Abstract
High proliferation rates are characteristic of cancer, and proliferation markers make up the majority of genes included in RNA-based prognostic gene signatures applied for breast cancer patients. Based on prior data on differences in molecular subgroups of breast cancer, we hypothesized that the significance of single proliferation markers might differ in luminal, Her2-positive and triple-negative subtypes. Therefore, we compared mRNA expression data of Ki67, TOP2A, and RacGAP1 using a pool of 562 Affymetrix U133A microarrays from breast cancer samples. “Luminal,” “triple-negative,” and “Her2-positive” subcohorts were defined by ESR1 and ERBB2 mRNA expression using pre-defined cut-offs. The analysis of the three potential proliferation markers revealed subtype-specific differences: in luminal carcinomas, expression of all three markers was a significant indictor of early recurrence in univariate and multivariate analysis, but RacGAP1 was superior to Ki67 and TOP2A in significance. In triple-negative tumors, only Ki67 was a significant and independent marker, whereas none of the markers showed a significant prognostic impact in Her2-positive cases. Within the group of luminal carcinomas, the proliferation markers had different impact depending on the treatment of patients: in untreated patients, Ki67, TOP2A, and RacGAP1 were significant and independent prognostic markers. In chemotherapy-treated patients, overexpression of all three markers was predictive for early recurrence, but only RacGAP1 retained significance in multivariate analysis. In contrast, RacGAP1 was the only predictive proliferation marker in the endocrine treatment group. These data point to subtype-specific differences in the relevance of proliferation-associated genes, and RacGAP1 might be a strong prognostic and predictive marker in the luminal subgroup.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the major characteristics of malignant cancer is enhanced and limitless cell division, resulting in high proliferation rates [1]. Since cell division rate might be a useful prognostic indicator, estimation of proliferation rate has been performed by clinicians and pathologists by various methods including determination of the mitotic index, S-phase fraction and immunocytochemistry for nuclear antigens involved in cell cycle regulation and mitosis, i.e., Ki67 [2–4].
The technical progress in RNA-based methods resulted in the identification of various prognostic gene expression signatures or panels of 16–512 genes that can be applied in routine care of breast cancer patients [5–9]. In large meta-analyses, Wirapati et al. [10] and Desmedt et al. [11] could show that the prognostic power of these signatures is mostly due to the presence of proliferation markers which make up the majority of genes in these sets.
Since the ground-breaking publications of Perou and workers [12, 13], it is widely accepted that mammary carcinomas can be divided into at least four subtypes which differ in biology and prognosis, namely, the Her2-positive tumors, the basal-like carcinomas (mostly identical to triple-negative tumors (TNT)), and two groups of ER-positive tumors: luminal A and luminal B. Desmedt et al. [11] could show that the prognostic markers in these subgroups can be attributed to different expression modules: In the luminal subgroups, proliferation and histological grade were significantly associated with outcome, whereas in the basal and Her2-positive groups, the immune response module was associated with prognosis, and tumor invasion had a prognostic impact in Her2-positive tumors only. In other studies of prognostic and predictive gene sets in ER-positive and ER-negative carcinomas, it was shown that both groups differ strongly in the biological processes which influence prognosis and chemotherapy response [14], and prognostic gene signatures which mainly consist of genes involved in proliferation failed to predict the risk of tumor recurrence in non-luminal carcinomas [10].
Although a large number of studies has shown a significant prognostic value of proliferation markers in terms of recurrence-free survival (RFS) and overall survival (OAS), especially in ER-positive tumors, they were not included as routine investigation in most guidelines. Possible reasons are the poor reproducibility of the results of these studies, time-consuming procedures and difficulties in standardization and evaluation of the results [15]. In 2011, the St. Gallen guidelines dealing with strategies for molecular breast cancer subtypes recommended the use of the immunohistochemical Ki67 labeling index for discrimination of luminal B and luminal A tumors [16] with a cut-off point of 14 %. In clinical routine, this cut-point is problematic: a recent study on the reproducibility of Ki67 immunohistochemistry in moderately differentiated carcinomas revealed a high inter- and intraobserver variability [17].
Compared to immunohistochemical assays, RNA-based methods might allow more precise quantification of gene expression. Since proliferation has been shown to be an important factor in progression of luminal tumors [11], we asked if RNA expression of a single proliferation marker instead of a proliferation module might be suitable as prognostic or predictive marker in this subgroup, which might be much more practical for routine purposes. In order to find out which proliferation-associated marker might have the best prognostic impact, we analyzed mRNA expression of three candidate genes, Ki67, TOP2a, and RacGAP1, in breast cancer samples. In a recent study on protein–protein interaction networks in breast cancer samples, all three markers are members of a cell cycle-associated sub-network associated with a poor prognosis, although their position within this network indicates regulatory differences [18].
