Abstract
Metastases are the major cause of cancer-related deaths, but the mechanisms of the metastatic process remain poorly understood. In recent years, the involvement of microRNAs (miRNAs) in cancer has become apparent, and the objective of this study was to identify miRNAs associated with breast cancer progression. Global miRNA expression profiling was performed on 47 tumor samples from 14 patients with paired samples from primary breast tumors and corresponding lymph node and distant metastases using LNA-enhanced miRNA microarrays. The identified miRNA expression alterations were validated by real-time PCR, and tissue distribution of the miRNAs was visualized by in situ hybridization. The patients, in which the miRNA profile of the primary tumor and corresponding distant metastasis clustered in the unsupervised cluster analysis, showed significantly shorter intervals between the diagnosis of the primary tumor and distant metastasis (median 1.6 years) compared to those that did not cluster (median 11.3 years) (p < 0.003). Fifteen miRNAs were identified that were significantly differentially expressed between primary tumors and corresponding distant metastases, including miR-9, miR-219-5p and four of the five members of the miR-200 family involved in epithelial-mesenchymal transition. Tumor expression of miR-9 and miR-200b were confirmed using in situ hybridization, which also verified higher expression of these miRNAs in the distant metastases versus corresponding primary tumors. Our results demonstrate alterations in miRNA expression at different stages of disease progression in breast cancer, and suggest a direct involvement of the miR-200 family and miR-9 in the metastatic process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is one of the leading causes of cancer deaths worldwide and the most common cancer among women [1]. The mortality of breast cancer is primarily caused by metastatic spread to distant organs, rendering the ability to predict, detect, and eliminate metastases one of the most important challenges in breast cancer treatment. Metastases are established through a complex multistep process for which the molecular and regulatory mechanisms are not fully elucidated.
MicroRNAs (miRNAs) are short non-coding single-stranded RNA molecules of approximately 22 nucleotides in length that bind to complementary sequences on target messenger RNA transcripts (mRNAs), usually resulting in translational repression and gene silencing [2]. miRNAs regulate various biological functions in the normal cell, such as cell growth, differentiation, and apoptosis, and they also play critical roles in diseases, including cancer [3]. A single miRNA has the potential to regulate expression of many different target genes, making miRNAs promising therapeutic targets [4]. Several studies have reported amplification or loss of miRNAs in cancer tissue compared to normal, suggesting that miRNAs can function as either “oncogenes” or “tumor suppressor genes” in cancer development [5]. miRNA expression has also been correlated to cancer outcome, making miRNAs promising prognostic markers as well [6]. Furthermore, miRNA expression profiling has been proposed as a strategy to classify tumors of unknown origin, e.g., Lu et al. [6] developed a miRNA classifier based on 217 miRNAs that correctly classified the origin of the majority of poorly differentiated cancers, while a mRNA-based classifier was not able to do so. Newer miRNA classifiers of cancer of unknown origin has also been developed based on less than 50 miRNAs [7, 8].
The role of miRNAs in the development of metastasis in breast cancer is just beginning to be explored, and analyses have primarily been performed on cell lines and by comparison of primary breast cancers with subsequent disease recurrence or the lack thereof. A few miRNAs, such as miR-31 [9] and miR-335 [10] have been identified to be anti-metastatic, while miR-21 [11], miR-373 [12], and miR-520c [12] have been found to be pro-metastatic.
The miR-200 family, which consists of the 5 miRNAs, miR-200a, miR-200b, miR-200c, miR-141, and miR-429, is a group of miRNAs that has been related to disease progression in breast cancer and other types of cancers. These miRNAs cluster on two different chromosomes and are divided into two subgroups by a single nucleotide difference in the seed region [13, 14]. The miR-200 family has been shown to inhibit the epithelial-to-mesenchymal transition (EMT), suggested to be important for one of the initial steps in the metastatic spread of cancer cells [15–17] (reviewed in [18]). During EMT, cells downregulate the cell–cell adhesion molecule E-cadherin, and the miR-200 family has been shown to inhibit the E-cadherin transcriptional repressors ZEB1 and ZEB2 (Zinc finger E-box-binding homeobox), thereby inducing up-regulation of E-cadherin and EMT inhibition [15, 16, 19]. The miR-200 family therefore suppresses migration, and has been found to be down-regulated during EMT [16, 19]. The reversal of EMT, mesenchymal-to-epithelial transition (MET), is considered to be critical for the late stages of the metastatic process, enabling the tumor cells to colonize and grow at distant sites (reviewed in [20]). This suggests that the dynamic ability first to undergo EMT and subsequently MET is an important feature of metastatic cells.
