Abstract
Tumor recurrence and metastasis result in an unfavorable prognosis for cancer patients. Recent studies have suggested that specific microRNAs (miRNAs) may play important roles in the development of cancer cells. However, prognostic markers and the outcome prediction of the miRNA signature in breast cancer patients have not been comprehensively assessed. The aim of this study was to identify miRNA biomarkers relating to clinicopathological features and outcome of breast cancer. A miRNA microarray analysis was performed on breast tumors of different lymph node metastasis status and with different progression signatures, indicated by overexpression of cyclin D1 and β-catenin genes, to identify miRNAs showing a significant difference in expression. The functional interaction between the candidate miRNA, miR-30a, and the target gene, Vim, which codes for vimentin, a protein involved in epithelial–mesenchymal transition, was examined using the luciferase reporter assay, western blotting, and migration and invasion assays. The association between the decreased miR-30a levels and breast cancer progression was examined in a survival analysis. miR-30a negatively regulated vimentin expression by binding to the 3′-untranslated region of Vim. Overexpression of miR-30a suppressed the migration and invasiveness phenotypes of breast cancer cell lines. Moreover, reduced tumor expression of miR-30a in breast cancer patients was associated with an unfavorable outcome, including late tumor stage, lymph node metastasis, and worse progression (mortality and recurrence) (p < 0.05). In conclusion, these findings suggest a role for miR-30a in inhibiting breast tumor invasiveness and metastasis. The finding that miR-30a downmodulates vimentin expression might provide a therapeutic target for the treatment of breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In contrast to the total abolition of gene function by permanent mutation, the reversibility of epigenetic modification allows gene expression to be switched on and off and thus is suggested to provide selective advantage for clonal evolution during tumor progression [1, 2]. Tumor recurrence and metastasis result in an unfavorable prognosis for patients with cancer. Tumor cell metastasis is characterized by an unstable phenotypic heterogeneity, in which the phenotype fluctuates too frequently for the changes to be mediated exclusively by irreversible genetic alterations. During tumor metastasis, cancer cells undergo epithelial–mesenchymal transition (EMT) in which epithelial tumor cells in the primary cancer site are converted into aggressive and metastatic tumor cells. One of the main characteristics of EMT is the combination of loss of cell–cell contact, reduced E-cadherin expression, and enhanced expression of mesenchymal markers, such as vimentin [3–5]. The mesenchymal-like cells generated are transported to metastatic sites, where they may undergo mesenchymal–epithelial transition by regaining E-cadherin expression, allowing cell–cell adhesion and connecting adjacent cells to form new foci [6]. For this reason, epigenetic regulation appears to be more important than genetic level regulation in maintaining flexibility, and, in the case of E-cadherin, hypermethylation of the promoter region is seen in various human carcinomas [7–10]. In cancer metastasis, there is evidence for a significant contribution of epigenetic regulation of EMT by microRNAs (miRNAs) [11, 12], a novel class of short non-coding RNA molecules consisting of 19–25 nucleotides with the potential to inhibit gene expression by binding to complementary sequences in the 3′-untranslated region (UTR) of target mRNA transcripts. miRNAs are an emerging class of negative regulators, and deregulation of miRNAs in tumors affects the expression of oncogenes and/or tumor suppressor genes and leads to malignant transformation in human cancer [13–16]. However, knowledge of prognostic markers and the effectiveness of outcome prediction of the miRNA signature in breast cancer patients is limited.
We recently investigated a gene expression signature that is an important predictor of an unfavorable outcome in patients with invasive ductal carcinoma (IDC) and found that overexpression of the genes making up this expression signature, namely cyclin D1 and β-catenin, is highly associated with tumor cells of an advanced stage, lymph node metastasis (LNM), and a decreased survival rate (manuscript submitted). To examine the mechanism responsible for the changes associated with this signature, the present study investigated whether and how deregulation of specific miRNAs associated with epigenetic aberrance was involved in driving the invasiveness and metastasis of tumor cells. The miRNA expression profile was then compared between breast tumors showing overexpression of cyclin D1 and β-catenin and LNM and those with a normal signatures of both genes and no LNM. Among the miRNAs showing a significant difference in expression in tumor cells, particular attention was focused on miR-30a, as in silico prediction of target genes possibly regulated by this miRNA molecule resulted in the identification of the gene, Vim, coding for vimentin, a mesenchymal marker implicated in EMT during breast tumorigenesis [17–19]. We explored whether miR-30a regulated vimentin expression and showed that increased expression of miR-30a resulted in downregulation of vimentin expression and a subsequent decrease in the vimentin-mediated migration and invasiveness of breast cancer cells. Moreover, reduced levels of miR-30a expression in the primary breast tumor tissue were found to predict an unfavorable outcome, including advanced stage, LNM, and a decreased survival rate in IDC patients. These findings identify the role of miR-30a in breast cancer and shed light on a novel fundamental mechanism with clinical significance and translational implications.
Materials and methods
Study population
The present study is part of an ongoing cooperative study aimed at discovering markers for the evaluation of breast cancer progression in Taiwan, where breast cancer is characterized by low incidence [20], early tumor onset [21], reproductive hormone dependency [22, 23], and novel genomic alteration [23–26]. The study was approved by the Ethics Committees of the Institutional Review Boards of the Tri-Service General Hospital, Taipei, and the Chung Shan Medical University Hospital, Taichung, Taiwan. The enrolled female patients pathologically confirmed primary IDC of the breast were a subset of women randomly selected from the ongoing hospital-based breast cancer cohort collected in the Surgery Department of the Tri-Service General Hospital, collected between July 1997 and April 2006. Informed consent was obtained from each participant prior to specimen acquisition. The resected breast cancer tissues were immediately frozen in liquid nitrogen until analysis. Tumor grade in each patient was categorized as I, II, or III according to the Nottingham modification of the Scarff–Bloom–Richardson system, and the pathology of these tumors was classified according to the sixth edition of the AJCC Cancer Staging Manual. None of the patients received neoadjuvant treatment before primary surgery, thus avoiding any effects on gene expression. The histological diagnosis of all specimens was reviewed by a certified pathologic physician, and the clinicopathological findings are summarized in Table 1.