Ki67 was described in 1991 as a nuclear non-histone protein which is absent in quiescent cells but strongly expressed in proliferating cells in all phases of the cell cycle [19]. After development of an antibody which is suitable for immunohistochemistry on formalin-fixed, paraffin-embedded tissues (MIB1), it was widely introduced into histopathology, where the percentage of Ki67-positive cells is used as a marker of proliferative activity. In breast cancer, high Ki67 levels are associated with poor prognosis, but lack independent significance in most studies [4, 20]. In addition, high Ki67 positivity has been implicated in chemotherapy response [21].
TOPisomerse IIα (TOP2a) reduces DNA supercoiling and twisting and is important for chromosome segregation and condensation in dividing cells. Its expression is highest in the G2/M phase of exponentially growing cells, and overexpression predicted shorter DFS and OAS in some studies [2, 22]. TOP2a is located on the same chromosome as Her2/neu and is frequently co-amplified with this receptor. Since TOP2a is a target of anthracycline, its amplification is a marker of sensitivity to these drugs [23, 24]. High TOP2a RNA levels are significantly associated with shorter metastasis-free survival in node-negative breast cancer, but associated with a high frequency of pathological complete response (pCR) in anthracycline-treated patients [25].
RacGAP1 (MgcRacGAP) is a GTPase activating protein (GAP) which co-localizes with the mitotic spindle in the metaphase and is essential for cytokinesis during the normal cell cycle [26]. During this phase, it is phosphorylated by Aurora B in the midbody which is essential for its GAP activity toward RhoA [27]. In addition, it acts as a nuclear chaperone and is essential for nuclear transportation of STAT transcription factors [28]. Up-regulation of RacGAP1 was observed in various malignant tumors, i.e., ovarian cancer [29], bladder cancer [30] and hepatocellular carcinoma [31].
Since proliferation is a prerequisite of tumor growth in all carcinomas, but the prognostic gene signatures are not equally prognostic in different breast cancer types [10], we assumed that the regulation of proliferation in mammary carcinomas differs depending on molecular subtypes. Thus, single proliferation markers might not be equally informative in luminal, Her2-positive and triple-negative breast cancer. Based on this hypothesis, we studied mRNA expression of all three proliferation markers in a retrospective analysis based on microarray data of 562 breast cancer samples with long-term follow-up information.
Materials and methods
Breast cancer samples
For this retrospective analysis, RNA expression data obtained from primary breast cancer tissue samples from Hamburg, Frankfurt, and Mainz were combined. Informed consent for the scientific use of tissue materials, which was approved by the local ethics committees, was obtained from all patients. The study was performed in accordance to the principles of the declaration of Helsinki and remark criteria [32]. The clinical and histological characteristics of these cohorts are given in Table 1. No radiotherapy, neoadjuvant chemotherapy or endocrine therapy had been administered before surgery. Adjuvant therapy was applied according to international recommendations.
Patients from Hamburg (n = 191) were treated at the University Medical Center Hamburg-Eppendorf, Germany, Department of Gynecology between 1991 and 2002 and selected on the basis of tissue availability. One hundred and five patients received anthracycline-based adjuvant chemotherapy regimens (mainly epirubicin/cyclophosphamide (EC) or cyclophosphamide/methotrexate/fluorouracil (CMF)). Seventy-five patients received endocrine therapy only, six patients were treated by radiation without any systemic therapy, and five patients remained untreated after surgery. The median follow-up time was 132 months.
The second cohort consisted of 200 consecutive lymph node-negative breast cancer patients, treated at the Department of Obstetrics and Gynecology of the Johannes Gutenberg University, Mainz between 1988 and 1998 [33]. The median time of follow-up was 92 months. Patients did not receive any systemic therapy in the adjuvant setting. Patients were treated either with modified radical mastectomy (n = 75) or with breast-conserving surgery followed by irradiation (n = 125), and did not show evidence of regional lymph node or distant metastases at the time of surgery.