miR-9 has also recently been shown to be associated with metastasis formation in several cancer types, such as breast, colon, ovarian, cervix, liver, and gastric cancer [21–25]. Recent studies have shown that miR-9 promotes metastasis formation [21, 22, 25], however, in contrast, other studies have suggested that increased expression of miR-9 suppresses metastasis formation [23, 24] and that miR-9 inhibits tumor growth through inhibition of NF-kappa B1 [26, 27]. Therefore, miR-9 might have different functions in different cancers.
In this study, we compared global miRNA expression in primary breast cancers with the corresponding local lymph node metastasis and distant metastasis to delineate the changes in miRNA expression in the process of spreading from the primary tumor to distant sites. We identified 15 miRNAs that exhibited significantly altered expression between the primary breast cancer tumor and distant metastasis, and selected miRNAs were further analyzed by quantitative real-time PCR and their expression pattern visualized by in situ hybridization.
Materials and methods
Patients and tissues
Breast cancer patients from whom both the primary and metastatic tumors in either the brain or liver were available as formalin-fixed, paraffin-embedded (FFPE) tissue blocks were identified. When lymph node metastases were available, these tissues were also retrieved. All primary tumors were invasive ductal carcinomas and all samples had been formalin-fixed using the same standard procedure during the entire period of sample collection. The interval between diagnosis of the primary tumor and metastasis varied from 0 to 26.8 years (median 8.1 years). Pathology records were reviewed for each patient and 4 μm sections were cut from each tissue block and stained with hematoxylin and eosin (H&E). The slides were examined by a pathologist to select representative tumor foci and to determine the percentage of tumor content. The blocks containing the highest content of tumor tissue were chosen to a total of 47 that all included tumor tissues with more than 50% cancer cells except three that contained between 35 and 45%. In addition, cores (3 mm) of morphologically normal tissue of breast, lymph node, skin, ovary, and brain, respectively, were extracted from 10 of the tissue blocks of the surgical cancer specimens to be used as controls. The cores were cut into small pieces with a scalpel and used for RNA purification. Pathology and clinical data for each of the patients were obtained from the Pathology Data Bank and the Danish Breast Cancer Cooperative Group (DBCG) databases. The project was approved by the Ethics committee for the Southern Region of Denmark (S-20070061) and the Danish Data Protection Agency (2007–54–0094). Additional patient information is available at Supplemental Materials and methods.
Details of applied methodologies are available at Supplemental Materials and methods. In brief, total RNA was isolated from 5 × 10 μm FFPE tissue using the High Pure miRNA Isolation kit (Roche Applied Science, Indianapolis, IN) and used for miRNA expression profiling using the miRCURY LNA microRNA Arrays version 11.0 Extended Version (Exiqon, Vedbæk, DK). miRNA microarray results were analyzed using an ANOVA statistical test, followed by a post hoc analysis to identify significant differences between tumor sites (p < 0.1). Differences between primary tumors and metastases were analyzed using a Wilcoxon paired sample test (p < 0.01) with Benjamini–Hochberg False Discovery Rate (BH-FDR) correction. The real-time PCR results were analyzed using Student’s t test (p < 0.05).
Results
miRNAs differentially expressed in primary breast cancer versus corresponding distant metastasis as identified by global LNA-based miRNA microarray analysis
To compare the miRNA expression profiles of primary tumors and corresponding distant metastasis, 14 breast cancer patients from whom both the primary tumor and distant metastasis to either liver (n = 5) or brain (n = 9) were available, were identified. Ten of the patients also had lymph node metastases, which were also analyzed (Table S1). For five of the patients, two tissue samples from the primary breast cancer or brain metastasis were analyzed, and for three patients additional metastases other than lymph node, liver or brain were also analyzed (Table S1). Clinical data for the included breast cancer patients are listed in Table 1, as well as the estrogen receptor (ER), progesterone receptor (PgR), and human epidermal growth factor receptor 2 (HER2) status (Table 1). Interestingly, the ER and PgR status of the primary tumors and the distant metastasis was not always identical at the two locations.
