Abstract
Dietary methyl donors might influence DNA methylation during carcinogenesis of colorectal cancer (CRC). Among 609 CRC cases and 1,663 subcohort members of the Netherlands Cohort Study on diet and cancer (n = 120,852), we estimated CRC risk according to methyl donor intake across genotypes of folate metabolizing enzymes and methyltransferases.
Although diet–gene interactions were not statistically significant, methionine intake was inversely associated with CRC among subjects having both common rs2424913 and rs406193 DNMT3B C > T genotypes (highest versus lowest tertile: RR = 0.44; p trend = 0.05). Likewise, vitamin B2 was modestly inversely associated among individuals with the MTHFR c.665CC (rs1801133) genotype (RR = 0.66; p trend = 0.08), but with a significant reduced risk when ≤ 1 rare allele occurred in the combination of folate metabolizing enzymes MTHFR, MTRR and MTR (RR = 0.30; p trend = 0.005). Folate or vitamin B6 were neither inversely associated with CRC nor was methyl donor intake associated with the CpG island methylator phenotype (CIMP).
Despite the absence of heterogeneity across genotypes, might an effect of methyl donors on CRC be more pronounced among individuals carrying common variants of folate metabolizing enzymes or DNA methyltransferases. Combining genotypes may assist to reveal diet associations with CRC, possibly because rare variants of related genes may collectively affect specific metabolic pathways or enzymatic functions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hypermethylation of CpG islands in gene promoters is an important epigenetic alteration involved in carcinogenesis [1]. In colorectal cancer (CRC), the CpG island methylator phenotype (CIMP) is characterized by frequent promoter CpG island hypermethylation [2]. However, little is known about potential determinants of this type of aberrant DNA methylation in CRC.
Folate and methionine are dietary methyl group donors that may be hypothesized to influence DNA methylation, whereas vitamins B2 and B6 potentially modulate the bioavailability of methyl groups [3, 4]. Low folate status or intake was suggested to decrease genomic methylation [5–8], while folate supplementation resulted in increased global DNA methylation in the colonic mucosa [9]. Adequate methyl donor intake possibly also prevents aberrant CpG island promoter hypermethylation. In this respect, a weak inverse association with gene promoter hypermethylation was suggested [10], although methyl donor intake and alcohol consumption, which may reduce the bioavailability of folate, were not associated with CIMP in CRC [11]. Conversely, folate supplementation was suggested to increase promoter hypermethylation of multiple genes in colorectal mucosa [12], and circulating folate concentration was associated with increased gene promoter hypermethylation in colorectal tumors [13]. In addition, high vitamin B6 intake may be associated with increased MutL homologue 1 (MLH1) promoter methylation in CRC [14]. Apparently, the precise effect of methyl group bioavailability on gene promoter hypermethylation is still unclear and should be investigated further.
A potential effect of methyl donor intake on DNA methylation may be modified by polymorphisms in folate metabolizing enzymes. For example, the catalytic activity of the methylene tetrahydrofolate reductase (MTHFR) enzyme may be reduced in individuals carrying rare variants of the MTHFR c.665C > T (rs1801133) and c.1286A > C (rs1801131) polymorphisms [15, 16], which were also associated with the CIMP phenotype in colorectal cancer [17–19]. We previously observed inverse associations between methionine synthase (MTR) c.2756A > G (rs1805087) and CIMP, and between methionine synthase reductase (MTRR) c.66A > G (rs1801394) with MLH1 hypermethylation [20]. Other enzymes involved in epigenetic regulation of gene expression are DNA methyltransferases (DNMTs) and histone methyltransferases (HMTs). However, whether an influence of methyl donor intake is modified by polymorphisms in such epigenetic regulators has not previously been studied in relation to CRC.
Here, we aimed to investigate associations between dietary folate, methionine, vitamins B2 and B6 with overall CRC, and risk of CRCs harboring CIMP, accounting for the occurrence of any, or combinations of rare variants of folate metabolizing enzymes MTHFR, MTR and MTRR, the DNA methyltransferase DNMT3B, and histone methyltransferases Euchromatin histone methyltransferase 1 (EHMT1), Euchromatin histone methyltransferase 2 (EHMT2) and PR domain zinc finger protein 2 (PRDM2) in the Netherlands Cohort Study on diet and cancer.
