Abstract
The estrogen receptor (ER) is a key predictive biomarker in the treatment of breast cancer. There is uncertainty regarding the use of hormonal therapy in the setting of weakly positive ER by immunohistochemistry (IHC). We report intrinsic subtype classification on a cohort of ER weakly positive early-stage breast cancers. Consecutive cases of breast cancer treated by primary surgical resection were retrospectively identified from 4 centers that engage in routine external proficiency testing for breast biomarkers. ER-negative (Allred 0 and 2) and ER weakly positive (Allred 3–5) cases were included. Gene expression profiling was performed using qRT-PCR. Intrinsic subtype prediction was made based upon the PAM50 gene expression signature. 148 cases were included in the series: 60 cases originally diagnosed as ER weakly positive and 88 ER negative. Of the cases originally assessed as ER weakly positive, only 6 (10 %) were confirmed to be of luminal subtype by gene expression profiling; the remaining 90 % of cases were classified as basal-like or HER2-enriched subtypes. This was not significantly different than the fraction of luminal cases identified in the IHC ER-negative cohort (5 (5 %) luminal, 83(95 %) non-luminal). Recurrence-free, and overall, survival rates were similar in both groups (p = 0.4 and 0.5, respectively) despite adjuvant hormonal therapy prescribed in the majority (59 %) of weakly positive ER cases. Weak ER expression by IHC is a poor correlate of luminal subtype in invasive breast cancer. In the setting of highly sensitive and robust IHC methodology, cutoffs for ER status determination and subsequent systemic therapy should be revisited.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Accurate determination of estrogen receptor (ER) status in breast cancer is a crucial component of standard pathology assessment, forming the foundation for key systemic treatment decisions. The link between estrogen deprivation and anti-breast cancer effect was first demonstrated over 120 years ago [1]. The evolution of the ER story over the last century, from empiric oophorectomy to modern-era immunohistochemistry (IHC), and targeted endocrine systemic therapy is complex and still evolving [2].
Dextran-coated charcoal (DCC) or ‘ligand binding assay’ is a forerunner to current ER assessment techniques [3], and was the predominant methodology that was used in the pivotal trials documenting the benefits of adjuvant hormone therapy for ER-positive breast cancer [4]. Ligand binding assays, despite offering quantitative information, are fraught with technical challenges including the requirement of fresh, immediately frozen tissue without a direct morphologic correlate. These limitations led to its widespread replacement by IHC for determination of ER status [5]. Since the introduction of IHC in the early 1990s, many additional advances have improved the sensitivity of these techniques including antigen retrieval, [6] polymer-based detection systems [7], rabbit monoclonal antibodies [8], and external proficiency testing [9].
Endeavoring to ascribe quantitative attributes to ER IHC, Harvey et al. [10] retrospectively performed ER IHC on residual frozen breast cancer tissue with available DCC results. This group applied an Allred score (average intensity + proportion score [quantified 0, 2–8]) and found that tumors scoring 3 out of 8 or higher benefited from adjuvant hormonal therapy, whereas those with scores 0 or 2 did not. This and similar studies led to the American Society of Clinical Oncology/College of American Pathologists guidelines indicating that IHC expression of ER in 1 % of cells or more should be reported positively [11].
Little is known about how weakly ER-positive breast cancers respond to hormonal therapy. Subgroup analysis in large trials demonstrating benefits of hormone therapy in ER-positive cancers has not been possible as the incidence of ER weakly positive cancer is low (ranging from 1 to 6.7 % of breast cancers) [12–15]. Further uncertainty is added by the reported variability in IHC results across laboratories [16, 17]. Currently, there is doubt among oncologists and pathologists as to how to treat and report ER ‘weakly positive’ breast cancers. The balance between benefit from hormonal therapy and unnecessary exposure to risks of serious side effects such as osteoporosis, thromboembolic disease, and endometrial carcinoma remains to be elucidated for this group of patients.