Global miRNA expression profiling was performed on a total of 57 samples using the miRCURY LNA™ microRNA Array v. 11.0 Extended Version. Raw data has been deposited in the Gene Expression Omnibus (GEO) database with accession number GPL13703. A principal component analysis of the samples and the common reference channels showed only small variances between the common reference channels, indicating that the observed variances of the tissue samples were likely related to biological differences between the tumor samples and not technical variances (Supplementary Fig. S1). When only the tissue samples were included in the principal component analysis, it was found that the three RNA samples from patient 7 were distinctively different from the other samples. This patient also differed from the others by having an ER- primary tumor. The principal component analysis also showed that four of the included normal tissues also differed from the rest of the samples (Supplementary Fig. S2).
An unsupervised cluster analysis of all cancer samples did not readily separate the samples into groups based on being either primary tumors or distant metastases, but for five of the patients, the primary tumor and corresponding distant metastasis clustered together (Pearson correlation ≥0.25), indicating that these tissues had very similar miRNA expression, as was not the case for the remaining nine patients (Fig. 1a). Seven of 10 lymph node metastasis clustered with their corresponding primary tumor. Interestingly, it was observed that the median interval between the diagnosis of the primary tumor and detection of the distant metastasis was only 1.6 years (range 0–4.8 years) for patients, where the primary tumor and corresponding metastasis clustered together, while it was 11.3 years (range 2.2–26.8 years) for patients where the primary tumor and corresponding metastasis did not cluster (p < 0.003), indicating a correlation between the extent of miRNA similarity between the primary tumor and the corresponding metastasis and time to distant recurrence.
Hierarchical cluster analysis of the global miRNA tumor data. a Unsupervised clustering of the 57 samples based on global miRNA expression analysis including primary tumors, lymph node, and distant metastasis, and morphologically normal tissues from breast cancer patients. For the sample names, the initial number denotes the patient number, while the number following the tissue name indicates whether more than one sample was analyzed from the same tissue. The number in parenthesis denotes if the same sample was applied to two different LNA arrays. N denotes normal tissue samples. b Supervised hierarchical clustering with the significantly differentially-expressed miRNAs (p < 0.1) between each tumor location. Overall, brain and liver metastasis could be separated from lymph node metastasis and breast tumors
To identify miRNA differences dependent on the localization of the tumors, an ANOVA followed by a Post Hoc analysis was applied (p < 0.1). A one-way ANOVA with cancer localization as a factor was performed, and the pair-wise comparison of significant miRNAs between all locations was conducted using a Student–Newman–Keuls Post Hoc test (Table 2). This analysis identified 122 miRNAs that exhibited significantly altered expression between tumor locations. More miRNAs exhibited altered expression between the primary breast tumors and the distant metastases (liver or brain) than between the primary tumor and lymph node metastasis (Table 2). There were also fewer miRNAs exhibiting altered expression between the liver and brain metastases than between the lymph node and liver/brain metastases (Table 2).
When applying the identified differentially-expressed miRNAs in a hierarchical cluster analysis, brain and liver metastasis could be distinguished from lymph nodes and primary breast cancers with very few exceptions (Fig. 1b). The clustering also showed that duplicate samples from the same patient and same site were highly similar, and therefore the data from duplicate samples were pooled. A new ANOVA analysis with additional Post Hoc test was performed with the pooled duplicate samples, again with localization of the tumors as factor. This resulted in 74 miRNAs that were significantly differentially expressed (p < 0.1) between the localizations of cancer, of which 68 were the same as seen in the equivalent ANOVA without pooled duplicates. A hierarchical clustering based on these miRNAs also separated the distant metastasis from the lymph nodes and primary tumors (Supplementary Fig. S3).