Methods
Study population
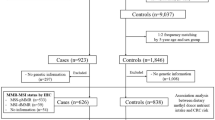
The participants of this study were incident CRC patients from the Netherlands Cohort Study on diet and cancer (NLCS), which has been described in detail elsewhere [21]. Briefly, this prospective cohort study was initiated in September 1986 and includes 58,279 men and 62,573 women aged 55–69 years and free of disease at baseline. The cohort is followed for cancer occurrence by annual record linkage to the Netherlands Cancer Registry (NCR) and to the Pathologisch Anatomisch Landelijk Geautomatiseerd Archief (PALGA), a nationwide network and registry of histopathology and cytopathology reports [22, 23]. A subcohort of 5,000 subjects was randomly selected after baseline exposure measurement, to estimate accumulation of person-time in the cohort through biennial follow-up of vital status. Cases with prevalent cancer other than non-melanoma skin cancer were excluded from this subcohort, which left 4,774 men and women eligible for analysis.
Food frequency questionnaire
At baseline, participants filled out a self-administered, 150-item semi-quantitative food frequency questionnaire (FFQ), which concentrated on habitual consumption of food and beverages during the year preceding the start of the study, and also contained questions about age, sex, body weight and length, smoking status and family history of CRC. Daily mean nutrient intakes were calculated as the cumulated product of the frequencies and portion sizes of all food items and their tabulated nutrient contents from the Dutch Food Composition Table (NEVO table, 1986) [24]. The questionnaire was validated through comparison with a 9-day diet record [25]. Reproducibility and stability of dietary habits were determined by five annually repeated measurements [26]. In order to minimize observer bias in coding and interpretation of the data, questionnaire data were key-entered twice for all incident cases in the cohort and for all subcohort members in a blinded manner with respect to case/subcohort status.
Folate data were derived from a validated liquid chromatography trienzyme method [27] used to analyze the 125 most important Dutch foods contributing to folate intake [28]. Dietary supplement data were also obtained via the food frequency questionnaire. However, the use of B-vitamin supplements was low (7%) and folic acid was generally not included in these supplements in the Netherlands in the late 1980s. Therefore, folic acid supplement use most likely plays a very minor role in our study population, and supplement use was not further accounted for in the analyses.
Sample collection
Subcohort members still alive in December 2000 (n = 3,579) were contacted and asked to collect mouth swabs, of whom 1,929 (54%) responded and returned the mouth swab with informed consent. In total, DNA could successfully be isolated of 1,829 subcohort members who also had complete follow-up information [20].
Tumor material of the CRC patients was collected after approval by the ethical review boards of Maastricht University, the NCR and PALGA. During a follow-up period of 7.3 years after baseline, 734 incident CRC patients were identified who had an available PALGA report of the lesion as well as a sufficient amount of isolated DNA needed for molecular analyses.
Genotyping analyses
MTHFR (rs1801133 and rs1801131), MTR (rs1805087), MTRR (rs1801394), DNMT3B (rs2424913 and rs406193), EHMT1 (rs4634736), EHMT2 (rs535586) and PRDM2 (rs2235515) genotypes were determined using multiplex polymerase chain reaction (PCR) amplification and single base extension (SBE) reactions as described previously [20, 29]. Genotype data were validated by sequencing of fragments containing specific SNPs, which were similar to the main results for all but one (99.6%) of the 9 SNPs within a subset of 30 samples [20]. Reproducibility of the analysis was established among 93 samples, and we observed that the analyses could be reproduced in 99.5% of these cases [20]. In total, genotyping analyses were successful from 1,736 subcohort members and 659 CRC patients.
Promoter methylation analyses
The CpG island methylator phenotype (CIMP) was defined by promoter hypermethylation of at least 3 out of 5 methylation markers (CACNA1G, IGF2, NEUROG1, RUNX3, and SOCS1), as suggested by Weisenberger et al. [2]. Hypermethylation of the CpG islands of these five CIMP markers and of the MLH1 gene was determined by Methylation Specific PCR (MSP) [30] and described in detail by de Vogel et al [20]. The MSP analyses were successful of 81, 79, 79, 90, 83, and 93% out of the 734 patients for CACNA1G, IGF2, NEUROG1, RUNX3, SOCS1, and MLH1, respectively.
