalities and deep intronic variants [5,6,7]. In hypertrophic cardiomyopathy (HCM), WGS has been observed to increase the yield by 20% compared with standard genetic testing using targeted panels, with approximately half of such increment due to deep intronic variants [8]. However, while WGS possesses the intrinsic technical advantage of not relying on an upstream exon capture method compared with any type of targeted sequencing (including WES), routine usage of WGS implies the need for computational infrastructures suited to store and analyse terabytes of data.
In any case, the technology enabling sequencing of DNA has evolved at a much faster pace than the ability to correctly interpret genetic variants, and the “narrative potential” of genome-wide sequencing data represents a high risk for erroneous (and potentially harmful) interpretation of variants as pathogenic [9]. The potential extent of such mistakes is highlighted by the fact that every individual exome contains ~ 54 variants previously (mis)classified as causal for a rare disease [10], and additional false positive interpretations of variants’ pathogenicity are probable if the analysis is not restricted to genes with a robustly validated association with disease.
In this review, we analyse advantages and perils associated with the routine adoption of a genome-wide sequencing approaches as first-tier genetic test, which will likely occur in many diagnostic laboratories in the short term, and how this will influence the interplay between diagnostics and research. While this review broadly focuses on cardiomyopathies, the issues discussed here are relevant to all Mendelian diseases (especially those with a partially unexplained genetic basis) in both the diagnostics and research contexts.
Genome-Wide Diagnostic Sequencing and the Potential for Gene Discovery
One advantage in the adoption of WES/WGS as a routine approach for clinical genetic testing is the possibility to use it as a tool for gene discovery when no (potentially) relevant variants (pathogenic/likely pathogenic/of uncertain significance) are detected in the known disease genes. While only genes with a proven role in disease should be routinely screened for diagnostic purposes (to minimize the probability of reporting false positive findings), the availability of sequencing data from all other genes enables research on potentially disease-causing alleles beyond validated disease genes. This can be done by means of co-segregation analysis in the patient’s family (if the pedigree is sufficiently large/informative) and/or by means of case-control burden testing (if a sufficiently large cohort is available) (Fig. 2).
Interplay between the diagnostic and research contexts relative to clinical genetic testing. Periodic re-analysis of variant evidence (pink loop) characterizes especially variants of uncertain significance (VUS), which can be upgraded to (likely) pathogenic or downgraded to benign as new evidence emerges, but can affect also variants previously classified as pathogenic or benign (dashed pink line) due to new evidence or, for example, updated interpretation guidelines. Publication of new robust evidence for genetic association with disease generally causes an upgrade in classification of variants previously believed to be benign to either VUS or (likely) pathogenic, constituting a new diagnostic finding
However, rigour and caution are necessary, especially in light of recent findings that demonstrated the need for strict and robust approaches in demonstrating new genetic associations with disease. The release of the Exome Aggregation Consortium (ExAC) database in October 2014—comprising publicly accessible WES data of > 60,000 individuals from various populations and common disease sequencing studies, now embedded in the larger Genome Aggregation Database, counting > 200,000 WES/WGS samples)—revealed how rare variants are collectively more common than previously estimated [10, 11]. Subsequently, gene-based rare variant association analyses demonstrating an actual lack of rare variation enrichment in cardiomyopathy patients compared with ExAC for many genes implicated in disease [12, 13]. More recently, the ClinGen consortium (established with the specific aim of assessing the clinical relevance of genes and genetic variants in a range of diseases [14]) classified 22 of the 33 genes (67%) evaluated for HCM as with limited or de facto non-existent evidence in support of their role in disease [15]. Furthermore, ClinGen curation efforts published thus far suggest how cardiovascular diseases are characterized by particularly high proportions of dubious gene-disease associations (Fig. 3), further underscoring the importance of stringent approaches to genetic association analyses in cardiomyopathies.
Proportions of genes implicated in 10 different diseases/groups of diseases classified in terms of the strength of evidence in support of their association with disease by ClinGen. The three cardiovascular conditions curated so far (panel on the left) are characterized by the lowest proportions of definitively validated gene-disease associations, all below 25%. In the case of HCM, genes classified as “definitive” by Clingen are MYH7, MYBPC3, TNNT2, TNNI3, MYL2, MYL3, ACTC1 and TPM1. Of the other seven genes listed in the main text for which the role in HCM is supported by robust segregation evidence, only CSRP3 and JPH2 are included in the figure in the “moderate evidence” group. The others have been either classified by Clingen as “definitive” for disease entities different from (but related to) HCM (ACTN2 and PLN for “intrinsic cardiomyopathy”, FLNC and ALPK3 for “syndromic conditions where isolated HCM may be seen”) or have been associated with HCM after the Clingen curation effort for HCM was published (FHOD3). Symbols between brackets (# and §) indicate conditions part of the same published curation effort by ClinGen
Taking HCM—the only cardiomyopathy curated by ClinGen so far—as an example (Fig. 3, left hand-side), the 8 sarcomeric gene-disease associations classified as definitive by ClinGen (MYBPC3, MYH7, TNNI3, TNNT2, MYL3, MYL2, ACTC1 and TPM1, reported to explain 30–50% of HCM cases [13, 16, 17]) were all originally based on strong segregation evidence in large pedigrees alone (18–96 individuals each) [18,19,20,21,22,23,24] and published in the decade following the first association study (MYH7) in 1990. Since 2000, seven additional genetic associations (CSRP3, PLN, ACTN2, FLNC, ALPK3, JPH2 and FHOD3) [25,26,27,28,29,30] with robust evidence of variant co-segregation with disease, alone or combined with other types of evidence, were proposed in the space of 19 years. However, these genes are estimated to explain at most 1% of HCM cases, with some of them being extremely rarely mutated. Accordingly, the increase in diagnostic yield given by the screening of 51 extra genes beyond those with a fully proven role in disease is, at most, negligible [31]. Taking this into account, on one hand, the existence of unknown Mendelian HCM genes is plausible (hence the utility of WES/WGS as gene discovery tools), but on the other hand, it is unlikely that such genes will be major contributors to the disease burden. In dilated cardiomyopathy (DCM), the main disease gene—TTN, explaining up to 20% of disease cases—has been discovered less than one decade ago [32], and it may be more likely that major genetic contributors to disease are still to be identified.
In addition to segregation analysis of large family pedigrees, gene-centric case-control association analysis is another effective strategy to gather statistical evidence in support of association with disease for dominant disease genes. In some cases, such the associations of FLNC and FHOD3 with HCM, family segregation evidence was combined with cohort-based case-control analyses, but in absence of sufficiently informative pedigrees, case-control association analysis on large cohorts can be a valid alternative. This is demonstrated for example by the case of PLN in HCM, for which evidence of variant segregation in large families is not available, but multiple cohort-based approaches provided statistical evidence in support of the association with disease [12, 13, 17, 33], ultimately classified as definitive by ClinGen [15].