To determine whether the miRNAs that differed in the primary breast tumors versus brain metastases were tissue-specific, and thus reflected the surrounding non-malignant tissue rather than the metastases, miRNA expression in two adjacent normal brain tissues was correlated to that of all brain metastases (Supplementary Fig. S4). miRNA expression in the primary tumor was further subtracted to eliminate patient-specific miRNAs. This correlation scatter plot showed that many of the miRNAs up-regulated in the brain metastasis were also up-regulated in the normal tissue, suggesting that these miRNAs could be tissue-specific and related to the surrounding brain tissue. However, it has to be kept in mind that this “normal tissue” was located adjacent to the metastases and therefore might not represent true normal tissue. A few miRNAs exhibited clear expression differences between brain metastasis and normal brain tissue (Supplementary Fig. S4).
To identify the specific miRNAs differentially expressed between primary breast cancer and distant metastasis, three different criterions were chosen. A non-parametric Wilcoxon paired sample test (p < 0.01) with Benjamini–Hochberg False Discovery Rate (FDR) correction [28] identified 89 miRNAs that exhibited significantly altered expression between the primary breast cancers and corresponding distant metastasis. The second criteria was a variance cut-off of 0.3, and the third criteria was a twofold up- or down-regulation between the primary tumor and metastasis in more than one patient. As the lymph node metastases were observed to be very similar to the breast cancer tumors in both the unsupervised cluster analysis and the ANOVA analysis, lymph node metastases were not included.
A total of 15 miRNAs that fulfilled all three analysis criteria were identified. Thirteen of these miRNAs had higher expression in the metastases compared to the primary tumors and included four members of the miR-200 family: hsa-miR-200a, hsa-miR-200b, hsa-miR-200c, and hsa-miR-141. Hsa-miR-9, hsa-miR-219-5p, hsa-miR-1274a (now a tRNA [29]), hsa-miRPlus-E1136, hsa-miRPlus-E1088, hsa-miR-525-pre, hsa-miRPlus-G1248-5p, hsa-miRPlus-G1307-pre, and hsa-miRPlus-G1249-5p were also more highly expressed in the metastases compared to the primary breast cancer tumors, while hsa-miR-202 and hsa-miRPlus-E1133 had lower expression. The log2 ratio of miR-200b, miR-141, miR-9, and miR-219-5p in the metastasis versus primary tumor is shown for each patient in Fig. 2A.
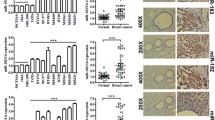
Comparison of miR-200b, miR-141, miR-9, and miR-219-5p expression in primary breast cancers versus corresponding distant metastases (brain, liver, ovary or skin) as determined by a global miRNA array analysis log 2 ratio and b quantitative real-time PCR analysis ∆Ct values. asterisk Indicates p < 0.05. X-axis represents each patient number. _N denotes normal tissue adjacent to the metastasis or primary tumor. Positive values correspond to a higher expression in the distant metastasis versus corresponding primary breast cancer in both analyses
miRNAs differentially expressed in primary breast tumors versus corresponding distant metastasis as analyzed by quantitative PCR analysis
Expression of miR-200b, miR-141, miR-9, and miR-219-5p were further analyzed in the primary breast tumors and corresponding distant metastases for 12 of the 14 patients using real-time PCR. miR-200b was found to be more highly expressed in the metastasis versus the primary tumor in 8 of 12 patients, and 6 of these reached significance (p < 0.05) (Fig. 2b). miR-141 was also more highly expressed in the metastasis versus the primary tumor in 9 of 12 patients, and 3 of these reached significance (p < 0.05) (Fig. 2b). Ten out of 12 patients had higher expression of miR-9 in the metastasis compared to the primary tumor, and 8 of these reached significance (p < 0.05) (Fig. 2b) and finally, 11 of 12 patients had higher miR-219-5p expression in the metastasis versus the primary tumor and 10 of these reached significance (p < 0.05) (Fig. 2b).
