Abstract
Purpose
Few targeted treatment options currently exist for patients with advanced, often recurrent breast cancers, both triple-negative breast cancer (TNBC) and hormone receptor-positive breast cancer. Forkhead box M1 (FOXM1) is an oncogenic transcription factor that drives all cancer hallmarks in all subtypes of breast cancer. We previously developed small-molecule inhibitors of FOXM1 and to further exploit their potential as anti-proliferative agents, we investigated combining FOXM1 inhibitors with drugs currently used in the treatment of breast and other cancers and assessed the potential for enhanced inhibition of breast cancer.
Methods
FOXM1 inhibitors alone and in combination with other cancer therapy drugs were assessed for their effects on suppression of cell viability and cell cycle progression, induction of apoptosis and caspase 3/7 activity, and changes in related gene expressions. Synergistic, additive, or antagonistic interactions were evaluated using ZIP (zero interaction potency) synergy scores and the Chou–Talalay interaction combination index.
Results
The FOXM1 inhibitors displayed synergistic inhibition of proliferation, enhanced G2/M cell cycle arrest, and increased apoptosis and caspase 3/7 activity and associated changes in gene expression when combined with several drugs across different pharmacological classes. We found especially strong enhanced effectiveness of FOXM1 inhibitors in combination with drugs in the proteasome inhibitor class for ER-positive and TNBC cells and with CDK4/6 inhibitors (Palbociclib, Abemaciclib, and Ribociclib) in ER-positive cells.
Conclusion
The findings suggest that the combination of FOXM1 inhibitors with several other drugs might enable dose reduction in both agents and provide enhanced efficacy in treatment of breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The past few decades have seen significant advancements in the treatment of breast cancer thanks to the discovery of specific, molecularly targeted therapies like estrogen receptor (ER) modulators and human epidermal growth factor 2 (HER2) inhibitors that effectively suppress tumor growth in patients with hormone receptor-positive (HR +) cancers [1]. Despite these advances, few treatment options exist for patients with hormone receptor-positive tumors that do not respond to hormone therapy, acquire resistance to targeted therapy or become metastatic [2, 3]. There are even fewer options for patients with triple-negative breast cancer (TNBC), a particularly aggressive group of breast cancers, where the lack of expression of the usual druggable molecular targets makes cytotoxic chemotherapy the standard systemic treatment option [4]. TNBCs are usually associated with worse patient clinical outcomes compared to the hormone receptor-positive (HR +) subtypes, and many with metastatic TNBC will unfortunately die of this disease [5]. TNBCs are very heterogeneous and have only recently been classified into 6 subtypes that possess unique biological, histological, molecular, and transcriptional profiles [4, 6]. As a result of this diversity, the development of targeted molecular therapies for TNBC as a whole has been difficult [7].
Forkhead box M1 (FOXM1) is an oncogenic transcription factor that is overexpressed in all breast cancer subtypes compared to normal tissue and its overexpression is associated with poor clinical outcomes in patients [8]. FOXM1 is most highly elevated in TNBC compared to other subtypes of breast cancer and has a distinct gene expression signature that is enriched in pathways driving cancer cell growth and cell cycle progression, making it an important marker in TNBC and a potential target for novel therapies [9, 10]. In ER + cancers, FOXM1 regulates ER expression and acts as a binding partner at genomic estrogen-responsive elements (EREs) to promote mitogenic tumor growth, metastasis, and treatment resistance [11,12,13]. Similarly, FOXM1 is a downstream target of HER2 that helps promote cell cycle progression and deregulation of mitotic checkpoints [14].
We recently reported on our development of small-molecule inhibitors of FOXM1 that interact directly with the FOXM1 protein and decrease its cellular mRNA and protein levels, resulting in potent suppression of breast cancer cell and tumor growth in pre-clinical models [13, 15, 16]. Furthermore, these inhibitors downregulated expression of FOXM1-controlled gene networks to induce cell cycle arrest, suppress proliferation, and stimulate apoptosis in several breast cancer subtypes [13] and also in other aggressive cancers, including some multiple myelomas [17] and high-grade serous ovarian cancer [18, 19].
To further investigate the potential of these inhibitors to suppress breast cancer, we sought to combine them with other therapies currently being used in the treatment of breast and other cancers or with compounds in early clinical trials that show promise in the treatment of breast cancer. We found that our inhibitors synergize with several cancer drugs in different pharmacological classes. We measured strong synergy when our compounds and other FOXM1 inhibitors were combined with drugs in the proteasome inhibitor class, with induction of G2/M cell cycle arrest and apoptosis. In ER-positive breast cancer cells, the combination of FOXM1 inhibitors with CDK4/6 inhibitors also resulted in synergistic suppression of growth and the expression of genes regulating cell proliferation.
Methods
Cell lines and cell culture methods, FOXM1 inhibitors, and cancer drugs studied
All breast cancer cell lines (MDA-MB-231; MDA-MB-468; MCF7; ZR75-1) were obtained from the ATCC and were maintained and cultured as described [11, 20]. Cells were STR profiled to confirm identity and routinely tested for mycoplasma using Real-Time PCR Mycoplasma Detection Kit (Akron Biotech, Boca Raton, FL).
FOXM1 inhibitors, NB55, NB73, and NB115, were synthesized in our laboratories as previously reported [13]. FDI6 was obtained from Millipore Sigma. The CDK4/6 inhibitors, Abemaciclib, Palbociclib, and Ribociclib, were obtained from Selleckchem (Houston, TX). The proteasome inhibitors, Bortezomib and TAK-243, were obtained from Selleckchem or MedChemExpress (MCE, Monmouth Junction, NJ). The PI3K inhibitor, Alpelisib, was from Selleckchem; the ALK inhibitor, Crizotinib, was from Selleckchem; and the AURKA inhibitor, Alisertib, was from Selleckchem.