Case-control comparisons across all genes through the utilization of WES/WGS is potentially a powerful approach to identify novel dominant disease genes. However, it can pose issues around multiple testing correction, given that adjustment for the testing of ~ 20,000 genes would require very large sample sizes to detect significant associations for rarely mutated disease genes. Gene prioritization approaches, such as the one proposed by Roca et al. [34] (based on scores measuring gene-specific (in)tolerance to variation) or by Zaidi et al. [35] (based on the pre-selection of genes highly expressed in the heart), can be applied ahead of variant filtration to exclude from the effective analysis those genes unlikely to play any role in cardiovascular disease for example, especially if cohorts are of moderate size. Large-scale efforts to create unprecedentedly large cohorts of rare disease patients without identified causative variants are underway to address these limitations in sample size and power. For example, Genomics England [36] have sequenced 100,000 genomes from rare disease and cancer patients, with plans to extend this to 5 million in the next few years.
In contrast to dominant inheritance, recessive inheritance enables analysis on smaller pedigrees, as stricter filtering based on genotypes and variant segregation in the family can be applied. Examples are recessive early-onset HCM caused by variants in ALPK3 and the recently published association between recessive DCM and the PPCS gene, demonstrated using 3 and 2 pedigrees of 2–10 sequenced individuals, respectively [28, 37]. There are also rare examples of recessive disease caused by specific variants in the established (and usually dominant) disease genes such as the protein-truncating variant p.Arg69Alafs*8 in TNNI3, observed to cause DCM only when homozygous [38]. In the case of de novo dominant variant occurrence, case-parents trios are per se sufficient to identify the pathogenic variant, given the expectation of only 1–2 de novo coding alleles per exome [34, 39]. This trio-based approach often translates into the identification of variants causing early-onset disease, as in the case of the p.His33Asn variant in ILK in arrhythmogenic right-ventricular cardiomyopathy (ARVC) (14 years old proband). Notably, WES has been reported to achieve a diagnostic yield of 50% in 18 parent-offspring pedigrees where early-onset DCM was observed [40], and overall, recessive cardiomyopathy and de novo dominant variant occurrence represent the cases in which routine usage of WES/WGS will likely be most productive in terms of Mendelian cardiomyopathy gene discovery.
Dominant or recessive inheritance models imply that causative variant(s), characterized by a large effect size, are sufficient for phenotype manifestation. However, cardiomyopathies are characterized by variable disease expressivity, with disease often presenting with different severity in different individuals also when caused by the same pathogenic variant. One hypothesis to explain this variability is the presence of genetic modifiers, i.e. variants that are not pathogenic in isolation but able to exert an effect on the phenotype in presence of a primary pathogenic allele, worsening or ameliorating disease severity.
For this reason, large or multiple families that share a common pathogenic Mendelian variant, and are characterized by variable penetrance and disease severity among variant carriers, represent a rare and valuable resource to study genetic modifiers of the disease phenotype, given the reduced phenotype variability associated with the primary disease variant. Such families are usually encountered in areas with genetic founder effects, such as the Netherlands [41,42,43,44], Italy [45] and South Africa [46]. Mouton et al. reported variants in the sarcomeric MYBPH gene to modify the degree of hypertrophy in HCM using 27 families carrying pathogenic alleles in MYH7 and TNNT2 [47]. Similar effects have been reported also for variants in non-sarcomeric genes, such as the regulator of cardiac development XIRP2, shown to exacerbate the effects of DCM caused by a variant in TNNT2 [48]. Research on modifier alleles is complex and generally requires particular datasets, and it is still preliminary even within validated disease genes, where some modifier alleles modulating the risk of adverse events and the disease severity have been identified for example in the main genes associated with Long QT syndrome (KCNQ1) and Brugada syndrome (SCN5A) [49, 50].
Some studies estimated the overall prevalence of carriers of multiple Mendelian (potentially) pathogenic variants (hence able to cause disease in isolation and not modifiers per se) and described a more severe cardiomyopathy phenotype in such patients [51,52,53,54] proposing an additive effect of Mendelian disease variants on disease risk and severity.
Our limited current knowledge on variable disease expressivity and penetrance in cardiomyopathies advocates for research efforts on validated disease genes (included in targeted gene panels) in the first instance, but adoption of WES/WGS for diagnostic purposes in centres where clinical evaluation is also performed would enable research to be also expanded to novel genes. This would represent a key development toward a deeper understanding of disease variability in the future, with the potential for improved risk prediction in cardiomyopathy patients and their families.
However, it is increasingly considered unlikely that the answer to the > 50% cardiomyopathy patients in whom the genetic cause of disease still remains elusive lies in Mendelian models of disease. Gene discovery efforts remain important as demonstrated by the novel gene-disease associations discovered in the last years such as FLNC, recently associated with both HCM and DCM [27, 55], but their impact on the yield of genetic testing will likely remain limited. In this respect, gene discovery aimed at finding modifier genes could have a greater impact on the effectiveness of the clinical care offered to patients through augmented understanding of disease variability, translating into more precise prognostication.
Genome-Wide Sequencing and Non-Mendelian Inheritance Models
As far as the high proportion of genotype-negative cardiomyopathy is concerned, innovative approaches to study non-Mendelian models of disease are much needed, as these more complex modes of inheritance are likely holding important answers. HCM in patients with no overt family history and no identifiable genetic causes has been shown to progress with a particularly favourable clinical course and has been defined as “non-familial”, with a hypothetically different and more complex underlying genetic aetiology [56]. Reports of cardiomyopathy caused by the co-occurrence of multiple variants and concurrent proof of each allele not being pathogenic in isolation are still very sparse, even for variants in validated disease genes. A rare example is that of p.Asn83His in TNNT2 and p.Asp955Asn in MYH7, both currently classified as variants of uncertain significance (VUS) but observed to cause severe DCM in a consanguineous family only when co-inherited [57].
Evidence of the contribution to cardiomyopathies of more common variants with lower effect sizes dates back to about a decade ago, with the published association of a common intronic variant in MYBPC3 with HCM in South Asians [58]. Subsequently, moderately-sized case-control genome-wide association studies (GWAS) have been performed both in HCM and DCM [59, 60] and detected significant associations for variants in FHOD3 and BAG3, respectively, with rare variants in both genes associated with the respective disease later on. Other GWAS studies have also highlighted how common variants influence cardiomyopathy-related phenotypes such as left-ventricular hypertrophy [61] and heart failure development [62, 63]. One area of research trying to address complex inheritance models and directly related to GWAS is that of polygenic risk scores (PRS), quantifying disease susceptibility based on large numbers of common variants of individually small effect sizes. Usually, PRS are developed using large cohorts of patients included in GWAS and genotyped with SNP arrays, as done in atherosclerosis or coronary artery disease [64, 65]. Routine adoption of WGS in the diagnostic setting could help to define patients’ family members disease risk (once PRS have been developed) and also further validate the role of PRS both in potentially non-Mendelian disease and in quantifying the effect of modifiers if in presence of a Mendelian pathogenic variant. However, such application in the diagnostic setting is still highly speculative for cardiomyopathies.
