Abstract
Purpose
Previous studies indicate that breast cancer molecular subtypes differ with respect to their dependency on autophagy, but our knowledge of the differential expression and prognostic significance of autophagy-related biomarkers in breast cancer is limited.
Methods
Immunohistochemistry (IHC) was performed on tissue microarrays from a large population of 3992 breast cancer patients divided into training and validation cohorts. Consensus staining scores were used to evaluate the expression levels of autophagy proteins LC3B, ATG4B, and GABARAP and determine the associations with clinicopathological variables and molecular biomarkers. Survival analyses were performed using the Kaplan–Meier function and Cox proportional hazards regression models.
Results
We found subtype-specific expression differences for ATG4B, with its expression lowest in basal-like breast cancer and highest in Luminal A, but there were no significant associations with patient prognosis. LC3B and GABARAP levels were highest in basal-like breast cancers, and high levels were associated with worse outcomes across all subtypes (DSS; GABARAP: HR 1.43, LC3B puncta: HR 1.43). High ATG4B levels were associated with ER, PR, and BCL2 positivity, while high LC3B and GABARAP levels were associated with ER, PR, and BCL2 negativity, as well as EGFR, HER2, HER3, CA-IX, PD-L1 positivity, and high Ki67 index (p < 0.05 for all associations). Exploratory multi-marker analysis indicated that the combination of ATG4B and GABARAP with LC3B could be useful for further stratifying patient outcomes.
Conclusions
ATG4B levels varied across breast cancer subtypes but did not show prognostic significance. High LC3B expression and high GABARAP expression were both associated with poor prognosis and with clinicopathological characteristics of aggressive disease phenotypes in all breast cancer subtypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Macroautophagy (herein autophagy) is an evolutionarily conserved lysosome-mediated degradation and recycling process shown to contribute to both tumor suppression and tumor progression, depending primarily on the stage of tumorigenesis [1,2,3]. In advanced malignancies, autophagy promotes cancer cell survival and contributes to cancer progression and drug resistance, hence becoming a promising target for anticancer therapy [1,2,3]. Recent preclinical and clinical trial findings have led to a resurgent interest in anticancer autophagy inhibition strategies [4], driving an accompanying need to better understand the potential biomarker utility of autophagy proteins. Autophagy is a multi-step process that involves more than 30 core autophagy-related (ATG) proteins [5,6,7]. These proteins act to initiate, elongate, and complete the formation of double-membrane structures called autophagosomes that encapsulate portions of cytoplasm. Autophagosomes fuse with lysosomes to form autolysosomes where the contents are degraded, released and recycled by the cell [1, 8, 9].
Among the key autophagy proteins are the Atg8 and ATG4 families. In mammals, the Atg8 family of ubiquitin-like proteins is represented by two subfamilies: the microtubule-associated protein 1 light chain 3 (MAP1LC3, or LC3) subfamily, consisting of LC3A, LC3B, LC3B2, and LC3C, and the gamma-aminobutyric acid receptor-associated protein (GABARAP) subfamily that includes GABARAP, GABARAPL1, and Golgi-associated ATPase enhancer of 16 kDa (GATE-16)/GABARAPL2 [10]. Each family member, while demonstrating some functional redundancy with other members, plays distinct roles in the autophagy process [11, 12]. For example, LC3s are required for the elongation step [11, 13], whereas GABARAPs were shown to play important roles in autophagy initiation [13] and autophagosome closure [11]. The LC3 and GABARAP proteins are processed by the ATG4B cysteine protease at two different steps in the autophagy process. First, ATG4B cleaves the carboxyl terminus of newly synthesized pro-LC3 to generate LC3-I [14] enabling its conjugation to phosphatidylethanolamine (PE) to form membrane-bound LC3-II, which is required for autophagosome elongation. Second, ATG4B functions in delipidation of LC3-II from the autophagosome membrane to ensure recycling of LC3-I in the cell [15]. Other ATG4 family members (ATG4A, ATG4C, and ATG4D) have been described in mammalian cells, though ATG4B displays the broadest substrate specificity and greatest affinity for pro-LC3B [16,17,18] and can also process other family members including GABARAP. ATG4B has become an attractive therapeutic target [19,20,21,22,23,24,25], and several preclinical studies reported ATG4B inhibition as an effective strategy to sensitize cancer cells to chemotherapy [19, 26,27,28].
A variety of strategies to develop clinical biomarkers for monitoring autophagy in cancer patients are under investigation. Due to the dynamic nature of autophagy, complexity of its regulation, and autophagy-independent roles of ATG proteins, the assessment of several different markers in the same patient has been recommended. The most common biomarker approach explored to date in various cancers is the immunohistochemical analysis of autophagy proteins, particularly LC3B, in tumor tissue (reviewed in [3, 29]). While the overall levels of LC3B were investigated previously in many different tumor types, only a few more recent studies [30,31,32,33] distinguished between the processed forms of LC3B, namely cytosolic (LC3B-I) and membrane-bound (LC3B-II, or LC3B “puncta”) forms. Only an LC3B punctate pattern in cells represents the membrane-bound LC3B-PE form and can be used to infer levels of autophagosomes [34, 35].