The identification of intrinsic molecular subtypes based upon gene expression profiles has enlightened the understanding of the biology and treatment of breast cancers [18, 19]. Individual cases can be characterized as 5 “intrinsic” subtypes of breast cancer that have been repeatedly observed [20, 21]. Luminal intrinsic subtypes are driven by ERα signaling, potentially benefit from hormonal therapy, and typically have positive ER IHC. HER2-enriched subtypes are highly proliferative and aggressive tumors driven by amplification of the ERBB2 region and are clinically detectable by IHC for HER2 overexpression and/or ERBB2 amplification status by in situ hybridization (ISH) techniques. Basal-like subtypes are also highly proliferative and aggressive tumors characterized by genetic instability, and typically lack expression of hormone receptors and HER2 (so-called ‘triple-negative’ phenotype). Additionally, these tumors may also be identified by their expression of certain basal keratins (e.g., keratin 5) or epidermal growth factor receptor (EGFR) [22]. Basal-like breast cancers are a heterogeneous group of tumors. Currently, systemic cytotoxic chemotherapy is the mainstay of treatment; however, other targeted therapies (e.g., PARP inhibitors, immune check-point inhibitors) are being assessed [23].
Gene expression studies are steadily defining their utility in the clinical realm. The PAM50 gene signature uses expression data from 50 key genes to identify the major molecular subtype and to assign a risk of recurrence (ROR) score to surgically resected breast cancer specimens [24]. PAM50 luminal subtype designation has been shown to be more discriminatory than IHC (using a conventional >1 % cutoff), to predict adjuvant tamoxifen benefit in premenopausal women [25].
In this study, we apply the PAM50 classifier to a series of IHC ER weakly positive and ER-negative breast cancers. We compare the intrinsic subtype distribution and outcomes, presenting evidence that ER weakly positive tumors are more similar in clinical behavior to ER-negative than to ER-positive breast cancer, questioning the benefit of hormonal therapy in this population.
Methods
Approval by the research ethics review board of the University of British Columbia/British Columbia Cancer Agency and all affiliated centers was obtained prior to the commencement of this study.
Samples and clinical data
The records of four academic centers in the metropolitan Vancouver region were retrospectively reviewed for the period from 1/9/2010 to 1/9/2013. Consecutive cases of invasive breast cancer were included based upon clinically reported ER status and the following criteria: primary diagnosis of breast cancer, ER IHC weakly positive (defined as Allred 3, 4, or 5 of 8), primary surgical treatment without neoadjuvant systemic therapy, and sufficient tumor tissue available for qRT-PCR analysis (greater than 5 mm invasive disease present in archived FFPE material after a complete diagnostic workup). Additionally, a similar number of control cases were identified (Allred 0 or 2) meeting the same tumor size inclusion criteria. Exclusion criteria consisted of locoregional recurrence or second diagnosis of breast cancer, core needle biopsy only, cytology specimens, and cases not directly meeting the inclusion criteria.
ER status was extracted directly from the patient charts and was not necessarily determined on the specimen collected for study (i.e., performed on prior core biopsy). In cases with discordant ER results (i.e., from a core needle biopsy and ensuing resection), the most positive (highest Allred score) was documented. All participating laboratories are accredited and engage in ongoing external proficiency testing for breast biomarkers including (a minimum of) triennial exercises through the Canadian Immunohistochemistry Quality Control (CIQC) program, and have maintained high-quality, reproducible staining throughout the entire study period [26]. Individual staining protocols were documented as follows (antigen retrieval time [minutes], clone, dilution, vendor, incubation time [minutes], detection system): institution 1 (32, SP1, 1:50, Thermo Scientific, 32, OptiView), institution 2 (36, SP1, pre-diluted, Ventana, 16, ultraview DAB), institution 3 (32, 1:50, Thermo Scientific, 16, OptiView), and institution 4 (32, SP1, 1:50, Ventana, 32, Ventana iView).
Each of the 4 participating institutions routinely reports ER status as Allred scores, defined as the sum of average intensity (0: none, 1: weak, 2: moderate, 3: strong) and proportion (0:none, 1:<1 %, 2:1–10 %, 3:10–33 %, 4: 33–66 %, 5:>67 %) scores [10]. Allred scores were retrieved directly as originally reported at the time of diagnosis.