E-cadherin and ZEB1 expression in primary breast cancer versus corresponding distant metastasis analyzed by immunohistochemistry
Since we found that the miR-200 family and miR-9 were more highly expressed in the metastasis versus primary tumors, combined with the fact that the miR-200 family is known to inhibit the E-cadherin transcriptional repressors ZEB1 and ZEB2 leading to up-regulation of E-cadherin [15, 16, 19, 30], while miR-9 reportedly inhibits E-cadherin expression [21], we examined the protein expression of E-cadherin and ZEB1 in the primary breast tumors and distant metastases in 12 of the 14 included patients using immunohistochemistry. All samples showed high E-cadherin expression and stronger or equal staining intensity of E-Cadherin was observed in all but one of the metastasis when compared to the corresponding primary breast tumors (Fig. 3d and Supplementary Table S2) likely as a result of diminished repression by members of the miR-200 family, while miR-9 seemed to have a lesser impact on E-cadherin expression. The single patient (patient 7) with lower E-cadherin expression intensity in the metastasis compared to the primary tumor was the same patient that had an ER- primary tumor and in whom the miRNA profiles were significantly different than the remaining patients in the microarray analysis. ZEB1 expression was also examined; however, none of the cancer cells in the sections of the primary breast cancers or metastases exhibited ZEB1 staining, likely due to very low expression levels of ZEB1 in these cells as a result of high E-cadherin expression. ZEB1 staining was seen in some of the stroma cells of the same tissue sections, indicating that the staining procedure was successful.
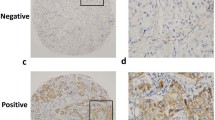
Comparison of miR-200b, miR-9, and E-cadherin expression in primary breast cancers versus corresponding distant metastases reveal higher expression in the metastases. In situ hybridization of miR-200b (a) and miR-9 (b) in primary breast tumor and corresponding metastasis (patient 12 and 16, respectively). Left two columns the original color image (Blue stain in situ hybridization signal, red stain nuclear red counterstain, and purple stain in situ hybridization signal located over nuclear red stain). Right two columns are identical to the left two columns, but the images have undergone color segmentation for clearer visualization (White stain in situ hybridization signal, red stain nucleus, and black stain background structures). Scrambled miRs were used as negative controls. c Normal colon epithelia and normal cerebellum was used as positive control tissue for miR-200b and miR-9 staining, respectively. Arrows indicate some of the positively-stained cells. Magnification (a–c) ×40, scale bar = 100 μm. d E-cadherin expression in primary breast tumor and corresponding distant metastasis of two representative patients (patient 3 and 10). Magnification x40
miR-9 and miR-200b exhibit higher expression in cancer cells of distant metastasis versus corresponding primary breast tumors as determined by in situ hybridization
To determine the cell types within the primary breast cancers and corresponding metastases that expressed miR-9 and miR-200b, these tissues from 9 of the 14 patients were analyzed by in situ hybridization (Table 3). miR-200b was found to be expressed in cancer cells of the distant metastases and not, or to a lesser extent, in cancer cells of the corresponding primary breast tumor in 5 of 9 patients (Table 3 and Fig. 3a), consistent with the higher expression of miR-200b observed in the metastases versus primary breast cancers found in the miRNA microarray analyses. In addition, for 3 patients, 100% of the cancer cells in both the primary breast tumor and the corresponding metastases expressed miR-200b. The staining for miR-200b was primarily seen in the cytoplasm of the cancer cells, as clearly visualized in the color segmentation images (Fig. 3a, right two columns). In the colon tissue used as positive control tissue for miR-200b, the staining for miR-200b was observed in a subpopulation of epithelial cells in the normal colon mucosa (Fig. 3c).
miR-9 expression was observed in cancer cells from 4 patients, and in all of these, expression was higher or was exclusively seen in the metastatic cells (Table 3, Fig. 3b). Interestingly, the staining for miR-9 was primarily seen in the nucleus of the cancer cells, and only to a lesser extent in the cytoplasm, as clearly visualized in the color segmentation images (Fig. 3b, right two columns). Nuclear miR-9 staining was also observed in the normal cerebellum used as positive control tissue for miR-9 (Fig. 3c). The staining was mostly observed in granule cells located in the molecular layer of the cerebellar cortex in close proximity to the granular layer (Fig. 3c).
Discussion
The dissemination of cancer cells from the primary cancer to form metastasis at distant sites is a multistep process that has not yet been fully elucidated. To identify miRNAs involved in the development of distant metastases, global miRNA profiling was performed on primary tumors, corresponding local lymph node metastases and distant metastases on a panel of patients.