By RNA expression profile, breast cancer is classified into four main molecular subtypes, namely Luminal A (Lum A), Luminal B (Lum B), basal-like, and HER2-enriched. Alternative clinical and immunohistochemical sub-classifications have been described, including triple-negative breast cancer (TNBC), which is immunohistochemically negative for ER, PR, and HER2 receptors. Although often used interchangeably with basal-like, it is only approximately 70–75% of TNBC that has a basal-like expression profile [36, 37]. These distinct breast cancer subtypes have shown differential sensitivity to autophagy modulation [26, 38, 39], but our knowledge of autophagy protein expression and biomarker potential in breast cancer is still limited.
Here, we report the differential expression levels of ATG4B, GABARAP, and LC3B in the major breast cancer subtypes. We demonstrate significant associations between these autophagy-related proteins, clinicopathological characteristics of the disease, and established biomarkers. In addition, following the reporting recommendations for tumor marker prognostic studies (REMARK) [40], we report the prognostic values of ATG4B, GABARAP, and LC3B alone and in combination in a large cohort of breast cancer patients.
Results
Patient demographics and pathologic data
In our cohort of 3992 breast cancer patients, split into training and validation sets (see Sect. 2), the mean age at diagnosis was 59.1 years, and the median follow-up time (the time of diagnosis to time of event or last follow-up) was 12.6 years; the range for follow-up was 1 month to 18.5 years (Supplementary Table S1). The median tumor size was 2 cm; 53.5% of patients had grade 3 tumors, 43% were node positive, 70% were ER positive, and 13% were HER2 positive. Adjuvant systemic therapy was given to 58% (Supplementary Table S1). Tables 1, 2, 3 and Supplementary Table S2 show summary statistics of clinicopathological variables of interest split by ATG4B, GABARAP, and LC3B status. The values highlighted below are those from the validation cohort and include only those results that were independently significant in both training and validation sets unless otherwise stated (see Sect. 2). The separate results for training and combined cohorts are provided in Supplementary Tables S3, S4, and S6.
Differential ATG4B expression across breast cancer subtypes and biomarker associations
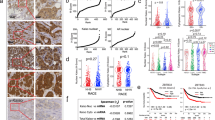
To determine the clinical relevance of ATG4B expression in breast cancer, we examined ATG4B cytoplasmic staining (Fig. 1) in our cohort and evaluated its association with breast cancer biomarkers. As shown in Figs. 2 and 3, ATG4B was detected at significantly lower levels in ER−/PR−/HER2− (triple-negative, TNP) vs. non-triple-negative breast cancers (p < 0.0001), and basal-like vs. non-basal molecular subtypes (p = 0.003). ATG4B expression did not significantly correlate with patient age at diagnosis, tumor size, or nodal status. However, as shown in Table 1, Fig. 3, and Supplementary Table S3, there was a significant association between ATG4B protein expression and lower tumor grade (p < 0.0001) and ER and PR positivity (p < 0.0001). In addition, there was a significant positive association between ATG4B and BCL2 (p = 0.02). There was a negative association between ATG4B and EGFR (p = 0.02), as well as ATG4B and Ki67 (p = 0.008).
Immunohistochemical staining showing expression of LC3B, ATG4B, and GABARAP in breast cancer patient samples. Representative images for LC3B cytoplasmic (H-score) and puncta (categorical) expression: a 100E; b 200D; c 200C; d 200B; e 300B; f 300A with zoomed-in image showing cells with high concentrations of individual puncta (circle) and clusters (long arrow). The short arrows in b–e point to puncta in tumor cells. Representative images for ATG4B expression (H-score): g 100; h 200; i 300. Representative images for GABARAP expression (H-score): j 100; k 200; l 300
In our patient cohort, the common clinicopathologic variables including patient age at diagnosis, tumor size, histologic grade, nodal status, ER/PR/HER2 status, and Ki67 expression were all statistically significant predictors of overall survival (OS), disease-specific survival (DSS), and relapse-free survival (RFS) on univariate survival analysis (Supplementary Table S5). ATG4B expression was not found to be significantly associated with patient prognosis (Supplementary Table S4).
Differential GABARAP expression across breast cancer subtypes, biomarker associations, and association with poor prognosis
To determine the clinical relevance of GABARAP expression in breast cancer, we scored its cytoplasmic staining pattern (Fig. 1, Supplementary Fig. S1) and evaluated its association with known biomarkers and patient survival. As shown in Fig. 2, basal-like and triple-negative breast cancers had the highest expression levels of GABARAP across all breast cancer subtypes. High GABARAP expression was associated with markers of a more aggressive phenotype, such as larger tumor size, negative ER and PR status, positive EGFR and CK5/6 expression, HER2 and HER3 positivity, high Ki67, low BCL2, high IGF1R expression, as well as high PD-L1 and CA-IX expression (Table 2, Fig. 4, Supplementary Table S3). All of these marker associations were statistically significant (p < 0.05 for all associations) in the validation cohort. High GABARAP expression levels were associated with poor overall survival and disease-specific survival in the validation cohort at 5- and 10-years cutpoints of follow-up time (Fig. 5a, Table 4): HR 1.27 (95% CI 1.06–1.52), p = 0.009 for OS at 10 years; HR 1.48 (95% CI 1.15–1.92), p = 0.002 for OS at 5 years; HR 1.43 (95% CI 1.15–1.80), p = 0.001 for DSS at 10 years; HR 1.69 (95% CI 1.25–2.32), p < 0.001 for DSS at 5 years. There were no breast cancer subtype-specific survival associations with GABARAP that were statistically significant in both training and validation cohorts (Supplementary Table S4). To determine if GABARAP expression was an independent marker of prognosis, we performed multivariable analyses with all clinicopathological parameters. GABARAP expression was not found to be an independent prognostic factor (p > 0.05) (Table 5, Supplementary Table S6).