The contribution of environmental factors to cardiomyopathy has started to be surveyed in relation to the presence of rare genetic variants using large-scale cohorts only recently, for example, in the context of DCM and exposure to chemotherapy, alcohol consumption and pregnancy, revealing an interplay between genetic and environmental exposures [66,67,68]. However, larger studies are needed to elucidate such interplay between non-genetic factors and common variants (rather than rare ones) in determining non-Mendelian cardiomyopathy.
Overall, in the case of research on non-Mendelian models of disease, the main issues are neither related to a restricted potential for discovery through the usage of targeted panels nor to the lack of particular datasets, but rather to the paucity of sufficiently large and well characterized cohorts and bespoke strategies to investigate these inheritance models. In conclusion, the contribution of routine genome-wide sequencing for diagnostic purposes is unlikely to contribute to discovery or identification of non-Mendelian disease in cardiomyopathies at present but could indirectly support the validation of PRS through the creation of increasingly large sequenced cohorts.
Complex Phenotypes: Where Genome-Wide Sequencing Means Genotype-First Diagnostics
Cardiomyopathies can be observed as isolated conditions or in conjunction with extra-cardiac features if occurring as one of the manifestations of a syndromic disease. While several syndromes, such as Friedrich ataxia and Barth syndrome, always present with extra-cardiac involvement, for others (like Fabry disease and Danon disease) cardiomyopathy can be the only manifestation.
HCM is characterized by the existence of metabolic genocopies such as Fabry disease (GLA gene), Danon disease (LAMP2) and PRKAG2-cardiomyopathy (PRKAG2) that can mimic its phenotype, sometimes in absence of extra-cardiac manifestations. In such cases, making a reliable diagnostic differentiation based on imaging techniques alone can be very difficult for the cardiologist [69]. Potentially pathogenic variants in the genes associated with these conditions are observed in 1.5–5% of patients referred for HCM genetic testing [13, 17, 70], and routine inclusion of these genes in targeted panels for HCM genetic testing enables a “genotype-first” diagnostic approach that is directly informative on the underlying molecular cause of disease and can inform the radically different clinical management that these patients require compared with patients affected by “classical” HCM.
While genes associated with HCM genocopies are today routinely included in targeted panels in use on HCM patients, many other genes associated with complex syndromic conditions characterized by cardiomyopathy are grouped in different targeted panels used for testing pre-specified disease categories. In some cases, this can translate into long, costly and distressing “diagnostic odysseys”, with patients undergoing multiple genetic tests and clinical evaluations before reaching a definitive diagnosis [71, 72]. In such cases, the usage of WES as first-tier diagnostic tool can translate into a decrease in medical interventions and incremental savings estimated in the range USD 1727–4140 per diagnosis [73,74,75], with earlier reports also anticipating significant time and resource optimization in case of adoption of WES at first genetic test appointment [76, 77].
Irrespective of the disease in question, such a comprehensive approach to diagnostic sequencing at first evaluation, will require the downstream adoption of specific strategies to minimize the risks of erroneous variant interpretation and false positive findings, as proposed for example in the context of paediatric DCM [78]. In addition, savings brought by the application of a single genetic test to all patients could be used to fund multi-disciplinary teams evaluating case selection and aiding correct variant interpretation, with an estimated cost of GBP ~ 400 (USD 500) per case [79]. In other words, if the advantages of genome-wide sequencing as a tool for genotype-first diagnostics will be handled with the necessary caution to avoid “over-interpretation” of the estimated ~ 25,000 genetic variants detected in each individual, first-tier WES (or WGS, in the future) will bring undeniable benefits, especially to a subset of patients.
Exome vs Genome Sequencing: 2% VS 100%
The rationale for utilizing WES is that ~ 2% of the genome— the portion comprising protein-coding regions of all genes—harbours ~ 85% of genetic variants with large effects on disease-related traits [80]. Therefore, WES represents a powerful strategy to comprehensively evaluate protein-altering variation in the genome at lower costs and less computationally intensive labour compared with WGS.
However, WES suffers from two main limitations. First, like targeted gene panels, it relies on an exon-capture step upstream of the sequencing. Different approaches to exon capture have been developed and enhanced, but no method currently exhibits perfect uniformity of target coverage [81, 82]. Second, implicit in the rationale of WES is the fact that non-coding regions of the genome harbour an estimated 15% variants with large effect sizes on disease traits, and any pathogenic variant in such regions would be missed [80].
The former drawback translates into some exons or genes being only sub-optimally covered by WES, with GC content significantly and inversely associated with gene coverage and potentially creating bias toward variant identification in a subset of better-covered genes [83]. In this respect, multiple studies have demonstrated WGS—especially if not requiring DNA enrichment—to be more powerful than WES in detecting exome variants, including (but not limited to) copy number variants (CNV) expanding beyond protein-coding regions and providing hitherto unprecedented exome coverage [84,85,86].
WGS virtually enables detection of all types of variants occurring in a genome (with some caveats e.g. lower depth of coverage implies lower costs but also lower variant detection sensitivity), but the production, processing and analysis of huge amounts of data currently renders its routine adoption in many small diagnostic facilities unrealistic. In spite of the drop of sequencing prices over the last decade, estimates of sequencing costs per individual sample often exclude consumables costs, which are particularly high for WGS [87]. However, costs of hardware and computing power have also dropped quickly over the years, making the acquisition of appropriate sequencing and informatics infrastructure increasingly feasible for large diagnostic laboratories.
The main diagnostic advantage in choosing WGS lies in the detection of (potentially) causative variants not readily detectable by WES–CNVs and variants in the non-coding regions of the genome. In the context of cardiomyopathies, there have been limited studies on such variants so far, with analysis hampered by access to sequencing datasets and the still limited variant interpretation capabilities. As far as CNVs are concerned, large insertions or deletions have been reported in ~ 1% of HCM cases in several cohort-based studies, with such estimates including CNVs affecting non-validated disease genes [88,89,90], suggesting a minor contribution of CNVs to the disease burden in HCM. The contribution of these variants may be higher in other conditions such as ARVC and DCM, with reported detection rates of 4–7% in two recently published studies [91, 92], consistent with the more prominent role for loss of function variants in these conditions. Of note, a diagnostic pipeline including both routine usage of WES and detection of CNV has been recently proposed for DCM but implies additional costs for a SNP array specifically used for CNV detection [78]. Usage of WES would however prevent screening of non-coding regions of the genome. Studies on such variants in cardiomyopathies are still limited in number but suggest that deep-intronic variation may contribute substantially to the disease burden. For example, WGS identified pathogenic variants in 9% of a selected subset of patients (with HCM/left-ventricular hypertrophy/surgical interventions/family history of HCM) in which the genetic cause was elusive [8] and deep-intronic variants in MYBPC3 alone are estimated to explain 6.5% of HCM cases [93].
