Abstract
DNA methylation as part of the epigenetic gene-silencing complex is a universal occurring change in lung cancer. Numerous studies investigated methylation of specific genes in primary tumors, in serum or plasma samples, and in specimens from the aerodigestive tract epithelium of lung cancer patients. In most studies, single genes or small numbers of genes were analyzed. Moreover, it has been observed that methylation of certain genes can already be detected in samples from the upper aerodigestive tract epithelium of cancer-free heavy smokers. These findings indicated that methylation of certain genes may be a useful biomarker for prognosis, disease recurrence, early detection, and lung cancer risk assessment. So far, several genes were identified which seem to be of worse prognostic relevance when they were found to be methylated. In addition, it has been shown that a panel of markers may be relevant to predict disease recurrence after surgery. In comparison to analysis of single or small numbers of genes, methods for genome-wide detection of methylation were developed recently. These approaches are focused on either pharmacological re-activation of methylated genes followed by expression microarray analysis or on microarray analysis of sodium bisulfite-treated or affinity-enriched methylated DNA sequences. With currently available methods for the simultaneous detection of methylation, up to 28,000 CpG islands can be analyzed. Overall, we are just at the beginning of translating these findings into the clinic and there is hope that future patients will benefit from these results.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Despite enormous effort in early detection programs and development of new drugs and innovative treatment strategies in recent years, lung cancer is still very often a deadly disease with 5-year survival rates of about 14% [1]. While numerous genetic abnormalities have been described in lung carcinomas during the past decades, the detection of epigenetic changes in lung cancer started significantly later. At the beginning of this era, methylation analysis of only single genes was performed in a small number of primary tumors of lung cancer patients. However, rapidly, an increasing number of genes were investigated for their methylation status and researchers reported that methylation can also be detected in blood samples and in exfoliative material of the aerodigestive tract epithelium from lung cancer patients. Moreover, it became evident that methylation of certain genes can already be detected in cancer-free smokers. In the meantime, data about worse prognostic impact of methylation of certain genes have been reported. Moreover, it has been demonstrated that the methylation status of certain genes may predict disease recurrence in non-small cell lung cancer (NSCLC) patients. Recently, high throughput approaches to detect methylation have been developed. These methods provide researchers with a huge amount of information which will help to better understand the role of epigenetic changes in the pathogenesis of lung cancer. We now need to translate our findings about DNA methylation in lung cancer into clinical practice which hopefully will result in a better outcome for these patients.
In this review, we summarize results from single-gene methylation analysis as well as from high throughput methylation approaches in lung cancer cell lines and in specimens from lung cancer patients and cancer-free smokers.
2 DNA methylation and regulation of gene expression
Epigenetic mechanisms largely contribute to the regulation of the transcriptional activity of a gene. Among these mechanisms, DNA methylation and various chemical modifications of histone proteins (acetylation, methylation, ubiquitination, phosphorylation, and ADP-ribosylation) are key regulators which affect the binding of transcription factors to DNA and change the structure of chromatin resulting either in gene activation or gene silencing [2, 3]. In recent years, the interaction between DNA methylation and histone modifications resulting in gene silencing became more and more evident [4]. Methyl-cytosines bind methyl-CpG binding domain proteins. Through interactions of them with histone deacetylases, histone methyltransferases, and chromatin-remodeling enzymes, methylated DNA is translated into compacted chromatin which is repressive for transcription. Several studies revealed that DNA methylation directly influences both the acetylation and the methylation state of histone proteins [5–7].
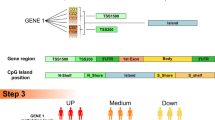
In mammalian cells, DNA is methylated by DNA methyltransferases (DNMTs). DNMT1 is associated with the replication complex and responsible for both methylating fully unmethylated DNA (de novo methylation) and maintaining pre-existing DNA methylation patterns in a cell [8]. DNMT3A and DNMT3B are de novo DNMTs [9, 10]. The target position for the addition of a methyl-group is the 5′-carbon of cytosines within CpG dinucleotides. The CpG dinucleotide is relatively rare in the mammalian genome [11]. However, certain regions of the genome, 0.5-4 kb in length, harbor CpG dinucleotides at high density and are called CpG islands (CGIs). CGIs are found in approximately 60% of the human gene-promoter regions and may extend into the 5′ coding end of the gene [12]. Computational analysis predicts 29,000 CGIs in the human genome and most chromosomes harbor 5-15 islands per Mb [13, 14]. While 70-80% of all non-CGI CpG dinucleotides in the human genome are methylated, CGI CpG dinucleotides usually remain unmethylated. Exceptions are CGIs associated with imprinted, X-linked, and tissue-specific expressed genes [15–17]. Alterations in CGI methylation and histone modifications leading to silencing of certain tumor suppressor genes (TSGs) frequently occur in cancer cells [18, 19].
DNA methylation is a reversible process and expression of methylated genes can be restored by the use of DNMT inhibitors such as 5-aza-2′-deoxycytidine (Aza-dC). In addition, histone deacetylase (HDAC) inhibitors such as trichostatin A (TSA) synergistically enhance the re-activating effect of DNMT inhibitors [20].
3 Interaction between microRNAs and DNA methylation in lung cancer
Recently, it has become apparent that microRNAs (miRNAs) are involved in tumorigenesis [21]. miRNAs are noncoding, 18-25 nt long RNAs which regulate a variety of biological processes including development, cellular differentiation, proliferation, and apoptosis by silencing specific target genes [22]. miRNAs induce translational repression by binding to complementary sequences in the 3′ untranslated regions (3′ UTR) of target mRNAs or by directing mRNA degradation [23]. Deregulation of miRNA expression resulting in abnormal activation or repression of oncogenes and TSGs is a common event in various human cancers [21]. It has been shown that DNA methylation is one mechanism for miRNA gene silencing [24–27]. Examples for methylated miRNA genes in lung cancer are miR-34a and miR124a [27, 28]. Interestingly, both of these miRNAs are part of molecular pathways whose depletion may contribute to the development of a malignant phenotype. While miR34a is part of the p53 network, miR124a is supposed to regulate levels of the cell cycle progression factor CDK6 [27, 29].
Interestingly, Fabbri et al. [30] demonstrated recently that miRNAs are not only targets but also regulators of DNA methylation. They reported that DNMT3A and DNMT3B are regulated by miR29 and that increased expression of miR-29 in lung cancer cells results in restoration of normal DNA methylation and expression patterns of the silenced TSGs FHIT and WWOX.
4 DNA methylation of single or small numbers of genes in lung cancer patients
Different methods were developed to determine the methylation status of genes. The most frequently used methods include bisulfite genomic sequencing and methylation-specific polymerase chain reaction (MSP) [31–33]. In addition, quantitative real-time polymerase chain reaction (RT-PCR) approaches are being used [31, 34, 35]. These methods are based on the treatment of genomic DNA with sodium bisulfite which converts unmethylated cytosine bases into uracil while methylated cytosines remain unchanged. Subsequently, methylation can be detected using specific primer sets for either methylated or unmethylated target sites.
To date, a large number of genes have been investigated for their methylation status in lung cancer patients. Most of the studies investigated surgically obtained tumor samples from NSCLC patients. Genes which were intensively studied in primary NSCLC samples include p16, RASSF1A, APC, RARβ-2, CDH1, CDH13, DAPK, and MGMT [36–43]. While p16 and RASSF1A are involved in cell cycle regulation, CDH1 and CDH13 play a role in regulation of cell adhesion, APC inhibits β-catenin, RARβ-2 is involved in growth regulation, DAPK acts in regulation of apoptosis and MGMT functions in DNA repair. Other genes investigated in primary NSCLCs include for instance ASC/TMS1, FHIT, hSRBC, TSLC1, and DAL-1 [44–49]. ASC/TMS1 and FHIT have pro-apoptotic function, hSRBC is a cell cycle regulator and TSLC1 and DAL-1 are putative TSGs involved in a cell adhesion cascade. The frequencies of methylation of these genes in primary NSCLCs were detected in up to 96%. Details about some of these genes are shown in Table 1.