Microsatellite instability
MSI was determined by a pentaplex PCR, using the MSI markers BAT-26, BAT-25, NR-21, NR-22 and NR-24, as described in detail by Suraweera et al. [31]. MSI analyses were successful on 662 (90%) out of the 734 available samples.
Statistical analyses
Cox proportional hazards regression models were used to estimate multivariate-adjusted incidence rate ratios (RR) and corresponding 95% confidence intervals (CI) over tertiles of dietary folate, methionine, vitamins B2 and B6, using the lowest tertiles as reference. Tests for dose response trends over the tertiles of intake were estimated by fitting the ordinal exposure variables as continuous variables and evaluated using the Wald test. Standard errors of the RR were estimated using the robust Huber-White sandwich estimator to account for additional variance introduced by sampling from the cohort [32]. The proportional hazards assumption was tested using the scaled Schoenfeld residuals [33] and by fitting the main determinants as time-dependent variables. The dietary variables were adjusted for total energy intake by calculating nutrient residuals from the regression of nutrient intake on total energy intake, as described by Willett et al. [34]. The analyses were stratified according to genetic status of individuals, i.e. among those homozygous to common genetic variants and among subjects carrying rare alleles. Interactions were tested between dietary folate, methionine, vitamins B2 and B6, and each of the genetic variants. Associations between dietary factors and CRC were also estimated for combinations of genotypes per functional group (i.e. based on the number of rare alleles in any of the folate metabolizing enzymes MTHFR, MTR and MTRR, in the DNA methyltransferase DNMT3B, or in any of the histone methyltransferases EHMT1, EHMT2 and PRDM2.
To investigate whether dietary methyl donors have an effect on promoter hypermethylation in CRC, associations of folate, methionine, vitamins B2, and B6 with the CIMP phenotype were estimated. The associations with MLH1 hypermethylation and MSI were reported previously [14]. Furthermore, it was investigated whether the associations with MLH1 hypermethylation, MSI, or CIMP would be modified by genetic status, by estimating the associations with methylation endpoints within genotypes of folate metabolizing enzymes, DNMT3B and histone methyltransferases.
All models included the co-variates dietary folate, methionine, vitamin B2 and B6 and were additionally adjusted for age, sex, family history of CRC, smoking status, body mass index (BMI), alcohol consumption, and energy intake. After excluding subjects with missing information on these covariates or subjects who did not completely filled out the questionnaire, 1,663 subcohort members and 609 CRC cases remained for statistical analyses. All analyses were performed with the Stata statistical software package (version 10).
Results
CRC risk was estimated over tertiles of folate intake, methionine, vitamins B2 and B6, among subjects homozygous for common alleles and among carriers of rare alleles. Folate or methionine intakes were not associated with CRC within either common homozygotes or within heterozygotes and rare homozygotes of any of the genotypes (Table 1). However, we observed a non-significant inverse association between vitamin B2 intake and CRC risk among subjects with the MTHFR c.665CC (rs1801133) common genotype (RR for the highest versus the lowest tertile of intake = 0.66, p trend = 0.08, Table 2), and an inverse association among subjects with the common GG genotype of PRDM2 G > A (rs2235515, RR = 0.67, p trend = 0.05). In addition, vitamin B2 was associated with reduced CRC risk in individuals carrying the variant allele of DNMT3B C > T (rs2424913, RR = 0.69, p trend = 0.05). Conversely, subjects in the third tertile of vitamin B6 intake were at increased CRC risk when they carried the rare allele of DNMT3B C > T (rs406193, RR = 1.90, p trend = 0.04), or the common allele of PRDM2 G > A (rs2235515, RR = 1.49, p trend = 0.03). However, interactions between these dietary factors and genotypes were not statistically significant.
We also investigated the associations between methyl donor intake and CRC risk according to the number of rare alleles within each functional group (i.e. folate metabolizing enzymes, DNMT3B and histone methyltransferases). It appeared that methionine was inversely associated with CRC if subjects were homozygous to both of the common variants of the DNMT3B rs2424913 and rs406193 C > T SNPs (RR = 0.44, p trend = 0.05, P interaction = 0.07, Table 3). Moreover, relatively high vitamin B2 intake was associated with reduced CRC risk in subjects carrying less than one rare variant of folate metabolizing enzymes (RR = 0.30, p trend = 0.005, P interaction = 0.36, Table 4). No dietary associations were observed according to the number of rare alleles in the studied histone methyltransferases (Table 5).