Laser capture microdissection
To ensure that the tissue samples assayed consisted of >95 % pure breast tumor epithelial cells, laser capture microdissection (LCM) was performed on routinely immunostained slides using a PixCell laser capture microscope (Arcturus Engineering, Mountain View, CA, USA) as described previously [27, 28]. The dehydrated tissue section was overlaid with a thermoplastic film mounted on an optically transparent cap, and the visually selected areas (tumor cells) were bound to the membrane by short, low-energy laser pulses, resulting in focal melting of the polymer. On average, 2,500–3,000 LCM shots were performed on a single tumor to obtain sufficient tumor cells for comparative qRT-PCR analysis. The laser-captured tumor cells were immersed in 50–100 μl of digestion buffer (10 mM Tris–HCl, pH 8.0, 1 mM EDTA, 400 μg/ml of proteinase K, and 1 % Tween 20) and digested at 55 °C overnight. After digestion, the enzyme was heat-inactivated (95 °C for 10 min) and the extract used directly for RNA isolation.
RNA isolation
Total RNA was isolated from paired LCM-dissected tumor and non-tumor cells from each patient using an RNAqueous®-Micro Kit (Ambion Inc., Austin, TX, USA) and the yield of RNA determined by spectrophotometry at 260 nm. miRNAs from tumor and non-tumor cells were extracted from tissue sections using a mirVana miRNA isolation kit according to the manufacturer’s instructions (Ambion Inc., Austin, TX, USA) and the RNA concentration in each sample quantified on a NanoDrop 1000 spectrophotometer (NanoDrop Technologies, Waltham, MA, USA).
miRNA microarray
A human miRNA microarray contained probes for 381 human miRNAs from TaqMan® Low Density Array Human MicroRNA Panel v1.0 (Applied Biosystems) and levels of mature miRNAs were analyzed on an Applied Biosystems 7900HT Fast Real-Time PCR System (Foster City, CA, USA). Expression level of each miRNA was measured using the same amount of template RNA in each well with small nuclear RNA U48 (RNU48) as an endogenous negative control and RNU6B as a positive control. Relative quantification of miRNA expression was performed using the 2-ddCt method. The threshold cycle, Ct was automatically calculated by the SDS2.2 software package (Applied Biosystems, Foster City, CA, USA). Ct values for all targets were determined using the automatic threshold in RQ Manager v1.1 analysis software.
Cell culture
The breast cancer cell lines Hs578T and MDA-MB-231 were cultured in DMEM (Life Technologies) containing 0.1 mM sodium pyruvate, 10 % fetal bovine serum (FBS), 2 mmol/l of l-glutamine, 100 IU/ml of penicillin, and 100 mg/ml of streptomycin (all from Biosource, Rockville, MD, USA) in a humidified 5 % CO2 atmosphere at 37 °C. Transfection with different miRNAs and plasmid constructs was performed using DOTAP (Biontex, Laboratories, GmbH) and Turbofect™ (Fermentas, Germany) in vitro transfection reagents according to the manufacturer’s recommendations. Transfectants were cultured and selected for 2 weeks in the medium containing 3 μg/ml of puromycin.
Western blotting analysis
Cell extracts were prepared in ice-cold RIPA lysis buffer. After whole cell protein extracts were quantified by BCA protein assay, equivalent amounts of cell lysates were resolved by 8–12 % SDS polyacrylamide gel electrophoresis and transferred onto a polyvinylidene difluoride membrane, which was then blocked in 5 % non-fat milk in PBST and probed overnight at 4 °C with a monoclonal antibody against human vimentin (1:500, Santa Cruz Biotechnology). Anti-β-actin (Sigma-Aldrich Corp., St. Louis, MO, USA) was used to normalize for protein loading. The blot was incubated with an appropriate horseradish peroxidase conjugated second antibody and immunoreactive proteins visualized by the enhanced chemiluminescence assay (western blotting luminal reagent; Santa Cruz Biotechnology) and the band intensities quantified by densitometry (Digital Protein DNA Imagineware, Huntington Station, NY, USA).
Dual luciferase reporter assay
The 3′UTR sequence of the human vimentin gene (Vim) was cloned into plasmid pGL4.13 (Promega, Madison, WI), yielding the recombinant vector pGL4.13_1, containing the firefly luciferase open reading frame under the control of the SV40 promoter. Two miR-30a complementary sites with the sequence GTTTAC in the Vim 3′UTR were mutated singly or together to remove complementarity to miR-30a using a QuikChange II XL site-directed mutagenesis kit (Stratagene, La Jolla, CA, USA) with pGL4.13_1/Vim-WT as the template, and the mutants were named Vim 3′UTR/mut1, Vim 3′UTR/mut2, and Vim 3′UTR/mut12. The mutated nucleotides capitalized, and the sequences of the mismatch primers used to generate different Vim 3′UTR mutants are shown in Table 2. MDA-MB-231 cells were cotransfected with the reporter construct, the control or mutated Vim 3′UTR constructs, and pCDNA3/miR-30a, then the cells were lysed 24 h later, and the firefly luciferase/Renilla luciferase activity ratio of each sample measured in a dual-luciferase assay (Promega, Madison, WI, USA). The experiments were performed in triplicate. Data are presented as mean ± standard deviation (SD) for each transfection condition. Significance of comparisons was assessed by using the two-sided unpaired t test for means. A p value <0.05 is taken to indicate statistical significance.