Representative formalin-fixed paraffin-embedded (FFPE) tissue blocks were selected for the study. H&E-stained sections were reviewed by anatomical pathologists (BSS, ZK, NM) for confirmation of diagnosis and selection of tumor-rich areas for molecular studies. The chart of each patient was reviewed and relevant information was extracted including age, tumor staging criteria, grade, breast biomarker profile, treatment, and outcome in the form of disease recurrence and death due to disease.
Nucleic acid extraction, reverse transcription, and qPCR
From each block, duplicate 1-mm core samples from tumor-rich areas were removed and de-paraffinized. RNA was recovered using the High Pure RNA Paraffin kit including an on-column DNase I treatment (Roche Applied Science, Indianapolis IN). RNA yields were assessed using an ND-1000 spectrophotometer (NanoDrop Technologies, Rockland DE, USA).
Complementary DNA (cDNA) synthesis was completed using a mixture of random hexamers and gene-specific primers, and real-time quantitative PCR (qRT-PCR) was performed with the Roche LightCycler 480 instrument using SYBR Green I dye as previously described [24, 27]. Each 384-well plate contained samples and a calibrator in triplicate with 2.5 ng cDNA and 10 ng cDNA, respectively, per reaction. A tumor sample was considered of insufficient quality if any of the reference controls (ACTB, PSMC4, RPLP0, MRPL19, or SF3A1) failed.
Sample subtype prediction
The, previously validated [17], PAM50 assay has been constructed to provide stable and highly reproducible subtype classification in FFPE and frozen specimens. The qRT-PCR assay consists of 50 discriminatory genes relating to ER signaling, growth factor signaling, proliferation, invasion, and basal phenotypes) and an additional 5 housekeeping genes for sample normalization [28]. Analysis by qRT-PCR is performed by normalizing the raw Ct values to gene-specific technical controls, followed by normalization to sample controls [29]. The distance to each centroid is calculated using Spearman’s rank correlation. The centroid associated with the largest positive correlation value is assigned as the subtype of the sample.
Identification of comparison cohort of ER strongly positive cancers
For comparative purposes, but not included as study samples, a cohort of 26 ER strongly positive (Allred 6,7,8) cases was assembled from an available set with both IHC and PAM50 data. The cases originated from the same four centers, during a similar time period, and PAM50 intrinsic subtyping was performed using an identical qRT-PCR methodology. An additional set of 447 ER strongly positive cases was identified within a larger published series of cases with PAM50 intrinsic subtype data, also performed with identical methodology; however, these cases were from different centers and time periods [30].
Statistical analysis
Primary clinical outcomes were relapse-free survival (RFS) and overall survival (OS). RFS was defined as the time from diagnosis to any recurrence, including local or distant recurrence by metastasis, and OS was defined as the time from diagnosis to death from any cause. The primary outcomes of RFS and OS as defined above were quantified using Kaplan–Meier curves and compared by log-rank and Wilcoxon tests. Clinicopathologic variables were compared by χ 2 analysis. All statistical analyses were performed using SPSS version 19. Test characteristics were calculated as sensitivity ≡ true positives/(false negatives + true positives) and specificity ≡ true negatives/(false positives + true negatives). Associated 95 % confidence intervals were calculated as s ± 1.96 x √s*((1 − s)/n)), where s represents either sensitivity or specificity and n is the total number of cases.
Results
In total, 148 cases were identified from the participating centers. Sixty cases were ER weakly positive (Allred 3,4,5) and 88 cases were ER negative (Allred 0, 2) by IHC. The ER weakly positive and ER-negative groups showed a similar distribution of patient age, tumor grade, and adjuvant chemotherapy treatment. Both groups had similar proportions of HER2 positivity as determined clinically by IHC/ISH (37 % of ER weakly positive and 30 % of ER-negative cases) and similar distributions of PR staining. Six cases (4 %) showed an ER-negative, PR-positive IHC phenotype, and each of these cases showed weak (Allred 3–5) PR staining. Adjuvant hormonal therapy was prescribed to 58 % of the ER weakly positive cohort. The demographic data of the study population are summarized in Table 1.