With respect to CpG island promoter hypermethylation, we observed no overall associations between folate, methionine, vitamins B2 or B6 with CIMP (Table 6). Moreover, there were no clear associations between methyl donor intake and CIMP, MLH1 hypermethylation or MSI when accounting for genetic status of individuals (data not shown).
Discussion
In the current prospective case-cohort study, we observed no clear associations between dietary folate and vitamin B6 with CRC risk when accounting for genetic variants of folate metabolizing enzymes, DNA methyltransferases, or histone methyltransferases. However, relatively high methionine intake may protect against CRC if enzymatic activity of DNMT3B is not affected by two C > T SNPs in its encoding gene. In addition, subjects with high vitamin B2 intake may be at reduced CRC risk in combination with optimal MTHFR activity in individuals homozygous for the common c.665CC (rs1801133) variant, with common PRDM2 GG (rs2235515) genotype and among those with the variant allele of DNMT3B C > T (rs2424913). We observed a strong inverse association between vitamin B2 intake and CRC risk among individuals carrying ≤ 1 rare allele in the combination of any of the folate metabolizing enzymes MTHFR, MTR, or MTRR. There were no associations with the CIMP phenotype overall, or within strata of the studied genotypes.
The MTHFR c.665C > T (rs1801133) polymorphism reduces binding of the MTHFR enzyme to its cofactor flavin adenine dinucleotide (FAD), a metabolite of vitamin B2, resulting in loss of enzymatic activity [15]. The potentially resulting reduced bioavailability of methyl groups may induce DNA hypomethylation in for example blood cells [35, 36] or CpG island promoter hypermethylation in CRC [19, 37]. We observed an inverse association between vitamin B2 and CRC risk, predominantly among subjects homozygous for the MTHFR c.665CC (rs1801133) variant, suggesting that vitamin B2 may maximize the catalytic activity of MTHFR when binding to FAD is optimal. Similarly, it was recently observed that high vitamin B2 plasma concentrations, in combination with MTHFR c.665CC or CT genotypes, may reduce risk of CRA recurrence, whereas such an inverse association was not observed among individuals with the MTHFR c.665TT variant [38].
The rare variant of another MTHFR polymorphism, MTHFR c.1286A > C (rs1801131), may also reduce enzymatic MTHFR activity [16], and was associated with CIMP in colorectal cancer [18], possibly in combination with low folate and methionine intakes and high alcohol consumption [17]. However, we previously observed that this polymorphism was neither associated with overall CRC or with the CIMP phenotype [39], nor when methyl donor intake was accounted for in the current study. Possibly, the use of different panels to identify CIMP-high (a “classic” panel [17] or a new panel [18, 39] which may be more robust [2]) may have contributed to this inconsistency. Moreover, different essays to measure DNA methylation were used, i.e. MSP [17, 39] or a quantitative method [18]. However, in addition to this variety of approaches, it is also important to realize that the one-carbon metabolism is involved in both DNA synthesis as well as DNA methylation, both of which may have an effect on colorectal carcinogenesis [40]. The relative contribution of each of these biological processes in carcinogenesis remains to be established and may not have been similar in the investigated study populations. Furthermore, global DNA hypomethylation and CIMP are possibly inversely associated in CRC [41], and methyl group donors may have an effect on both of these potentially distinct methylation-associated pathways in colorectal carcinogenesis. In this respect, low folate and high alcohol intakes were associated with LINE-1 hypomethylation as an indicator for global DNA hypomethylation [8], which is in agreement with in vivo experimental data [7].
We did not observe associations between methyl donor intake and the CIMP phenotype in CRC, either overall or after stratifying the analyses for the genetic variants of folate metabolizing enzymes or methyltransferases. Moreover, overall associations between methyl donor intake and CIMP in CRC were not observed in another population-based study [11], while in the same cohort, a diet-gene association with CIMP in CRC was observed for only one out of thirteen one-carbon metabolism genes [17]. However, these studies, as well as our study, may have lacked adequate power to demonstrate such associations. The effect of methyl donor intake on gene promoter hypermethylation may indeed be weak though, and to demonstrate whether such an effect is modified by genetic variability of the methyl metabolism requires studies with large numbers of cases. Nonetheless, we did observe an inverse association between vitamin B2 and CRC risk in individuals carrying ≤1 variant allele out of the four studied SNPs of folate metabolizing enzymes, suggesting that the combination of common wild-type genotypes, which possibly results in higher bioavailability of methyl groups, protects against CRC in these people.