Cell viability assay and assessment of synergistic efficacy of test compounds
The WST-1 assay (Roche, Basel, Switzerland) was used to quantify cell viability as described [20]. Absorbance was measured at 450 nm using a VICTOR X5 PerkinElmer 2030 Multilabel Plate Reader. All assays were performed in triplicate and analyzed using Graph Pad Prism 8.0. As detailed further in Results, assessments of synergy in the effectiveness of FOXM1 inhibitors and other cancer drugs used Zero Interaction Potency (ZIP) synergy score determinations, as well as the Bliss and Loewe model assessments [21,22,23,24], and evaluation using the Chou–Talalay combination index (CI) where a CI less than 1 implies synergy [25].
Cell cycle analysis
Cells were synchronized by double-thymidine block prior to cell cycle analysis by Propidium Iodide (PI) staining. Briefly, the cells were blocked with 2-mM thymidine for 18 h and then released with fresh media for 8 h followed by re-blocking with thymidine for another 16 h. Following the second block, the cells were released with or without treatments with the NB compounds and/or other drugs and cells were collected and fixed in 70% ethanol at the time points indicated. After alcohol fixation for 2 h, the cells were washed with cold 1XPBS followed by staining with 50-µg/ml PI solution in PBS with addition of 5-µg/ml RNAse A for at least 4 h. The cells were then analyzed using the Flow Cytometry analyzer BD LSR II for percentage of cells in G0/G1, S, and G2/M phases of the cell cycle.
Apoptosis analysis
The cells were analyzed for percentage of apoptotic cells following vehicle or compound treatments by staining with the Alexa Fluor® 488 Annexin V/Dead Cell Apoptosis Kit (Thermo Fisher) and flow cytometry according to the manufacturer’s protocol. Caspase activity in the cells after treatment with control vehicle or compounds was determined using the Caspase-Glo 3/7 Assay system (Promega) in a 96-well format following the manufacturer’s instructions.
RNA isolation and real-time PCR
Total RNA was isolated using TRIzol (Invitrogen) and reverse transcribed using MMTV reverse transcriptase (New England BioLabs). Real-time PCR was performed using SYBRgreen PCR Master Mix (Quantabio) as described [13]. Relative mRNA levels of genes were normalized to the housekeeping gene 36B4 and fold change calculated relative to the vehicle-treated samples. Results are the average ± SD from at least two independent experiments carried out in triplicate. Primer sequences for the genes studied were obtained from the Harvard Primer Bank. Sequences are available on their website.
Proteasome activity analysis
Cells were exposed to 25 µM of the indicated test compounds or control vehicle prior to addition of the 20S proteasome substrate LLVY-AMC (7-amino-4-methylcoumarin) and free AMC fluorescence was monitored after 2 h at 380 nm/460 nm (excitation/emission). Values are mean ± SEM of 4 replicate determinations.
Statistical analyses
Statistics were calculated using analysis of variance (ANOVA), 2-way ANOVA with Dunnett’s multiple comparisons, or Student’s t test, as appropriate, using GraphPad Prism 8.0 software. Significance was designated as * for p < 0.05, ** for p < 0.01, *** for p < 0.001, and **** for p < 0.0001.
Results
Identification of potential combination candidates
In this study, we looked only at drugs that are currently being used as clinical anti-cancer agents or are in pre-clinical development for the treatment of cancer. To narrow down potential candidates, the landscape of potential cancer drugs was divided into (1) FDA-approved cancer drugs and (2) pre-clinical compounds of interest (Table 1). From the group of FDA-approved drugs, first priority was given to those already being used in the treatment of breast cancer, focusing on drugs that can inhibit either HR + breast cancer (such as the CDK4/6 inhibitors) or both HR + and TNBC. To further address the need for new molecular targeted therapies in TNBC, the next level of priority was given to FDA-approved drugs currently used to treat other cancers but shown to be highly effective in TNBC pre-clinical models, such as Crizotinib, Alpelisib, and Bortezomib. Pre-clinical compounds of interest were chosen on the basis of rational selection of targets related to FOXM1, expression of the target in HR + and/or TNBC, and effectiveness in pre-clinical cancer studies. An overview of the synergy studies pipeline is presented in Fig. 1.
We used triple-negative MDA-MB-231 cells and ER-positive (MCF7, ZR75-1) breast cancer cells as our initial screening cell lines. To select doses for combination studies in the cell lines, we first characterized the dose–response curves of each compound alone in suppression of cell viability and chose NB73, which displayed good FOXM1-specific activity in our previous studies [13, 15, 16], as the first FOXM1 inhibitor to screen in combination studies. We implemented a 3 × 3 matrix design with NB73 and each combination compound (Fig. 2), estimating 3 doses for each compound that would fall at or under its IC50. Percent inhibition matrices for four of these combinations (NB73 with Bortezomib, Crizotinib, Alisertib, or Alpelisib) are shown in Fig. 2. We quantified synergy using the Zero Interaction Potency (ZIP) model with the SynergyFinder web application, which produces a delta score at each dose pair to represent the level of synergy between two compounds and allows for visualization of all delta scores in a combination matrix as an interaction landscape [23]. The magnitude of the delta score is symmetrically centered around zero, where delta scores that are more negative are more antagonistic, while delta scores that are more positive are more synergistic. Generally, a delta score ranging from 0 to 10 is representative of additivity and scores above 10 are representative of synergistic interactions [21]. Negative scores (below zero) indicate compound antagonism. In some cases, synergy was also evaluated using the Chou–Talalay method [25].
ZIP synergy matrices of combinations of NB73 and candidate compounds. ZIP synergy score is average synergy score of all dose pairs in the matrix with 95% confidence interval, n = 4 at each treatment. MDA-MB-231 cells were treated for 72 h with the doses shown of NB73 or candidate compound alone and in combination at each dose pair
Our findings showed combinations of NB73 with Bortezomib had high delta scores, suggesting synergy (Fig. 2). Combination of Crizotinib or Alisertib with NB73 gave delta scores in the additivity range, while combinations with Alpelisib were neither additive nor synergistic (Fig. 2). Additional analysis of Crizotinib and Alisertib to quantify possible synergy confirmed that these combinations with NB73 were likely to be additive or only weakly synergistic, whereas NB73 with Bortezomib gave a high ZIP synergy score (Fig. 2), and we therefore continued with Bortezomib and also evaluated another proteasome inhibitor, TAK-243, in further studies.