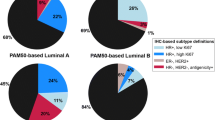
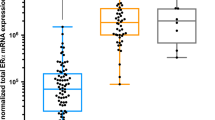
Of the 60 ER weakly positive cases, 6 (10 %) were of luminal subtype (luminal A/B), 24 (40 %) were HER2 enriched, and 30 (50 %) were basal like by the PAM50 signature. In 88 ER-negative cases, 5 (6 %) were luminal, 34 (39 %) were HER2 enriched, and 49 (56 %) were basal like. The distributions of PAM50 subtype predictions are shown in Table 2 and Fig. 1a and b.
Distribution of intrinsic subtype predictions for ER-negative (a), ER weakly positive (b), and ER strong positive (c, d) breast cancers. Data in part D are extracted from previously published results [30]
For comparison, 26 ER strongly positive (Allred 6,7,8) cases with available PAM50 intrinsic subtype data were identified from the same centers and time period. Subtype predictions on this set were 25 (96 %) luminal, 1 (4 %) HER2 enriched, and 0 basal like. Similarly, ER IHC strongly positive cases identified from the previously published independent series by Nielsen et al. [30] that were assessed with the same PAM50 qRT-PCR assay showed the following intrinsic subtype predictions: 426 (95 %) luminal, 20 (5 %) HER2 enriched, and 1 (0.2 %) basal like (Fig. 1c and d).
Adjuvant hormonal therapy was prescribed to 35 (58 %) of patients in the ER weakly positive group and to 4 (5 %) of patients in the ER-negative group. Adjuvant chemotherapy was prescribed to 58 (97 %) of patients in the ER weakly positive group and to 73 (83 %) in the ER-negative group (Table 1). Clinical outcomes over a median 54-month (range 5–58 month) period included 8 (14 %) recurrences and 8 (14 %) deaths in the ER weakly positive group and 15 (17 %) recurrences and 8 (9 %) deaths in the ER-negative group. No statistical differences in recurrence (p = 0.53) or death (p = 0.41) were identified between the ER weakly positive and ER-negative groups (Fig. 2). For comparison, the 26 strongly positive ER cases displayed a median survival time of 66 months, higher than the 51-month median survival among the ER weakly positive cases.
Using PAM50 intrinsic subtype as a reference standard, we computed test characteristics of ER weakly positive IHC for diagnosing luminal-type breast cancer. In this cohort, weak ER positivity showed a sensitivity of 55 % (47–3 %) and a specificity of 61 % (53–69 %) for identifying true luminal cases [values shown as mean (95 % confidence interval), Table 3].
Discussion
This study is a retrospective review of consecutive cases of ER weakly positive breast cancer at 4 hospitals that participate in breast cancer EQA testing programs. The data indicate that breast cancer cases considered ER “weakly positive” by IHC, when assessed at an RNA expression profile level, show a virtually identical intrinsic subtype distribution as breast cancers that are considered ER negative by IHC in the same laboratories.
Previous work by Chia et al. [25] has demonstrated that luminal subtype prediction by PAM50 is superior to IHC in predicting benefit from adjuvant tamoxifen treatment in a randomized trial. As intrinsic subtyping is not routinely performed at the participating institutions, clinicians make treatment decisions based upon IHC status. In the ER weakly positive cohort, as many as 90 % of the patients may have been incorrectly subtyped as luminal-type breast cancers by IHC. A large proportion (58 %) of patients in the ER weakly positive group were treated with adjuvant hormonal therapy, a treatment pattern for ER weakly positive cases also reported elsewhere [31]. Outcome data show no significant differences in overall survival or relapse-free survival between ER weakly positive and ER-negative patients, although the number of events and surveillance times are inadequately powered to draw definitive conclusions on clinical outcome. These data support the conclusion that ER weak positivity by IHC does not correlate with luminal biology. Although not entirely conclusive based only on the data presented here, the findings do suggest that these ER weakly positive (by IHC) patients are unlikely to benefit from hormonal therapy and are being exposed to their potential side effects.