Establishment of breast tumor cells stably expressing miR-30a
Using a lentivirus expression system (Thermo Fisher Scientific; Waltham, MA, USA) and the Trans-Lentiviral™ GIPZ Packaging System (Open Biosystems, Huntsville, AL, USA), breast cancer cells were transduced with plasmid pLemiR expressing primary-miR-30a (pri-miR-30a) transcripts under the control of the CMV promoter (Open Biosystems, Huntsville, AL, USA). TurboRed Fluorescent Protein and a puromycin-resistance selectable marker were used to allow screening for non-transduced cells, and cell lines stably expressing miR-30a were established.
Invasion and migration assays
For the invasion assay, matrigel (Collaborative Biomedical Products, Bedford, MA, USA) was applied to 8-μm pore size polycarbonate membrane filters, and then cells (1.0 × 105) cultured in DMEM were seeded into the upper section of the Boyden chamber (Neuro Probe, Cabin John, MD, USA). The lower chamber contained the same medium plus 10 % FBS, and the chamber was incubated overnight at 37 °C, after which non-invading cells were removed from the interior of the insert using a cotton-tip applicator, and the invasive cells attached to the lower surface of the membrane were fixed with methanol and stained with Giemsa. Invading cells were quantified by counting five random high-powered fields using a Olympus Ckx41 light microscope. For the migration assay, the cells were seeded into the Boyden chamber on membrane filters that were not coated with matrigel and incubated for 16 h at 37 °C, then non-migrating cells were removed from the upper membrane surface, and the invading cells on the lower membrane surface quantified as described above.
Comparatively quantitative real-time PCR analysis
The LCM was performed on 221 breast cancer tissue slides, and the single-tube TaqMan miRNA assay (Applied Biosystems, Foster City, CA, USA) was used to detect and quantify the mature miRNA on an Applied Biosystems instruments. The level of expression of the miRNA biomarker was determined using the TaqMan real-time PCR assay and normalized to that for RNU6B. Triplicate qPCR experiments were performed on each breast carcinoma to determine the levels of the target mRNA in the isolated tumor and non-tumor cells. The comparative CT method (-ddCt) was used to estimate the relative expression (fold change) of the miR-30a transcript in the tumor and non-tumor cells in each case (2-ddCt, where ddCt = dCt miR-30a − dCt RNU6B).
Statistical analysis
The Chi-squared test was used to examine whether there was an association between decreased miR-30a transcript levels in cancer tissue and the pathological features of tumor (stage and LNM status). The miR-30a transcript was quantified using the comparative CT method as described previously [29, 30]. To examine whether miR-30a could be used as a prognostic biomarker in breast cancer patients, Kaplan–Meier survival analysis (p value of the log-rank test) and Cox regression analysis [hazard ratio (HR) and 95 % confidence interval (95 % CI)] were used to explore the association between the 5-year recurrence-free/disease-free survival rate and miR-30a transcript levels in tumors of breast cancer patients. Recurrence-free survival was measured as the time from surgery to recurrence or the end of the study, while disease-free survival was defined as the time from surgery to recurrence or death or the end of the study.
Results
A miRNA expression signature in the primary tumor predicts its metastatic potential
To search for miRNAs that might play a role in breast cancer progression, we used miRNA chips to compare the miRNA expression profiles of three groups of patients with differential expression of the cyclin D1 and β-catenin genes and different LNM status. We compared the profiles of five patients with Stage I/II cancer (LNM−) and normal expression of cyclin D1 and β-catenin (group I), four tumors with Stage I/II (LNM−) and increased expression of cyclin D1 and β-catenin (group II), and five tumors with Stage III/IV (LNM+) and increased expression of cyclin D1 and β-catenin (group III). None of the fourteen patients had received neoadjuvant treatment before surgery, thus avoiding any confounding effect on gene expression. Cyclin D1 and β-catenin make up an expression signature that is highly associated with poor pathological features and a worse clinical outcome (manuscript submitted). It should be noted that the determination of the miRNA expression pattern was based on cancerous cells microdissected from tumor tissues to ensure that the sample tested consisted of >95 % pure breast tumor epithelial cells (Fig. 1a), supporting the validity of our measurement. Using a cutoff of a greater than twofold difference in levels, we identified 52 miRNAs that were significantly deregulated in the primary tumor of patients with metastatic breast cancer than in those with no metastases, 25 of which were upregulated (red section of Fig. 1b) in group III or II compared with group I and 27 downregulated (green section of Fig. 1b). The five highly downregulated miRNAs were miR-502, miR-485, miR-328, miR-30a, and miR-519e that may affect vimentin expression due to their being predicted to bind 3′UTR of the Vim gene in an in silico analysis, and the results for the first four are shown in Fig. 1c. Those identified 52 miRNAs that were differentially expressed between the metastatic tumors and the non-metastatic samples are summarized in Fig. 1d.