FOXM1 inhibitors interact synergistically with proteasome inhibitors
The ubiquitin–proteasome pathway (UPP) is highly dysregulated and contributes to the pathobiology of many cancers, and small-molecule inhibition of 20S proteasome activity has become a successful treatment strategy in hematological malignancies, like mantle cell lymphoma and multiple myeloma [26]. Proteasome inhibitors act as potent cancer-suppressing agents through regulation of cell cycle proteins, inhibition of the NFkB pathway, induction of endoplasmic reticulum stress and the unfolded protein response (UPR) and importantly, induction of pro-apoptotic proteins [26, 27]. It has been shown that the proteasome inhibitor Bortezomib is a potent inducer of apoptosis in breast cancer cells [28]. Furthermore, proteasome addiction was identified as a vulnerability of triple-negative breast cancer cells relative to other breast cancer subtypes [29]. In previous studies, we found that the FOXM1 inhibitors we developed induce cell death and apoptosis in breast cancer cells [13], leading to the hypothesis that the combination of FOXM1 inhibitors with proteasome inhibitors may enhance this process. Thus, we hypothesized that combining our FOXM1-targeted compounds with proteasome inhibitors might effectively inhibit breast cancer cells through several potential mechanisms, including enhancement of cell cycle inhibition, UPR induction, and pro-apoptotic signaling.
Effects of FOXM1 inhibitor and proteasome inhibitors alone and together on the cell cycle and on apoptosis
As seen in Fig. 3 and Supp. Fig. S1, we evaluated the combination of FOXM1 inhibitors NB73, or NB55, or NB115 with Bortezomib or with TAK-243 (MLN7243). TAK-243 is a compound entering human clinical trials that inhibits the ubiquitin proteasome system (UPS) by inhibiting ubiquitin activating enzyme (UAE) rather than the 26S proteasome [30]. Bortezimib or TAK-243 with FOXM1 inhibitors synergistically inhibited estrogen receptor-positive MCF7 cell proliferation as quantified by the Zero Interaction Potency (ZIP) and Chou–Talalay models, as did FDI6, a different inhibitor of FOXM1 [31]. To further evaluate this, we examined a broader range of proteasome inhibitor concentrations both in the ER + MCF7 cells and in TNBC MDA-MB-231 cells (Fig. 4). We observed synergy between the FOXM1 inhibitor and proteasome inhibitor in MCF7 cells (Fig. 4a and b), but notably we found that viability of the TNBC cell lines MDA-MB-231 and MDA-MB-468 (data not shown) was not effectively suppressed by cotreatment with FOXM1 inhibitor and TAK-243 (Fig. 4c). In fact, cotreatments consistently gave negative ZIP synergy scores, indicating that the two types of inhibitors are likely acting antagonistically in these cells. This suggests that the 26S-directed more general proteasome inhibition by Bortezomib may be more broadly compatible with FOXM1 inhibition across cell types than is the novel UAE-targeting mechanism of TAK-243.
Assessment of synergistic interaction in growth suppression between FOXM1 inhibitors (NB73; FDI6) and proteasome inhibitors. MCF7 cells were treated for 72 h with the doses shown of FOXM1 inhibitor and proteasome inhibitor alone or in combination at each dose pair. Inhibition matrices, ZIP synergy matrices, 3D ZIP synergy plots, and Combination Index (CI) plots are shown for all combinations. a NB73 + TAK-243. b FDI6 + Bortezomib. c FDI6 + TAK-243. Inhibition matrices show the average of 4 replicates. ZIP Synergy score is average synergy score of all dose pairs in the matrix with 95% confidence interval, 4 replicates per treatment. Combination Index (CI) scores were calculated using the average of 4 replicates at each dose pair using CompuSyn software
Effect of FOXM1 inhibitor and proteasome inhibitor alone and together in HR + and in TNBC cells, studied over a broad range of concentrations. All cell treatments were for 72 h. a NB73 and TAK-243, MCF7 cells; b NB73 and Bortezomib, MDA-MB-231 cells; and c NB73 and TAK-243, MDA-MB-231 cells. Inhibition matrices show the average of 4 replicates. ZIP synergy score is average score of all dose pairs in the matrix
To further investigate the FOXM1 inhibitor/proteasome inhibitor combination, we performed cell cycle analysis with cells synchronized at the G1/S boundary with double-thymidine block and then treated with NB73 and Bortezomib or each compound alone. While both compounds alone arrested cells at the G0 and G1 phases, the combination of the compounds resulted in dramatic G2/M arrest of the majority of the cell population, indicating enhanced inhibition of cell cycle progression (Fig. 5a). Next, to investigate the effect of the combination on the apoptosis pathway, we tested the effect of the combination or of the compounds alone on caspase 3/7 activity (Fig. 5b). We treated MDA-MB-231 cells with doses of NB73 and Bortezomib that were expected to elicit little apoptotic activity alone. At these doses, NB73 alone did not increase caspase activity, while Bortezomib alone displayed a dose-dependent increase in caspase 3/7 activity, reaching a maximum of fourfold over Vehicle at the highest dose. When the compounds were combined, we saw a tenfold increase in caspase 3/7 activity over Vehicle at the 1.5-µM NB73 + 30-nM Bortezomib dose, which was 2.8-fold greater than the effect of Bortezomib alone at 30 nM. Thus, NB73, while having no effect on its own at the 0.375 or 1.5 µM dose, significantly enhanced the pro-apoptotic effect of Bortezomib.