The data we present here are supported by similar studies reported in the literature. Prabhu et al. [32] performed gene expression analysis for ER-related gene expression (including ESR1, PgR, GATA3, TFF1, FOXA1, and XBP1) compared with IHC on 240 tumors, finding that, in 21 ER weakly positive tumors (IHC proportion between 1 and 10 %), gene expression was more similar to their ER-negative group. Deyarmin et al. [33] showed that 54 consecutive breast cancers with weak ER positivity (proportion between 1 and 10 %) shared similar outcomes and molecular intrinsic subtype predictions (by GeneChip and Breast PRS algorithm) to ER-negative cancers. Iwamoto et al. [14] performed gene expression profiling, including PAM50 intrinsic subtype prediction on 25 ER weakly positive breast cancers (proportion between 1 and 9 %) and showed that these had a similar intrinsic subtype distribution and clinical outcome as ER-negative tumors.
The data presented here are in complete agreement with previous reports. This study represents the largest cohort of ER weakly positive cases reported in the literature to date with associated molecular data. Here, ER weak positivity is defined as Allred 3–5, which includes cancers that may have weak intensity staining in up to 66 % of tumor cells, as well as tumors which may have strong staining in <1 % of tumor cells. This definition is broader than the definition of 1–10 % used elsewhere. ER status was derived directly from the patient chart, representing the result upon which treatment decisions were made, and comes from a combination of biopsy and surgical specimens. Intrinsic subtype prediction was performed using the validated PAM50 algorithm on a robust qRT-PCR platform. Our results add to the body of evidence that ER weak positivity by IHC is not a sufficient surrogate for luminal intrinsic subtype and questions the use of hormonal therapy in this subgroup of patients.
Previous studies of ER weakly positive cancers have been criticized for biasing data with biopsy-only results, or using outdated IHC techniques [34]. We used a combination of biopsy and resection specimen IHC ER results, depending on which specimen type was reported for clinical biomarker assessment. Central ER re-testing or resection specimen re-testing (in cases of ER weakly positive results on core biopsy) was not part of the study design. All ER-negative results on core biopsy were clinically repeated on the subsequent excision specimens as part of standard of care. We specifically opted to use the clinically reported ER results (as performed by accredited pathology laboratories), as these were the bases for clinical decision making. We limited our study to contemporary cases, using current, sensitive IHC protocols (as well as current standards for cold ischemic time and fixation length).
Due to the use of contemporary cases, one limitation of the present work is the relatively short clinical follow-up time, with limited number of events available for statistical analyses. Another limitation of the study is the lack of a parallel analysis of ER strong positive cases. To address this issue, we collected data from 26 cases of ER strong positive cancers reported during a similar time period (2009–2013) from the same centers as the 148 study cases that had undergone PAM50 subtype analysis by qRT-PCR, and found that the intrinsic subtype predictions were overwhelmingly (97 %) luminal. These findings are entirely congruent with the ER and PAM50 results previously reported by Nielsen et al. [30]. This large published cohort included 447 cases of ER strong positive cancers, 95 % of which were found to be of luminal intrinsic subtype. Samples for the Nielsen cohort were originally collected from 1986 to 1992, originate from a wider range of healthcare facilities, and were subsequently assessed for ER by IHC centrally.
A particular strength of the data we report is the clinical relevance. ER status on the study cohort was performed in several accredited clinical laboratories, all using modern IHC techniques. All treatment decisions recorded in this study are likely predicated, in part, on the clinically reported ER results. Despite the fact that no difference was demonstrated between overall and relapse-free survival between the ER weakly positive and ER-negative groups, the study was underpowered to prove definitively that ER weakly positive cancers retain similar clinical behavior to ER-negative cancers or to rule out some benefit of hormonal therapy for this subgroup.
A question often raised is whether the 1 % IHC cutoff endorsed by ASCO/CAP guidelines for ER positivity is too low [11]. Our data indicate that, by gene expression profiling, 90 % of patients with IHC ER weakly positive breast cancers are indeed not of luminal subtype, despite usually being treated as such. In light of our study, and other similar reports cited above, the optimal cutoff for ER positivity (as assessed by modern IHC techniques) should be revisited and may need to be revised.