miR-30a expression is decreased in breast tumors with overexpression of cyclin D1 and β-catenin or lymph node metastasis (LNM). a Illustration of the laser capture-microdissection technique. The left, center, and right panels show the tissue before laser capture-microdissection LCM (i.e., HE staining), the LCM-captured cells, and the tumor after LCM, respectively. b and c Total RNA was extracted from LCM tumor cells from low-stage (Stage I/II) breast tumors (n = 5) showing normal expression of cyclin D1 and β-catenin and no LNM (group I); low-stage tumors (n = 4) showing increased expression of cyclin D1 and β-catenin, but no LNM (group II); and high-stage (Stage III/IV) tumors (n = 5) showing increased expression of cyclin D1 and β-catenin and LNM (Group III). RNAs from tumors in the same group were pooled and subjected to TaqMan-LDA miRNA microarray analysis. b shows a comparison of the results for group II and I (left panel) and for group III and I (right panel); 52 miRNAs showing a greater than twofold change in expression were associated with tumor stage and LNM, of which 25 were upregulated (red) and 27 downregulated (green) during tumor progression. c The five highly downregulated miRNAs were miR-328, miR-485, miR-502, miR-30a, and miR-519e; the quantitative data for the first four are shown. d 52 miRNAs that were differentially expressed between the metastatic tumors and the non-metastatic samples. (Color figure online)
To gain an insight into the functional consequences of differential miRNA expression, we used the miRNA target searching program TargetScan 5.1 (www.microrna.org) and a computational algorithm to explore whether the expression of any EMT-associated genes was regulated by these five miRNAs, and the results suggested that expression of the Vim gene, encoding an intermediate filament normally expressed in cells of mesenchymal origin, might be associated with decreased expression of these miRNAs. Vimentin, an important EMT-associated marker, is involved in linking the cytoskeleton to the membrane, and deregulation of vimentin has been suggested to be associated with increased invasiveness or migration of cells [18, 31].
miR-30a directly targets the vimentin 3′UTR and downregulates vimentin expression
To test the in silico prediction that vimentin expression might be modulated by specific miRNAs, we measured vimentin levels by immunoblotting after transfection of the human breast cancer cell lines Hs578T and MDA-MB-231 with the precursors of the above five miRNAs predicted to affect vimentin expression or with vector alone (Fig. 2a). As shown in Fig. 2b, no significant difference in Vim mRNA levels was found in breast cancer cells transfected with these five miRNAs, whereas, in the same cells, vimentin protein expression was inhibited by more than 60 % by miR-30a; however, the other four miRNAs had only a minor, or a non-significant, effect (Fig. 2c, d).
miR-30a represses vimentin expression. a A computational algorithm was used to predict five miRNAs that might affect vimentin expression. b–d miRNAs that were predicted to impact vimentin expression were transfected into breast cancer cell lines Hs578T and MDA-MB-231; then, 24 h later, vimentin mRNA levels were measured by RT-PCR in (b) and vimentin protein levels by western blotting in (c) and compared to those in cells transfected with control vector. d Quantitative results for three independent experiments showing suppression of vimentin expression on western blots by overexpression of miR-30a in Hs578T cells (left panel) or MDA-MB-231 cells (right panel). *p < 0.05
Furthermore, to map putative interaction sites between vimentin 3′UTR and miR-30a, three algorithms, miRanda, PicTar, and TargetScan, were used to predict the mRNA targets for miR-30a, and two potential sites were identified within the 3′UTR of Vim in different species (Fig. 3a). To determine whether miR-30a directly targets Vim mRNA and the relative importance of these two sites in repressing vimentin expression, we inserted the full-length 3′UTR of Vim into the pGL4.13-luciferase reporter (Vim 3′UTR-luc) and examined the effect of miR-30a on the luciferase activity. In addition, we generated three 3′UTR mutants, Vim 3′UTR/Mut1-luc, and Vim 3′UTR/Mut2-luc with a mismatched version of the miR-30a complementary sequence within site 1 or 2, and Vim 3′UTR/Mut12-luc containing both mismatched sequences (Fig. 3b). We found that miR-30a significantly reduced the activity of the luciferase gene fused to the Vim 3′UTR by more than 45 %, supporting the notion that miR-30a can modulate vimentin expression by binding to its 3′UTR (Fig. 3c). In addition, a significant reduction (54 %) in luciferase activity was observed in the presence of pre-miR-30a using the reporter construct containing the Vim 3′UTR/Mut1 clone, but not the Vim 3′UTR/Mut2 or Vim 3′UTR/Mut12 clones. These results indicate that the region from 178 to 184 is the important site within the 3′UTR of the Vim gene that is required for miR-30a binding.
miR-30a as a vimentin-targeting miRNA. a Sequence of the 3′UTR of vimentin mRNA showing the two predicted seed regions for the binding of miR-30a in different vertebrate species. b The sequences at sites 1 and 2 of the three mutants with a mismatch of the miR-30a complementary sequence at site 1 (Vim 3′UTR/Mut1) or site 2 (Vim 3′UTR/Mut2) or both (Vim 3′UTR/Mut12). c Constructs containing the reporter gene luciferase 3′-linked to the wild-type or a mutant 3′UTR of Vim (luc-Vim 3′UTR/WT) and miR-30a were cotransfected into MDA-MB-231 cells and the firefly luciferase activity of the reporter measured and normalized to that of the internal Renilla luciferase, The values are the mean ± SD for three independent experiments. **p < 0.01
Overexpression of miR-30a suppresses the motility and invasiveness of breast cancer cells
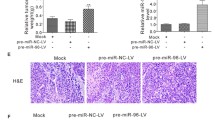
Overexpression of vimentin has been associated with enhanced metastatic potential in breast cancer [32, 33]. To confirm the role of miR-30a in inhibiting breast cancer progression, we studied the effect of miR-30a on the migration and invasion of breast cancer cell lines. In two stable clones of Hs578T and MD-MBA-231 lentivirally transduced with miR-30a, we first confirmed that vimentin expression was inhibited by miR-30a (Fig. 4a), and then found that the migration (Fig. 4b) and invasiveness (Fig. 4c) of these cells were significantly inhibited compared with control oligonucleotide transfected cells. Moreover, decreased vimentin protein levels in the miR-30a-transduced breast cancer cells were restored by transfection of the Anti-miR™ miRNA inhibitor of miR-30a (anti-miR-30a) (Ambion, Inc), which also significantly enhanced cell migration (Fig. 4b, d) and invasion (Fig. 4c, d) compared with transfected cell lines with negative control. In parallel, we investigated a dramatic decrease in the migration, and invasion of breast cancer cells was detected following vimentin silencing (Fig. 4b–d), in which it was verified that the inhibitory effect of miR-30a on tumor motility was mediated by downregulation of vimentin.