The Frankfurt cohort (n = 171) included patients who underwent surgical breast cancer resection between 1996 and 2003 at the Department of Gynecology and Obstetrics at the Goethe-University of Frankfurt [34]. Patients were selected based on tissue availability, median follow-up time was 45 months. All of them had received adjuvant endocrine therapy (n = 52) or chemotherapy (CMF or EC; n = 119).
As clinical endpoint, we used RFS which was defined as the period (in months) from the date of diagnosis to the date at which relapse was clinically identified.
Analysis of Ki67, TOP2A, and RacGAP1 expression
All tissue samples were snap-frozen after surgery and stored in liquid nitrogen until use. RNA extraction, cDNA synthesis, and microarray analysis were performed in the same laboratory for all samples as described [35]. Tumor cell content exceeded 40 % in all the samples, as shown by H&E staining of cryo-cut sections.
The Affymetrix (Santa Clara, CA, USA) HG-U133A array and GeneChip System were used to quantify the relative transcript abundance in the breast cancer tissues. Hybridization intensity data were automatically acquired and processed by Affymetrix Microarray Suite 5.0 software. Arrays were analyzed using MAS5 algorithm. Arrays from different cohorts were adapted using magnitude normalization as described [36]. MAS5 expression values were log transformed and median centered over arrays.
For evaluation of ESR1 expression data, we used probeset 205225_at because of its highest concordance with biochemical ER status in comparison to the other ESR1 probesets present on the Affymetrix array [36]. Similarly, out of the two probesets for HER2 which are present on the U133A array, probeset 216836_at was chosen which worked best in ROC analysis in comparison with IHC/FISH results [36]. For Ki67, four probesets are included on the genechip (212020_s_at; 212021_s_at; 212022_s_at and 212023_s_at) which showed similar expression levels. Due to its broadest range of expression values, the probeset 212022_s_at was used for further analysis. Out of the two probesets for TOP2A, 201291_s_at and 201292_at, the latter probeset was chosen because of its higher expression values. For RacGAP1, only one probeset (222077_s_at) was present on the Affymetrix array.
Statistical analysis
We performed univariate Cox regression analysis using continuous variables and different cut-off points (see below) and constructed Kaplan–Meier curves and used log-rank tests to determine the significance of the three putative proliferation markers. A multivariate Cox proportional hazards regression model was used to examine simultaneously the effects of multiple covariates on survival including nodal involvement (positive vs. negative), tumor stage (T1, T2, T3, T4), histological grading (G1/G2 vs. G3) and the tested proliferation marker. The effect of each variable was assessed with the use of the Wald test and described by the hazard ratio, with a 95 % confidence interval. Correlations of Ki67, TOP2A, and RacGAP1 (quartiles) with clinical and histological variables were calculated by Chi-square tests using the groups given above, and correlations with age by Pearson test using the continuous data. All reported p-values are two sided and p-values of less than 0.05 were considered to indicate a significant result. All analyses were performed using the SPSS 20 software.
Results
Definition and characterization of molecular subgroups
Combining microarray datasets from three institutions, we were able to analyze mRNA data from a total of 562 breast cancer samples. According to Sorlie et al. [13], molecular subtypes include the ER-positive luminal A and B tumors, Her2-positive and basal-like tumors, the latter being generally ER- and Her2-negative. In order to analyze the prognostic or predictive impact of proliferation markers in different molecular subtypes of breast cancer, an approximated definition of these subtypes in our cohort was performed, based on estrogen receptor (ESR1) and Her2/neu (ErbB2) mRNA expression data (Fig. 1). The Affymetrix probesets which we used for this definition and the cut-off values for ESR1 and ERBB2 expression were chosen according to the results of a prior meta-analysis of ESR1 and ERBB2 mRNA expression in >3000 Affymetrix datasets [36]. Thus, tumors were defined as Her2-positive if the normalized expression of probeset 216836_at was above 0.0135. Similarly, tumors were regarded as ESR1-negative if expression levels of probeset 205225_at were below 0.0075.