A majority of the studies also investigated if methylation of certain genes occurs in corresponding non-malignant lung tissue samples. It has been shown that most genes were found to be methylated only in tumors but not at all or only in a very low percentage in the corresponding non-malignant lung tissues [38, 41, 43, 47]. Overall, these findings demonstrate that methylation of cancer-related genes is a frequent event in the pathogenesis of lung cancer.
However, DNA methylation cannot only be detected in primary tumors but also in blood, sputum, bronchial brushings, and bronchioloalveolar lavage (BAL) samples from lung cancer patients. A major advantage is that these specimens can be obtained in a non-invasive or minimally invasive way. Methylation in blood can usually only be detected if the primary tumor is methylated. Sputum can be obtained from smokers. BAL samples represent lung-specific material, however, a bronchoscopy needs to be performed. Overall, detecting methylation in these samples is potentially useful for predicting disease recurrence after surgery, for monitoring response to therapy, and for early detection of lung cancer.
Numerous manuscripts have been published which describe the frequency of methylation of genes (for instance MGMT, p16, DAPK, APC, CDH13, FHIT, RARβ-2, RASSF1A) in serum or plasma samples and in the corresponding primary tumors [50–56]. In most studies, the frequencies of methylation in serum or plasma samples were lower compared to primary tumors. Usually, methylation in serum or plasma samples was detected only if the corresponding tumors were methylated [50, 52]. In the majority of the studies, methylation was not detected in serum or plasma samples from healthy controls [51, 54].
Several genes were found to be methylated in sputum samples from lung cancer patients. These genes include hMLH1, p16, CDH13, DAPK, LAMC2, MGMT, PAX5ά PAX5β, RASSF1A [55–59]. Another gene which is frequently methylated in NSCLC patients is the ASC/TMS1 gene. Interestingly, ASC/TMS1 methylation was observed in 14% of primary tumors and in 17% of sputum samples of stage I NSCLC patients, but in 60% of primary tumors and in 41% of sputum samples of later stages NSCLC patients suggesting that methylation of ASC/TMS1 is a marker for late-stage lung cancer [60]. The majority of studies describe similar or slightly lower frequencies of methylation of certain genes in sputum samples compared to primary tumors.
Methylation of eight genes was evaluated in different kinds of material including primary tumor biopsies, serum, and sputum samples from stage III NSCLC patients [59]. The prevalence for methylation of the genes in sputum was similar to that seen in tumors, but was higher than that in serum. The authors conclude that sputum can be used as a surrogate for tumor tissue to predict the methylation status of NSCLCs. Shivapurkar et al. [61] analyzed the methylation status of 11 genes in primary NSCLCs, in adjacent non-malignant lung tissues, in peripheral blood mononuclear cells from cancer-free patients, in sputum samples of cancer patients and in controls. The gene 3-OST-2 showed the highest levels of methylation in tumors combined with lowest levels of methylation in control tissues. Moreover, the authors reported that quantitative analysis of 3-OST-2, RASSF1A, p16, and APC methylation in sputum samples may have excellent potential as a biomarker assay for lung cancer patients.
Methylation of genes, for instance, p16, APC, RASSF1A, CDH13, DAPK, MGMT, RASSF1A, and RARβ-2, was also detected in bronchial lavage samples [56, 62–66]. However, the frequency of methylation in this kind of samples was usually lower compared to the frequencies in primary tumors and sputum samples. In addition, the frequencies of methylation were lower in peripherally located tumors compared to centrally located tumors [62].
Recently, Han et al. [67] investigated methylation of DAPK, RASSF1A, and PAX5β in exhaled breath condensate from lung cancer patients and control individuals. Both groups contained current, former, and never smokers. The RASSF1A methylation density was different regarding the smoking status. Former smokers had higher methylation density versus never smokers and current smokers. Methylation of specific CpG sites of DAPK and PAX5β promoter were associated with lung cancer status. The authors concluded that DNA methylation can be detected in exhaled breath condensate, however, additional studies are necessary to define the potential impact of this observation.
5 Differences in the methylation patterns due to histology and genetic abnormalities
Toyooka et al. [68] investigated the frequencies of methylation of eight genes in a large number of lung tumors consisting of small cell lung cancers (SCLC), NSCLCs and bronchial carcinoids. The frequencies of methylation of APC, CDH13 and p16 were significantly higher in NSCLCs compared to neuroendocrine tumors. On the other hand, the frequencies of methylation of the genes RASSF1A, CDH1 and RARβ-2 were higher in SCLCs compared to carcinoids. In addition, the methylation index, a parameter which reflects the methylation status of all genes tested, was significantly higher in SCLCs compared to carcinoids. However, differences in the frequencies of methylation were seen not only between NSCLCs, SCLCs and carcinoids, but also between NSCLC subtypes. The frequencies of APC and CDH13 methylation were significantly higher in adenocarcinomas compared to squamous cell carcinomas. On the contrary, methylation of p16 was more frequently detected in squamous cell carcinomas compared to adenocarcinomas. Methylation of nine genes including p16, CDH1, TIMP3, RASSF1A, FHIT, APC, DAPK, MGMT, and GSTP1 was analyzed in primary NSCLCs by Gu et al. [69]. Overall, the methylation index was significantly higher in adenocarcinomas than in squamous cell carcinomas. A comparison of the methylation pattern of 47 gene-promoter regions between primary NSCLCs and corresponding adjacent normal lung tissues was performed by Ehrich et al. [70]. Six genes with statistically significant differences in methylation between tumor and normal tissue were identified. Moreover, using a hierarchical clustering algorithm normal lung could be distinguished from lung cancer tissue with >95% sensitivity and specificity. These studies indicate that DNA methylation is tissue type-specific. Distinct methylation signatures distinguish the major histologic lung cancer types and also tumor tissues from non-malignant lung tissues.
Differences in the methylation patterns were also found between lung tumors with certain genetic abnormalities. Toyooka et al. [71] reported that in lung adenocarcinomas the probability of having EGFR mutations was significantly lower among tumors with p16 and CDH13 methylation than in tumors without. In addition, the methylation index was significantly lower in EGFR mutant cases compared to wild-type cases. In contrast, the frequency of K-RAS mutations was significantly higher in p16 methylated cases than in unmethylated cases. The methylation index was marginally higher in K-RAS mutant cases compared to wild-type cases. Overall, the authors concluded that there are differences in the involvement of methylation between EGFR- and K-RAS-mediated tumorigenesis. These findings suggest that genetic and epigenetic changes interact systematically to promote tumorigenesis of lung adenocarcinomas.
6 DNA methylation as biomarker
In general, detecting methylation of certain genes seems to be a powerful biomarker for prognosis of patients, disease recurrence, early detection and risk assessment. In addition, it also might be a therapeutic target. However, there are some open questions and limitations which still need to be discussed, e.g., is there sufficient tumor DNA in blood, especially in early stages of disease? Tumor markers in blood are not organ-specific and DNA methylation detected in blood may indicate a tumor other than lung cancer. Satisfactory sputum specimens can usually only be obtained from smokers but not from non-smokers or long-term former smokers. Therefore, this sample type is useful only for a subgroup of patients. Sputum which is obtained from the central lung, however, may not detect peripheral tumors. Another point is that there are several methods by which methylation can be detected and the question is which of them is optimal. Moreover, it has been shown that the sensitivity and the specificity are higher if panel of markers are investigated rather than single genes. However, which panel of markers has the highest sensitivity and specificity? In the following sections, we describe findings about how DNA methylation can function as a biomarker.