Kaplan–Meier survival curves according to GABARAP expression (a), LC3B cytoplasmic expression (b), LC3B puncta expression (c), LC3B cytoplasmic and puncta expression combined (d) in the validation set. Exploratory analyses of the three markers (LC3B puncta, GABARAP, and ATG4B) combined (e) in the entire cohort. L− low (scored as D/E) LC3B expression, L+ high (scored as A/B/C) LC3B expression, G− low (H-score < 175) GABARAP expression, G+ high (H-score > 175) GABARAP expression, A− low (H-score < 150) ATG4B expression, A+ high (H-score > 150) ATG4B expression
High diffuse or punctate LC3B expression is associated with predictors of more aggressive phenotype and poor prognosis in breast cancer
To evaluate LC3B biomarker potential in this large cohort, we scored both diffuse (H-score) and punctate staining patterns (Fig. 1). Among all intrinsic subtypes, basal-like breast cancers had the highest levels of both punctate and diffuse LC3B (Fig. 2). Higher LC3B expression (H-score and puncta score) was significantly associated with several clinicopathological variables characteristic of a more aggressive phenotype (Table 3, Fig. 6, Supplementary Table S3), including higher tumor grade, lymphovascular invasion, ER/PR negativity, positive HER2, HER3, CK5/6, EGFR, and CA-IX expression, as well as high Ki67 index. There was a statistically significant positive association between LC3B and GABARAP (p ≤ 0.0001); no statistically significant association between LC3B and ATG4B, or between GABARAP and ATG4B was established. As shown in Fig. 5b–d and Table 4, high diffuse expression of LC3B was associated with poor DSS at 18 years (HR 1.31 (95% CI 1.07–1.61), p = 0.009) and poor OS, DSS and RFS at 10 years and 5 years (p < 0.05 and p ≤ 0.003; Table 4); high puncta levels of LC3B were associated with poor RFS at 5 years (HR 1.41 (95% CI 1.13–1.76), p = 0.002) and with poor OS and DSS at 5-, 10- and 18-year cutpoints of follow-up time (Fig. 5a, Table 4): HR 1.22 (95% CI 1.02–1.46), p = 0.027 for OS at 10 years; HR 1.56 (95% CI 1.21–2.03), p < 0.001 for OS at 5 years; HR 1.43 (95% CI 1.14–1.79), p = 0.001 for DSS at 10 years; HR 1.99 (95% CI 1.46–2.75), p < 0.001 for DSS at 5 years. There were no subtype-specific survival associations with LC3B that were statistically significant in both training and validation cohorts (Supplementary Table S4). To determine if LC3B was an independent prognostic marker, we performed multivariable analyses with all clinicopathological parameters. We found that LC3B puncta, but not LC3B H-score, was significantly associated with OS and DSS outcomes in the training cohort (p < 0.001 at 18-, 10- and 5-years) and combined cohort (p < 0.006 at 18, 10, and 5 years), but did not independently validate (p > 0.05) (Table 5, Supplementary Table S6).
Multi-marker expression exploratory analysis: patients with high LC3B puncta, high GABARAP, and low ATG4B expression have the poorest prognosis
To determine if the combination of ATG4B, LC3B, and GABARAP can provide additional information regarding patient survival, we performed exploratory multi-marker analyses in the whole cohort. After combining all three markers (Fig. 5e), patients with high LC3B puncta expression (A,B,C puncta expression; denoted L+) tumors had an inferior prognosis compared to those with low LC3B puncta expression (D,E puncta expression; denoted L−) tumors, which prompted us to further analyze the patients within the L+ and L− subgroups according to GABARAP and ATG4B expression. Within the L+ subgroup, high GABARAP expression (GABARAP H-score > 175; denoted G+) patients had significantly worse prognosis than low GABARAP expression (GABARAP H-score ≤ 175; denoted G-) patients (HR for 10-year DSS was 1.3 (1.05–1.61), p = 0.0144; HR for 10-year PFS was 1.48 (1.13–1.96), p = 0.0034), while patients with high ATG4B expression (ATG4B H-score > 150 denoted A +) demonstrated a better 10-year DSS compared to patients with low ATG4B expression (ATG4B H-score ≤ 150; denoted A−) (HR 0.78 (0.63–0.97), p = 0.0250). When all markers were combined, the most significant difference in survival in the L + subset of patients was reached between the G−/L+/A− and G+/L+/A− groups, with the latter group demonstrating the worst prognosis (HR for 10-year DSS was 1.51 (1.13–2.01), p = 0.0038; HR for 10-year PFS was 1.50 (1.06–2.19), p = 0.0172). Together, these analyses suggest that even though LC3B puncta has the strongest association with patient survival, further patient stratification may be possible by combining LC3B with GABARAP and ATG4B expression.