Of note, our ability to interpret these types of variants is still limited and hampered by the lack of public catalogues of clinically classified non-coding variants, although efforts to characterize the effect of non-coding variants at the genome-wide level are being carried out [94]. Currently, definitive interpretation of deep-intronic or regulatory variation requires RNA as well as DNA sequencing to investigate the effects of such variants on splicing patterns or gene expression levels [8, 93, 95], with access to myocardial tissue rarely available for routine genetic testing. However, in the near future, WGS will plausibly be employed by a larger number of laboratories thanks to the combination of lower costs and more accessible informatics infrastructure. While research on variants on non-coding regions is still in the early stages, WGS will enable much needed research on such variant classes which, along with improved CNV detection through the optimal coverage of WGS, promises a substantial impact on the yield of genetic tests in cardiomyopathy.
Not All that Glitters Is Gold: Dangers of Genome-Wide Diagnostic Sequencing
As mentioned above, in the last 5 years, new population genetics resources such as ExAC contributed to raise awareness in the scientific and clinical communities about the dangers of a permissive approach to genetic associations and variant interpretation. Resources of fundamental importance to clinical geneticists such as ClinVar [96] are partially contaminated from inconsistent variant interpretations, although enhancements in variant classification frameworks—such as the release of the current variant interpretation guidelines [97]—and efforts to identify variants that have been misclassified or are characterized by low penetrance aid more accurate and concordant results [98, 99].
The expansion of clinical genetic sequencing facilitates a deeper understanding of genetic variation and its role in health and disease, enabling in turn enhanced variant detection and interpretation in the diagnostic setting. However, routine adoption of genome-wide sequencing assays for clinical genetic testing in cardiomyopathies—as well as in other conditions—will also pose some challenges (Fig. 4).
It is key that researchers, physicians and geneticists remain aware of the dangers that false positive variant interpretation (i.e. a variant incorrectly classified as (likely) pathogenic) can have on patients and—often especially—on their family members. Labelling a healthy individual as intrinsically affected can have serious clinical, psychological and financial consequences, for example, by unnecessary implants of defibrillators, life-long anxiety and insurance discrimination [100,101,102]. Uncertain test results—irrespective of the classification as “VUS” being correct or not—also bring negative consequences such as concern and poor understanding of the implications [103].
Both WES and WGS imply the detection of extremely high numbers of variants compared with targeted gene panels, many of which may be plausible candidates for disease causation as characterized by rarity in population databases and therein classifiable as VUS. Data sharing efforts and strategies based on the intersection of public exome/genome variation databases with private cohorts will doubtlessly help in limiting the number of candidate pathogenic variants, for example through the creation of “non-pathogenic variants blacklists” [104].
Rigorous variant interpretation frameworks applied by diagnostic laboratories and recommended by regulating bodies are of little utility if not paired with accurate gene selection for the disease in question. The main risk of not applying both strategies lies, in the best-case scenario, in a substantial increase in the number of uncertain genetic findings, as shown on a smaller scale with the adoption of extended targeted gene panels compared with those including only validated disease genes [105]. As reflected by the complex gene classification scheme adopted by ClinGen [15], the degree of robustness of a given gene-disease association is not easy to define and it may be difficult to establish, in the clinical genetics setting, which genes with moderate evidence of association to include in diagnostic panels. With the routine adoption of WES or WGS for diagnostic sequencing, guidelines defining which genes to include in virtual panel testing are warranted.
Of note, restricting the testing to a limited subset of genes can also pose ethical dilemmas, as findings of potential medical value, unrelated to the disease being tested for, can occur in other genes. For this reason, the American College of Medical Genetics and Genomics has issued dedicated guidelines, identifying 59 genes in which clinically relevant variants should always be reported to the patients [106]. Naturally, when WES/WGS sequencing will be routinely used for diagnostics, incidental findings will be encountered at much higher frequencies, likely posing further challenges in establishing the optimal trade-off between the risks of “over-reporting” variants in absence of the related phenotype and often characterized by low penetrance, and not informing about the increased susceptibility to a specific, potentially life-threatening condition.
On the research side, approaches that investigate Mendelian protein-altering genetic causes of disease beyond validated disease genes are necessary, although those investigating non-coding variants and more complex inheritance models are plausibly holding the answer to a greater proportion of the genotype-negative cardiomyopathy patients. In this respect, genome-wide sequencing assays can be used in analysing families where the observed disease segregation is incompatible with Mendelian inheritance. This has been done in several studies that, using WES, proposed the cause of disease to lie in the combined effect of multiple variants segregating in the pedigree [53, 57, 107,108,109]. Such analyses are largely needed to elucidate potential oligo-genic inheritance in cardiomyopathies and could also serve the creation of catalogues of variants with intermediate effect sizes that in combination (perhaps also with non-genetic factors) can cause disease, as currently done for single pathogenic alleles e.g. in ClinVar, besides potentially representing validation datasets for PRS development. However, while investigations on these poorly explored disease models are absolutely warranted, they pose difficult challenges related to discerning variants that contribute to the disease onset from those that are instead benign bystanders or that contribute only in some cases due, for example, to low penetrance. Of note, only some of these studies include robust complementary evidence demonstrating that any subset of the proposed pathogenic combination of variants cannot cause disease in absence of the other allele(s). In some instances, WES-based studies on families hypothesise a decisive contributory role in reaching disease onset for variants in genes that are far from validated for the disease in question, such as TTN in HCM, or report very high diagnostic yields including many such genes and hypothesising oligo-genic inheritance [107, 110].
Despite cardiovascular genetics professionals willing to pay approximately USD 150 for every 1% increase in diagnostic yield, the lower number of VUS was a significant determinant for the uptake of a specific test over the others, and cardiovascular geneticists are currently still reported to have an overall strong preference for targeted panel testing over WES or WGS [111]. This reflects the fact that, irrespective of the disease in question, the mode of inheritance and the potential proportion of cases explained by the hypothesised genetic cause, it is fundamental that clinical genetics keeps following an “innocent-until-proven-guilty” philosophy, also to protect geneticists from having to cope with unmanageable amounts of uninterpretable variants.
This implies that what remains hypothetical in research—pending definitive validation, which always constitutes research progress—should not be incorporated into diagnostic practice and that the relevant bodies issue standardized guidelines concerning the diagnostic workflow in relation to genes’ and variants’ association with disease. Carefully addressing ethical aspects related to the potential discovery of pathogenic variants or risk alleles for conditions unrelated to the one being tested will also be a priority, given that incidental findings will most likely be encountered increasingly often.
References
The Cost of Sequencing a Human Genome [Internet]. Genome.gov. [citato 16 ottobre 2019]. Recuperato da: https://www.genome.gov/about-genomics/fact-sheets/Sequencing-Human-Genome-cost
van Dijk EL, Auger H, Jaszczyszyn Y, Thermes C. Ten years of next-generation sequencing technology. Trends Genet. 2014;30:418–26.
Walsh R, Cook SA. Issues and challenges in diagnostic sequencing for inherited cardiac conditions. Clin Chem. 2017;63:116–28.