6.1 DNA methylation markers with unfavorable prognostic relevance
A worse prognostic impact of methylation in primary tumors of NSCLC patients has been reported for several genes. Several years ago, Tang et al. [72] reported that methylation of DAPK is strongly associated with survival in stage I NSCLC patients. This finding was confirmed later by Lu et al. [73]. Methylation of RASSF1A as unfavorable prognostic parameter in NSCLC patients was reported by Burbee et al. [43]. Similar findings were observed by Kim et al. [74] and by Tomizawa et al. [75] in patients with stage I lung adenocarcinomas. Toyooka et al. [76] reported that p16 methylation was significantly related to unfavorable prognosis in patients with lung adenocarcinomas. Two genes (RASSF1A, RUNX3) out of ten tested were identified to be of poor prognostic relevance when they were found to be methylated in stage I NSCLC patients [77]. The prognostic impact of methylation of five genes located at the chromosomal region 3p was studied by Seng et al. [78] in primary NSCLCs. Interestingly, methylation of DLEC1 was found to be an independent marker of poor survival in both the whole study population as well as in patients with squamous cell carcinomas. hMLH1 methylation was also of prognostic impact particularly in patients with large cell carcinomas. Concordant methylation of DLEC1 and hMLH1 was a strong indicator for poor prognosis in the whole study population. Genes with a poor prognostic relevance are listed in Table 1.
6.2 DNA methylation markers relevant for disease recurrence
A potential association between DNA methylation in the blood and disease recurrence in stage I NSCLC patients after surgery was investigated by Brock et al. [79]. Methylation of the genes p16, MGMT, DAPK, RASSF1A, CDH13, APC, and ASC was determined in tumor and corresponding lymph nodes of the patients. Interestingly, methylation of p16, CDH13, RASSF1A, and APC in tumors and in histologically tumor-negative lymph nodes was associated with disease recurrence. If p16 and CDH13 were methylated in tumor and mediastinal lymph nodes, the odds ratio for recurrence was 15.5. The authors discussed that methylation of these genes in histologically normal lymph nodes probably indicates the presence of microscopically undetectable micrometastasis. Because the current method of risk assessment for recurrence in stage I NSCLC patients is imprecise, the authors suggested that detection of methylation of certain genes may identify cells with a potential for metastatic spread and could be helpful to predict disease recurrence.
6.3 Smoking, early detection of lung cancer, and risk assessment
Smoking is the major risk factor for lung cancer and about 85-90% of all lung cancers occur in smokers [80]. In addition to the major genetic differences between lung cancers arising in smokers and never smokers [81], it has been shown that differences in the frequencies of methylation of certain genes occur in these two types of lung cancer. Toyooka et al. [82] investigated the methylation status of seven genes in a large number of lung adenocarcinomas. The authors observed higher rates of methylation of p16 and APC as well as a significantly higher methylation index in ever smokers compared to never smokers. In addition, the mean methylation index of tumors arising in former smokers was significantly lower than the mean of current smokers. A comparison of methylation of certain genes in bronchial epithelium between current and former smokers was performed by Belinsky et al. [83]. Interestingly, p16 and DAPK methylation was found frequently in bronchial epithelium from current smokers as from former smokers, suggesting that methylation changes can persist long after smoking cessation.
Recently, researchers tried to elucidate the interaction of tobacco carcinogens and DNA methylation during lung carcinogenesis. Damiani et al. [84] developed an in vitro cell transformation model to identify critical mediators of premalignancy. Using this model, they observed increased levels of DNMT1 protein during exposure of immortalized bronchial epithelial cells to the carcinogens methyl-nitrosurea, benzo(a)pyrene-diolepoxide 1 or both, and an association with methylation of certain genes. The authors conclude that carcinogen-induced methylation is mediated by DNMT1 and is causal for transformation of immortalized bronchial epithelial cells. In addition, it has been shown that carcinogens induce single- and double-strand breaks (DSB) in DNA and reduced capacity for repair of DNA damage has been associated with lung cancer [85]. Thus, Leng et al. [86] investigated the hypothesis that DSB repair capacity and sequence variations in genes involved in this pathway are associated with a high methylation index in current and former cancer-free smokers. A 50% reduction in the mean level of DSB repair capacity was observed in lymphocytes from smokers with a high methylation index compared to smokers with no genes methylated. In addition, single nucleotide polymorphisms within several genes were highly associated with the methylation index. Overall, this study identified an association between DSB DNA repair capacity and alterations of certain genes within this pathway as an important factor for gene methylation in sputum, which finally results in an increased risk for lung cancer.
It has been observed that methylation of certain genes can already be detected in aerodigestive tract samples from cancer-free smokers. Methylation of several genes, for instance p16, DAPK, GSTP1, and RASSF1A, were detected in different sample types from this population [83, 87–90]. In most studies, the frequency of methylation of these genes was significantly lower in samples from cancer-free smokers compared to lung tumors. Methylation of four genes was analyzed in oropharyngeal brushes, sputum samples, bronchial brushes, and BAL samples of the upper aerodigestive tract epithelium from heavy smokers without evidence of cancer but with morphometric evidence of sputum atypia [91]. Interestingly, methylation occurred more frequently in samples from the central airways (sputum, bronchial brushes) compared to the peripheral airways (BAL), and only occasionally in the oropharynx. RARβ-2 was the most frequently methylated gene. Results from sputum samples and bronchial brushes were comparable.
Bhutani et al. [92] investigated if the oral epithelium may be used as a surrogate tissue for assessing tobacco-induced molecular alterations in the lungs. They determined the methylation status of p16 and FHIT in oral and bronchial brush specimens from 127 smokers enrolled in a chemoprevention trial. Specimens were obtained at baseline and three months after intervention. At baseline, methylation in bronchial tissue was present in 23% of samples for p16 and in 17% of samples for FHIT. Comparable results were found for oral tissue. Of these samples, 19% were p16 methylated and 15% were FHIT methylated. The mean bronchial methylation index was far higher in patients with oral tissue methylation compared to patients without oral tissue methylation. Similar findings were observed after three months. The authors conclude that oral epithelium has some potential as surrogate tissue for assessing tobacco-induced molecular damage in the lungs.
Methylation of seven genes was studied in atypical adenomatous hyperplasia (AAH) of the lung (a putative preneoplastic lesion), adjacent normal lung tissue, and in synchronous lung adenocarcinomas [93]. The authors observed a significant increase in the frequency of methylation during the histologic progression from normal to AAH and finally to adenocarcinoma. Multifocal AAHs had distinct patterns of methylation suggesting divergent epigenetic field defects.
Using a highly sensitive PCR approach, Palmisano et al. [94] found methylation of p16 and/or MGMT in DNA from sputum in 100% of patients with squamous cell lung carcinomas up to three years before clinical diagnosis. The prevalence for methylation of several genes in sputum and plasma samples from women at different risk for lung cancer (lung cancer survivors, clinically cancer-free smokers, and never smokers) was investigated by Belinsky et al. [55]. The authors reported that concomitant methylation of multiple genes in sputum is strongly associated with lung cancer risk. In another project from this research group, six genes were identified which are associated with a 50% increased lung cancer risk in a high-risk cohort [58]. The concomitant methylation of three or more of these six genes was associated with a 6.5-fold increased risk for lung cancer.
Overall, these results demonstrate that methylation of certain genes can be an early event in the pathogenesis of lung cancer and that methylation can increase during malignant progression. A panel of certain methylated genes may be useful as biomarker for early detection of lung cancer and risk assessment. In addition, oral epithelium has some potential to serve as surrogate tissue for tobacco-induced methylation changes in the lungs.
7 DNA methylation as potential therapeutic target
While methylation of single genes cannot be reversed at the present time, global approaches for demethylation are currently being tested as therapeutic modalities. The effect of therapies that reverse epigenetic changes in certain malignant diseases has been reported in the past few years. DNMT inhibitors such as Aza-dC or 5-azacytidine (AzaC) have been approved by the Food and Drug Administration of the USA in myelodysplatic syndrome [95, 96], whereas HDAC inhibitors have shown to be active in cutaneous and peripheral T-cell lymphoma [97, 98]. However, in general, the efficacies of these drugs in solid tumors are disappointing. Several reasons for these findings have been discussed: it is not precisely understood how these drugs work, it is not known how to optimally select patients for this therapy, and it is not clear how to find the optimal dosing. Thus, we know very little about the actual or potential efficacy of these drugs in lung cancer. Recently, Juergens et al. [99] reported preliminary results from a phase II study investigating the efficacy of AzaC in combination with the HDAC inhibitor entinostat in pretreated NSCLC patients. Interestingly, the authors observed a benefit from this treatment in several patients. Overall, this treatment was well tolerated. However, additional clinical studies are necessary for complete evaluation.