Effect of NB73 and Bortezomib alone and in combination on the cell cycle and on caspase 3/7 activity. a MDA-MB-231 cells were synchronized by double-thymidine block and were released for 24 h with or without treatment with Vehicle, NB73 (1.5 µM), Bortezomib (5 or 15 nM) or the indicated combinations. The percent of cells in different phases of the cell cycle were monitored by flow cytometry. n = 2 experiments, mean + SD. b Caspase 3/7 activity was monitored after cell treatment with Veh, NB73 (at 0.375 or 1.5uM), Bortezomib (7.5 or 30 nM), or the indicated combinations for 24 h before using the Caspase-Glo assay; n = 3 experiments, mean + SD. *p < 0.05 and ****p < 0.0001 for combinations versus Bortezomib alone
We also evaluated whether there was possible proteasome-modulating activity of our NB compounds using a fluorescence-based proteasome substrate activity assay. While thiostrepton, lactacystin, and MG132 reduced proteasome activity as expected [14, 32], NB55, NB73, and FDI6 at quite high concentrations (25 µM) did not elicit any suppression of proteasome activity (Supp. Fig. S2). Thus, the NB and FDI6 FOXM1 inhibitors do not themselves have proteasome-modulating activity.
FOXM1 inhibitors act synergistically with CDK4/6 inhibitors to inhibit ER-positive breast cancer cells
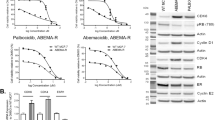
CDK4/6 inhibitors are currently used to treat HR +, HER2- breast cancers, providing a significant improvement in progression-free survival in these patients [33,34,35]. The cyclin-D-CDK4/6 complex is important for cell growth in response to mitogenic growth signals and is necessary for phosphorylation of checkpoint protein retinoblastoma (Rb), lifting its inhibition of entry into S phase [36, 37]. In addition, multi-site phosphorylation of FOXM1 by CDK4/6 in G1 and G1/S stabilizes FOXM1 against proteasome-mediated degradation and activates FOXM1 transcriptional activity, providing a rationale for the combination of FOXM1 inhibitors with CDK4/6 inhibitors [38]. In our extended dose studies we identified that the combination of NB73 or NB115 with Abemaciclib, Palbociclib, or Ribociclib gave very good synergy scores in ER-positive MCF7 cells (Fig. 6) and also in ER-positive ZR75-1 cells (Supp. Fig. S3).
Assessment of enhanced growth suppression of ER-positive breast cancer cells by FOXM1 inhibitors and CDK4/6 inhibitors. MCF7 cells were treated for 72 h with the doses shown of FOXM1 inhibitor or CDK4/6 inhibitor alone or in combination at each dose pair. Cell viability dose–response inhibition matrices, ZIP synergy matrices, and 3D ZIP synergy plots are shown for all combinations. a Cells treated with NB73 and Abemaciclib. b Cells treated with NB73 and Palbociclib. c Cells treated with NB73 and Ribociclib. Inhibition matrices show the average of 4 replicates. ZIP Synergy score is average synergy score of all dose pairs in each matrix
Regulation of gene expression by combined FOXM1 and CDK4/6 or proteasome inhibition
To begin to understand how the combination of NB73 and CDK4/6 inhibitor might be enhancing suppression of cell proliferation, we conducted gene expression studies in MCF7 cells treated with different dose combinations of NB73 and Palbociclib. In these studies, we found that combining the two compounds resulted in greater suppression of the expression of genes involved in cell cycle progression (AURKB, E2F1, PLK1, CCNB1, CENPF, and MCM3) than either compound alone (Fig. 7a–f). Likewise, the combination of NB73 and Bortezomib in TNBC MDA-MB-231 cells generally elicited greatly reduced expression of proliferation-related genes (E2F1, PCNA), survivin/BIRC5 gene, and of the FOXM1 gene itself at both 24 h and 48 h of treatment (Fig. 8).
Enhanced suppression of cell cycle genes by the combination of NB73 with Palbociclib. MCF7 cells were treated for 24 h with the nM doses shown of NB73 or Palbociclib alone or in combination as indicated. RNA was extracted from cells and gene expression was monitored by qRT-PCR. Assays were run in triplicate. Values are mean ± SEM. *p < 0.05; **p < 0.01; ***p < 0.001; and ****p < 0.0001 for combinations versus compounds alone
Enhanced up or down regulation of key gene expressions by combined treatment with NB73 plus Bortezomib. MDA-MB-231 cells were treated for 24 h or 48 h with Bortezomib (Bort, 6 nM) alone or NB73 alone (850 nM) or with both in combination. RNA was extracted from cells and gene expression was monitored by qRT-PCR. Assays were run in triplicate. Values are mean ± SEM. *p < 0.05; **p < 0.01; ***p < 0.001; and ****p < 0.0001 for combinations versus compounds alone
Discussion
We have identified several compounds that display synergistic effects when combined with FOXM1 inhibitors, with proteasome inhibitors and CDK4/6 inhibitors showing especially robust enhanced efficacy in suppressing breast cancer viability and related gene expressions.
Marked synergy between FOXM1 inhibitors and proteasome inhibitors
NB73 and Bortezomib synergistically inhibited cell proliferation, as did NB55 and NB115, two other FOXM1 inhibitors we developed [13]. Importantly, the combination of Bortezomib with FDI6, a FOXM1 inhibitor of a different chemical class [31], was also highly synergistic, implicating the FOXM1-specific effect of this interaction. Furthermore, we observed that the combination of NB73 plus Bortezomib resulted in dramatic arrest of cells at G2/M and induced more robust caspase 3/7 activity than either compound alone. Since our FOXM1 inhibitors suppress the expression of genes promoting cell cycle progression and enhance apoptosis stimulating genes, as do proteasome inhibitors [13, 26, 27], it appears that combined targeting of FOXM1 and proteasome activity may effectively inhibit breast cancer cells by several possible mechanisms, including enhancement of cell cycle inhibition, induction of the unfolded protein response, and stimulation of pro-apoptotic signaling [13].