Inhibitory effect of miR-30a on the migration and invasiveness of breast cancer cell lines. a Expression of miR-30a suppresses vimentin expression in the breast cancer cell lines Hs578T (upper panel) and MDA-MB-231 (lower panel), and this effect is overcome in cells treated with an inhibitor of miR-30a (anti-miR-30a). b and c Reduced vimentin levels caused by miR-30a are significantly associated with lower migration and invasiveness in Hs578T cells, and these effects are inhibited by introduction of anti-miR-30a. In addition, a dramatic decrease in the migration/invasion of breast cancer cells was detected following knockdown of Vim gene. Representative micrographs of migration/invasion filter membranes after crystal violet staining. d Quantitative analysis of migration/invasion. The values are the mean ± SD for three separate experiments. **p < 0.01, ***p < 0.001
Downregulation of miR-30a is associated with an unfavorable outcome in breast cancer
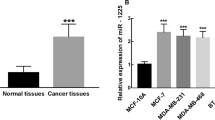
Since miR-30a suppresses tumor cell invasion and metastasis by downregulation of vimentin expression, we then measured miR-30a levels in IDCs. By using the comparative CT method (-ddCt), relative expression levels of the miR-30a compared between tumor and non-tumor cells of 221 patients are shown in Fig. 5a. The results showed that the percentage of patients with reduced miR-30a levels, measured by a greater than twofold decrease in microdissected tumor cells compared to adjacent non-tumor breast cells, was significantly higher in patients with tumors of advanced stage or with LNM (Fig. 5b). We then examined whether decreased miR-30a levels were associated with an unfavorable outcome in IDC patients. In our cohort of breast cancer patients followed up for 5 years, there was a trend toward a decreased recurrence-free survival (Fig. 5c) and decreased disease-free survival (Fig. 5d) in patients with lower miR-30a levels in tumor cells compared with the corresponding non-tumor region. When stage and estrogen receptor status were taken into consideration in the Cox regression model, breast cancer patients with decreased miR-30a levels in the primary cancerous site had an increased HR for recurrence (HR = 3.96; Log-rank p value = 0.005) or recurrence plus death (OR = 1.94; Log-rank p value = 0.01) during the follow-up period. Our study confirms the clinical relevance of our experimental data on miR-30a in breast cancer.
Decreased expression of miR-30a is associated with poor clinical features and decreased survival rates. a Expression levels of miR-30a in tumor as compared with non-tumor in the different stages and LNM in patients with breast IDC. b Percentage of patients showing decreased miR-30a expression levels in tumors, defined as a twofold decrease in expression in the microdissected cancerous tissue (T) compared to adjacent non-cancerous breast epithelium (N), in tumors of different stages or different lymph node metastasis (LNM) status. c Kaplan–Meier analysis of the difference in 5-year breast cancer recurrence-free survival rate, and in d 5-year disease-free survival rate (i.e., breast cancer recurrence plus death) between tumors with high or low miR-30a levels. The hazard ratios (HRs) and 95 % confidence intervals (95 % CIs) shown in (c) and (d) were estimated using the Cox regression model adjusted for the effects of age, tumor stage, and estrogen receptor status
Discussion
In the present study, to unravel the mechanisms by which miRNAs affect breast cancer progression, a combination of miRNA microarray analysis of tumor cells microdissected from the primary tumor site, three matching algorithms, and gene expression data was used to reveal associations between miRNAs and biological functions. Our findings in breast cell models showed that inhibition of the EMT-promoting migration of breast cancer was causally linked to downregulation of vimentin by miR-30a. Further, tumor cells of decreased miR-30a expression with increased metastatic activity was confirmed in breast cancer patients, in whom reduced expression of miR-30a was significantly associated with LNM, advanced stage, and decreased recurrence-free survival and disease-free survival.
Vimentin is required to maintain the architecture of the cytoplasm [34] and aberrant vimentin expression during EMT is suggested to be an essential element for epithelial plasticity and tumor cell metastasis [19, 33, 35]. EMT trans-differentiation processes involve the conversion of adherent epithelial cells into individual migratory cells, leading to changes in cell phenotype into more loose mesenchymal-like cells, and promoting local invasion and metastatic dissemination of tumor cells [33, 36]. miR-30a is involved in hepatobiliary, prefrontal cortex, and renal pronephros development during organogenesis in vertebrates [37–39]. Recently, elevated miR-30a expression has been suggested to inhibit motility of lung cancer cells by decreasing the expression of Snail, a transcriptional regulator that represses E-cadherin expression during MET [40]. In the present study, we showed that the suppressive effect of miR-30a on vimentin expression led to decreased migration and invasiveness of breast cancer cells. Moreover, using the luciferase reporter assay, a significant increase in luciferase activity was seen in cells transfected with the mutated 3′UTR motif of Vim (Fig. 3c), confirming the binding of miR-30a to the vimentin gene.