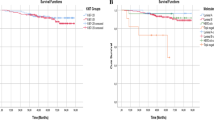
Characterization of the cohort with respect of normalized ErbB2 (a) and ESR1 (b) mRNA expression. For both markers, bimodal distributions of expression values were found. c Kaplan–Meier analysis of recurrence-free survival in HER2-positive patients (cut-off 0.0135), luminal tumors (cut-off 0.0075), and triple-negative tumors
By this approach, we defined a group of HER2high tumors, later referred to as “HER2-positive” (n = 69; 12 %) and, among the HER2-negative tumors, a cohort with low ESR1 expression levels (n = 96; 17 %). Since most ESR1-negative tumors also show absent or low expression of the progesterone receptor (PGR), the latter group is referred to as “TNT”). The majority of the cases of our cohort (n = 397; 71 %) belong to the ESR1high/HER2low (“luminal”) subgroup. To determine the clinical outcome of these molecular subgroups, Kaplan–Meier analysis and log-rank tests were performed using the RFS data. As shown in Fig. 1c, cases of the luminal subgroup had a significantly better prognosis compared to HER2-positive or ESR1low/HER2low (“triple-negative”) tumors in the first 10 years after surgery, but the probability of late recurrence is higher in the luminal carcinomas.
Prognostic value of the proliferation markers Ki67, TOP2A, and RacGAP1 in molecular breast cancer subtypes
Before analysis of the prognostic role of different proliferation markers, we first compared their expression patterns in our total cohort of 562 tumors. As shown in Fig. 2, all markers showed an approximately normal distribution. Regarding the expression data, the median expression values of TOP2A (median value = 0.0036) and RacGAP1 (median value = 0.0040) were clearly higher compared to those of Ki67 (median value = 0.0002). By Spearman correlation, we found significant positive correlations between RacGAP1 and TOP2A (ρ = 0.747), RacGAP1 and Ki67 (ρ = 0.734), and TOP2A and Ki67 (ρ = 0.741; all p < 0.0001; not shown). Regarding clinical and histological characteristics, none of the three markers correlated significantly with age, stage or nodal involvement, whereas all of them correlated with high grading (p < 0.001; not shown). Based on the hypothesis that proliferation markers behave differently in molecular subgroups, we next analyzed the prognostic and predictive impact of Ki67, TOP2A, and RacGAP1 separately in the luminal, triple-negative, and HER2-positive subcohorts.
Since proliferation has been shown to be a dominant prognostic factor in luminal (ER-positive, HER2-negative) tumors [11], we first used univariate Cox regression analysis to analyze the prognostic value of Ki67, TOP2A, and RacGAP1 in this group (n = 397). Using the continuous expression values, all three markers were significant prognostic indicators (Ki67, p = 0.040; TOP2A, p = 0.004; RacGAP1, p < 0.001; not shown). Since categorization is required in routine settings, the subcohort was then divided into four groups of similar size (quartiles, Q1–Q4), and different cut-offs for low versus high expression were applied. Interestingly, the cut-off value Q3 (quartiles 1–3 vs. Q4) gave the best prognostic information for all three markers (Table 2). Hazard ratios for high expression of Ki67, TOP2A, and RacGAP1 were 1,657, 1,657, and 2,053, respectively. For RacGAP1, a significant difference in outcome was also observed when the median (Q2) was used as cut-point, with a hazard ratio of 1,629 (p = 0,017; Table 2). In Fig. 3, the corresponding Kaplan–Meier plots for RFS and p-values after log-rank test are shown.
For identification of the most suitable marker in this group, we also performed a multivariate Cox regression analysis including all three proliferation markers, each with cut-off Q3 (quartiles Q1–3 vs. Q4). Interestingly, RacGAP1 expression was a significant prognostic indicator in this setting (p = 0.012), whereas Ki67 and TOP2A lost significance (Table S1). Similar results were obtained if the continuous expression data of Ki67, TOP2A, and RacGAP1 were analyzed in multivariate analysis (MVA). Only RacGAP1 expression retained its prognostic significance (p = 0.016; not shown). As additional prognostic variables, the histological grading, nodal involvement, and FIGO stage were included in the analysis of single proliferation markers. As shown in Table 3, high RacGAP1 expression was an independent, highly significant prognostic indicator in luminal tumors (HR = 2.136; p = 0.001), in addition to nodal involvement (HR = 1.844; p = 0.014) and advanced stage (p = 0.029). High expression of Ki67 and TOP2A also had a significant but weaker prognostic impact in MVA in this molecular subgroup (Ki67: HR 1.631; p = 0.038; TOP2A: HR 1.631; p = 0.038; Table 3).