The success of proteasome inhibitors in hematological malignancies, such as malignant myeloma, has made them an important area of investigation for other cancer types, particularly those that display dependence on the ubiquitin–proteasome system [39]. Triple-negative breast cancers are an example of such malignancies, and proteasome inhibitors have been identified in several pre-clinical drug screens as agents that have cytotoxic and pro-apoptotic activity in this subtype, which is generally resistant to many other treatments [29, 40, 41]. Despite its promise in pre-clinical solid tumor animal models, however, clinical studies of Bortezomib have failed to demonstrate efficacy in solid tumors [42]. This is likely due to the non-optimal pharmacokinetic properties of Bortezomib, which appear to result in impaired distribution to solid tumors [43]. Therefore, we conducted some studies with TAK-243, a novel inhibitor of the unfolded protein system (UPS) that targets ubiquitin-activating enzyme (UAE) and thus is mechanistically and chemically distinct from other proteasome inhibitors which are targeted to the 26S proteasome [44]. TAK-243 has shown effective tumor suppression in pre-clinical solid tumor studies and thus is a potential future avenue for UPS-inhibiting therapy in cancer. In addition, by inhibiting the ubiquitin-activating enzyme (UAE), TAK-243 treatment impairs the function of multiple E2 enzymes needed for DNA damage repair, resulting in enhanced killing of lung carcinoma cells by UV irradiation due to unresolved DNA damage repair [44]. Notably, we found that ER-positive breast cancer cells (MCF7 and ZR75-1) showed synergy in suppression of cell viability with FOXM1 inhibitors and Bortezomib or TAK-243, whereas the TNBC cells examined showed synergy only with FOXM1 inhibitor and Bortezomib but not with TAK-243.
Synergy of FOXM1 inhibitors with CDK4/6 inhibitors in impeding growth of ER-containing breast cancer cells
The cyclin D-CDK4/6 complex regulates the G1/S transition through phosphorylation of Rb and FOXM1 in cells entering the cell cycle after mitogenic growth signaling [45, 46]. Hence, blockade of cyclin-dependent kinase phosphorylation and activation of FOXM1 by CDK4/6 inhibitors may well contribute in further enhancing the suppression of FOXM1 activity by FOXM1 inhibitors. Most luminal breast cancers display upregulation of the Rb pathway, and CDK4/6 inhibitors like Abemaciclib or Palbociclib in combination with hormone therapy are an important treatment avenue for advanced-stage or metastatic HR +, HER2- breast cancer [47]. Nonetheless, resistance to these treatments frequently develops. Thus, we examined the rational combination of CDK4/6 inhibition with FOXM1 inhibition in ER-positive breast cancer cell lines, which theoretically modulates three important regulators of the G1/S transition (FOXM1, CDK4/6, and Rb). Our studies with ER-positive cells (MCF7 and ZR75-1) revealed enhanced effectiveness when Abemaciclib, Palbociclib, or Ribociclib were combined with NB73 or NB115, as well as with FDI6. Indeed, combinations of CDK4/6 inhibitors with other agents, such as PI3K inhibitor or other antiestrogens, are being explored clinically [48]. Because TNBC tumors frequently exhibit functional loss of Rb, triple-negative breast cancers are currently considered poor candidates for CDK4/6 inhibition [49]. Still, precision medicine approaches may make it possible to identify if specific subsets of TNBC patients can benefit from CDK4/6-targeted therapy alone or in combination with FOXM1 inhibitors in the future. Recent studies have demonstrated the sensitivity of the luminal androgen receptor (LAR) TNBC subtype to CDK4/6 inhibition [50]. In addition, other cyclin-dependent kinase inhibitors are currently in clinical development that show promise as possible therapeutic approaches in TNBC, including the CDK1/5/9 inhibitor Dinaciclib [51].
Possible benefits from combination therapies
Maintenance of efficacy with possible reduced toxicity is a key consideration in the study of any combination therapies [52]. The toxicity profile of Bortezomib is characterized by a high incidence of polyneuropathy (80%) in multiple myeloma (MM) patients, but this finding is complicated by the fact that neuropathy is a common symptom of MM in itself, as well as from other drugs used to treat MM, like vinca alkaloids [53]. Other side effects include thrombocytopenia, neutropenia, anemia, fatigue, and diarrhea [26]. Nevertheless, Bortezomib has been combined with lenalidomide and dexamethasone in an MM treatment regimen that was well tolerated in phase II clinical studies and is frequently used as initial therapy in the USA [53]. In addition, next-generation proteasome inhibitors like Carfilzomib appear to have improved toxicity profiles compared to Bortezomib, providing an additional avenue for future investigation in combination treatments with FOXM1 inhibitors [54].
As reported in the MONARCH 2 phase III clinical trial, the most common toxicities for Abemaciclib as a monotherapy are diarrhea, nausea, and fatigue, with most patients reporting no or grade 1–2 diarrhea; grade 3–4 neutropenia occurred in 10% of patients [55]. When Abemaciclib was combined with aromatase inhibitors (AI) in the MONARCH 3 phase III trial, the most increased adverse effect was grade 3 or 4 neutropenia, which is reversible and can be clinically managed without compromising treatment efficacy [56]. Although our FOXM1 inhibitors showed no obvious toxicity in pre-clinical in vivo test systems [13, 17], as we learn more about the possible potential side effects of our FOXM1 inhibitors in vivo, it will be important to select combination candidates with toxicity profiles that are not exacerbated by the addition of a FOXM1 inhibitor.
Conclusion
The present studies show that the FOXM1 inhibitors work synergistically with several different compounds to suppress breast cancer cells. From profiles of the compounds having the most effective combined activity, it appears that FOXM1 inhibitors may synergize best with compounds that directly affect the cell cycle and apoptotic pathways. More specifically, our findings suggest that FOXM1 inhibition, along with CDK4/6 or proteasome inhibition might ultimately prove to be a useful approach for breast cancer treatment because synergy is especially noticeable at low doses of each inhibitor, suggesting that good suppression of cancer cell viability might still be maintained with reduced doses of both agents with the additional benefit of reduced side effects. Clearly, future studies will be needed to further characterize the mechanisms behind these synergistic combinations, as well as explore other novel combinations that have the potential to complement the biological effects of FOXM1 inhibition.