In miRNA expression assays, the miR-30 family has been frequently found to be downregulated in diverse types of tumors [41, 42]. Interestingly, studies on the genetic loss of chromosome 6q13 hinted at a role for miR-30a in human breast cancer tumorigenesis [43, 44]. qRT-PCR results herein showed that reduced miR-30a mRNA levels in breast cancer cells microdissected from the primary tumor tissue were significantly associated with poor clinical features (advanced stage and LNM) and a worse outcome, consistent with the results of previous cancer cohort studies [45–47]. It is noteworthy that miRNAs bind to specific sequences of target mRNAs to eventually suppress protein translation. It has been reported that protein expression level of a gene is not uniquely modulated by one miRNA; alternatively, the identified miRNAs and their downstream target mRNAs may have different extents of less-than perfect complementarity, which allow one miRNA binding various mRNA transcripts [48, 49]. Similar to the results in vimentin silencing study in this study also, we found that introduction of miR-30a resulted in reduced vimentin expression and decreased migration and invasiveness in a breast tumor cell model. In addition, a good practice to achieve association study is to compare differences of miR-30a levels between paired laser capture microdissected tumor- and non-tumor cells to ensure the validity of this biomarker in reflecting of poor prognosis of breast cancer. More importantly, an investigation of miR-30a expression levels in a group of vimentin-ablated tissues with microdissecting treatment is the best solution to resolve the existence of specific correlation between levels of vimentin and miR-30a regarding in our female breast cancer cohort.
On the basis of databases for miRNA target prediction, members of the miR-30 family have been shown to share the same seed sequence. It has been suggested that downregulation of miR-30 family members decreases metastatic potential; using β-cell development of human fetal pancreatic islets as a model, miR-30d and miR-30a were shown to be involved in downregulating vimentin during EMT of primary pancreatic epithelial cells [50]. In addition, miR-30a and miR-30e share a common seed sequence in the Snail 3′UTR, and reduced expression of Snail is essential for maintaining epithelial-like cells, and, as a result, inhibits the invasiveness and metastasis of non-small cell lung cancers [40]. Recently, it was shown that transforming growth factor-β (TGF-β) acts as a positive regulator of EMT by stimulating normal mammary epithelial cells to adopt mesenchymal- and stem cell-like features and promoting invasion and metastasis [51, 52]. Expression of miR-30 family members is reduced during TGF-β-promoted tumor metastasis, in which increased expressions of invasion/metastasis-associated mesenchymal markers, including N-cadherin, Slug, Snail, and Twist, are seen [51]. Moreover, elevated miR-30c and miR-30a levels may help predict clinical benefit in patients with advanced breast cancer who receive tamoxifen therapy [53]. These results suggest that miR-30 family members may interact synergistically with each other in inhibiting tumor growth and regulating tumor progression. Given the importance of the miR-30 family in predicting progression and outcome of breast cancer, further investigations are required to extend this approach from a single miRNA to multiple miRNAs to understand the cross-talk among miR-30 members in etiological pathways, such as EMT-wide networks.
In conclusion, using microarray analysis of miRNAs expressed in breast cancers with different LNM status, the present study demonstrates, for the first time, a role of miR-30a in inhibiting breast tumor invasiveness and metastasis by directly targeting the vimentin gene. Significantly reduced levels of miR-30a were found in tumors compared with normal mammary ductal epithelium. Moreover, miR-30a expression was inversely correlated with prognosis in patients with IDC, supporting the notion that this miRNA may serve as a tumor biomarker for predicting the outcome of breast cancer. Finally, the finding that miR-30a inhibits vimentin expression may help in developing a potential therapeutic target for treating breast cancer.
Abbreviations
- IDC:
-
Invasive ductal carcinoma
- EMT:
-
Epithelial–mesenchymal transition
- miRNA:
-
microRNA
- 3′UTR:
-
3′-untranslated region
- LNM:
-
Lymph node metastasis
- LCM:
-
Laser capture microdissection
- qRT-PCR:
-
Quantitative real-time reverse transcription polymerase chain reaction
- DFS:
-
Disease-free survival
- OS:
-
Overall survival
- HR:
-
Hazard ratio
- OR:
-
Odds ratio
- 95 % CI:
-
95 % confidence interval
References
Orlando FA, Brown KD (2009) Unraveling breast cancer heterogeneity through transcriptomic and epigenomic analysis. Ann Surg Oncol 16(8):2270–2279
Polyak K (2007) Breast cancer: origins and evolution. J Clin Invest 117(11):3155–3163
Guarino M, Rubino B, Ballabio G (2007) The role of epithelial–mesenchymal transition in cancer pathology. Pathology 39(3):305–318
Micalizzi DS, Farabaugh SM, Ford HL (2010) Epithelial–mesenchymal transition in cancer: parallels between normal development and tumor progression. J Mammary Gland Biol Neoplasia 15(2):117–134