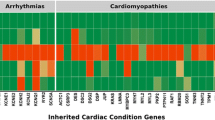
Pua CJ, Bhalshankar J, Miao K, Walsh R, John S, Lim SQ, et al. Development of a comprehensive sequencing assay for inherited cardiac condition genes. J Cardiovasc Transl Res. 2016;9:3–11.
Scocchia A, Wigby KM, Masser-Frye D, Campo MD, Galarreta CI, Thorpe E, et al. Clinical whole genome sequencing as a first-tier test at a resource-limited dysmorphology clinic in Mexico. NPJ Genom Med. 2019;4:1–12.
Clark MM, Stark Z, Farnaes L, Tan TY, White SM, Dimmock D, et al. Meta-analysis of the diagnostic and clinical utility of genome and exome sequencing and chromosomal microarray in children with suspected genetic diseases. NPJ Genom Med. 2018;3:1–10.
Lionel AC, Costain G, Monfared N, Walker S, Reuter MS, Hosseini SM, et al. Improved diagnostic yield compared with targeted gene sequencing panels suggests a role for whole-genome sequencing as a first-tier genetic test. Genet Med. 2018;20:435–43.
Bagnall RD, Ingles J, Dinger ME, Cowley MJ, Ross SB, Minoche AE, et al. Whole genome sequencing improves outcomes of genetic testing in patients with hypertrophic cardiomyopathy. J Am Coll Cardiol. 2018;72:419–29.
Goldstein DB, Allen A, Keebler J, Margulies EH, Petrou S, Petrovski S, et al. Sequencing studies in human genetics: design and interpretation. Nat Rev Genet. 2013;14:460–70.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
ExAC project pins down rare gene variants. Nature. 2016;536:249.
Walsh R, Thomson KL, Ware JS, Funke BH, Woodley J, McGuire KJ, et al. Reassessment of Mendelian gene pathogenicity using 7,855 cardiomyopathy cases and 60,706 reference samples. Genet Med [Internet]. 2016 [citato 25 novembre 2016]; Recuperato da: http://www.nature.com/gim/journal/vaop/ncurrent/full/gim201690a.html
Walsh R, Buchan R, Wilk A, John S, Felkin LE, Thomson KL, et al. Defining the genetic architecture of hypertrophic cardiomyopathy: re-evaluating the role of non-sarcomeric genes. Eur Heart J. 2017;38:3461–8.
Rehm HL, Berg JS, Brooks LD, Bustamante CD, Evans JP, Landrum MJ, et al. ClinGen--the clinical genome resource. N Engl J Med. 2015;372:2235–42.
Ingles J, Goldstein J, Thaxton C, Caleshu C, Corty EW, Crowley SB, et al. Evaluating the clinical validity of hypertrophic cardiomyopathy genes. Circ Genom Precis Med. 2019;12:e002460.
Maron BJ, Maron MS, Semsarian C. Genetics of hypertrophic cardiomyopathy after 20 years: clinical perspectives. J Am Coll Cardiol. 2012;60:705–15.
Mazzarotto F, Girolami F, Boschi B, Barlocco F, Tomberli A, Baldini K, et al. Defining the diagnostic effectiveness of genes for inclusion in panels: the experience of two decades of genetic testing for hypertrophic cardiomyopathy at a single center. Genet Med. 2018.
Geisterfer-Lowrance AA, Kass S, Tanigawa G, Vosberg HP, McKenna W, Seidman CE, et al. A molecular basis for familial hypertrophic cardiomyopathy: a beta cardiac myosin heavy chain gene missense mutation. Cell. 1990;62:999–1006.
Thierfelder L, Watkins H, MacRae C, Lamas R, McKenna W, Vosberg HP, et al. Alpha-tropomyosin and cardiac troponin T mutations cause familial hypertrophic cardiomyopathy: a disease of the sarcomere. Cell. 1994;77:701–12.
Watkins H, Conner D, Thierfelder L, Jarcho JA, MacRae C, McKenna WJ, et al. Mutations in the cardiac myosin binding protein-C gene on chromosome 11 cause familial hypertrophic cardiomyopathy. Nat Genet. 1995;11:434–7.
Poetter K, Jiang H, Hassanzadeh S, Master SR, Chang A, Dalakas MC, et al. Mutations in either the essential or regulatory light chains of myosin are associated with a rare myopathy in human heart and skeletal muscle. Nat Genet. 1996;13:63–9.
Kimura A, Harada H, Park JE, Nishi H, Satoh M, Takahashi M, et al. Mutations in the cardiac troponin I gene associated with hypertrophic cardiomyopathy. Nat Genet. 1997;16:379–82.
Flavigny J, Richard P, Isnard R, Carrier L, Charron P, Bonne G, et al. Identification of two novel mutations in the ventricular regulatory myosin light chain gene (MYL2) associated with familial and classical forms of hypertrophic cardiomyopathy. J Mol Med. 1998;76:208–14.
Mogensen J, Klausen IC, Pedersen AK, Egeblad H, Bross P, Kruse TA, et al. Alpha-cardiac actin is a novel disease gene in familial hypertrophic cardiomyopathy. J Clin Invest. 1999;103:R39–43.
Geier C, Gehmlich K, Ehler E, Hassfeld S, Perrot A, Hayess K, et al. Beyond the sarcomere: CSRP3 mutations cause hypertrophic cardiomyopathy. Hum Mol Genet. 2008;17:2753–65.
Girolami F, Iascone M, Tomberli B, Bardi S, Benelli M, Marseglia G, et al. Novel α-Actinin 2 variant associated with familial hypertrophic cardiomyopathy and juvenile atrial ArrhythmiasCLINICAL PERSPECTIVE. Circ Cardiovasc Genet. 2014;7:741–50.
Valdés-Mas R, Gutiérrez-Fernández A, Gómez J, Coto E, Astudillo A, Puente DA, et al. Mutations in filamin C cause a new form of familial hypertrophic cardiomyopathy. Nat Commun. 2014;5:5326.
Almomani R, Verhagen JMA, Herkert JC, Brosens E, van Spaendonck-Zwarts KY, Asimaki A, et al. Biallelic truncating mutations in ALPK3 cause severe pediatric cardiomyopathy. J Am Coll Cardiol. 2016;67:515–25.
Vanninen SUM, Leivo K, Seppälä EH, Aalto-Setälä K, Pitkänen O, Suursalmi P, et al. Heterozygous junctophilin-2 (JPH2) p.(Thr161Lys) is a monogenic cause for HCM with heart failure. PLoS One. 2018;13:e0203422.
Ochoa JP, Sabater-Molina M, García-Pinilla JM, Mogensen J, Restrepo-Córdoba A, Palomino-Doza J, et al. Formin homology 2 domain containing 3 (FHOD3) is a genetic basis for hypertrophic cardiomyopathy. J Am Coll Cardiol. 2018;72:2457–67.
Thomson KL, Ormondroyd E, Harper AR, Dent T, McGuire K, Baksi J, et al. Analysis of 51 proposed hypertrophic cardiomyopathy genes from genome sequencing data in sarcomere negative cases has negligible diagnostic yield. Genet Med. 2019;21:1576–84.