8 High throughput DNA methylation profiling in lung cancer
Recently, a new era in detecting DNA methylation has begun by performing genome-wide methylation analysis. Since it has been reported by Costello et al. [100] that on average 600 CpG islands are targets for methylation in tumor cells, researchers have expressed increasing interest in developing techniques to investigate not only methylation of single genes but also large-scale and genome-wide analyses of methylation. The first approach for large-scale DNA methylation analysis of CGIs was Restriction Landmark Genomic Scanning (RLGS). By RLGS, Dai et al. [101] analyzed 1,184 CGIs and discovered 11 genes which are differentially methylated in lung cancer compared to matching non-malignant lung tissue samples. Two of these genes (GNAL and BMP3B) were methylated in more than 50% of tumors analyzed. Brena et al. [102] used RLGS to identify methylated genes in lung adeno- and squamous cell carcinomas. They found a 47-gene methylation signature which could distinguish these two histologic subgroups. In addition, they identified methylation of OLIG1 as new prognostic parameter for lung cancer patients.
Other frequently used assays for high throughput methylation analysis include expression microarray analysis of cell lines before and after treatment with DNMT inhibitors +/− HDAC inhibitors (“pharmacological re-activation”), BeadArray-based methylation analysis of a panel of cancer-related genes (Illumina GoldenGate™ methylation assay), and microarray analysis in combination with immunoprecipitation of methylated DNA (5-methylcytidine antibody; MeDIP-chip) [103]. These methods are discussed in detail below.
8.1 Gene-expression profiling of pharmacologically re-activated genes
As already mentioned above, DNA methylation is reversible by the use of DNMT inhibitors such as Aza-dC. In vitro, Aza-dC has been shown to efficiently induce re-expression of genes silenced by methylation [41, 43, 45, 47, 104]. Moreover, a synergistic effect in gene re-expression between Aza-dC and the HDAC inhibitor TSA has been reported [20]. Based on this observation, an indirect approach for the identification of a broad range of so far unknown methylated genes has been developed [105]. In this assay, gene expression patterns of cells treated with Aza-dC and/or TSA and of untreated counterparts are compared by expression microarray analysis. Using a similar approach, Shames et al. [106] compared gene expression in seven NSCLC and three human bronchial epithelial cell lines before and after Aza-dC treatment. They identified 132 tumor-specifically methylated genes and some of them were involved in specific molecular pathways whose loss may contribute to the development of a malignant phenotype. Thirty-one out of 45 genes which were further investigated by MSP in primary lung carcinomas and matching non-malignant lung tissue samples were found to be methylated only in tumors. Out of these seven genes with potential clinical relevance, ALDH1A3, BNC1, CCNA1, CTSZ, LOX, MSX1, and NRCAM were identified. Similarly, Zhong et al. [107] performed gene-expression microarray analysis of nine NSCLC cell lines before and after treatment with Aza-dC and TSA and identified 214 genes whose expression was up-regulated after drug treatment. Interestingly, some of these genes (NNAT, RASGRP2, MT3, ATF3, and CYLD) have strong growth suppressing properties suggesting an important role of these genes in lung cancer pathogenesis. A summary of results from these two studies are shown in Table 2.
Overall, these data suggest that gene expression microarray analysis in cells treated with demethylating agents is a powerful approach to identify unknown methylated genes. A major disadvantage of this assay is that it can be performed only in cell lines but not in tissue samples. Because it has been suggested that cancer cell lines can acquire methylation in culture, cell lines may not accurately reflect the methylation patterns of tumors in vivo [108, 109]. Therefore, it is necessary to validate data obtained from cancer cell lines in tumor samples of patients.
8.2 High throughput DNA methylation analysis using BeadArrays
As described above, treatment of genomic DNA with sodium bisulfite converts unmethylated cytosine bases to uracil while 5-methylcytosine remains unchanged. Specific primers can then distinguish whether the sequence of interest is methylated or not. Using a sodium bisulfite conversion approach combined with the BeadArray technology (Illumina GoldenGate™ methylation assays), Bibikova et al. [110] analyzed methylation profiles of 1,536 CpG sites from 371 cancer-related genes in cancer cell lines of different histologic types and in primary lung adenocarcinomas and non-malignant lung tissue samples. Several genes of potential clinical interest including ASCL2, CDH13, HOXA5, HOXA11, NPY, RUNX3, TERT, and TP73 were identified and further examined by bisulfite genomic sequencing. Moreover, the authors demonstrated that methylation signatures were different between the cancer cell lines of various histologic types. In addition, 55 CpG sites were found which methylation pattern distinguished lung carcinomas from non-malignant lung tissue samples [110]. Similar results were reported by Christensen et al. [111]. This research group analyzed lung adenocarcinomas, mesotheliomas, and non-malignant lung tissue samples for methylation of 1,413 CpG loci associated with 773 cancer-related genes using Illumina’s GoldenGate™ methylation assay. Differences in methylation patterns were found between lung tumors and non-malignant lung tissue samples and between lung adenocarcinomas and mesotheliomas.
Recently, Illumina released the new generation of BeadArrays for assessment of DNA methylation where 27,578 CpG loci covering more than 14,000 genes can be analyzed at single-nucleotide resolution. However, data derived from this new generation of BeadArrays have not been reported yet.
8.3 Methylome profiling using tiling microarrays
In the last few years, tiling arrays were developed which are a powerful tool to perform whole-genome analysis including DNA methylation [112]. Recently, Weber et al. [103] established a strategy to isolate methylated DNA fragments by immunoprecipitation (MeDIP) using an anti-5-methylcytidine antibody. Moreover, they combined the MeDIP assay with the microarray technology (MeDIP-chip). Using this approach in SW48 colon carcinoma cells and normal colon mucosa, they obtained genome-wide DNA methylation patterns and identified about 200 genes which are differently methylated between these sample types [103]. However, MeDIP-chip analysis in lung cancer has not been reported yet. A very similar approach is the methylated CpG island recovery assay, which isolates methylated DNA fragments using recombinant MBD2b/MBD3L proteins. These fragments are subsequently hybridized to microarrays. Using this approach, Rauch et al. [113] investigated the methylation status of 27,800 CGIs in primary tumors of five patients with stage I lung squamous cell carcinomas. With a range from 216 to 848 methylated CGIs, the number of methylated genes in this study is in concordance with previously reported data [100]. Further methylation analysis in 20 primary squamous cell carcinomas and non-malignant lung tissue samples identified eight genes (OTX1, BARHL2, MEIS1, OC2, PAX6, IRX2, TFAP2A, and EVX2) which might be of clinical interest for these patients. Results from high throughput DNA methylation analysis are summarized in Table 3.
In conclusion, using these high throughput approaches, hundreds of genes were found to be methylated in tumors. Methylation patterns between tumors and corresponding non-malignant lung tissues and also between tumors of various histologies are different confirming previously obtained data that DNA methylation is tissue type-specific. In addition, a large number of genes unknown to be methylated in lung cancer so far were identified.
However, it seems to be of interest that genes which are known to be frequently methylated in lung cancer by single-gene methylation analysis are infrequently found to be methylated by these high-throughput approaches. Well-known methylated genes originally identified from single-gene analysis were also found to be methylated by genome-wide approaches (CDH1 and TIMP3 [106], FHIT [111], p16 [113], SLIT2 [111, 113] and CDH13 [110]). Although Bibikova et al. [110] observed methylation of RASSF1A and ASC/TMS1 in a panel of lung cancer cell lines, methylation of these genes was not detected in primary lung adenocarcinomas. A possible explanation for differences in results obtained by single-gene and high-throughput DNA methylation approaches could be that, so far, mainly small numbers of samples were tested using high-throughput analyses. Other problems with genomic approaches include a continuous scale, as opposed to plus or minus values, multiple probes for single genes which often result in different answers and multiple isoforms, not all of which are analyzed. Further studies addressing these issues are necessary.