In conclusion, this report documents the intrinsic subtype distribution of ER weakly positive breast cancers in a large metropolitan area over a 3-year period. These data indicate that, using modern highly sensitive IHC techniques, ER weak positivity is not a biomarker for luminal biology as defined by expression profiling. These results raise doubts as to whether ER weakly positive breast cancers are likely to benefit from hormonal therapy. Additional studies linking contemporary estrogen receptor immunohistochemistry results to patient outcomes, preferably within the context of randomized clinical trials, are needed to better define optimal diagnostic and treatment algorithms for this group of patients.
References
Beatson GT (1896) On the treatment of inoperable cases of carcinoma of the mamma: suggestions for a new method of treatment, with illustrative cases. Lancet 148(3803):162–165
Wolff AC, Dowsett M (2011) Estrogen receptor: a never ending story? J Clin Oncol 29(22):2955–2958
McGuire WL (1973) Estrogen receptors in human breast cancer. J Clin Invest 52(1):73–77
Early Breast Cancer Trialists’ Collaborative Group (2011) Relevance of breast cancer hormone receptors and other factors to the efficacy of tamoxifen: patient-level meta-analysis of randomized trials. Lancet 378(9793):771–784
Dowsett M (2006) Estrogen receptor: methodology matters. J Clin Oncol 24(36):5626–5628
Gown AM (2004) Unmasking the mysteries of antigen or epitope retrieval and formalin fixation. Am J Clin Pathol 121(2):172–174
Sabattini E, Bisgaard K, AScani S, Poggi S, Piccioli M, Ceccarelli C, Pieri F, Fraternali-Orcioni G, Pileri SA (1998) The EnVision++ system: a new immunohistochemical method for diagnostics and research. Critical comparison with the APAAP, ChemMate, CSA, LABC, and SABC techniques. J Clin Pathol 51(7):506–511
Cheang MC, Treaba DO, Speers CH, Olivotto IA, Bajdik CD, Chia SK, Goldstein LC, Gelmon KA, Huntsman D, Gilks CB, Nielsen TO, Gown AM (2006) Immunohistochemical detection using the new rabbit monoclonal antibody SP1 of estrogen receptor in breast cancer is superior to mouse monoclonal antibody 1D5 in predicting survival. J Clin Oncol 24(36):5637–5644
Terry J, Tolakovic EE, Garrat J et al (2009) Implementation of a Canadian external quality assurance program for breast cancer biomarkers: an initiative of Canadian Quality Control in immunohistochemistry (cIQc) and Canadian Association of Pathologists (CAP) National Standards Committee/immunohistochemistry. Appl Immunohistochem Mol Morphol 17(5):375–382
Harvey JM, Clark GM, Osborne CK, Allred DC (1999) Estrogen receptor status by immunohistochemistry is superior to the ligand-biding assay for predicting response to adjuvant endocrine therapy in breast cancer. J Clin Oncol 17(5):1474–1481
Hammond ME, Hayes DF, Dowsett M et al (2010) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol 28(16):2784–2795
Nadji M, Gomez-Fernandez C, Ganjei-Azar P, Morales AR (2005) Immunohistochemistry of estrogen and progesterone receptors reconsidered: experience with 5993 breast cancers. Am J Clin Pathol 123(1):21–27
Khoshnoud MR, Löfdahl B, Fohlin H et al (2011) Immunohistochemistry compared to cytosol assays for determination of estrogen receptor and prediction of the long-term effect of adjuvant tamoxifen. Breast Cancer Res Treat 126(2):421–430
Iwamoto T, Booser D, Valero V et al (2012) Estrogen receptor (ER) mRNA and ER-related gene expression in breast cancers that are 1–10 % ER-positive by immunohistochemistry. J Clin Oncol 30(7):729–734
Allred DC, Carlson RW, Berry DA et al (2009) NCCN task force report: estrogen receptor and progesterone receptor testing in breast cancer by immunohistochemistry. J Natl Compr Canc Netw 7(S6):s1–s21