The findings of our study may indicate that relatively high methionine intake protects against CRC if enzymatic DNMT3B activity is not affected by two polymorphisms. DNMT3B activity, which may be increased by the DNMT3B C > T (rs2424913) polymorphism [42], was associated with CIMP-high in CRC [43], and with increased risk of various other types of cancer [42, 44, 45]. In addition, experimental research suggested that DNMT3B overexpression induced formation of tumors with promoter hypermethylation [46]. DNMT3B C > T (rs2424913) was also associated with increased colorectal adenoma risk in individuals with low folate and methionine intakes [47], suggesting a nutrient–gene interaction in colorectal carcinogenesis. In view of the function of the DNMT3B enzyme of incorporating methyl groups into DNA, an interaction between methionine intake, DNMT3B polymorphisms and CpG island hypermethylation may be expected, but we did not observe clear associations between methyl donor intake and CIMP, MLH1 hypermethylation or MSI when accounting for DNMT3B genotypes.
The potential protective effects of vitamin B2 or methionine may only be present among individuals with ≤1 polymorphism in folate metabolizing enzymes or among those with common wild-type genotypes of DNMT3B, respectively. This suggests that the occurrence of only one rare variant may be compensated for, but that the combination of several polymorphic genes may lead to disruption of a particular metabolic or regulatory function and to the abolishment of beneficial effects of nutrients. However, we should be careful in drawing definite conclusions because the sample size of our study may have been insufficient to conduct stratified analyses with adequate precision. Moreover, the P-values for interaction were not statistically significant, suggesting the absence of heterogeneity of diet associations with CRC across genotypes. In addition, we conducted several stratified analyses, and these multiple comparisons do not exclude the possibility of reporting chance findings. Nonetheless, although these observations are based on subgroup analyses, and thus have to be interpreted with some caution, this study may indicate that combining genotypes is important to reveal associations of dietary factors with cancer risk. Such an approach has not been followed in previous studies investigating associations between genetic factors and cancer risk, and we recommend that combinations of genotypes should be considered in addition to overall analyses in future studies.
Subgroup analyses in the present study indicated that vitamin B2 and methionine may protect against CRC among individuals who do not carry rare variants of folate metabolizing enzymes and a DNA methyltransferase. However, larger studies are needed to investigate a potential interaction between dietary methyl donor intake, genetic variation of folate metabolizing enzymes and epigenetic regulators, and methylation endpoints in CRC with more precision. Because multiple genes may collectively affect the folate metabolism, combining genotypes of related genes is a useful approach of investigating associations of dietary methyl donors and CRC.
References
Herman JG, Baylin SB (2003) Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 349:2042–2054
Weisenberger DJ, Siegmund KD, Campan M et al (2006) CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet 38:787–793
Kim YI (2005) Nutritional epigenetics: impact of folate deficiency on DNA methylation and colon cancer susceptibility. J Nutr 135:2703–2709
Ulrich CM (2005) Nutrigenetics in cancer research–folate metabolism and colorectal cancer. J Nutr 135:2698–2702
Kim YI (2004) Folate and DNA methylation: a mechanistic link between folate deficiency and colorectal cancer? Cancer Epidemiol Biomarkers Prev 13:511–519
Pufulete M, Al-Ghnaniem R, Rennie JA et al (2005) Influence of folate status on genomic DNA methylation in colonic mucosa of subjects without colorectal adenoma or cancer. Br J Cancer 92:838–842
Linhart HG, Troen A, Bell GW et al. (2009) Folate deficiency induces genomic uracil misincorporation and hypomethylation but does not increase DNA point mutations. Gastroenterology 136: 227–235 e3