9 Summary
Numerous genes have been identified which are involved in different pathways relevant for lung cancer pathogenesis and which are frequently methylated leading together with other epigenetic mechanisms to gene silencing. In addition, it has been shown that methylation of certain genes can be of clinical relevance. While at the beginning of the DNA methylation era mainly single genes were investigated for their methylation status, recently, methods for high-throughput DNA methylation analysis were developed. These approaches provide researchers with a huge amount of information about methylation patterns in lung cancer cell lines and samples from lung cancer patients. However, because of differences in methodology, it is difficult to compare results obtained from these studies. In addition, most studies have analyzed relatively small numbers of samples. Thus, which high throughput approach is the most suitable one needs to be evaluated. It can be hypothesized that using such methods, hundreds of genes could be identified which are of clinical interest. For instance, disease recurrence in NSCLC patients after surgery could be probably much more precisely predicted than today. It could also be helpful in selecting patients who may benefit from an epigenetic therapy. Overall, findings suggest that high throughput methylation profiling could be of enormous help to improve prognosis of lung cancer patients.
9.1 Key unanswered questions
The key unanswered question remains whether it is possible to translate the findings from global methylation profiling to patient care—diagnosis, prognosis, and therapy.
References
Travis, W. D., Travis, L. B., & Devesa, S. S. (1995). Lung cancer. Cancer, 75, 191–202.
Shogren-Knaak, M., Ishii, H., Sun, J. M., Pazin, M. J., Davie, J. R., & Peterson, C. L. (2006). Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science, 311, 844–847.
Vettese-Dadey, M., Grant, P. A., Hebbes, T. R., Crane- Robinson, C., Allis, C. D., & Workman, J. L. (1996). Acetylation of histone H4 plays a primary role in enhancing transcription factor binding to nucleosomal DNA in vitro. Embo Journal, 15, 2508–2518.
Esteller, M. (2008). Epigenetics in cancer. New England Journal of Medicine, 358, 1148–1159.
Espada, J., Ballestar, E., Fraga, M. F., Villar-Garea, A., Juarranz, A., Stockert, J. C., et al. (2004). Human DNA methyltransferase 1 is required for maintenance of the histone H3 modification pattern. Journal of Biological Chemistry, 279, 37175–37184.
Ikegami, K., Ohgane, J., Tanaka, S., Yagi, S., & Shiota, K. (2009). Interplay between DNA methylation, histone modification and chromatin remodeling in stem cells and during development. International Journal of Developmental Biology, 53, 203–214.
Eden, S., Hashimshony, T., Keshet, I., Cedar, H., & Thorne, A. W. (1998). DNA methylation models histone acetylation. Nature, 394, 842.
Rhee, I., Jair, K. W., Yen, R. W., Lengauer, C., Herman, J. G., Kinzler, K. W., et al. (2000). CpG methylation is maintained in human cancer cells lacking DNMT1. Nature, 404, 1003–1007.
Gowher, H., & Jeltsch, A. (2001). Enzymatic properties of recombinant Dnmt3a DNA methyltransferase from mouse: the enzyme modifies DNA in a non-processive manner and also methylates non-CpG [correction of non-CpA] sites. Journal of Molecular Biology, 309, 1201–1208.
Okano, M., Xie, S., & Li, E. (1998). Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nature Genetics, 19, 219–220.
Matsuo, K., Clay, O., Takahashi, T., Silke, J., & Schaffner, W. (1993). Evidence for erosion of mouse CpG islands during mammalian evolution. Somatic Cell and Molecular Genetics, 19, 543–555.
Wang, Y., & Leung, F. C. (2004). An evaluation of new criteria for CpG islands in the human genome as gene markers. Bioinformatics, 20, 1170–1177.
Lander, E. S., Linton, L. M., Birren, B., Nusbaum, C., Zody, M. C., Baldwin, J., et al. (2001). Initial sequencing and analysis of the human genome. Nature, 409, 860–921.
Venter, J. C., Adams, M. D., Myers, E. W., Li, P. W., Mural, R. J., Sutton, G. G., et al. (2001). The sequence of the human genome. Science, 291, 1304–1351.
Bird, A. P. (1986). CpG-rich islands and the function of DNA methylation. Nature, 321, 209–213.
Gardiner-Garden, M., & Frommer, M. (1987). CpG islands in vertebrate genomes. Journal of Molecular Biology, 196, 261–282.
Razin, A., & Cedar, H. (1994). DNA methylation and genomic imprinting. Cell, 77, 473–476.
Jones, P. A., & Baylin, S. B. (2002). The fundamental role of epigenetic events in cancer. Nature Reviews. Genetics, 3, 415–428.
Fraga, M. F., Ballestar, E., Villar-Garea, A., Boix-Chornet, M., Espada, J., Schotta, G., et al. (2005). Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nature Genetics, 37, 391–400.
Cameron, E. E., Bachman, K. E., Myohanen, S., Herman, J. G., & Baylin, S. B. (1999). Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nature Genetics, 21, 103–107.
Calin, G. A., & Croce, C. M. (2006). MicroRNA signatures in human cancers. Nature Reviews. Cancer, 6, 857–866.
He, L., & Hannon, G. J. (2004). MicroRNAs: small RNAs with a big role in gene regulation. Nature Reviews. Genetics, 5, 522–531.
Eulalio, A., Huntzinger, E., & Izaurralde, E. (2008). Getting to the root of miRNA-mediated gene silencing. Cell, 132, 9–14.
Bueno, M. J., Perez de Castro, I., Gomez de Cedron, M., Santos, J., Calin, G. A., Cigudosa, J. C., et al. (2008). Genetic and epigenetic silencing of microRNA-203 enhances ABL1 and BCR-ABL1 oncogene expression. Cancer Cell, 13, 496–506.
Brueckner, B., Stresemann, C., Kuner, R., Mund, C., Musch, T., Meister, M., et al. (2007). The human let-7a-3 locus contains an epigenetically regulated microRNA gene with oncogenic function. Cancer Research, 67, 1419–1423.
Fazi, F., Racanicchi, S., Zardo, G., Starnes, L. M., Mancini, M., Travaglini, L., et al. (2007). Epigenetic silencing of the myelopoiesis regulator microRNA-223 by the AML1/ETO oncoprotein. Cancer Cell, 12, 457–466.
Lujambio, A., Ropero, S., Ballestar, E., Fraga, M. F., Cerrato, C., Setien, F., et al. (2007). Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Research, 67, 1424–1429.
Lodygin, D., Tarasov, V., Epanchintsev, A., Berking, C., Knyazeva, T., Korner, H., et al. (2008). Inactivation of miR-34a by aberrant CpG methylation in multiple types of cancer. Cell Cycle, 7, 2591–2600.
Corney, D. C., Flesken-Nikitin, A., Godwin, A. K., Wang, W., & Nikitin, A. Y. (2007). MicroRNA-34b and MicroRNA-34c are targets of p53 and cooperate in control of cell proliferation and adhesion-independent growth. Cancer Research, 67, 8433–8438.
Fabbri, M., Garzon, R., Cimmino, A., Liu, Z., Zanesi, N., Callegari, E., et al. (2007). MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proceedings of the National Academy of Sciences of the United States of America, 104, 15805–15810.
Shames, D. S., Minna, J. D., & Gazdar, A. F. (2007). Methods for detecting DNA methylation in tumors: from bench to bedside. Cancer Letters, 251, 187–198.
Herman, J. G., & Baylin, S. B. (2001). Methylation-specific PCR. Current Protocols in Human Genetics, Chapter 10, Unit 10 16.
Herman, J. G., Graff, J. R., Myohanen, S., Nelkin, B. D., & Baylin, S. B. (1996). Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proceedings of the National Academy of Sciences of the United States of America, 93, 9821–9826.
Fraga, M. F., & Esteller, M (2002). DNA methylation: a profile of methods and applications. Biotechniques, 33, 632, 634, 636-649.
Campan, M., Weisenberger, D. J., Trinh, B., & Laird, P. W. (2009). MethyLight. Methods in Molecular Biology, 507, 325–337.
Dammann, R., Li, C., Yoon, J. H., Chin, P. L., Bates, S., & Pfeifer, G. P. (2000). Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3. Nature Genetics, 25, 315–319.