Discussion
We explored the differential expression and prognostic potential of three functionally related autophagy proteins, LC3B, GABARAP, and ATG4B, using tissue microarrays from a cohort of 3992 breast cancer patients. This is the first analysis of GABARAP and ATG4B in breast cancer patients and also the first study to quantitatively evaluate the punctate and diffuse staining patterns of LC3B with respect to prognostic value. Other groups previously evaluated biomarker potential of LC3B in breast cancer [30,31,32,33, 41, 42]. Only some of these studies [30,31,32,33] evaluated LC3B puncta in tumor cells, while others did not specify the pattern (diffuse or punctate) of LC3B staining that was assessed. Here, we analyzed both patterns of LC3B expression: diffuse (H-score), representing the cytoplasmic form of LC3B (LC3B-I), and punctate, representing the membrane-bound form of LC3B (LC3B-II). We established that both LC3B staining patterns were associated with poor prognosis.
The highest LC3B and GABARAP expression levels were observed in basal-like breast cancers. It is possible that these patterns are a reflection of the previously reported high autophagy levels in basal-like breast cancers [38]. However, caution must be taken when interpreting these results with respect to autophagy status: high LC3B puncta expression has been linked to both autophagy induction (i.e., increased autophagosome formation) and autophagy inhibition (i.e., decreased lysosomal turnover). While additional assays evaluating autophagic flux [43,44,45] would be required to clarify the functional status of autophagy, this does not limit the potential biomarker utility of the observed staining patterns.
Overexpression of ATG4B was shown previously to cause extensive delipidation and interfere with efficient lysosomal fusion [14, 46], thereby suppressing autophagic flux. This observation suggests that relatively low levels of ATG4B may instead reflect active or functional autophagic flux. If so, then the low ATG4B expression coupled with the high LC3B/GABARAP expression in basal-like tumors may be indicative of increased autophagy levels in these tumors. It should be noted, however, that ATG4B expression levels are not necessarily equivalent to ATG4B activity levels. Recent reports evaluating ATG4B function in autophagy showed that post-translational modifications of ATG4B, such as phosphorylation, play important roles in regulating ATG4B autophagic activity [47, 48]. Autophagy-independent roles of ATG4B have also been described [27, 28, 49]. One of the limitations of our study was the assessment of total, rather than phosphorylated, ATG4B levels in breast tumors, due to the unavoidable technical variability in surgical collection of human tumor tissues and the unreliability of phospho-antigens in formalin-fixed/paraffin-embedded specimens. In the future, it will be valuable to identify phosphorylation sites of ATG4B regulated in the context of breast cancer, and to assess their prognostic relevance, ideally in fresh-frozen tissue sections.
We found a number of significant associations among autophagy proteins and known breast cancer biomarkers. Both high LC3B expression and high GABARAP expression were associated with clinicopathological characteristics of aggressive disease phenotypes: high-grade tumors, ER/PR negativity, HER2 and HER3 positivity, high Ki67 (proliferation) index, and positive CK5/6 and EGFR expression. In contrast to LC3B and GABARAP, we found a negative correlation between the expression levels of ATG4B and the expression of poor prognostic markers such as Ki67, EGFR, and HER3. Each of these variables and markers has been previously described in relation to patient prognosis and cancer progression [50,51,52,53,54,55,56,57,58], but their association with LC3B, GABARAP, and ATG4B protein expression has not been reported previously.
We found statistically significant associations of LC3B and GABARAP expression with patient survival in univariable analyses. A number of previous reports [30,31,32, 41, 42] have also described an association of high LC3B expression with poor prognosis in breast cancer patients and the triple-negative (ER−, PR−, and HER2-negative) subtype, consistent with our findings on a much larger series. Other groups, however, indicated an opposite trend where LC3B negative expression correlated with adverse patient outcomes [32, 59]. Differences in patient cohort characteristics, including treatments received, intratumoral heterogeneity of LC3B expression, as well as staining and scoring techniques, may explain this discrepancy. Although LC3B puncta had the strongest association with patient prognosis, our exploratory multi-marker analysis indicated that further stratification by ATG4B and GABARAP expression may provide additional information regarding patient survival. Further studies to validate these marker combinations are required.
We overcame some of the known hurdles in the biomarker discovery process [60]—namely, sample size and follow-up time, but there are other limitations to our study. This large set of breast cancer TMAs was constructed from samples obtained from patients between 1986 and 1992. Even though many clinicopathological characteristics, such as tumor size, histological grade, ER/PR/HER2 status, Ki67 index, and intrinsic molecular subtype, remain relevant today, the treatment options available to patients have changed over time and shaped disease outcomes. This treatment aspect should be taken into consideration when interpreting the associations of GABARAP and LC3B with patient survival outcomes. To further validate these findings and determine independent prognostic marker potential, future studies should be performed on independent cohorts of patients, including those who received more modern treatments.
In summary, this is the largest cohort of breast cancer patients analyzed for LC3B biomarker value, and the first study to quantitatively evaluate its punctate versus diffuse staining patterns with respect to prognostic value. This is also the first analysis of GABARAP and ATG4B in breast cancer patients. Overall, we found that the differential expression of ATG4B did not associate with patient prognosis. High LC3B (punctate or diffuse) and high GABARAP expression associated with poor prognosis among all breast cancer subtypes, although multivariable analyses did not support either marker as an independent prognostic factor in this cohort. We also identified unique associations between known breast cancer biomarkers and these key autophagy proteins. Multi-marker analyses suggest that the combination of ATG4B and GABARAP with LC3B could be useful for further stratifying patient outcomes. This multi-marker approach and its biomarker potential warrant further investigation.