By a similar approach, the relevance of the three proliferation markers was analyzed in “triple-negative” carcinomas (n = 96). Using the continuous variables, only Ki67 had a significant impact on prognosis in this subtype (Ki67, p = 0.017; TOP2A, p = 0.082; RacGAP1, p = 0.161). With the categorized variables, Ki67 expression above median (cut-off Q2) was a significant prognostic indicator in this group (HR 2.248; 95 % CI 1.111-4.546; p = 0.024; Table 2B). The negative correlation with RFS could also be shown in Kaplan–Meier analysis (Fig. 4a). In MVA including the three continuous markers, only Ki67 was of borderline significance (p = 0.050; not shown) which confirms its superior value in comparison with TOP2A and RacGAP1 in this molecular subtype. In MVA including stage, nodal status and histological grading, Ki67 lost its significance (p = 0.072; Table S2), and none of the other proliferation markers had an impact on RFS (not shown).
In HER2-positive tumors (n = 69), none of the proliferation markers had a significant prognostic impact in log-rank tests or univariate Cox regression analysis using continuous data or categorized variables (Table 2C). Yet, there was a tendency pointing to a longer RFS in tumors with low Ki67 expression (cut-off Q1; Table 2C; Fig. 4b). In MVA including all three markers (continuous expression values) or clinical and histological parameters, neither Ki67 nor TOP2A or RacGAP1 had a significant influence on RFS in this molecular subgroup (not shown).
Predictive value of the proliferation markers Ki67, TOP2A, and RacGAP1 in luminal breast cancer
Our cohort of luminal (ESRhigh/HER2low) carcinomas included patients who had undergone chemotherapy (n = 115), endocrine treatment without chemotherapy (n = 106) and patients who did not receive any systemic treatment after surgery (n = 176). In order to find out if the proliferation markers Ki67, TOP2A, or RacGAP1 might be predictive of therapy response, we performed log-rank tests and univariate Cox regression analysis in these subgroups using the cut-off values described above. The results of this stratified analysis are shown in Table 4.
In patients without systemic treatment, all three markers had a significant prognostic impact if the median expression values or the cut-off Q3 were used. In contrast, only RacGAP1 was a significant predictor of early recurrence in patients treated with endocrine therapy (p = 0.041). In chemotherapy-treated patients, high expression (upper quartile) of any of these markers was predictive of worse outcome. Yet, in contrast to Ki67 and TOP2A (p = 0.039 after log-rank test), RacGAP1 had a stronger influence with p = 0.003 (Table 4).
The influence of high RACGAP expression in chemotherapy-treated patients was further analyzed in MVA including stage, grading and nodal involvement. Here, RacGAP1 retained its predictive influence indicating that it might be an independent predictive marker for response to chemotherapy (p = 0.047; Table 5). In contrast, high Ki67 or high TOP2A expression (both markers were identical if the cut-off Q3 was used) lost significance.
In patients treated with endocrine therapy, none of the proliferation markers was an independent predictor in multivariate Cox regression analysis (not shown). Interestingly, in patients without systemic treatment, all three markers turned out as independent prognostic indicators after MVA (Table 6). In addition, only advanced stage was a significant predictor of recurrence.
Discussion
Studies based on gene expression profiling led to a reliable definition of molecular subgroups of breast cancer which differ in biological behavior, prognosis, and treatment options [12, 13]. Since these subtypes also use distinct intracellular signaling pathways, we suspected that there might also be discrepancies in proliferation markers which are of prognostic or predictive value in these subcohorts. For clinical routine, a single marker should be more practical than multigene classifiers.
In our analysis of three potential proliferation markers, Ki67, TOP2A, and RacGAP1, we found clear subtype-specific differences. Interestingly, high mRNA expression of the mitosis-associated gene RacGAP1 is a strong unfavorable prognostic marker in ER-positive (luminal), but not in HER2-positive or TNT. This prognostic value in luminal tumors is independent of histological grading, nodal involvement or clinical stage and is more pronounced than the prognostic value of Ki67 or TOP2A which behave similarly in this group. The correlation of Ki67 and TOP2A expression has been already described on mRNA basis [22] and in an immunohistochemical study [37]. Although RacGAP1 expression correlates with both TOP2A and Ki67, their prognostic impact in breast cancer subgroups is unequal. Regarding the biological role of RacGAP1, there is still little information, but previous data suggest that this protein is essential for cytokinesis during mitosis [26, 27]. Thus, its expression might reflect the proliferation rate in luminal cancer cells. In contrast, Ki67 is the only significant prognostic proliferation marker in “triple-negative” carcinomas in our cohort. In HER2-positive carcinomas, none of the three proliferation markers had a significant prognostic impact. These results confirm our assumption that the molecular pathways leading to strong proliferation of the tumor cells might differ between molecular subtypes.