Esteller, M., Corn, P. G., Baylin, S. B., & Herman, J. G. (2001). A gene hypermethylation profile of human cancer. Cancer Research, 61, 3225–3229.
Zöchbauer-Müller, S., Fong, K. M., Virmani, A. K., Geradts, J., Gazdar, A. F., & Minna, J. D. (2001). Aberrant promoter methylation of multiple genes in non-small cell lung cancers. Cancer Research, 61, 249–255.
Virmani, A. K., Rathi, A., Sathyanarayana, U. G., Padar, A., Huang, C. X., Cunnigham, H. T., et al. (2001). Aberrant methylation of the adenomatous polyposis coli (APC) gene promoter 1A in breast and lung carcinomas. Clinical Cancer Research, 7, 1998–2004.
Virmani, A. K., Rathi, A., Zöchbauer-Müller, S., Sacchi, N., Fukuyama, Y., Bryant, D., et al. (2000). Promoter methylation and silencing of the retinoic acid receptor-beta gene in lung carcinomas. Journal of the National Cancer Institute, 92, 1303–1307.
Toyooka, K. O., Toyooka, S., Virmani, A. K., Sathyanarayana, U. G., Euhus, D. M., Gilcrease, M., et al. (2001). Loss of expression and aberrant methylation of the CDH13 (H-cadherin) gene in breast and lung carcinomas. Cancer Research, 61, 4556–4560.
Toyooka, S., Toyooka, K. O., Miyajima, K., Reddy, J. L., Toyota, M., Sathyanarayana, U. G., et al. (2003). Epigenetic down-regulation of death-associated protein kinase in lung cancers. Clinical Cancer Research, 9, 3034–3041.
Burbee, D. G., Forgacs, E., Zöchbauer-Müller, S., Shivakumar, L., Fong, K. M., Gao, B., et al. (2001). Epigenetic inactivation of RASSF1A in lung and breast cancers and malignant phenotype suppression. Journal of the National Cancer Institute, 93, 691–699.
Virmani, A., Rathi, A., Sugio, K., Sathyanarayana, U. G., Toyooka, S., Kischel, F. C., et al. (2003). Aberrant methylation of TMS1 in small cell, non small cell lung cancer and breast cancer. International Journal of Cancer, 106, 198–204.
Zöchbauer-Müller, S., Fong, K. M., Maitra, A., Lam, S., Geradts, J., Ashfaq, R., et al. (2001). 5′ CpG island methylation of the FHIT gene is correlated with loss of gene expression in lung and breast cancer. Cancer Research, 61, 3581–3585.
Zöchbauer-Müller, S., Fong, K. M., Geradts, J., Xu, X., Seidl, S., End-Pfutzenreuter, A., et al. (2005). Expression of the candidate tumor suppressor gene hSRBC is frequently lost in primary lung cancers with and without DNA methylation. Oncogene, 24, 6249–6255.
Heller, G., Fong, K. M., Girard, L., Seidl, S., End-Pfützenreuter, A., Lang, G., et al. (2006). Expression and methylation pattern of TSLC1 cascade genes in lung carcinomas. Oncogene, 25, 959–968.
Kikuchi, S., Yamada, D., Fukami, T., Masuda, M., Sakurai-Yageta, M., Williams, Y. N., et al. (2005). Promoter methylation of DAL-1/4.1B predicts poor prognosis in non-small cell lung cancer. Clinical Cancer Research, 11, 2954–2961.
Kikuchi, S., Yamada, D., Fukami, T., Maruyama, T., Ito, A., Asamura, H., et al. (2006). Hypermethylation of the TSLC1/IGSF4 promoter is associated with tobacco smoking and a poor prognosis in primary nonsmall cell lung carcinoma. Cancer, 106, 1751–1758.
Esteller, M., Sanchez-Cespedes, M., Rosell, R., Sidransky, D., Baylin, S. B., & Herman, J. G. (1999). Detection of aberrant promoter hypermethylation of tumor suppressor genes in serum DNA from non-small cell lung cancer patients. Cancer Research, 59, 67–70.
Usadel, H., Brabender, J., Danenberg, K. D., Jeronimo, C., Harden, S., Engles, J., et al. (2002). Quantitative adenomatous polyposis coli promoter methylation analysis in tumor tissue, serum, and plasma DNA of patients with lung cancer. Cancer Research, 62, 371–375.
Hsu, H. S., Chen, T. P., Hung, C. H., Wen, C. K., Lin, R. K., Lee, H. C., et al. (2007). Characterization of a multiple epigenetic marker panel for lung cancer detection and risk assessment in plasma. Cancer, 110, 2019–2026.
Fujiwara, K., Fujimoto, N., Tabata, M., Nishii, K., Matsuo, K., Hotta, K., et al. (2005). Identification of epigenetic aberrant promoter methylation in serum DNA is useful for early detection of lung cancer. Clinical Cancer Research, 11, 1219–1225.
Wang, Y., Yu, Z., Wang, T., Zhang, J., Hong, L., & Chen, L. (2007). Identification of epigenetic aberrant promoter methylation of RASSF1A in serum DNA and its clinicopathological significance in lung cancer. Lung Cancer, 56, 289–294.
Belinsky, S. A., Klinge, D. M., Dekker, J. D., Smith, M. W., Bocklage, T. J., Gilliland, F. D., et al. (2005). Gene promoter methylation in plasma and sputum increases with lung cancer risk. Clinical Cancer Research, 11, 6505–6511.
Anglim, P. P., Alonzo, T. A., & Laird-Offringa, I. A. (2008). DNA methylation-based biomarkers for early detection of non-small cell lung cancer: an update. Mol Cancer, 7, 81.
Wang, Y. C., Lu, Y. P., Tseng, R. C., Lin, R. K., Chang, J. W., Chen, J. T., et al. (2003). Inactivation of hMLH1 and hMSH2 by promoter methylation in primary non-small cell lung tumors and matched sputum samples. Journal of Clinical Investigation, 111, 887–895.
Belinsky, S. A., Liechty, K. C., Gentry, F. D., Wolf, H. J., Rogers, J., Vu, K., et al. (2006). Promoter hypermethylation of multiple genes in sputum precedes lung cancer incidence in a high-risk cohort. Cancer Research, 66, 3338–3344.
Belinsky, S. A., Grimes, M. J., Casas, E., Stidley, C. A., Franklin, W. A., Bocklage, T. J., et al. (2007). Predicting gene promoter methylation in non-small-cell lung cancer by evaluating sputum and serum. British Journal of Cancer, 96, 1278–1283.
Machida, E. O., Brock, M. V., Hooker, C. M., Nakayama, J., Ishida, A., Amano, J., et al. (2006). Hypermethylation of ASC/TMS1 is a sputum marker for late-stage lung cancer. Cancer Research, 66, 6210–6218.
Shivapurkar, N., Stastny, V., Suzuki, M., Wistuba, I. I., Li, L., Zheng, Y., et al. (2007). Application of a methylation gene panel by quantitative PCR for lung cancers. Cancer Letter, 247, 56–71.
Schmiemann, V., Bocking, A., Kazimirek, M., Onofre, A. S., Gabbert, H. E., Kappes, R., et al. (2005). Methylation assay for the diagnosis of lung cancer on bronchial aspirates: a cohort study. Clinical Cancer Research, 11, 7728–7734.
Grote, H. J., Schmiemann, V., Geddert, H., Rohr, U. P., Kappes, R., Gabbert, H. E., et al. (2005). Aberrant promoter methylation of p16(INK4a), RARB2 and SEMA3B in bronchial aspirates from patients with suspected lung cancer. International Journal of Cancer, 116, 720–725.
Grote, H. J., Schmiemann, V., Kiel, S., Bocking, A., Kappes, R., Gabbert, H. E., et al. (2004). Aberrant methylation of the adenomatous polyposis coli promoter 1A in bronchial aspirates from patients with suspected lung cancer. International Journal of Cancer, 110, 751–755.