Herman DS, Lam L, Taylor MRG, Wang L, Teekakirikul P, Christodoulou D, et al. Truncations of Titin causing dilated cardiomyopathy. N Engl J Med. 2012;366:619–28.
Walsh R, Mazzarotto F, Whiffin N, Buchan R, Midwinter W, Wilk A, et al. Quantitative approaches to variant classification increase the yield and precision of genetic testing in Mendelian diseases: the case of hypertrophic cardiomyopathy. Genome Med. 2019;11:5.
Roca I, Fernández-Marmiesse A, Gouveia S, Segovia M, Couce ML. Prioritization of variants detected by next generation sequencing according to the mutation tolerance and mutational architecture of the corresponding genes. Int J Mol Sci [Internet]. 2018 [citato 5 novembre 2019];19. Recuperato da: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6032105/
Zaidi S, Choi M, Wakimoto H, Ma L, Jiang J, Overton JD, et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature. 2013;498:220–3.
Turnbull C, Scott RH, Thomas E, Jones L, Murugaesu N, Pretty FB, et al. The 100 000 genomes project: bringing whole genome sequencing to the NHS. BMJ. 2018;361:k1687.
Iuso A, Wiersma M, Schüller H-J, Pode-Shakked B, Marek-Yagel D, Grigat M, et al. Mutations in PPCS, encoding phosphopantothenoylcysteine synthetase, cause autosomal-recessive dilated cardiomyopathy. Am J Hum Genet. 2018;102:1018–30.
Kühnisch J, Herbst C, Al-Wakeel-Marquard N, Dartsch J, Holtgrewe M, Baban A, et al. Targeted panel sequencing in pediatric primary cardiomyopathy supports a critical role of TNNI3. Clin Genet. 2019.
Veltman JA, Brunner HG. De novo mutations in human genetic disease. Nat Rev Genet. 2012;13:565–75.
Long PA, Evans JM, Olson TM. Diagnostic yield of whole exome sequencing in pediatric dilated cardiomyopathy. J Cardiovasc Dev Dis. 2017;4.
Claes GRF, van Tienen FHJ, Lindsey P, Krapels IPC, Helderman-van den Enden AT, Hoos MB, et al. Hypertrophic remodelling in cardiac regulatory myosin light chain (MYL2) founder mutation carriers. Eur Heart J. 2016;37:1815–22.
van Velzen HG, Schinkel AFL, Oldenburg RA, van Slegtenhorst MA, Frohn-Mulder IME, van der Velden J, et al. Clinical characteristics and Long-term outcome of hypertrophic cardiomyopathy in individuals with a MYBPC3 (myosin-binding protein C) founder mutation. Circ Cardiovasc Genet. 2017;10.
Hoorntje ET, Bollen IA, Barge-Schaapveld DQ, van Tienen FH, Te Meerman GJ, Jansweijer JA, et al. Lamin A/C-related cardiac disease: late onset with a variable and mild phenotype in a large cohort of patients with the lamin A/C p.(Arg331Gln) founder mutation. Circ Cardiovasc Genet. 2017;10.
Hoorntje ET, van Spaendonck-Zwarts KY, Te Rijdt WP, Boven L, Vink A, van der Smagt JJ, et al. The first titin (c.59926 + 1G > A) founder mutation associated with dilated cardiomyopathy. Eur J Heart Fail. 2018;20:803–6.
Calore C, De Bortoli M, Romualdi C, Lorenzon A, Angelini A, Basso C, et al. A founder MYBPC3 mutation results in HCM with a high risk of sudden death after the fourth decade of life. J Med Genet. 2015;52:338–47.
Moolman-Smook JC, De Lange WJ, Bruwer EC, Brink PA, Corfield VA. The origins of hypertrophic cardiomyopathy-causing mutations in two South African subpopulations: a unique profile of both independent and founder events. Am J Hum Genet. 1999;65:1308–20.
Mouton JM, van der Merwe L, Goosen A, Revera M, Brink PA, Moolman-Smook JC, et al. MYBPH acts as modifier of cardiac hypertrophy in hypertrophic cardiomyopathy (HCM) patients. Hum Genet. 2016;135:477–83.
Long Pamela A., Larsen Brandon T., Evans Jared M., Olson Timothy M. Exome sequencing identifies pathogenic and modifier mutations in a child with sporadic dilated cardiomyopathy. Journal of the American Heart Association. 4:e002443.
Duchatelet S, Crotti L, Peat RA, Denjoy I, Itoh H, Berthet M, et al. Identification of a KCNQ1 polymorphism acting as a protective modifier against arrhythmic risk in long QT syndrome. Circ Cardiovasc Genet [Internet]. 2013 [citato 7 novembre 2019];6. Recuperato da: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3864834/
Bezzina CR, Barc J, Mizusawa Y, Remme CA, Gourraud J-B, Simonet F, et al. Common variants at SCN5A-SCN10A and HEY2 are associated with Brugada syndrome, a rare disease with high risk of sudden cardiac death. Nat Genet. 2013;45:1044–9.
Xu T, Yang Z, Vatta M, Rampazzo A, Beffagna G, Pilichou K, et al. Compound and digenic heterozygosity contributes to arrhythmogenic right ventricular cardiomyopathy. J Am Coll Cardiol. 2010;55:587–97.
Rigato I, Bauce B, Rampazzo A, Zorzi A, Pilichou K, Mazzotti E, et al. Compound and digenic heterozygosity predicts lifetime arrhythmic outcome and sudden cardiac death in desmosomal gene-related arrhythmogenic right ventricular cardiomyopathy. Circ Cardiovasc Genet. 2013;6:533–42.
Cowan JR, Kinnamon DD, Morales A, Salyer L, Nickerson DA, Hershberger RE. Multigenic disease and Bilineal inheritance in dilated cardiomyopathy is illustrated in nonsegregating LMNA pedigrees. Circ Genom Precis Med. 2018;11:e002038.
Girolami F, Ho CY, Semsarian C, Baldi M, Will ML, Baldini K, et al. Clinical features and outcome of hypertrophic cardiomyopathy associated with triple sarcomere protein gene mutations. J Am Coll Cardiol. 2010;55:1444–53.
Begay RL, Graw SL, Sinagra G, Asimaki A, Rowland TJ, Slavov DB, et al. Filamin C truncation mutations are associated with arrhythmogenic dilated cardiomyopathy and changes in the cell-cell adhesion structures. JACC Clin Electrophysiol. 2018;4:504–14.
Ingles J, Burns C, Bagnall RD, Lam L, Yeates L, Sarina T, et al. Nonfamilial hypertrophic cardiomyopathy: prevalence, natural history, and clinical implications. Circ Cardiovasc Genet. 2017;10.
Petropoulou E, Soltani M, Firoozabadi AD, Namayandeh SM, Crockford J, Maroofian R, et al. Digenic inheritance of mutations in the cardiac troponin (TNNT2) and cardiac beta myosin heavy chain (MYH7) as the cause of severe dilated cardiomyopathy. Eur J Med Genet. 2017;60:485–8.