Schernhammer ES, Giovannucci E, Kawasaki T, Rosner B, Fuchs CS, Ogino S (2010) Dietary folate, alcohol and B vitamins in relation to LINE-1 hypomethylation in colon cancer. Gut 59:794–799
Pufulete M, Al-Ghnaniem R, Khushal A et al (2005) Effect of folic acid supplementation on genomic DNA methylation in patients with colorectal adenoma. Gut 54:648–653
van Engeland M, Weijenberg MP, Roemen GM et al (2003) Effects of dietary folate and alcohol intake on promoter methylation in sporadic colorectal cancer: the Netherlands cohort study on diet and cancer. Cancer Res 63:3133–3137
Slattery ML, Curtin K, Sweeney C et al (2007) Diet and lifestyle factor associations with CpG island methylator phenotype and BRAF mutations in colon cancer. Int J Cancer 120:656–663
van den Donk M, Pellis L, Crott JW et al (2007) Folic acid and vitamin B-12 supplementation does not favorably influence uracil incorporation and promoter methylation in rectal mucosa DNA of subjects with previous colorectal adenomas. J Nutr 137:2114–2120
Kawakami K, Ruszkiewicz A, Bennett G, Moore J, Watanabe G, Iacopetta B (2003) The folate pool in colorectal cancers is associated with DNA hypermethylation and with a polymorphism in methylenetetrahydrofolate reductase. Clin Cancer Res 9:5860–5865
de Vogel S, Bongaerts BW, Wouters KA et al (2008) Associations of dietary methyl donor intake with MLH1 promoter hypermethylation and related molecular phenotypes in sporadic colorectal cancer. Carcinogenesis 29:1765–1773
Frosst P, Blom HJ, Milos R et al (1995) A candidate genetic risk factor for vascular disease: a common mutation in methylenetetrahydrofolate reductase. Nat Genet 10:111–113
van der Put NM, Gabreels F, Stevens EM et al (1998) A second common mutation in the methylenetetrahydrofolate reductase gene: an additional risk factor for neural-tube defects? Am J Hum Genet 62:1044–1051
Curtin K, Slattery ML, Ulrich CM et al (2007) Genetic polymorphisms in one-carbon metabolism: associations with CpG island methylator phenotype (CIMP) in colon cancer and the modifying effects of diet. Carcinogenesis 28:1672–1679
Hazra A, Fuchs CS, Kawasaki T, Kirkner GJ, Hunter DJ, Ogino S (2010) Germline polymorphisms in the one-carbon metabolism pathway and DNA methylation in colorectal cancer. Cancer Causes Control 21:331–345
Oyama K, Kawakami K, Maeda K, Ishiguro K, Watanabe G (2004) The association between methylenetetrahydrofolate reductase polymorphism and promoter methylation in proximal colon cancer. Anticancer Res 24:649–654
de Vogel S, Wouters KA, Gottschalk RW et al. (2009) Genetic variants of methyl metabolizing enzymes and epigenetic regulators: associations with promoter CpG island hypermethylation in colorectal cancer. Cancer Epidemiol Biomarkers Prev (in press)
van den Brandt PA, Goldbohm RA, van ‘t Veer P, Volovics A, Hermus RJ, Sturmans F (1990) A large-scale prospective cohort study on diet and cancer in The Netherlands. J Clin Epidemiol 43:285–295
Van den Brandt PA, Schouten LJ, Goldbohm RA, Dorant E, Hunen PM (1990) Development of a record linkage protocol for use in the Dutch Cancer Registry for Epidemiological Research. Int J Epidemiol 19:553–558
Casparie M, Tiebosch AT, Burger G et al (2007) Pathology databanking and biobanking in The Netherlands, a central role for PALGA, the nationwide histopathology and cytopathology data network and archive. Cell Oncol 29:19–24
Nevo Table (1986) Dutch food composition table 1986–1987, Voorlichtingsbureau voor de voeding, The Hague, The Netherlands, 1986
Goldbohm RA, van den Brandt PA, Brants HA et al (1994) Validation of a dietary questionnaire used in a large-scale prospective cohort study on diet and cancer. Eur J Clin Nutr 48:253–265
Goldbohm RA, van ‘t Veer P, van den Brandt PA et al (1995) Reproducibility of a food frequency questionnaire and stability of dietary habits determined from five annually repeated measurements. Eur J Clin Nutr 49:420–429
Konings EJ (1999) A validated liquid chromatographic method for determining folates in vegetables, milk powder, liver, and flour. J AOAC Int 82:119–127
Konings EJ, Roomans HH, Dorant E, Goldbohm RA, Saris WH, van den Brandt PA (2001) Folate intake of the Dutch population according to newly established liquid chromatography data for foods. Am J Clin Nutr 73:765–776