Kim, H., Kwon, Y. M., Kim, J. S., Lee, H., Park, J. H., Shim, Y. M., et al. (2004). Tumor-specific methylation in bronchial lavage for the early detection of non-small-cell lung cancer. Journal of Clinical Oncology, 22, 2363–2370.
Chan, E. C., Lam, S. Y., Tsang, K. W., Lam, B., Ho, J. C., Fu, K. H., et al. (2002). Aberrant promoter methylation in Chinese patients with non-small cell lung cancer: patterns in primary tumors and potential diagnostic application in bronchoalevolar lavage. Clinical Cancer Research, 8, 3741–3746.
Han, W., Wang, T., Reilly, A. A., Keller, S. M., & Spivack, S. D. (2009). Gene promoter methylation assayed in exhaled breath, with differences in smokers and lung cancer patients. Respiratory Research, 10, 86.
Toyooka, S., Toyooka, K. O., Maruyama, R., Virmani, A. K., Girard, L., Miyajima, K., et al. (2001). DNA methylation profiles of lung tumors. Mol Cancer Therapeutics, 1, 61–67.
Gu, J., Berman, D., Lu, C., Wistuba, I. I., Roth, J. A., Frazier, M., et al. (2006). Aberrant promoter methylation profile and association with survival in patients with non-small cell lung cancer. Clinical Cancer Research, 12, 7329–7338.
Ehrich, M., Field, J. K., Liloglou, T., Xinarianos, G., Oeth, P., Nelson, M. R., et al. (2006). Cytosine methylation profiles as a molecular marker in non-small cell lung cancer. Cancer Research, 66, 10911–10918.
Toyooka, S., Tokumo, M., Shigematsu, H., Matsuo, K., Asano, H., Tomii, K., et al. (2006). Mutational and epigenetic evidence for independent pathways for lung adenocarcinomas arising in smokers and never smokers. Cancer Research, 66, 1371–1375.
Tang, X., Khuri, F. R., Lee, J. J., Kemp, B. L., Liu, D., Hong, W. K., et al. (2000). Hypermethylation of the death-associated protein (DAP) kinase promoter and aggressiveness in stage I non-small-cell lung cancer. Journal of the National Cancer Institute, 92, 1511–1516.
Lu, C., Soria, J. C., Tang, X., Xu, X. C., Wang, L., Mao, L., et al. (2004). Prognostic factors in resected stage I non-small-cell lung cancer: a multivariate analysis of six molecular markers. Journal of Clinical Oncology, 22, 4575–4583.
Kim, D. H., Kim, J. S., Ji, Y. I., Shim, Y. M., Kim, H., Han, J., et al. (2003). Hypermethylation of RASSF1A promoter is associated with the age at starting smoking and a poor prognosis in primary non-small cell lung cancer. Cancer Research, 63, 3743–3746.
Tomizawa, Y., Kohno, T., Kondo, H., Otsuka, A., Nishioka, M., Niki, T., et al. (2002). Clinicopathological significance of epigenetic inactivation of RASSF1A at 3p21.3 in stage I lung adenocarcinoma. Clinical Cancer Research, 8, 2362–2368.
Toyooka, S., Suzuki, M., Maruyama, R., Toyooka, K. O., Tsukuda, K., Fukuyama, Y., et al. (2004). The relationship between aberrant methylation and survival in non-small-cell lung cancers. British Journal of Cancer, 91, 771–774.
Yanagawa, N., Tamura, G., Oizumi, H., Kanauchi, N., Endoh, M., Sadahiro, M., et al. (2007). Promoter hypermethylation of RASSF1A and RUNX3 genes as an independent prognostic prediction marker in surgically resected non-small cell lung cancers. Lung Cancer, 58, 131–138.
Seng, T. J., Currey, N., Cooper, W. A., Lee, C. S., Chan, C., Horvath, L., et al. (2008). DLEC1 and MLH1 promoter methylation are associated with poor prognosis in non-small cell lung carcinoma. British Journal of Cancer, 99, 375–382.
Brock, M. V., Hooker, C. M., Ota-Machida, E., Han, Y., Guo, M., Ames, S., et al. (2008). DNA methylation markers and early recurrence in stage I lung cancer. New England Journal of Medicine, 358, 1118–1128.
Parkin, D. M., Pisani, P., Lopez, A. D., & Masuyer, E. (1994). At least one in seven cases of cancer is caused by smoking. Global estimates for 1985. International Journal of Cancer, 59, 494–504.
Sun, S., Schiller, J. H., & Gazdar, A. F. (2007). Lung cancer in never smokers—a different disease. Nature Reviews Cancer, 7, 778–790.
Toyooka, S., Maruyama, R., Toyooka, K. O., McLerran, D., Feng, Z., Fukuyama, Y., et al. (2003). Smoke exposure, histologic type and geography-related differences in the methylation profiles of non-small cell lung cancer. International Journal of Cancer, 103, 153–160.
Belinsky, S. A., Palmisano, W. A., Gilliland, F. D., Crooks, L. A., Divine, K. K., Winters, S. A., et al. (2002). Aberrant promoter methylation in bronchial epithelium and sputum from current and former smokers. Cancer Research, 62, 2370–2377.
Damiani, L. A., Yingling, C. M., Leng, S., Romo, P. E., Nakamura, J., & Belinsky, S. A. (2008). Carcinogen-induced gene promoter hypermethylation is mediated by DNMT1 and causal for transformation of immortalized bronchial epithelial cells. Cancer Research, 68, 9005–9014.
Shen, H., Spitz, M. R., Qiao, Y., Guo, Z., Wang, L. E., Bosken, C. H., et al. (2003). Smoking, DNA repair capacity and risk of nonsmall cell lung cancer. International Journal of Cancer, 107, 84–88.
Leng, S., Stidley, C. A., Willink, R., Bernauer, A., Do, K., Picchi, M. A., et al. (2008). Double-strand break damage and associated DNA repair genes predispose smokers to gene methylation. Cancer Research, 68, 3049–3056.
Kersting, M., Friedl, C., Kraus, A., Behn, M., Pankow, W., & Schuermann, M. (2000). Differential frequencies of p16(INK4a) promoter hypermethylation, p53 mutation, and K-ras mutation in exfoliative material mark the development of lung cancer in symptomatic chronic smokers. Journal of Clinical Oncology, 18, 3221–3229.
Honorio, S., Agathanggelou, A., Schuermann, M., Pankow, W., Viacava, P., Maher, E. R., et al. (2003). Detection of RASSF1A aberrant promoter hypermethylation in sputum from chronic smokers and ductal carcinoma in situ from breast cancer patients. Oncogene, 22, 147–150.
Lamy, A., Sesboue, R., Bourguignon, J., Dautreaux, B., Metayer, J., Frebourg, T., et al. (2002). Aberrant methylation of the CDKN2a/p16INK4a gene promoter region in preinvasive bronchial lesions: a prospective study in high-risk patients without invasive cancer. International Journal of Cancer, 100, 189–193.
Soria, J. C., Rodriguez, M., Liu, D. D., Lee, J. J., Hong, W. K., & Mao, L. (2002). Aberrant promoter methylation of multiple genes in bronchial brush samples from former cigarette smokers. Cancer Research, 62, 351–355.
Zöchbauer-Müller, S., Lam, S., Toyooka, S., Virmani, A. K., Toyooka, K. O., Seidl, S., et al. (2003). Aberrant methylation of multiple genes in the upper aerodigestive tract epithelium of heavy smokers. International Journal of Cancer, 107, 612–616.
Bhutani, M., Pathak, A. K., Fan, Y. H., Liu, D. D., Lee, J. J., Tang, H., et al. (2008). Oral epithelium as a surrogate tissue for assessing smoking-induced molecular alterations in the lungs. Cancer Prevention Researcg (Philadelphia, PA), 1, 39–44.
Licchesi, J. D., Westra, W. H., Hooker, C. M., & Herman, J. G. (2008). Promoter hypermethylation of hallmark cancer genes in atypical adenomatous hyperplasia of the lung. Clinical Cancer Research, 14, 2570–2578.