Dhandapany PS, Sadayappan S, Xue Y, Powell GT, Rani DS, Nallari P, et al. A common MYBPC3 (cardiac myosin binding protein C) variant associated with cardiomyopathies in South Asia. Nat Genet. 2009;41:187–91.
Wooten EC, Hebl VB, Wolf MJ, Greytak SR, Orr NM, Draper I, et al. Formin homology 2 domain containing 3 variants associated with hypertrophic cardiomyopathy. Circ Cardiovasc Genet. 2013;6:10–8.
Villard E, Perret C, Gary F, Proust C, Dilanian G, Hengstenberg C, et al. A genome-wide association study identifies two loci associated with heart failure due to dilated cardiomyopathy. Eur Heart J. 2011;32:1065–76.
Arnett DK, Meyers KJ, Devereux RB, Tiwari HK, Gu CC, Vaughan LK, et al. Genetic variation in NCAM1 contributes to left ventricular wall thickness in hypertensive families. Circ Res. 2011;108:279–83.
Nay A, Vargas Jose D, Chaojie Y, Cabrera Claudia P, Warren Helen R, Kenneth F, et al. Genome-wide analysis of left ventricular image-derived phenotypes identifies fourteen loci associated with cardiac morphogenesis and heart failure development. Circulation. 2019;140:1318–30.
Shah S, Henry A, Roselli C, Lin H, Sveinbjörnsson G, Fatemifar G, et al. Genome-wide association and Mendelian randomisation analysis provide insights into the pathogenesis of heart failure. Nat Commun. 2020;11:1–12.
Natarajan P, Young R, Stitziel NO, Padmanabhan S, Baber U, Mehran R, et al. Polygenic risk score identifies subgroup with higher burden of atherosclerosis and greater relative benefit from statin therapy in the primary prevention setting. Circulation. 2017;135:2091–101.
Mega JL, Stitziel NO, Smith JG, Chasman DI, Caulfield M, Devlin JJ, et al. Genetic risk, coronary heart disease events, and the clinical benefit of statin therapy: an analysis of primary and secondary prevention trials. Lancet. 2015;385:2264–71.
Ware JS, Li J, Mazaika E, Yasso CM, DeSouza T, Cappola TP, et al. Shared genetic predisposition in peripartum and dilated cardiomyopathies. N Engl J Med. 2016;374:233–41.
Ware JS, Amor-Salamanca A, Tayal U, Govind R, Serrano I, Salazar-Mendiguchía J, et al. Genetic etiology for alcohol-induced cardiac toxicity. J Am Coll Cardiol. 2018;71:2293–302.
Garcia-Pavia P, Kim Y, Restrepo-Cordoba MA, Lunde IG, Wakimoto H, Smith AM, et al. Genetic variants associated with cancer therapy-induced cardiomyopathy. Circulation. 2019;140:31–41.
Sankaranarayanan R, Fleming EJ, Garratt CJ. Mimics of hypertrophic cardiomyopathy – diagnostic clues to aid early identification of phenocopies. Arrhythm Electrophysiol Rev. 2013;2:36–40.
Ho CY. Hypertrophic cardiomyopathy: for heart failure clinics: genetics of cardiomyopathy and heart failure. Heart Fail Clin. 2010;6:141–59.
Thevenon J, Duffourd Y, Masurel-Paulet A, Lefebvre M, Feillet F, El Chehadeh-Djebbar S, et al. Diagnostic odyssey in severe neurodevelopmental disorders: toward clinical whole-exome sequencing as a first-line diagnostic test. Clin Genet. 2016;89:700–7.
Sawyer SL, Hartley T, Dyment DA, Beaulieu CL, Schwartzentruber J, Smith A, et al. Utility of whole-exome sequencing for those near the end of the diagnostic odyssey: time to address gaps in care. Clin Genet. 2016;89:275–84.
Tan TY, Dillon OJ, Stark Z, Schofield D, Alam K, Shrestha R, et al. Diagnostic Impact and cost-effectiveness of whole-exome sequencing for ambulant children with suspected monogenic conditions. JAMA Pediatr. 2017;171:855–62.
Monroe GR, Frederix GW, Savelberg SMC, de Vries TI, Duran KJ, van der Smagt JJ, et al. Effectiveness of whole-exome sequencing and costs of the traditional diagnostic trajectory in children with intellectual disability. Genet Med. 2016;18:949–56.
Vrijenhoek T, Middelburg EM, Monroe GR, van Gassen KLI, Geenen JW, Hövels AM, et al. Whole-exome sequencing in intellectual disability; cost before and after a diagnosis. Eur J Hum Genet. 2018;26:1566–71.
van Nimwegen KJM, Schieving JH, Willemsen MAAP, Veltman JA, van der Burg S, van der Wilt GJ, et al. The diagnostic pathway in complex paediatric neurology: a cost analysis. Eur J Paediatr Neurol. 2015;19:233–9.
Valencia CA, Husami A, Holle J, Johnson JA, Qian Y, Mathur A, et al. Clinical Impact and Cost-effectiveness of whole exome sequencing as a diagnostic tool: a pediatric center’s experience. Front Pediatr [Internet]. 2015 [citato 29 ottobre 2019];3. Recuperato da: https://doi.org/10.3389/fped.2015.00067/full
Herkert JC, Abbott KM, Birnie E, Meems-Veldhuis MT, Boven LG, Benjamins M, et al. Toward an effective exome-based genetic testing strategy in pediatric dilated cardiomyopathy. Genet Med. 2018;20:1374–86.
Taylor J, Craft J, Blair E, Wordsworth S, Beeson D, Chandratre S, et al. Implementation of a genomic medicine multi-disciplinary team approach for rare disease in the clinical setting: a prospective exome sequencing case series. Genome Med. 2019;11:46.
Choi M, Scholl UI, Ji W, Liu T, Tikhonova IR, Zumbo P, et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci U S A. 2009;106:19096–101.
Samorodnitsky E, Jewell BM, Hagopian R, Miya J, Wing MR, Lyon E, et al. Evaluation of hybridization capture versus amplicon-based methods for whole-exome sequencing. Hum Mutat. 2015;36:903–14.
Chilamakuri CSR, Lorenz S, Madoui M-A, Vodák D, Sun J, Hovig E, et al. Performance comparison of four exome capture systems for deep sequencing. BMC Genomics. 2014;15:449.
Manase D, D’Alessandro LC, Manickaraj AK, Al Turki S, Hurles ME, Mital S. High throughput exome coverage of clinically relevant cardiac genes. BMC Med Genomics [Internet]. 2014 [citato 9 maggio 2016];7. Recuperato da: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4272796/
Belkadi A, Bolze A, Itan Y, Cobat A, Vincent QB, Antipenko A, et al. Whole-genome sequencing is more powerful than whole-exome sequencing for detecting exome variants. Proc Natl Acad Sci USA. 2015;112(17):5473–8.