Methods
Study population
The study cohort included n = 3992 (interpretable cases n = 2876, n = 2826, and n = 3122 for LC3B, ATG4B, and GABARAP, respectively) patients initially diagnosed and seen at the British Columbia Cancer Agency between 1986 and 1992. This large, well-characterized cohort was described previously [61]; see Supplementary Methods. The study cohort was split into training and validation sets (see Supplementary Table S1). The training set was randomly selected to include 50% of the total study population and was used to identify optimal cutoff points as well as biomarker associations and prognostic trends, which were then tested on the remaining 50% (validation set) of the study population. All values described in the text are based on the validation cohort and include only those results that were significant in both training and validation sets.
Immunohistochemistry
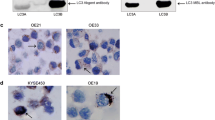
Slices from the tissue microarray [61] (Supplementary Methods) were stained previously with a number of markers including ER, PR, HER2, Ki67, CK5/6 and EGFR, as well as HER3, BCL2, IGF1R, CA-IX, and PD-L1 using procedures as described [50,51,52,53,54,55,56,57,58, 62, 63]. Intrinsic subtypes based on an immunohistochemical marker panel were assigned using the definitions previously published [50, 51]. ATG4B, GABARAP, and LC3B immunohistochemistry with anti-ATG4B (#A2981, Sigma), anti-GABARAP (#ab109364, Abcam), and anti-LC3B (#ab48394, Abcam) antibodies, respectively, was performed using the Ventana Discovery Ultra automated instrument (Ventana Medical Systems Inc, Tucson, USA) according to manufacturer's recommendations. Anti-ATG4B and anti-LC3B antibodies have been previously described [27, 30]; anti-GABARAP antibody was validated using GABARAP knock-out cell lines (See Supplementary Fig. S1 and Supplementary Methods).
Scoring and analysis of LC3B
Each sample was assessed by two independent observers for the intensity of cytoplasmic expression of LC3B in tumor cells under X20 objective magnification, as well as for the presence (i.e., positivity) of LC3B puncta under X40 objective magnification. LC3B cytoplasmic expression was scored using a H-score, consisting of multiplying cytoplasmic staining intensity of tumor cells (from 0 to 3) to the percentage of tumor cells staining (from 0 to 100%) which resulted in a score range of 0 to 300. The score was then categorized into 7 groups (0, 50, 100, 150, 200, 250, 300) by rounding to the closest multiple of 50. LC3B puncta expression was scored using a 5-point categorical scoring system: A. super high (clusters in > 50% of cells), B. high with some (< 50%) clusters, C. high puncta (in > 50% of cells), no clusters, D. low puncta (puncta in < 50% of cells), no clusters, E. negative (Fig. 1). For this purpose, a cluster refers to a dense accumulation of LC3B puncta (e.g., Fig. 1f) as opposed to more distinct individual puncta (e.g., Fig. 1c). All clinical and pathological data for each tumor sample were blinded to the scorers during scoring by two pathologists. The weighted Kappa values (0.71 for the H-score, and 0.66 for puncta scoring) indicated that substantial agreement between the two scores was achieved [64].
Scoring and analysis of ATG4B and GABARAP
Each sample was assessed for the presence (i.e., positivity) and intensity of cytoplasmic expression of ATG4B and GABARAP in tumor cells under X20 objective magnification. ATG4B and GABARAP cytoplasmic expression were scored using the same H-score, consisting of multiplying cytoplasmic staining intensity of tumor cells (from 0 to 3) to the percentage of tumor cells staining (from 0 to 100%) which resulted in a score range of 0 to 300. The score was then categorized into 7 groups (0, 50, 100, 150, 200, 250, 300) by rounding to the closest multiple of 50 (Fig. 1). All clinical and pathological data for each tumor sample were blinded to the two scorers during scoring. The weighted Kappa values (0.92 for ATG4B and 0.75 for GABARAP) indicated that almost perfect or substantial, respectively, agreement between the two scores was achieved [64]. GABARAP was also evaluated for a punctate pattern of expression, similar to LC3B. However, the GABARAP puncta were less distinct and independently deemed unreliable for scoring by two pathologists.
Statistical analysis
The associations of ATG4B and LC3B status with continuous variables (age, tumor size) were assessed using Welch’s t-test (one-way analysis of variance with no assumption on homogeneity of variances) and the associations with other binarized or categorical variables (grade, nodal status, LVSI, and other IHC markers) were done using a Chi-square test. Correlation was assessed by Spearman correlation test. Survival analyses were performed using Kaplan–Meier plots and Cox proportional hazards regression models. Multivariable Cox proportional hazard regression models were used to adjust the prognostic significance of ATG4B/GABARAP/LC3B with clinicopathological parameters. A split training/validation analysis approach was used as a mean to cross validate any findings and to mitigate against overfitting from data-driven cutpoint determination. The entire cohort was split into training (n = 2003) and validation (n = 1989) sets. Exploratory analyses were first performed on the training set to generate hypotheses that were subsequently tested on the validation set as internal cross validation. All effect estimates (such as hazard ratio) and p values reported in the text are based on the validation cohort and include only results that were found significant in both training and validation sets unless stated otherwise.