Data availability
All data are available within the manuscript or may be obtained from the Corresponding Author (B.S.K) upon reasonable request.
References
Harbeck N, Penault-Llorca F, Cortes J, Gnant M, Houssami N, Poortmans P, Ruddy K, Tsang J, Cardoso F (2019) Breast cancer. Nat Rev Dis Primers. https://doi.org/10.1038/s41572-019-0111-2
Anastasiadi Z, Lianos GD, Ignatiadou E, Harissis HV, Mitsis M (2017) Breast cancer in young women: an overview. Updates Surg 693:313–317. https://doi.org/10.1007/s13304-017-0424-1
Waks AG, Winer EP (2019) Breast cancer treatment: a review. JAMA 3213:288–300. https://doi.org/10.1001/jama.2018.19323
Lehmann BD, Pietenpol JA (2015) Clinical implications of molecular heterogeneity in triple negative breast cancer. Breast 24(2):S36-40. https://doi.org/10.1016/j.breast.2015.07.009
Abramson VGMI (2014) Molecular heterogeneity of triple negative breast cancer. Current Breast Cancer Reports 63:154–158
Wang DY, Jiang Z, Ben-David Y, Woodgett JR, Zacksenhaus E (2019) Molecular stratification within triple-negative breast cancer subtypes. Sci Rep 91:19107. https://doi.org/10.1038/s41598-019-55710-w
Lyons TG (2019) Targeted therapies for triple-negative breast cancer. Curr Treat Options Oncol 2011:82. https://doi.org/10.1007/s11864-019-0682-x
Lu XF, Zeng LWQ, Chen CF, Sun SM, Lin HY (2018) FoxM1 is a promising candidate target in the treatment of breast cancer. Oncotarget 91:842–852. https://doi.org/10.18632/oncotarget.23182
Huang S, Xu W, Hu P, Lakowski TM (2019) Integrative analysis reveals subtype-specific regulatory determinants in triple negative breast cancer. Cancers (Basel). https://doi.org/10.3390/cancers11040507
Tan Y, Wang Q, Xie Y, Qiao X, Zhang S, Wang Y, Yang Y, Zhang B (2019) Identification of FOXM1 as a specific marker for triplenegative breast cancer. Int J Oncol 541:87–97. https://doi.org/10.3892/ijo.2018.4598
Bergamaschi A, Madak-Erdogan Z, Kim YJ, Choi YL, Lu H, Katzenellenbogen BS (2014) The forkhead transcription factor FOXM1 promotes endocrine resistance and invasiveness in estrogen receptor-positive breast cancer by expansion of stem-like cancer cells. Breast Cancer Res 165:436. https://doi.org/10.1186/s13058-014-0436-4
Sanders DA, Ross-Innes CS, Beraldi D, Carroll JS, Balasubramanian S (2013) Genome-wide mapping of FOXM1 binding reveals co-binding with estrogen receptor alpha in breast cancer cells. Genome Biol. https://doi.org/10.1186/gb-2013-14-1-r6
Ziegler Y, Laws MJ, Sanabria Guillen V, Kim SH, Dey P, Smith BP, Gong P, Bindman N, Zhao Y, Carlson K et al (2019) Suppression of FOXM1 activities and breast cancer growth in vitro and in vivo by a new class of compounds. NPJ Breast Cancer 5:45. https://doi.org/10.1038/s41523-019-0141-7
Gartel AL (2017) FOXM1 in cancer: interactions and vulnerabilities. Cancer Res 7712:3135–3139. https://doi.org/10.1158/0008-5472.CAN-16-3566
Dey P, Wang A, Ziegler Y, Kim SH, El-Ashry D, Katzenellenbogen JA, Katzenellenbogen BS (2020) Suppression of tumor growth, metastasis, and signaling pathways by reducing FOXM1 activity in triple negative breast cancer. Cancers (Basel) 129:2677. https://doi.org/10.3390/cancers12092677
Ziegler Y, Guillen VS, Kim SH, Katzenellenbogen JA, Katzenellenbogen BS (2021) Transcription regulation and genome rewiring governing sensitivity and resistance to FOXM1 inhibition in breast cancer. Cancers (Basel) 1324:6282. https://doi.org/10.3390/cancers13246282
Cheng Y, Sun F, Thornton K, Jing X, Dong J, Yun G, Pisano M, Zhan F, Kim SH, Katzenellenbogen JA et al (2022) FOXM1 regulates glycolysis and energy production in multiple myeloma. Oncogene 4132:3899–3911. https://doi.org/10.1038/s41388-022-02398-4
Liu C, Barger CJ, Karpf AR (2021) FOXM1: a multifunctional oncoprotein and emerging therapeutic target in ovarian cancer. Cancers (Basel). https://doi.org/10.3390/cancers13123065
Liu C, Muñoz-Trujillo C, Kim SH, Katzenellenbogen J, Katzenellenbogen B, Karpf A: New 1,1-diarylethylene FOXM1 inhibitors potently reduce intracellular FOXM1 and suppress high-grade serous ovarian carcinoma cell viability. In.: Research Square; 2022.