Proliferation has been shown to be a strong and independent prognostic indicator in ER-positive (luminal) breast cancer [11, 14]. In MVA, the proliferation module had the highest HR and the most significant p value, followed by histological grade [11]. In a subset of patients with relatively high ER expression, Dai et al. identified another signature for definition of cases with an extremely poor prognosis. Most of the 50 prognostic genes of this signature are cell cycle-associated genes, among them RacGAP1 (ID-GAP) [38]. Among the group of luminal tumors, gene expression analysis identified two groups of different prognosis: a larger group (luminal A) which is characterized by relatively low proliferation [10] and a smaller group with higher Ki67 levels and poor outcome (luminal B). In our cohort, the upper quartile of the patients with the highest Ki67, TOP2A, or RacGAP1 levels had a significantly shorter RFS which might roughly correspond to the luminal B subtype.
Stratification of our analysis according to treatment strategy in luminal tumors revealed that RacGAP1 expression is not only a prognostic marker, but might also predict treatment response in Tamoxifen-treated patients and in those treated with adjuvant chemotherapy with or without endocrine treatment. In cases without any systemic treatment, all three markers (Ki67, TOP2A, RacGAP1) had a significant influence on RFS which is largely independent of the cut-off used for analysis. This confirms earlier results which indicated that outcome is largely dependent on tumor proliferation in untreated patients [33]. Since chemotherapy targets at mitotic cells, the negative prognostic influence of proliferation is partly abrogated by the higher sensitivity toward chemotherapy in this treatment group. Only tumors with high expression levels of Ki67, TOP2A, or RacGAP1 (cut-off Q3; upper 25 vs. lower 75 %) have a poor outcome in chemotherapy-treated patients, where the overall effect of these biomarkers probably is a mixture of prognostic and predictive impact. Among the three proteins, RacGAP1 turned out to be the most reliable predictive marker since it retained its significance in MVA.
Our results obtained by Ki67 analysis after stratification of the cohort are largely in line with those of a recent study where Ki67 was investigated by immunohistochemistry in G2 breast cancer patients: High Ki67 index was a significant prognostic indicator in untreated patients, whereas in Tamoxifen-treated cases, only small differences between tumors with high versus low Ki67 expression were observed [39]. In another study, Ki67 was a predictive marker for response to chemotherapy [21]. In our RNA-based analysis, Ki67 was prognostic and predictive for chemotherapy response in ER-positive tumors. Yet, its value is outperformed by RacGAP1 in our cohort, since this marker is highly prognostic even in MVA.
In conclusion, our results indicate that the molecular breast cancer subtypes differ in the relevance of the proliferation markers Ki67, TOP2a, and RacGAP1. To our knowledge, this is the first publication which shows that in luminal carcinomas, RacGAP1 mRNA expression is superior to Ki67 and TOP2a as prognostic and predictive marker. Thus, determination of this new marker on RNA level might be a simple alternative to multigene signatures in ER+/HER2− breast cancer. It should be evaluated if this marker can also applied by immunohistochemistry similar to Ki67. Limitations of this study are its retrospective design and the heterogeneous group of patients with relatively small numbers in the treatment groups. Therefore, further studies are needed to validate the prognostic and predictive value of RacGAP1 expression in luminal breast cancer.