Cutpoint determination
A cutpoint was defined for each marker in this study (ATG4B, GABARAP and LC3B H-score) using the Akaike Information Criterion (AIC). To determine the optimal cutpoint, a univariable Cox model fit was generated on the training set for all possible cutpoints and the corresponding AIC values calculated (see Supplementary Methods).
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- AIC:
-
Akaike information criterion
- ATG:
-
Autophagy-related
- ATG4B:
-
Autophagy-related 4B
- BCL2:
-
B-cell lymphoma 2
- CA-IX:
-
Carbonic anhydrase 9
- DSS:
-
Disease-specific survival
- EGFR:
-
Epidermal growth factor receptor
- ER:
-
Estrogen receptor
- FDA:
-
Food and Drug Administration
- GABARAP:
-
GABA type A receptor-associated protein
- GATE-16:
-
Golgi-associated ATPase enhancer of 16 kDa
- GABARAPL2:
-
GABA type A receptor-associated protein like 2
- HER2:
-
Human epidermal growth factor receptor 2
- HER3:
-
Human epidermal growth factor receptor 3
- HR:
-
Hazard ratio
- IHC:
-
Immunohistochemistry
- LC3B:
-
Microtubule associated protein 1 light chain 3 beta
- OS:
-
Overall survival
- PD-L1:
-
Programmed death-ligand 1
- PE:
-
Phosphatidylethanolamine
- PR:
-
Progesterone receptor
- RFS:
-
Relapse-free survival
- TMA:
-
Tissue microarray
- TNBC:
-
Triple-negative breast cancer
- TNP:
-
Triple-negative phenotype
References
Galluzzi L, Pietrocola F, Bravo-San Pedro JM et al (2015) Autophagy in malignant transformation and cancer progression. EMBO J 34:856–880
Amaravadi R, Kimmelman AC, White E (2016) Recent insights into the function of autophagy in cancer. Genes Dev 30:1913–1930
Levy JMM, Towers CG, Thorburn A (2017) Targeting autophagy in cancer. Nat Rev Cancer 17:528–542
Dolgin E (2019) Anticancer autophagy inhibitors attract “resurgent” interest. Nat Rev Drug Discov 18:408–410
Ravikumar B, Sarkar S, Davies JE et al (2010) Regulation of mammalian autophagy in physiology and pathophysiology. Physiol Rev 90:1383–1435
Yang Z, Klionsky DJ (2010) Mammalian autophagy: core molecular machinery and signaling regulation. Curr Opin Cell Biol 22:124–131
Itakura E, Mizushima N (2010) Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy 6:764–776
Feng Y, He D, Yao Z et al (2014) The machinery of macroautophagy. Cell Res 24:24–41
Mizushima N, Yoshimori T, Ohsumi Y (2011) The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol 27:107–132
Nguyen TN, Padman BS, Usher J et al (2016) Atg8 family LC3/GABARAP proteins are crucial for autophagosome–lysosome fusion but not autophagosome formation during PINK1/Parkin mitophagy and starvation. J Cell Biol 251:857
Weidberg H, Shvets E, Shpilka T et al (2010) LC3 and GATE-16/GABARAP subfamilies are both essential yet act differently in autophagosome biogenesis. EMBO J 29:1792–1802
Shpilka T, Weidberg H, Pietrokovski S et al (2011) Atg8: an autophagy-related ubiquitin-like protein family. Genome Biol 12:226
Schaaf MBE, Keulers TG, Vooijs MA et al (2016) LC3/GABARAP family proteins: autophagy-(un)related functions. FASEB J 30:3961–3978
Tanida I, Sou Y, Ezaki J et al (2004) HsAtg4B/HsApg4B/autophagin-1 cleaves the carboxyl termini of three human Atg8 homologues and delipidates microtubule-associated protein light chain 3- and GABAA receptor-associated protein-phospholipid conjugates. J Biol Chem 279:36268–36276
Satoo K, Noda NN, Kumeta H et al (2009) The structure of Atg4B–LC3 complex reveals the mechanism of LC3 processing and delipidation during autophagy. EMBO J 28:1341–1350
Fujita N, Hayashi-Nishino M, Fukumoto H et al (2008) An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol Biol Cell 19:4651–4659
Li M, Hou Y, Wang J et al (2011) Kinetics comparisons of mammalian Atg4 homologues indicate selective preferences toward diverse Atg8 substrates. J Biol Chem 286:7327–7338
Kauffman KJ, Yu S, Jin J et al (2018) Delipidation of mammalian Atg8-family proteins by each of the four ATG4 proteases. Autophagy 14:992–1010
Akin D, Wang SK, Habibzadegah-Tari P et al (2014) A novel ATG4B antagonist inhibits autophagy and has a negative impact on osteosarcoma tumors. Autophagy 10:2021–2035
Kurdi A, Cleenewerck M, Vangestel C et al (2017) ATG4B inhibitors with a benzotropolone core structure block autophagy and augment efficiency of chemotherapy in mice. Biochem Pharmacol 138:150–162
Rothe K, Lin H, Lin KBL et al (2014) The core autophagy protein ATG4B is a potential biomarker and therapeutic target in CML stem/progenitor cells. Blood 123:3622–3634
Tran E, Chow A, Goda T et al (2013) Context-dependent role of ATG4B as target for autophagy inhibition in prostate cancer therapy. Biochem Biophys Res Commun 441:726–731