Meienberg J, Zerjavic K, Keller I, Okoniewski M, Patrignani A, Ludin K, et al. New insights into the performance of human whole-exome capture platforms. Nucleic Acids Res. 2015;43:e76.
Meienberg J, Bruggmann R, Oexle K, Matyas G. Clinical sequencing: is WGS the better WES? Hum Genet. 2016;135:359–62.
Schwarze K, Buchanan J, Fermont JM, Dreau H, Tilley MW, Taylor JM, et al. The complete costs of genome sequencing: a microcosting study in cancer and rare diseases from a single center in the United Kingdom. Genet Med. 2019:1–10.
Lopes LR, Murphy C, Syrris P, Dalageorgou C, McKenna WJ, Elliott PM, et al. Use of high-throughput targeted exome-sequencing to screen for copy number variation in hypertrophic cardiomyopathy. Eur J Med Genet. 2015;58:611–6.
Mademont-Soler I, Mates J, Yotti R, Espinosa MA, Pérez-Serra A, Fernandez-Avila AI, et al. Additional value of screening for minor genes and copy number variants in hypertrophic cardiomyopathy. PLoS One. 2017;12:e0181465.
Ceyhan-Birsoy O, Pugh TJ, Bowser MJ, Hynes E, Frisella AL, Mahanta LM, et al. Next generation sequencing-based copy number analysis reveals low prevalence of deletions and duplications in 46 genes associated with genetic cardiomyopathies. Mol Genet Genomic Med. 2016;4:143–51.
Kalliopi P, Elisabetta L, Ilaria R, Rudy C, Marzia DB, Marina PM, et al. Large genomic rearrangements of desmosomal genes in italian arrhythmogenic cardiomyopathy patients. Circ Arrhythm Electrophysiol. 2017;10:e005324.
Mates J, Mademont-Soler I, del Olmo B, Ferrer-Costa C, Coll M, Pérez-Serra A, et al. Role of copy number variants in sudden cardiac death and related diseases: genetic analysis and translation into clinical practice. Eur J Hum Genet. 2018;26:1014–25.
Janin A, Chanavat V, Rollat-Farnier P-A, Bardel C, Nguyen K, Chevalier P, et al. Whole MYBPC3 NGS sequencing as a molecular strategy to improve the efficiency of molecular diagnosis of patients with hypertrophic cardiomyopathy. Human Mutation [Internet]. 2019 [citato 19 novembre 2019];n/a.
Wells A, Heckerman D, Torkamani A, Yin L, Sebat J, Ren B, et al. Ranking of non-coding pathogenic variants and putative essential regions of the human genome. Nat Commun. 2019;10:1–9.
Cummings BB, Marshall JL, Tukiainen T, Lek M, Donkervoort S, Foley AR, et al. Improving genetic diagnosis in Mendelian disease with transcriptome sequencing. Sci Transl Med [Internet]. 2017 [citato 19 novembre 2019];9. Recuperato da: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5548421/
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014;42:D980–5.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Shah N, Hou Y-CC YH-C, Sainger R, Caskey CT, Venter JC, et al. Identification of misclassified ClinVar variants via disease population prevalence. Am J Hum Genet. 2018;102:609–19.
Yang S, Lincoln SE, Kobayashi Y, Nykamp K, Nussbaum RL, Topper S. Sources of discordance among germ-line variant classifications in ClinVar. Genet Med. 2017;19:1118–26.
Lin G, Nishimura RA, Gersh BJ, Phil D, Ommen SR, Ackerman MJ, et al. Device complications and inappropriate implantable cardioverter defibrillator shocks in patients with hypertrophic cardiomyopathy. Heart. 2009;95:709–14.
Ader T, Susswein LR, Callanan NP, Evans JP. Attitudes and practice of genetic counselors regarding anonymous testing for BRCA1/2. J Genet Couns. 2009;18:606–17.
Kerruish N, Robertson S. Newborn screening: new developments, new dilemmas. J Med Ethics. 2005;31:393–8.
Burns C, Yeates L, Spinks C, Semsarian C, Ingles J. Attitudes, knowledge and consequences of uncertain genetic findings in hypertrophic cardiomyopathy. Eur J Hum Genet. 2017;25:809–15.
Maffucci P, Bigio B, Rapaport F, Cobat A, Borghesi A, Lopez M, et al. Blacklisting variants common in private cohorts but not in public databases optimizes human exome analysis. Proc Natl Acad Sci U S A. 2019;116:950–9.
Alfares AA, Kelly MA, McDermott G, Funke BH, Lebo MS, Baxter SB, et al. Results of clinical genetic testing of 2,912 probands with hypertrophic cardiomyopathy: expanded panels offer limited additional sensitivity. Genet Med. 2015;17:880–8.
Kalia SS, Adelman K, Bale SJ, Chung WK, Eng C, Evans JP, et al. Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SF v2.0): a policy statement of the American College of Medical Genetics and Genomics. Genet Med. 2017;19:249–55.
Li L, Bainbridge MN, Tan Y, Willerson JT, Marian AJ. A potential oligogenic etiology of hypertrophic cardiomyopathy, a classic single gene disorder. Circ Res. 2017;120:1084–90.
Gifford CA, Ranade SS, Samarakoon R, Salunga HT, de Soysa TY, Huang Y, et al. Oligogenic inheritance of a human heart disease involving a genetic modifier. Science. 2019;364:865–70.
Zaragoza MV, Fung L, Jensen E, Oh F, Cung K, McCarthy LA, et al. Exome sequencing identifies a novel LMNA splice-site mutation and multigenic Heterozygosity of potential modifiers in a family with sick sinus syndrome, dilated cardiomyopathy, and sudden cardiac death. PLoS One. 2016;11:e0155421.
Nguyen K, Roche S, Donal E, Odent S, Eicher J-C, Faivre L, et al. Whole exome sequencing reveals a large genetic heterogeneity and revisits the causes of hypertrophic cardiomyopathy. Circ Genom Precis Med. 2019;12:e002500.
Buchanan J, Blair E, Thomson KL, Ormondroyd E, Watkins H, Taylor JC, et al. Do health professionals value genomic testing? A discrete choice experiment in inherited cardiovascular disease. Eur J Hum Genet. 2019;27:1639–48.
Funding
This work was supported by the Italian Ministry of Health (RF-2013-02356787 and NET-2011-02347173), the European Union Horizon 2020 framework programme (SILICOFCM, GA 777204) and by Regione Toscana (Tuscany Registry of Sudden Cardiac Death [ToRSADE-FAS Salute 2014]). F.M. is supported by a post-doctoral research fellowship from the University of Florence. R.W. is supported by a post-doctoral grant from the Amsterdam Cardiovascular Sciences.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mazzarotto, F., Olivotto, I. & Walsh, R. Advantages and Perils of Clinical Whole-Exome and Whole-Genome Sequencing in Cardiomyopathy. Cardiovasc Drugs Ther 34, 241–253 (2020). https://doi.org/10.1007/s10557-020-06948-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10557-020-06948-4