Trimboli AJ, Fukino K, de Bruin A, Wei G, Shen L, Tanner SM, Creasap N, Rosol TJ, Robinson ML, Eng C, Ostrowski MC, Leone G (2008) Direct evidence for epithelial–mesenchymal transitions in breast cancer. Cancer Res 68(3):937–945
Wells A, Yates C, Shepard CR (2008) E-cadherin as an indicator of mesenchymal to epithelial reverting transitions during the metastatic seeding of disseminated carcinomas. Clin Exp Metastasis 25(6):621–628
Marsit CJ, Posner MR, McClean MD, Kelsey KT (2008) Hypermethylation of E-cadherin is an independent predictor of improved survival in head and neck squamous cell carcinoma. Cancer 113(7):1566–1571
Prasad CP, Mirza S, Sharma G, Prashad R, DattaGupta S, Rath G, Ralhan R (2008) Epigenetic alterations of CDH1 and APC genes: relationship with activation of Wnt/beta-catenin pathway in invasive ductal carcinoma of breast. Life Sci 83(9–10):318–325
Yates DR, Rehman I, Abbod MF, Meuth M, Cross SS, Linkens DA, Hamdy FC, Catto JW (2007) Promoter hypermethylation identifies progression risk in bladder cancer. Clin Cancer Res 13(7):2046–2053
Graziano F, Humar B, Guilford P (2003) The role of the E-cadherin gene (CDH1) in diffuse gastric cancer susceptibility: from the laboratory to clinical practice. Ann Oncol 14(12):1705–1713
Braun J, Hoang-Vu C, Dralle H, Huttelmaier S (2010) Downregulation of microRNAs directs the EMT and invasive potential of anaplastic thyroid carcinomas. Oncogene 29(29):4237–4244
Vetter G, Saumet A, Moes M, Vallar L, Le Bechec A, Laurini C, Sabbah M, Arar K, Theillet C, Lecellier CH, Friederich E (2010) miR-661 expression in SNAI1-induced epithelial to mesenchymal transition contributes to breast cancer cell invasion by targeting Nectin-1 and StarD10 messengers. Oncogene 29(31):4436–4448
Cho WC (2007) OncomiRs: the discovery and progress of microRNAs in cancers. Mol Cancer 6:60
Negrini M, Nicoloso MS, Calin GA (2009) MicroRNAs and cancer—new paradigms in molecular oncology. Curr Opin Cell Biol 21(3):470–479
Ortholan C, Puissegur MP, Ilie M, Barbry P, Mari B, Hofman P (2009) MicroRNAs and lung cancer: new oncogenes and tumor suppressors, new prognostic factors and potential therapeutic targets. Curr Med Chem 16(9):1047–1061
Shenouda SK, Alahari SK (2009) MicroRNA function in cancer: oncogene or a tumor suppressor? Cancer Metastasis Rev 28(3–4):369–378
Iwatsuki M, Mimori K, Fukagawa T, Ishii H, Yokobori T, Sasako M, Baba H, Mori M (2010) The clinical significance of vimentin-expressing gastric cancer cells in bone marrow. Ann Surg Oncol 17(9):2526–2533
Mendez MG, Kojima S, Goldman RD (2010) Vimentin induces changes in cell shape, motility, and adhesion during the epithelial to mesenchymal transition. FASEB J 24(6):1838–1851
Usami Y, Satake S, Nakayama F, Matsumoto M, Ohnuma K, Komori T, Semba S, Ito A, Yokozaki H (2008) Snail-associated epithelial–mesenchymal transition promotes oesophageal squamous cell carcinoma motility and progression. J Pathol 215(3):330–339
Yang PS, Yang TL, Liu CL, Wu CW, Shen CY (1997) A case-control study of breast cancer in Taiwan—a low-incidence area. Br J Cancer 75(5):752–756
Lo YL, Yu JC, Huang CS, Tseng SL, Chang TM, Chang KJ, Wu CW, Shen CY (1998) Allelic loss of the BRCA1 and BRCA2 genes and other regions on 17q and 13q in breast cancer among women from Taiwan (area of low incidence but early onset). Int J Cancer 79(6):580–587
Cheng TC, Chen ST, Huang CS, Fu YP, Yu JC, Cheng CW, Wu PE, Shen CY (2005) Breast cancer risk associated with genotype polymorphism of the catechol estrogen-metabolizing genes: a multigenic study on cancer susceptibility. Int J Cancer 113(3):345–353
Ming-Shiean H, Yu JC, Wang HW, Chen ST, Hsiung CN, Ding SL, Wu PE, Shen CY, Cheng CW (2010) Synergistic effects of polymorphisms in DNA repair genes and endogenous estrogen exposure on female breast cancer risk. Ann Surg Oncol 17(3):760–771
Ding SL, Sheu LF, Yu JC, Yang TL, Chen BF, Leu FJ, Shen CY (2004) Abnormality of the DNA double-strand-break checkpoint/repair genes, ATM, BRCA1 and TP53, in breast cancer is related to tumour grade. Br J Cancer 90(10):1995–2001
Hsu HM, Wang HC, Chen ST, Hsu GC, Shen CY, Yu JC (2007) Breast cancer risk is associated with the genes encoding the DNA double-strand break repair Mre11/Rad50/Nbs1 complex. Cancer Epidemiol Biomarkers Prev 16(10):2024–2032
Shen CY, Yu JC, Lo YL, Kuo CH, Yue CT, Jou YS, Huang CS, Lung JC, Wu CW (2000) Genome-wide search for loss of heterozygosity using laser capture microdissected tissue of breast carcinoma: an implication for mutator phenotype and breast cancer pathogenesis. Cancer Res 60(14):3884–3892
Lo YL, Shen CY (2002) Laser capture microdissection in carcinoma analysis. Methods Enzymol 356:137–144
Petroff BK, Phillips TA, Kimler BF, Fabian CJ (2006) Detection of biomarker gene expression by real-time polymerase chain reaction using amplified ribonucleic acids from formalin-fixed random periareolar fine needle aspirates of human breast tissue. Anal Quant Cytol Histol 28(5):297–302
Cheng CW, Yu JC, Wang HW, Huang CS, Shieh JC, Fu YP, Chang CW, Wu PE, Shen CY (2010) The clinical implications of MMP-11 and CK-20 expression in human breast cancer. Clin Chim Acta 411(3–4):234–241
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(9):e45
McInroy L, Maatta A (2007) Down-regulation of vimentin expression inhibits carcinoma cell migration and adhesion. Biochem Biophys Res Commun 360(1):109–114