Madak-Erdogan Z, Charn TH, Jiang Y, Liu ET, Katzenellenbogen JA, Katzenellenbogen BS (2013) Integrative genomics of gene and metabolic regulation by estrogen receptors alpha and beta, and their coregulators. Mol Syst Biol 9:676. https://doi.org/10.1038/msb.2013.28
Ianevski A, Giri AK, Aittokallio T (2020) SynergyFinder 2.0: visual analytics of multi-drug combination synergies. Nucleic Acids Res 48:W488–W493. https://doi.org/10.1093/nar/gkaa216
Loewe S (1953) The problem of synergism and antagonism of combined drugs. Arzneimittelforschung 36:285–290
Yadav B, Wennerberg K, Aittokallio T, Tang J (2015) Searching for drug synergy in complex dose-response landscapes using an interaction potency model. Comput Struct Biotechnol J 13:504–513. https://doi.org/10.1016/j.csbj.2015.09.001
Zhao W, Sachsenmeier K, Zhang L, Sult E, Hollingsworth RE, Yang H (2014) A new Bliss independence model to analyze drug combination data. J Biomol Screen 195:817–821. https://doi.org/10.1177/1087057114521867
Chou TC (2010) Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res 702:440–446. https://doi.org/10.1158/0008-5472.CAN-09-1947
Manasanch EE, Orlowski RZ (2017) Proteasome inhibitors in cancer therapy. Nat Rev Clin Oncol 147:417–433. https://doi.org/10.1038/nrclinonc.2016.206
Kambhampati S, Wiita AP (2020) Lessons learned from proteasome inhibitors, the paradigm for targeting protein homeostasis in cancer. Adv Exp Med Biol 1243:147–162. https://doi.org/10.1007/978-3-030-40204-4_10
Kretowski R, Borzym-Kluczyk M, Cechowska-Pasko M (2014) Efficient induction of apoptosis by proteasome inhibitor: bortezomib in the human breast cancer cell line MDA-MB-231. Mol Cell Biochem 3891–2:177–185. https://doi.org/10.1007/s11010-013-1939-5
Petrocca F, Altschuler G, Tan SM, Mendillo ML, Yan H, Jerry DJ, Kung AL, Hide W, Ince TA, Lieberman J (2013) A genome-wide siRNA screen identifies proteasome addiction as a vulnerability of basal-like triple-negative breast cancer cells. Cancer Cell 242:182–196. https://doi.org/10.1016/j.ccr.2013.07.008
Barghout SH, Patel PS, Wang X et al (2019) Preclinical evaluation of the selective small-molecule UBA1 inhibitor, TAK-243, in acute myeloid leukemia. Leukemia 33:37–51
Gormally MV, Dexheimer TS, Marsico G, Sanders DA, Lowe C, Matak-Vinkovic D, Michael S, Jadhav A, Rai G, Maloney DJ et al (2014) Suppression of the FOXM1 transcriptional programme via novel small molecule inhibition. Nat Commun 5:5165. https://doi.org/10.1038/ncomms6165
Guo N, Peng Z (2013) MG132, a proteasome inhibitor, induces apoptosis in tumor cells. Asia Pac J Clin Oncol 91:6–11. https://doi.org/10.1111/j.1743-7563.2012.01535.x
Cottu P, Ring A, Abdel-Razeq H, Marchetti P, Cardoso F, Salvador Bofill J, Martin M, Menon-Singh L, Wu J, De Laurentiis M (2022) Ribociclib plus letrozole in subgroups of special clinical interest with hormone receptor-positive, human epidermal growth factor receptor 2-negative advanced breast cancer: Subgroup analysis of the phase IIIb CompLEEment-1 trial. Breast 62:75–83. https://doi.org/10.1016/j.breast.2022.01.016
Eggersmann TK, Degenhardt T, Gluz O, Wuerstlein R, Harbeck N (2019) CDK4/6 inhibitors expand the therapeutic options in breast cancer: palbociclib, ribociclib and abemaciclib. BioDrugs 332:125–135. https://doi.org/10.1007/s40259-019-00337-6
Ha MJ, Raghavendra AS, Kettner NM, Qiao W, Damodaran S, Layman RM, Kelly KH, Shen Y, Tripathy D, Keyomarsi K (2022) Palbociclib plus endocrine therapy significantly enhances overall survival of HR+/HER2- metastatic breast cancer patients compared to endocrine therapy alone in the second-line setting-a large institutional study. Int J Cancer. https://doi.org/10.1002/ijc.33959
Matutino A, Amaro C, Verma S (2018) CDK4/6 inhibitors in breast cancer: beyond hormone receptor-positive HER2-negative disease. Ther Adv Med Oncol 10:1758835918818346. https://doi.org/10.1177/1758835918818346
Torres-Guzman R, Calsina B, Hermoso A, Baquero C, Alvarez B, Amat J, McNulty AM, Gong X, Boehnke K, Du J et al (2017) Preclinical characterization of abemaciclib in hormone receptor positive breast cancer. Oncotarget 841:69493–69507. https://doi.org/10.18632/oncotarget.17778
VanArsdale T, Boshoff C, Arndt KT, Abraham RT (2015) Molecular pathways: targeting the cyclin D-CDK4/6 axis for cancer treatment. Clin Cancer Res 2113:2905–2910. https://doi.org/10.1158/1078-0432.CCR-14-0816
Tsvetkov P, Adler J, Myers N, Biran A, Reuven N, Shaul Y (2018) Oncogenic addiction to high 26S proteasome level. Cell Death Dis 97:773. https://doi.org/10.1038/s41419-018-0806-4
Gautam P, Karhinen L, Szwajda A, Jha SK, Yadav B, Aittokallio T, Wennerberg K (2016) Identification of selective cytotoxic and synthetic lethal drug responses in triple negative breast cancer cells. Mol Cancer 151:34. https://doi.org/10.1186/s12943-016-0517-3
Wali VB, Langdon CG, Held MA, Platt JT, Patwardhan GA, Safonov A, Aktas B, Pusztai L, Stern DF, Hatzis C (2017) Systematic drug screening identifies tractable targeted combination therapies in triple-negative breast cancer. Cancer Res 772:566–578. https://doi.org/10.1158/0008-5472.CAN-16-1901
Roeten MSF, Cloos J, Jansen G (2018) Positioning of proteasome inhibitors in therapy of solid malignancies. Cancer Chemother Pharmacol 812:227–243. https://doi.org/10.1007/s00280-017-3489-0