References
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100:57–70
Colozza M, Azambuja E, Cardoso F, Sotiriou C, Larsimont D, Piccart MJ (2005) Proliferative markers as prognostic and predictive tools in early breast cancer: where are we now? Ann Oncol 16:1723–1739
Beresford MJ, Wilson GD, Makris A (2006) Measuring proliferation in breast cancer: practicalities and applications. Breast Cancer Res 8:216
Stuart-Harris R, Caldas C, Pinder SE, Pharoah P (2008) Proliferation markers and survival in early breast cancer: a systematic review and meta-analysis of 85 studies in 32,825 patients. Breast 17:323–334
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T et al (2004) A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351:2817–2826
Chang HY, Sneddon JB, Alizadeh AA, Sood R, West RB, Montgomery K, Chi JT, van de Rijn M, Botstein D, Brown PO (2004) Gene expression signature of fibroblast serum response predicts human cancer progression: similarities between tumors and wounds. PLoS Biol 2:E7
Foekens JA, Atkins D, Zhang Y, Sweep FC, Harbeck N, Paradiso A, Cufer T, Sieuwerts AM, Talantov D, Span PN et al (2006) Multicenter validation of a gene expression-based prognostic signature in lymph node-negative primary breast cancer. J Clin Oncol 24:1665–1671
van’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AAM, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT et al (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415:530–536
Wang Y, Klijn JG, Zhang Y, Sieuwerts AM, Look MP, Yang F, Talantov D, Timmermans M, Meijer-van Gelder ME, Yu J et al (2005) Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365:671–679
Wirapati P, Sotiriou C, Kunkel S, Farmer P, Pradervand S, Haibe-Kains B, Desmedt C, Ignatiadis M, Sengstag T, Schutz F et al (2008) Meta-analysis of gene expression profiles in breast cancer: toward a unified understanding of breast cancer subtyping and prognosis signatures. Breast Cancer Res 10:R65
Desmedt C, Haibe-Kains B, Wirapati P, Buyse M, Larsimont D, Bontempi G, Delorenzi M, Piccart M, Sotiriou C (2008) Biological processes associated with breast cancer clinical outcome depend on the molecular subtypes. Clin Cancer Res 14:5158–5165
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnson H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98:10869–10874
Iwamoto T, Bianchini G, Booser D, Qi Y, Coutant C, Ya-Hui Shiang C, Santarpia L, Matsuoka J, Hortobagyi GN, Symmans WF et al (2010) Gene pathways associated with prognosis and chemotherapy sensitivity in molecular subtypes of breast cancer. J Natl Cancer Inst 103:264–272
Dowsett M, Nielsen TO, A’Hern R, Bartlett J, Coombes RC, Cuzick J, Ellis M, Henry NL, Hugh JC, Lively T et al (2011) Assessment of Ki67 in breast cancer: recommendations from the International Ki67 in Breast Cancer Working Group. J Natl Cancer Inst 103:1656–1664
Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thurlimann B, Senn HJ (2011) Strategies for subtypes—dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol 22:1736–1747
Varga Z, Diebold J, Dommann-Scherrer C, Frick H, Kaup D, Noske A, Obermann E, Ohlschlegel C, Padberg B, Rakozy C et al (2012) How reliable is Ki-67 immunohistochemistry in grade 2 breast carcinomas? A QA study of the Swiss Working Group of Breast- and Gynecopathologists. PLoS One 7:e37379
van den Akker EB, Verbruggen B, Heijmans BT, Beekman M, Kok JN, Slagboom PE, Reinders MJ (2011) Integrating protein–protein interaction networks with gene–gene co-expression networks improves gene signatures for classifying breast cancer metastasis. J Integr Bioinform 8:188
Gerdes J, Li L, Schlueter C, Duchrow M, Wohlenberg C, Gerlach C, Stahmer I, Kloth S, Brandt E, Flad HD (1991) Immunobiochemical and molecular biologic characterization of the cell proliferation-associated nuclear antigen that is defined by monoclonal antibody Ki-67. Am J Pathol 138:867–873
Urruticoechea A, Smith IE, Dowsett M (2005) Proliferation marker Ki-67 in early breast cancer. J Clin Oncol 23:7212–7220
Yerushalmi R, Woods R, Ravdin PM, Hayes MM, Gelmon KA (2010) Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol 11:174–183
Rody A, Karn T, Ruckhaberle E, Muller V, Gehrmann M, Solbach C, Ahr A, Gatje R, Holtrich U, Kaufmann M (2009) Gene expression of topoisomerase II alpha (TOP2A) by microarray analysis is highly prognostic in estrogen receptor (ER) positive breast cancer. Breast Cancer Res Treat 113:457–466