Bosc D, Vezenkov L, Bortnik S et al (2018) A new quinoline-based chemical probe inhibits the autophagy-related cysteine protease ATG4B. Sci Rep 8:11653
Fu Y, Hong L, Xu J et al (2018) Discovery of a small molecule targeting autophagy via ATG4B inhibition and cell death of colorectal cancer cells in vitro and in vivo. Autophagy 15:1–17
Liu P-F, Tsai K-L, Hsu C-J et al (2018) Drug repurposing screening identifies tioconazole as an ATG4 inhibitor that suppresses autophagy and sensitizes cancer cells to chemotherapy. Theranostics 8:830–845
Bortnik S, Choutka C, Horlings HM et al (2016) Identification of breast cancer cell subtypes sensitive to ATG4B inhibition. Oncotarget 7:66970–66988
Liu P-F, Leung C-M, Chang Y-H et al (2014) ATG4B promotes colorectal cancer growth independent of autophagic flux. Autophagy 10:1454–1465
Liu P-F, Hsu C-J, Tsai W-L et al (2017) Ablation of ATG4B suppressed autophagy and activated AMPK for cell cycle arrest in cancer cells. Cell Physiol Biochem Int J Exp Cell Physiol Biochem Pharmacol 44:728–740
Bortnik S, Gorski SM (2017) Clinical applications of autophagy proteins in cancer: from potential targets to biomarkers. Int J Mol Sci 18:1496
Lazova R, Camp RL, Klump V et al (2012) Punctate LC3B expression is a common feature of solid tumors and associated with proliferation, metastasis, and poor outcome. Clin Cancer Res Off J Am Assoc Cancer Res 18:370–379
Ladoire S, Chaba K, Martins I et al (2012) Immunohistochemical detection of cytoplasmic LC3 puncta in human cancer specimens. Autophagy 8:1175–1184
Lefort S, Joffre C, Kieffer Y et al (2014) Inhibition of autophagy as a new means of improving chemotherapy efficiency in high-LC3B triple-negative breast cancers. Autophagy 10:2122–2142
Ladoire S, Penault-Llorca F, Senovilla L et al (2015) Combined evaluation of LC3B puncta and HMGB1 expression predicts residual risk of relapse after adjuvant chemotherapy in breast cancer. Autophagy 11:1878–1890
Holt SV, Wyspianska B, Randall KJ et al (2011) The development of an immunohistochemical method to detect the autophagy-associated protein LC3-II in human tumor xenografts. Toxicol Pathol 39:516–523
Martinet W, Schrijvers DM, Timmermans J-P et al (2013) Immunohistochemical analysis of macroautophagy. Autophagy 9:386–402
Bertucci F, Finetti P, Cervera N et al (2008) How basal are triple-negative breast cancers? Int J Cancer 123:236–240
Perou CM (2011) Molecular stratification of triple-negative breast cancers. Oncologist 16(Suppl 1):61–70
Maycotte P, Gearheart CM, Barnard R et al (2014) STAT3-mediated autophagy dependence identifies subtypes of breast cancer where autophagy inhibition can be efficacious. Cancer Res 74:2579–2590
Maycotte P, Thorburn A (2014) Targeting autophagy in breast cancer. World J Clin Oncol 5:224–240
Hayes DF, Ethier S, Lippman ME (2006) New guidelines for reporting of tumor marker studies in breast cancer research and treatment: REMARK. Breast Cancer Res Treat 100:237–238
Zhao H, Yang M, Zhao J et al (2013) High expression of LC3B is associated with progression and poor outcome in triple-negative breast cancer. Med Oncol Northwood Lond Engl 30:475
Chen S, Jiang Y-Z, Huang L et al (2013) The residual tumor autophagy marker LC3B serves as a prognostic marker in local advanced breast cancer after neoadjuvant chemotherapy. Clin Cancer Res Off J Am Assoc Cancer Res 19:6853–6862
Klionsky DJ, Abdelmohsen K, Abe A et al (2016) Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12:1–222
Chittaranjan S, Bortnik S, Gorski SM (2015) Monitoring autophagic flux by using lysosomal inhibitors and western blotting of endogenous MAP1LC3B. Cold Spring Harb Protoc 2015:743–750
He H, Yang Y, Xiang Z et al (2016) A sensitive IHC method for monitoring autophagy-specific markers in human tumor xenografts. J Biomark. https://doi.org/10.1155/2016/1274603
Yu Z-Q, Ni T, Hong B et al (2012) Dual roles of Atg8−PE deconjugation by Atg4 in autophagy. Autophagy 8:883–892
Yang Z, Wilkie-Grantham RP, Yanagi T et al (2015) ATG4B (autophagin-1) phosphorylation modulates autophagy. J Biol Chem 290:26549–26561
Huang T, Kim CK, Alvarez AA et al (2017) MST4 phosphorylation of ATG4B regulates autophagic activity, tumorigenicity, and radioresistance in glioblastoma. Cancer Cell 32(840–855):e8
Liu P-F, Chen H-C, Cheng J-S et al (2019) Association of ATG4B and phosphorylated ATG4B proteins with tumorigenesis and prognosis in oral squamous cell carcinoma. Cancers 11:1854
Cheang MCU, Voduc D, Bajdik C et al (2008) Basal-like breast cancer defined by five biomarkers has superior prognostic value than triple-negative phenotype. Clin Cancer Res Off J Am Assoc Cancer Res 14:1368–1376