Rhodes A, Jasani B, Barnes DM et al (2000) Reliability of immunohistochemical demonstration of estrogen receptors in routine practice: inter-laboratory variance in the sensitivity of detection and evaluation of scoring systems. J Clin Pathol 53:125–130
Parker RL, Huntsman DG, Lesack DW et al (2002) Assessment of inter-laboratory variation in immunohistochemical determination of estrogen receptor using a breast cancer tissue microarray. Am J Clin Pathol 117:723–728
Cm Perou, Sorlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752
Sørlie T, Perou CM, Tibshirani R et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98(19):10869–10874
Sorlie T, Tibshirani R, Parker J et al (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci 100:8418–8423
Hu Z, Fan C, Oh DS et al (2006) The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genom 7:96
Won JR, Gao D, Chow C et al (2013) A survey of immunohistochemical biomarkers for basal-like breast cancer against a gene expression profile gold standard. Mod Pathol 26(11):1438–1450
Park Y, Moryama A, Kithara T, Yoshida Y, Urita T, Kato R (2012) Triple-negative breast cancer and poly(ADP-ribose) polymerase inhibitors. Anticancer Agents Med Chem 12(6):672–677
Parker JS, Mullins M, Cheang MC et al (2009) Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 27(8):1160–1167
Chia SK, Bramwell VH, Tu D et al (2012) A 50-gene intrinsic subtype classifier for prognosis and prediction of benefit from adjuvant tamoxifen. Clin Cancer Res 18(16):4465–4472
Perez T, Makretsov N, Garatt J, Torlakovic E, Gilks CB, Mallett S. Modeling Canadian quality control test program for steroid hormone receptors in breast cancer: diagnostic accuracy study. Appl Immunohistochem Mol Morphol [Epub ahead of print] PMID2620083
Mullins M, Perreard L, Quackenbush JF et al (2007) Agreement in breast cancer classification between microarray and quantitative reverse transcription PCR from fresh-frozen and formalin-fixed, paraffin-embedded tissues. Clin Chem 53:1273–1279
Bernard PS (2012) Personalized medicine for breast cancer. In: Ginsburg G (ed) Genomic and personalized medicine, 2nd edn. Elsevier, Amsterdam, pp 707–713
Perreard L, Fan C, Quackenbush JF et al (2006) Classification and risk stratification of invasive breast carcinomas using a real-time quantitative RT-PCR assay. Breast Cancer Res 8:R23
Nielsen TO, Parker JS, Leung S et al (2010) A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer. Clin Cancer Res 16(21):5222–5232
Lin CM, Jaswal J, Vandenberg T, Tuck A, Brackstone M (2013) Weakly hormone receptor-positive breast cancer and use of adjuvant hormonal therapy. Curr Oncol 20(6):e612–e613
Prabhu JS, Korlimarla A, Desai K et al (2014) A majority of low (1–10%) ER positive breast cancers behave like hormone receptor negative tumors. J Cancer 5(2):156–165
Deyarmin B, Kane JL, Valente AL, van Laar R, Gallagher C, Shriver CD, Ellseworth RE (2013) Effect of ASCO/CAP guidelines for determining ER status on molecular subtype. Ann Surg Oncol 20(1):87–93
Rakha EA, Lee AH, Roberts J, Villena Salinas NM, Hodi Z, Ellis IO, Reis-Filho JS (2012) Low-estrogen receptor-positive breast cancer: the impact of tissue sampling, choice of antibody, and molecular subtyping. J Clin Oncol 30(23):2929–2930
Acknowledgments
The authors would like to thank the Canadian Breast Cancer Foundation—British Columbia/Yukon chapter for their support of this research. This work was supported by a grant from the Canadian Breast Cancer Foundation (British Columbia and Yukon).
Author information
Authors and Affiliations
Corresponding author
Additional information
B. S. Sheffield and Z. Kos have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Sheffield, B.S., Kos, Z., Asleh-Aburaya, K. et al. Molecular subtype profiling of invasive breast cancers weakly positive for estrogen receptor. Breast Cancer Res Treat 155, 483–490 (2016). https://doi.org/10.1007/s10549-016-3689-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-016-3689-z