Knaapen AM, Ketelslegers HB, Gottschalk RW et al (2004) Simultaneous genotyping of nine polymorphisms in xenobiotic-metabolizing enzymes by multiplex PCR amplification and single base extension. Clin Chem 50:1664–1668
Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB (1996) Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 93:9821–9826
Suraweera N, Duval A, Reperant M et al (2002) Evaluation of tumor microsatellite instability using five quasimonomorphic mononucleotide repeats and pentaplex PCR. Gastroenterology 123:1804–1811
Lin D, Wei L (1989) The robust inference for the Cox Proportional Hazards Model. JASA 84:1074–1078
Schoenfeld D (1982) Partial residuals for the proportional hazards regression models. Biometrika 69:239–241
Willett W, Stampfer MJ (1986) Total energy intake: implications for epidemiologic analyses. Am J Epidemiol 124:17–27
Friso S, Choi SW, Girelli D et al (2002) A common mutation in the 5, 10-methylenetetrahydrofolate reductase gene affects genomic DNA methylation through an interaction with folate status. Proc Natl Acad Sci USA 99:5606–5611
Stern LL, Mason JB, Selhub J, Choi SW (2000) Genomic DNA hypomethylation, a characteristic of most cancers, is present in peripheral leukocytes of individuals who are homozygous for the C677T polymorphism in the methylenetetrahydrofolate reductase gene. Cancer Epidemiol Biomarkers Prev 9:849–853
Mokarram P, Naghibalhossaini F, Saberi Firoozi M et al (2008) Methylenetetrahydrofolate reductase C677T genotype affects promoter methylation of tumor-specific genes in sporadic colorectal cancer through an interaction with folate/vitamin B(12) status. World J Gastroenterol 14:3662–3671
Figueiredo JC, Levine AJ, Grau MV et al (2008) Vitamins B2, B6, and B12 and risk of new colorectal adenomas in a randomized trial of aspirin use and folic acid supplementation. Cancer Epidemiol Biomarkers Prev 17:2136–2145
de Vogel S, Wouters KA, Gottschalk RW et al (2009) Genetic variants of methyl metabolizing enzymes and epigenetic regulators: associations with promoter CpG island hypermethylation in colorectal cancer. Cancer Epidemiol Biomarkers Prev 18:3086–3096
Kim YI (2007) Folate and colorectal cancer: An evidence-based critical review. Mol Nutr Food Res 51:267–292
Ogino S, Kawasaki T, Nosho K et al (2008) LINE-1 hypomethylation is inversely associated with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Int J Cancer 122:2767–2773
Shen H, Wang L, Spitz MR, Hong WK, Mao L, Wei Q (2002) A novel polymorphism in human cytosine DNA-methyltransferase-3B promoter is associated with an increased risk of lung cancer. Cancer Res 62:4992–4995
Nosho K, Shima K, Irahara N et al (2009) DNMT3B expression might contribute to CpG island methylator phenotype in colorectal cancer. Clin Cancer Res 15:3663–3671
Singal R, Das PM, Manoharan M, Reis IM, Schlesselman JJ (2005) Polymorphisms in the DNA methyltransferase 3b gene and prostate cancer risk. Oncol Rep 14:569–573
Wang L, Rodriguez M, Kim ES et al (2004) A novel C/T polymorphism in the core promoter of human de novo cytosine DNA methyltransferase 3B6 is associated with prognosis in head and neck cancer. Int J Oncol 25:993–999
Linhart HG, Lin H, Yamada Y et al (2007) Dnmt3b promotes tumorigenesis in vivo by gene-specific de novo methylation and transcriptional silencing. Genes Dev 21:3110–3122
Jung AY, Poole EM, Bigler J, Whitton J, Potter JD, Ulrich CM (2008) DNA methyltransferase and alcohol dehydrogenase: gene-nutrient interactions in relation to risk of colorectal polyps. Cancer Epidemiol Biomarkers Prev 17:330–338
Acknowledgments
The authors acknowledge Dr. M. Brink for the collection of the tissue samples. This study is funded by the Dutch Cancer Society (UM2004-3171 and UM99-1980).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
de Vogel, S., Wouters, K.A.D., Gottschalk, R.W.H. et al. Dietary methyl donors, methyl metabolizing enzymes, and epigenetic regulators: diet–gene interactions and promoter CpG island hypermethylation in colorectal cancer. Cancer Causes Control 22, 1–12 (2011). https://doi.org/10.1007/s10552-010-9659-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10552-010-9659-6