Cheang MCU, Chia SK, Voduc D et al (2009) Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J Natl Cancer Inst 101:736–750
Liu S, Chia SK, Mehl E et al (2010) Progesterone receptor is a significant factor associated with clinical outcomes and effect of adjuvant tamoxifen therapy in breast cancer patients. Breast Cancer Res Treat 119:53–61
Cheang MCU, Treaba DO, Speers CH et al (2006) Immunohistochemical detection using the new rabbit monoclonal antibody SP1 of estrogen receptor in breast cancer is superior to mouse monoclonal antibody 1D5 in predicting survival. J Clin Oncol Off J Am Soc Clin Oncol 24:5637–5644
Jensen KC, Turbin DA, Leung S et al (2008) New cutpoints to identify increased HER2 copy number: analysis of a large, population-based cohort with long-term follow-up. Breast Cancer Res Treat 112:453–459
Dawson S-J, Makretsov N, Blows FM et al (2010) BCL2 in breast cancer: a favourable prognostic marker across molecular subtypes and independent of adjuvant therapy received. Br J Cancer 103:668–675
Lou Y, McDonald PC, Oloumi A et al (2011) Targeting tumor hypoxia: suppression of breast tumor growth and metastasis by novel carbonic anhydrase IX inhibitors. Cancer Res 71:3364–3376
Chiu CG, Masoudi H, Leung S et al (2010) HER-3 overexpression is prognostic of reduced breast cancer survival: a study of 4046 patients. Ann Surg 251:1107–1116
Yerushalmi R, Gelmon KA, Leung S et al (2012) Insulin-like growth factor receptor (IGF-1R) in breast cancer subtypes. Breast Cancer Res Treat 132:131–142
Chang S-J, Ou-Yang F, Tu H-P et al (2016) Decreased expression of autophagy protein LC3 and stemness (CD44+/CD24−/low) indicate poor prognosis in triple-negative breast cancer. Hum Pathol 48:48–55
Goossens N, Nakagawa S, Sun X et al (2015) Cancer biomarker discovery and validation. Transl Cancer Res 4:256–269
Voduc D, Cheang M, Nielsen T (2008) GATA-3 expression in breast cancer has a strong association with estrogen receptor but lacks independent prognostic value. Cancer Epidemiol Biomark Prev Publ Am Assoc Cancer Res 17:365–373
Burugu S, Gao D, Leung S et al (2017) LAG-3+ tumor infiltrating lymphocytes in breast cancer: clinical correlates and association with PD-1/PD-L1+ tumors. Ann Oncol Off J Eur Soc Med Oncol 28:2977–2984
Chia S, Norris B, Speers C et al (2008) Human epidermal growth factor receptor 2 overexpression as a prognostic factor in a large tissue microarray series of node-negative breast cancers. J Clin Oncol Off J Am Soc Clin Oncol 26:5697–5704
McHugh ML (2012) Interrater reliability: the kappa statistic. Biochem Med 22:276–282
Acknowledgements
We thank Christine Chow, Angela Cheng, and Rebecca Wu for technical support and members of the Gorski laboratory for helpful comments and discussions.
Funding
This study was funded by CIHR in partnership with Avon Foundation for Women-Canada grant OBC127216 and CIHR PJT-159536 Grant to SMG. SBo was supported by a CIHR Frederick Banting and Charles Best Canada Graduate Scholarship Doctoral Award.
Author information
Authors and Affiliations
Contributions
SBo, KAG, and SMG conceived the study; SBo, BTC, and FD scored slides with assistance of JM and KG; SL, KA, and SBu performed statistical analyses; JX created KO lines and performed GABARAP antibody validation; SY and TN supervised scoring; KAG, TON, and SMG supervised and directed the study; SBo, BTC, and SL wrote the manuscript; all authors read, edited, and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
TON reports a licensing agreement with Veracyte, Inc. regarding the PAM50 intrinsic subtype classifier, not used in this study. The other authors declare that they have no conflict of interest.
Ethical approval
This study and our access to the de-identified human data were approved by the Research Ethics Board of the University of British Columbia, British Columbia Cancer Agency and Simon Fraser University. This article does not contain any studies with animals performed by any of the authors.
Informed consent
No informed consent was needed as anonymized archival specimens were used in this study, according to the Canadian Tri-Council Policy Statement for ethical research involving human subjects.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bortnik, S., Tessier-Cloutier, B., Leung, S. et al. Differential expression and prognostic relevance of autophagy-related markers ATG4B, GABARAP, and LC3B in breast cancer. Breast Cancer Res Treat 183, 525–547 (2020). https://doi.org/10.1007/s10549-020-05795-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-020-05795-z