Nielsen KV, Ejlertsen B, Moller S, Jorgensen JT, Knoop A, Knudsen H, Mouridsen HT (2008) The value of TOP2A gene copy number variation as a biomarker in breast cancer: update of DBCG trial 89D. Acta Oncol 47:725–734
O’Malley FP, Chia S, Tu D, Shepherd LE, Levine MN, Bramwell VH, Andrulis IL, Pritchard KI (2009) Topoisomerase II alpha and responsiveness of breast cancer to adjuvant chemotherapy. J Natl Cancer Inst 101:644–650
Brase JC, Schmidt M, Fischbach T, Sultmann H, Bojar H, Koelbl H, Hellwig B, Rahnenfuhrer J, Hengstler JG, Gehrmann MC (2010) ERBB2 and TOP2A in breast cancer: a comprehensive analysis of gene amplification, RNA levels, and protein expression and their influence on prognosis and prediction. Clin Cancer Res 16:2391–2401
Hirose K, Kawashima T, Iwamoto I, Nosaka T, Kitamura T (2001) MgcRacGAP is involved in cytokinesis through associating with mitotic spindle and midbody. J Biol Chem 276:5821–5828
Minoshima Y, Kawashima T, Hirose K, Tonozuka Y, Kawajiri A, Bao YC, Deng X, Tatsuka M, Narumiya S, May WS Jr et al (2003) Phosphorylation by aurora B converts MgcRacGAP to a RhoGAP during cytokinesis. Dev Cell 4:549–560
Kawashima T, Bao YC, Minoshima Y, Nomura Y, Hatori T, Hori T, Fukagawa T, Fukada T, Takahashi N, Nosaka T et al (2009) A Rac GTPase-activating protein, MgcRacGAP, is a nuclear localizing signal-containing nuclear chaperone in the activation of STAT transcription factors. Mol Cell Biol 29:1796–1813
Lu KH, Patterson AP, Wang L, Marquez RT, Atkinson EN, Baggerly KA, Ramoth LR, Rosen DG, Liu J, Hellstrom I et al (2004) Selection of potential markers for epithelial ovarian cancer with gene expression arrays and recursive descent partition analysis. Clin Cancer Res 10:3291–3300
Stone R 2nd, Sabichi AL, Gill J, Lee IL, Adegboyega P, Dai MS, Loganantharaj R, Trutschl M, Cvek U, Clifford JL (2010) Identification of genes correlated with early-stage bladder cancer progression. Cancer Prev Res (Phila) 3:776–786
Wang SM, Ooi LL, Hui KM (2011) Upregulation of Rac GTPase-activating protein 1 is significantly associated with the early recurrence of human hepatocellular carcinoma. Clin Cancer Res 17:6040–6051
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM (2006) Reporting recommendations for tumor MARKer prognostic studies (REMARK). Breast Cancer Res Treat 100:229–235
Schmidt M, Bohm D, von Torne C, Steiner E, Puhl A, Pilch H, Lehr HA, Hengstler JG, Kolbl H, Gehrmann M (2008) The humoral immune system has a key prognostic impact in node-negative breast cancer. Cancer Res 68:5405–5413
Ruckhäberle E, Rody A, Engels K, Gaetje R, von Minckwitz G, Schiffmann S, Grosch S, Geisslinger G, Holtrich U, Karn T et al (2008) Microarray analysis of altered sphingolipid metabolism reveals prognostic significance of sphingosine kinase 1 in breast cancer. Breast Cancer Res Treat 112:41–52
Ihnen M, Muller V, Wirtz RM, Schroder C, Krenkel S, Witzel I, Lisboa BW, Janicke F, Milde-Langosch K (2008) Predictive impact of activated leukocyte cell adhesion molecule (ALCAM/CD166) in breast cancer. Breast Cancer Res Treat 112(3):419–427
Karn T, Metzler D, Ruckhaberle E, Hanker L, Gatje R, Solbach C, Ahr A, Schmidt M, Holtrich U, Kaufmann M et al (2010) Data-driven derivation of cutoffs from a pool of 3,030 Affymetrix arrays to stratify distinct clinical types of breast cancer. Breast Cancer Res Treat 120:567–579
Fountzilas G, Valavanis C, Kotoula V, Eleftheraki AG, Kalogeras KT, Tzaida O, Batistatou A, Kronenwett R, Wirtz RM, Bobos M et al (2012) HER2 and TOP2A in high-risk early breast cancer patients treated with adjuvant epirubicin-based dose-dense sequential chemotherapy. J Transl Med 10:10
Dai H, van’t Veer L, Lamb J, He YD, Mao M, Fine BM, Bernards R, van de Vijver M, Deutsch P, Sachs A et al (2005) A cell proliferation signature is a marker of extremely poor outcome in a subpopulation of breast cancer patients. Cancer Res 65:4059–4066
Aleskandarany MA, Rakha EA, Macmillan RD, Powe DG, Ellis IO, Green AR (2011) MIB1/Ki-67 labelling index can classify grade 2 breast cancer into two clinically distinct subgroups. Breast Cancer Res Treat 127:591–599
Conflict of interest
R.M. Wirtz is employee of a diagnostic company. The other authors have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Milde-Langosch, K., Karn, T., Müller, V. et al. Validity of the proliferation markers Ki67, TOP2A, and RacGAP1 in molecular subgroups of breast cancer. Breast Cancer Res Treat 137, 57–67 (2013). https://doi.org/10.1007/s10549-012-2296-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-012-2296-x