Caravita T, de Fabritiis P, Palumbo A, Amadori S, Boccadoro M (2006) Bortezomib: efficacy comparisons in solid tumors and hematologic malignancies. Nat Clin Pract Oncol 37:374–387. https://doi.org/10.1038/ncponc0555
Hyer ML, Milhollen MA, Ciavarri J, Fleming P, Traore T, Sappal D, Huck J, Shi J, Gavin J, Brownell J et al (2018) A small-molecule inhibitor of the ubiquitin activating enzyme for cancer treatment. Nat Med 242:186–193. https://doi.org/10.1038/nm.4474
Anders L, Ke N, Hydbring P, Choi YJ, Widlund HR, Chick JM, Zhai H, Vidal M, Gygi SP, Braun P et al (2011) A systematic screen for CDK4/6 substrates links FOXM1 phosphorylation to senescence suppression in cancer cells. Cancer Cell 205:620–634. https://doi.org/10.1016/j.ccr.2011.10.001
Liu G, Sun Y, Ji P, Li X, Cogdell D, Yang D, Parker Kerrigan BC, Shmulevich I, Chen K, Sood AK et al (2014) MiR-506 suppresses proliferation and induces senescence by directly targeting the CDK4/6-FOXM1 axis in ovarian cancer. J Pathol 2333:308–318. https://doi.org/10.1002/path.4348
Sledge GW Jr, Toi M, Neven P, Sohn J, Inoue K, Pivot X, Burdaeva O, Okera M, Masuda N, Kaufman PA et al (2017) MONARCH 2: abemaciclib in combination with fulvestrant in women with HR+/HER2- advanced breast cancer who had progressed while receiving endocrine therapy. J Clin Oncol 3525:2875–2884. https://doi.org/10.1200/JCO.2017.73.7585
Rampioni Vinciguerra GL, Sonego M, Segatto I, Dall’Acqua A, Vecchione A, Baldassarre G, Belletti B (2022) CDK4/6 inhibitors in combination therapies: better in company than alone: a mini review. Front Oncol 12:891580. https://doi.org/10.3389/fonc.2022.891580
Patel JM, Torous V, Hacker MR, Nalven T, Garber JE, Alexander BM, Lee LJ, Collins LC, Schnitt SJ, Tung NM (2017) Retinoblastoma (Rb) protein expression in triple-negative breast cancer. J Clin Oncol 3515:1097–1097. https://doi.org/10.1200/JCO.2017.35.15_suppl.1097
Pernas S, Tolaney SM, Winer EP, Goel S (2018) CDK4/6 inhibition in breast cancer: current practice and future directions. Ther Adv Med Oncol 10:1758835918786451. https://doi.org/10.1177/1758835918786451
Nie L, Wei Y, Zhang F, Hsu YH, Chan LC, Xia W, Ke B, Zhu C, Deng R, Tang J et al (2019) CDK2-mediated site-specific phosphorylation of EZH2 drives and maintains triple-negative breast cancer. Nat Commun 101:5114. https://doi.org/10.1038/s41467-019-13105-5
Kahn AM, Blenman KRM, Sonis ST, Lustberg MB (2022) Strategies to mitigate the toxicity of cancer therapeutics. Adv Cancer Res 155:215–244. https://doi.org/10.1016/bs.acr.2022.02.006
Merin NM, Kelly KR (2014) Clinical use of proteasome inhibitors in the treatment of multiple myeloma. Pharmaceuticals (Basel) 81:1–20. https://doi.org/10.3390/ph8010001
Mushtaq A, Kapoor V, Latif A, Iftikhar A, Zahid U, McBride A, Abraham I, Riaz IB, Anwer F (2018) Efficacy and toxicity profile of carfilzomib based regimens for treatment of multiple myeloma: a systematic review. Crit Rev Oncol Hematol 125:1–11. https://doi.org/10.1016/j.critrevonc.2018.02.008
Martin JM, Goldstein LJ (2018) Profile of abemaciclib and its potential in the treatment of breast cancer. Onco Targets Ther 11:5253–5259. https://doi.org/10.2147/OTT.S149245
Cazzaniga ME, Danesi R, Girmenia C, Invernizzi P, Elvevi A, Uguccioni M, NetworkEr (2019) Management of toxicities associated with targeted therapies for HR-positive metastatic breast cancer: a multidisciplinary approach is the key to success. Breast Cancer Res Treat 1763:483–494. https://doi.org/10.1007/s10549-019-05261-5
Acknowledgements
This research was supported by grants from the Breast Cancer Research Foundation (BCRF-083 to BSK and BCRF-084 to JAK and BSK), the NIH/NCI (1R01 CA220284 to BSK and JAK), the Julius and Mary Landfield Cancer Research Fund (to BSK), and the NIH Training Program T32 GMO70421 Fellowship to VSG.
Funding
This research was supported by grants from the Breast Cancer Research Foundation (BCRF-083 to BSK and BCRF-084 to JAK and BSK) and the NIH/NCI (1R01 CA220284 to BSK and JAK), the Julius and Mary Landfield Cancer Research Fund (to BSK), and the NIH Training Program T32 GMO70421 Fellowship to VSG.
Author information
Authors and Affiliations
Contributions
BSK, VSG, YZ, and JAK conceived the project and provided leadership for the project. VSG, YZ, CG, SK, PD, BNP, NZD, SHK, JAK, and BSK carried out experiments and/or analyzed data. VSG, BSK, YZ, and JAK wrote the manuscript. All authors discussed the results and provided input and edits on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
BSK, JAK, and SHK are co-inventors on a Provisional Application filed by the University of Illinois to cover the FOXM1 inhibitor compounds described in this paper. The other authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or involving animals performed by any of the authors.
Informed consent
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guillen, V.S., Ziegler, Y., Gopinath, C. et al. Effective combination treatments for breast cancer inhibition by FOXM1 inhibitors with other targeted cancer drugs. Breast Cancer Res Treat 198, 607–621 (2023). https://doi.org/10.1007/s10549-023-06878-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-023-06878-3