Palmisano, W. A., Divine, K. K., Saccomanno, G., Gilliland, F. D., Baylin, S. B., Herman, J. G., et al. (2000). Predicting lung cancer by detecting aberrant promoter methylation in sputum. Cancer Research, 60, 5954–5958.
http://www.fda.gov/ Dacogen (decitabine), NDA# N21790.
Kaminskas, E., Farrell, A., Abraham, S., Baird, A., Hsieh, L. S., Lee, S. L., et al. (2005). Approval summary: azacitidine for treatment of myelodysplastic syndrome subtypes. Clinical Cancer Research, 11, 3604–3608.
Mann, B. S., Johnson, J. R., He, K., Sridhara, R., Abraham, S., Booth, B. P., et al. (2007). Vorinostat for treatment of cutaneous manifestations of advanced primary cutaneous T-cell lymphoma. Clinical Cancer Research, 13, 2318–2322.
Piekarz R, Wright J, Frye R, Allen SL, Craig M, Geskin L, et al. (2008). Results of a phase 2 NCI multicenter study of romidepsin in patients with relapsed peripheral T-cell lymphoma (PTCL). ASH Annual Meeting Abstracts, 112.
Juergens, R., Vendetti, F., Coleman, B., Sebree, R., Belinsky, S., Rudek, M., et al. (2009). A phase II-trial of 5-azacitidine (5AC) and entinostat (SNDX-275) in relapsed advanced lung cancer (NSCLC): an interim analysis. Abstract# A6.6, 13th World Conference on Lung Cancer (WCLC).
Costello, J. F., Fruhwald, M. C., Smiraglia, D. J., Rush, L. J., Robertson, G. P., Gao, X., et al. (2000). Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nature Genetics, 24, 132–138.
Dai, Z., Lakshmanan, R. R., Zhu, W. G., Smiraglia, D. J., Rush, L. J., Fruhwald, M. C., et al. (2001). Global methylation profiling of lung cancer identifies novel methylated genes. Neoplasia, 3, 314–323.
Brena, R. M., Morrison, C., Liyanarachchi, S., Jarjoura, D., Davuluri, R. V., Otterson, G. A., et al. (2007). Aberrant DNA methylation of OLIG1, a novel prognostic factor in non-small cell lung cancer. PLoS Med, 4, e108.
Weber, M., Davies, J. J., Wittig, D., Oakeley, E. J., Haase, M., Lam, W. L., et al. (2005). Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nature Genetics, 37, 853–862.
Dammann, R., Yang, G., & Pfeifer, G. P. (2001). Hypermethylation of the cpG island of Ras association domain family 1A (RASSF1A), a putative tumor suppressor gene from the 3p21.3 locus, occurs in a large percentage of human breast cancers. Cancer Research, 61, 3105–3109.
Suzuki, H., Gabrielson, E., Chen, W., Anbazhagan, R., van Engeland, M., Weijenberg, M. P., et al. (2002). A genomic screen for genes upregulated by demethylation and histone deacetylase inhibition in human colorectal cancer. Nature Genetics, 31, 141–149.
Shames, D. S., Girard, L., Gao, B., Sato, M., Lewis, C. M., Shivapurkar, N., et al. (2006). A genome-wide screen for promoter methylation in lung cancer identifies novel methylation markers for multiple malignancies. PLoS Medicine, 3, e486.
Zhong, S., Fields, C. R., Su, N., Pan, Y. X., & Robertson, K. D. (2007). Pharmacologic inhibition of epigenetic modifications, coupled with gene expression profiling, reveals novel targets of aberrant DNA methylation and histone deacetylation in lung cancer. Oncogene, 26, 2621–2634.
Bestor, T. H. (2003). Unanswered questions about the role of promoter methylation in carcinogenesis. Annals of the New York Academy of Sciences, 983, 22–27.
Sato, N., Fukushima, N., Maitra, A., Matsubayashi, H., Yeo, C. J., Cameron, J. L., et al. (2003). Discovery of novel targets for aberrant methylation in pancreatic carcinoma using high-throughput microarrays. Cancer Research, 63, 3735–3742.
Bibikova, M., Lin, Z., Zhou, L., Chudin, E., Garcia, E. W., Wu, B., et al. (2006). High-throughput DNA methylation profiling using universal bead arrays. Genome Research, 16, 383–393.
Christensen, B. C., Marsit, C. J., Houseman, E. A., Godleski, J. J., Longacker, J. L., Zheng, S., et al. (2009). Differentiation of lung adenocarcinoma, pleural mesothelioma, and nonmalignant pulmonary tissues using DNA methylation profiles. Cancer Research, 69, 6315–6321.
Mockler, T. C., Chan, S., Sundaresan, A., Chen, H., Jacobsen, S. E., & Ecker, J. R. (2005). Applications of DNA tiling arrays for whole-genome analysis. Genomics, 85, 1–15.
Rauch, T. A., Zhong, X., Wu, X., Wang, M., Kernstine, K. H., Wang, Z., et al. (2008). High-resolution mapping of DNA hypermethylation and hypomethylation in lung cancer. Proceedings of the National Academy of Sciences of the United States of America, 105, 252–257.
Tessema, M., Willink, R., Do, K., Yu, Y. Y., Yu, W., Machida, E. O., et al. (2008). Promoter methylation of genes in and around the candidate lung cancer susceptibility locus 6q23-25. Cancer Research, 68, 1707–1714.
Zhang, Z., Tan, S., & Zhang, L. (2006). Prognostic value of apoptosis-associated speck-like protein containing a CARD gene promoter methylation in resectable non-small-cell lung cancer. Clinical Lung Cancer, 8, 62–65.
Nakata, S., Sugio, K., Uramoto, H., Oyama, T., Hanagiri, T., Morita, M., et al. (2006). The methylation status and protein expression of CDH1, p16(INK4A), and fragile histidine triad in nonsmall cell lung carcinoma: epigenetic silencing, clinical features, and prognostic significance. Cancer, 106, 2190–2199.
Maruyama, R., Sugio, K., Yoshino, I., Maehara, Y., & Gazdar, A. F. (2004). Hypermethylation of FHIT as a prognostic marker in nonsmall cell lung carcinoma. Cancer, 100, 1472–1477.
Brabender, J., Usadel, H., Danenberg, K. D., Metzger, R., Schneider, P. M., Lord, R. V., et al. (2001). Adenomatous polyposis coli gene promoter hypermethylation in non-small cell lung cancer is associated with survival. Oncogene, 20, 3528–3532.
Agathanggelou, A., Honorio, S., Macartney, D. P., Martinez, A., Dallol, A., Rader, J., et al. (2001). Methylation associated inactivation of RASSF1A from region 3p21.3 in lung, breast and ovarian tumours. Oncogene, 20, 1509–1518.
Dammann, R., Takahashi, T., & Pfeifer, G. P. (2001). The CpG island of the novel tumor suppressor gene RASSF1A is intensely methylated in primary small cell lung carcinomas. Oncogene, 20, 3563–3567.
Kashiwabara, K., Oyama, T., Sano, T., Fukuda, T., & Nakajima, T. (1998). Correlation between methylation status of the p16/CDKN2 gene and the expression of p16 and Rb proteins in primary non-small cell lung cancers. International Journal of Cancer, 79, 215–220.
Kuramochi, M., Fukuhara, H., Nobukuni, T., Kanbe, T., Maruyama, T., Ghosh, H. P., et al. (2001). TSLC1 is a tumor-suppressor gene in human non-small-cell lung cancer. Nature Genetics, 27, 427–430.
Brabender, J., Usadel, H., Metzger, R., Schneider, P. M., Park, J., Salonga, D., et al. (2003). Quantitative O(6)-methylguanine DNA methyltransferase methylation analysis in curatively resected non-small cell lung cancer: associations with clinical outcome. Clinical Cancer Research, 9, 223–227.
Acknowledgment
This work was supported by the Vienna Science and Technology Fund (project number LS07-019).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Heller, G., Zielinski, C.C. & Zöchbauer-Müller, S. Lung cancer: From single-gene methylation to methylome profiling. Cancer Metastasis Rev 29, 95–107 (2010). https://doi.org/10.1007/s10555-010-9203-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-010-9203-x