The unsupervised hierarchical clustering of the tumor samples based on their global miRNA expression profiles lead to identification of two distinct patients groups, one in which the primary tumors and corresponding distant metastasis of a given patient clustered close together, and another in which primary tumors from multiple patients generally clustered in a group distinct from that of the distant metastasis. To assure that the observed differences were not due to variations in the percentage of tumor tissue in the analyzed samples, we carefully measured the tumor percentage in each sample. No such correlation between tumor % and clustering of primary tumors and metastasis was observed. In general, the tumor percentage was high in all samples (average 65%). These results indicate that larger differences between the patients than between the primary tumors and metastases exist in the former group. Importantly, these two patient groups could be distinguished with respect to time between diagnosis of the primary tumor and the distant metastasis. In the former patient group, the median time to relapse was only 1.6 years, while in the latter it was 11.3 years. This seems to indicate that in the patients in whom the metastasis was detected early, the metastases originated from the dominating subpopulation of the primary tumor, and these metastases have not, in the relatively short time span, developed very distinct features. In contrast, the patients in whom the distant metastasis was diagnosed late showed distinct differences between the primary tumor and corresponding distant metastasis, including miRNA alteration. The miRNA profiles of the local lymph node metastasis generally resembled those of the primary tumor, and since lymph node metastases generally were established at the same time as the diagnosis of the breast cancer, this is in line with our observation that metastases established early were very similar to the primary tumors. The similarity between the primary tumor and the corresponding metastasis has been an issue of intense debate, since it offers insight into how the metastases are established. If metastases are established through clonal selection, then only rare subpopulations in the primary tumor would have the ability to metastasize, meaning that primary tumors and metastases would exhibit very different molecular patterns [31]. On the other hand, global gene expression profiling analyses of primary tumors has revealed “metastatic signatures” expressed by the majority of the cancer cells in primary tumors, implying that the metastatic capacity of the primary tumor is acquired by mutations early during tumorigenesis [32, 33]. In this context, our results primarily indicate that primary tumors and metastases are very similar based on their miRNA expression profiles, but that alterations might occur in the distant metastasis due to mutations over time.
When examining the patient group as a whole, only a few miRNAs showed consistently altered expression between the primary tumor and the corresponding distant metastasis. Since the patient group was relatively heterogenous with respect to ER, PgR, and HER2 status, these potentially metastasis-related miRNAs should be general for invasive ductal breast carcinomas. The identified miRNAs included four of the five miR-200 family members and miR-9, all found to be significantly up-regulated in the distant metastasis versus the corresponding primary tumor. Interestingly, both the miR-200 family and miR-9 has been associated with EMT, a central process associated with metastasis formation. While several studies have shown that the miR-200 family induces up-regulation of E-cadherin and thereby inhibits EMT [15, 16, 19, 30], a recent study by Ma et al. [21] showed that miR-9 has the opposite effect, increasing breast cancer cell motility and invasiveness in vitro and metastasis formation in mice by inhibition of E-cadherin expression. This is quite interesting in relation to our results since we generally found higher expression of E-cadherin in the metastases versus primary tumors, although both miR-9 and the miR-200 family were expressed at higher levels in the metastases compared to primary tumors. This may indicate that the E-cadherin level is primarily influenced by miR-200 in the snapshot from which our samples were obtained, but it does not rule out very important functions of miR-9 on the intermediate stages of this process. Further, it illustrates the complex regulation of miRNAs and the difficulties in examining a dynamic process such as EMT/MET using clinical specimens. The higher level of E-cadherin in the metastases versus primary tumors supports the hypothesis that a MET transition occurs during the establishment of the distant metastasis. These results are in agreement with the study by Kowalski et al. [34] that also found higher E-cadherin expression in metastases versus primary breast tumors of ductal carcinomas. Other studies have reported association between loss of E-cadherin and different prognostic markers such as ER-negative [35] or lymph node-positive breast cancers [36], but in our study such an association was not observed, as high expression of E-cadherin was seen in both the ER-negative and in the lymph node positive samples. The two studies by Kowalski et al. [34] and Younis et al. [36] did not either find an association between the E-cadherin expression and the ER, PgR, or HER2 status.
The fact that miR-9 primarily is seen in the nuclei of both normal cerebellum and metastatic cancer cells is also interesting and is most likely important for the function of this miRNA. Even though miRNAs are usually localized in the cytoplasm other studies have reported specific miRNAs primarily expressed in the nucleus, for example miR-29b [37] and miR-219-5p [38]. Interestingly, the latter was one of the miRNAs that we found to be higher expressed in the metastases than in the primary tumors. These results could imply that the nuclear miRNAs may regulate transcription instead of the ascribed canonical translational regulatory function.