Satelli A, Li S (2011) Vimentin in cancer and its potential as a molecular target for cancer therapy. Cell Mol Life Sci 68(18):3033–3046
Vuoriluoto K, Haugen H, Kiviluoto S, Mpindi JP, Nevo J, Gjerdrum C, Tiron C, Lorens JB, Ivaska J (2011) Vimentin regulates EMT induction by Slug and oncogenic H-Ras and migration by governing Axl expression in breast cancer. Oncogene 30(12):1436–1448
Franke WW, Grund C, Kuhn C, Jackson BW, Illmensee K (1982) Formation of cytoskeletal elements during mouse embryogenesis. III. Primary mesenchymal cells and the first appearance of vimentin filaments. Differentiation 23(1):43–59
Dutsch-Wicherek M, Lazar A, Tomaszewska R (2010) The potential role of MT and vimentin immunoreactivity in the remodeling of the microenvironment of parotid adenocarcinoma. Cancer Microenviron 4(1):105–113
Sarrio D, Palacios J, Hergueta-Redondo M, Gomez-Lopez G, Cano A, Moreno-Bueno G (2009) Functional characterization of E- and P-cadherin in invasive breast cancer cells. BMC Cancer 9:74
Mellios N, Huang HS, Grigorenko A, Rogaev E, Akbarian S (2008) A set of differentially expressed miRNAs, including miR-30a-5p, act as post-transcriptional inhibitors of BDNF in prefrontal cortex. Hum Mol Genet 17(19):3030–3042
Hand NJ, Master ZR, Eauclaire SF, Weinblatt DE, Matthews RP, Friedman JR (2009) The microRNA-30 family is required for vertebrate hepatobiliary development. Gastroenterology 136(3):1081–1090
Agrawal R, Tran U, Wessely O (2009) The miR-30 miRNA family regulates Xenopus pronephros development and targets the transcription factor Xlim1/Lhx1. Development 136(23):3927–3936
Kumarswamy R, Mudduluru G, Ceppi P, Muppala S, Kozlowski M, Niklinski J, Papotti M, Allgayer H (2012) MicroRNA-30a inhibits epithelial-to-mesenchymal transition by targeting Snai1 and is downregulated in non-small cell lung cancer. Int J Cancer 130(9):2044–2053
Izzotti A, Calin GA, Arrigo P, Steele VE, Croce CM, De Flora S (2009) Downregulation of microRNA expression in the lungs of rats exposed to cigarette smoke. FASEB J 23(3):806–812
Volinia S, Galasso M, Costinean S, Tagliavini L, Gamberoni G, Drusco A, Marchesini J, Mascellani N, Sana ME, Abu Jarour R, Desponts C, Teitell M, Baffa R, Aqeilan R, Iorio MV, Taccioli C, Garzon R, Di Leva G, Fabbri M, Catozzi M, Previati M, Ambs S, Palumbo T, Garofalo M, Veronese A, Bottoni A, Gasparini P, Harris CC, Visone R, Pekarsky Y, de la Chapelle A, Bloomston M, Dillhoff M, Rassenti LZ, Kipps TJ, Huebner K, Pichiorri F, Lenze D, Cairo S, Buendia MA, Pineau P, Dejean A, Zanesi N, Rossi S, Calin GA, Liu CG, Palatini J, Negrini M, Vecchione A, Rosenberg A, Croce CM (2010) Reprogramming of miRNA networks in cancer and leukemia. Genome Res 20(5):589–599
Chappell SA, Walsh T, Walker RA, Shaw JA (1997) Loss of heterozygosity at chromosome 6q in preinvasive and early invasive breast carcinomas. Br J Cancer 75(9):1324–1329
Noviello C, Courjal F, Theillet C (1996) Loss of heterozygosity on the long arm of chromosome 6 in breast cancer: possibly four regions of deletion. Clin Cancer Res 2(9):1601–1606
Heinzelmann J, Henning B, Sanjmyatav J, Posorski N, Steiner T, Wunderlich H, Gajda MR, Junker K (2011) Specific miRNA signatures are associated with metastasis and poor prognosis in clear cell renal cell carcinoma. World J Urol 29(3):367–373
Li X, Zhang Y, Ding J, Wu K, Fan D (2010) Survival prediction of gastric cancer by a seven-microRNA signature. Gut 59(5):579–585
Tan X, Qin W, Zhang L, Hang J, Li B, Zhang C, Wan J, Zhou F, Shao K, Sun Y, Wu J, Zhang X, Qiu B, Li N, Shi S, Feng X, Zhao S, Wang Z, Zhao X, Chen Z, Mitchelson K, Cheng J, Guo Y, He J (2011) A 5-microRNA signature for lung squamous cell carcinoma diagnosis and hsa-miR-31 for prognosis. Clin Cancer Res 17(21):6802–6811
Brennecke J, Stark A, Russell RB, Cohen SM (2005) Principles of microRNA-target recognition. PLoS Biol 3(3):e85
Peter ME (2010) Targeting of mRNAs by multiple miRNAs: the next step. Oncogene 29(15):2161–2164
Ozcan S (2009) MiR-30 family and EMT in human fetal pancreatic islets. Islets 1(3):283–285
Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS, Cheng JQ (2008) MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol Cell Biol 28(22):6773–6784
Wendt MK, Allington TM, Schiemann WP (2009) Mechanisms of the epithelial–mesenchymal transition by TGF-beta. Future Oncol 5(8):1145–1168
Rodriguez-Gonzalez FG, Sieuwerts AM, Smid M, Look MP, Meijer-van Gelder ME, de Weerd V, Sleijfer S, Martens JW, Foekens JA (2011) MicroRNA-30c expression level is an independent predictor of clinical benefit of endocrine therapy in advanced estrogen receptor positive breast cancer. Breast Cancer Res Treat 127(1):43–51
Acknowledgments
We sincerely appreciate Ms. Show-Lin Yang for her assistance in organizing our study specimens. This study was supported by research grant NSC 98-2314-B-040-009-MY3 from the National Science Council, Taipei, Taiwan.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Cheng, CW., Wang, HW., Chang, CW. et al. MicroRNA-30a inhibits cell migration and invasion by downregulating vimentin expression and is a potential prognostic marker in breast cancer. Breast Cancer Res Treat 134, 1081–1093 (2012). https://doi.org/10.1007/s10549-012-2034-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-012-2034-4