Since miR-9 and miR-219-5p previously have been reported to be selectively expressed in the brain [39–42], and since we also found higher expression in the brain metastases and normal adjacent brain tissue compared to the liver metastases in our microarray study, it was important for us to investigate whether the observed increased expression of miR-9 and miR-219-5p in the metastasis were actually due to cancer cell expression or expression in adjacent normal brain tissue. In situ hybridization confirmed that both miR-9 and miR-200b were expressed in cancer cells and furthermore, generally verified higher expression of miR-9 and miR-200b in the cancer cells within the distant metastasis compared to those within the primary tumors (Fig. 3). This indicates that these miRNAs not only play a role in a specific tissue, but also in the metastastic process. It should be noted that miRNA in situ hybridization is a new technique and the results must be interpreted with some caution due to weak background staining of some of the negative controls.
Collectively, these results show that the miR-200 family and miR-9 might play important roles in the colonization of distant breast cancer metastases, and underscores the need for further investigation of whether, and how, cellular localization of the miRNA affects its function.
References
(IARC) IAfRoC (2010) Globocan 2008. http://globocaniarcfr/factsheets/populations/factsheetasp?uno=900#KEY
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Tsuchiya S, Okuno Y, Tsujimoto G (2006) MicroRNA: biogenetic and functional mechanisms and involvements in cell differentiation and cancer. J Pharmacol Sci 101(4):267–270
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S et al (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65(16):7065–7070
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al (2005) MicroRNA expression profiles classify human cancers. Nature 435(7043):834–838
Rosenfeld N, Aharonov R, Meiri E, Rosenwald S, Spector Y, Zepeniuk M et al (2008) MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol 26(4):462–469
Rosenwald S, Gilad S, Benjamin S, Lebanony D, Dromi N, Faerman A et al (2010) Validation of a microRNA-based qRT-PCR test for accurate identification of tumor tissue origin. Mod Pathol 23(6):814–823
Valastyan S, Reinhardt F, Benaich N, Calogrias D, Szasz AM, Wang ZC et al (2009) A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell 137(6):1032–1046
Tavazoie SF, Alarcon C, Oskarsson T, Padua D, Wang Q, Bos PD et al (2008) Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451(7175):147–152
Zhu S, Wu H, Wu F, Nie D, Sheng S, Mo YY (2008) MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res 18(3):350–359
Huang Q, Gumireddy K, Schrier M, le Sage C, Nagel R, Nair S et al (2008) The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nat Cell Biol 10(2):202–210
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36(Database issue):D154–D158
Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 39(Database issue):D152–D157
Park SM, Gaur AB, Lengyel E, Peter ME (2008) The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 22(7):894–907
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G et al (2008) The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10(5):593–601
Birchmeier C, Birchmeier W, Brand-Saberi B (1996) Epithelial-mesenchymal transitions in cancer progression. Acta Anat 156(3):217–226
Chaffer CL, Weinberg RA (2011) A perspective on cancer cell metastasis. Science 331(6024):1559–1564
Korpal M, Lee ES, Hu G, Kang Y (2008) The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 283(22):14910–14914
Hugo H, Ackland ML, Blick T, Lawrence MG, Clements JA, Williams ED et al (2007) Epithelial–mesenchymal and mesenchymal–epithelial transitions in carcinoma progression. J Cell Physiol 213(2):374–383
Ma L, Young J, Prabhala H, Pan E, Mestdagh P, Muth D et al (2010) miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat Cell Biol 12(3):247–256
Zhu L, Chen H, Zhou D, Li D, Bai R, Zheng S, et al. (2011) MicroRNA-9 up-regulation is involved in colorectal cancer metastasis via promoting cell motility. Med Oncol
Laios A, O’Toole S, Flavin R, Martin C, Kelly L, Ring M et al (2008) Potential role of miR-9 and miR-223 in recurrent ovarian cancer. Mol Cancer 7:35
Hu X, Schwarz JK, Lewis JS Jr, Huettner PC, Rader JS, Deasy JO et al (2010) A microRNA expression signature for cervical cancer prognosis. Cancer Res 70(4):1441–1448
Tan HX, Wang Q, Chen LZ, Huang XH, Chen JS, Fu XH et al (2010) MicroRNA-9 reduces cell invasion and E-cadherin secretion in SK-Hep-1 cell. Med Oncol 27(3):654–660
Guo LM, Pu Y, Han Z, Liu T, Li YX, Liu M et al (2009) MicroRNA-9 inhibits ovarian cancer cell growth through regulation of NF-kappaB1. FEBS J 276(19):5537–5546
Wan HY, Guo LM, Liu T, Liu M, Li X, Tang H (2010) Regulation of the transcription factor NF-kappaB1 by microRNA-9 in human gastric adenocarcinoma. Mol Cancer 9:16
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Statist 57(1):289–300
Schopman NC, Heynen S, Haasnoot J, Berkhout B (2010) A miRNA-tRNA mix-up: tRNA origin of proposed miRNA. RNA Biol 7(5):573–576
Korpal M, Kang Y (2008) The emerging role of miR-200 family of microRNAs in epithelial-mesenchymal transition and cancer metastasis. RNA Biol 5(3):115–119
Fidler IJ, Kripke ML (1977) Metastasis results from preexisting variant cells within a malignant tumor. Science 197(4306):893–895
van ‘t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M et al (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536
van de Vijver MJ, He YD, van’t Veer LJ, Dai H, Hart AA, Voskuil DW et al (2002) A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347(25):1999–2009
Kowalski PJ, Rubin MA, Kleer CG (2003) E-cadherin expression in primary carcinomas of the breast and its distant metastases. Breast Cancer Res 5(6):R217–R222
Nass SJ, Herman JG, Gabrielson E, Iversen PW, Parl FF, Davidson NE et al (2000) Aberrant methylation of the estrogen receptor and E-cadherin 5′ CpG islands increases with malignant progression in human breast cancer. Cancer Res 60(16):4346–4348
Younis LK, El Sakka H, Haque I (2007) The prognostic value of E-cadherin expression in breast cancer. Int J Health Sci 1(1):43–51
Hwang HW, Wentzel EA, Mendell JT (2007) A hexanucleotide element directs microRNA nuclear import. Science 315(5808):97–100
Jeffries CD, Fried HM, Perkins DO (2011) Nuclear and cytoplasmic localization of neural stem cell microRNAs. RNA 17(4):675–686
Nass D, Rosenwald S, Meiri E, Gilad S, Tabibian-Keissar H, Schlosberg A et al (2009) MiR-92b and miR-9/9* are specifically expressed in brain primary tumors and can be used to differentiate primary from metastatic brain tumors. Brain Pathol 19(3):375–383
Sempere LF, Freemantle S, Pitha-Rowe I, Moss E, Dmitrovsky E, Ambros V (2004) Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol 5(3):R13
Krichevsky AM, King KS, Donahue CP, Khrapko K, Kosik KS (2003) A microRNA array reveals extensive regulation of microRNAs during brain development. RNA 9(10):1274–1281
Lukiw WJ (2007) Micro-RNA speciation in fetal, adult and Alzheimer’s disease hippocampus. Neuroreport 18(3):297–300
Acknowledgments
We thank Lisbeth Mortensen and Ole Nielsen for excellent technical assistance with the immunohistochemical analysis and M. K. Occhipinti-Bender for editorial assistance. This study was supported in part by the Danish Cancer Society, the Danish Cancer Research Foundation, A Race Against Breast Cancer, Sino-Danish Breast Cancer Research Centre, NanoCAN Lundbeck Center of Excellence, and Danish Center for Translational Breast Cancer research (DCTB).
Ethical Standards
This manuscript complies with the currents laws in Denmark.
Conflict of interest
BSN is employee at Bioneer A/S. RS and TL are former employees of Exiqon A/S. All other authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gravgaard, K.H., Lyng, M.B., Laenkholm, AV. et al. The miRNA-200 family and miRNA-9 exhibit differential expression in primary versus corresponding metastatic tissue in breast cancer. Breast Cancer Res Treat 134, 207–217 (2012). https://doi.org/10.1007/s10549-012-1969-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-012-1969-9