Abstract
The identification of lobular carcinoma in situ (LCIS) in a patient’s specimen confers an appreciable increased risk of development of future invasive mammary carcinoma. However, the study of LCIS presents a challenge as it is usually only recognized in fixed specimens. Recent advances in high throughput genomics have made possible comprehensive copy number analysis of lesions such as this. Using array comparative genomic hybridization (aCGH), we characterized eight cases of lobular carcinoma (four invasive and four non-invasive) from microdissected samples of archival specimens and validated our results by quantitative real-time PCR (qRT-PCR). Immunohistochemistry (IHC) was performed on an independent set of 80 in situ ductal (DCIS) and lobular breast lesions to confirm our results. Amplification of the prolactin receptor gene (PRLr) was identified in 4/4 cases of LCIS by aCGH. We confirmed this amplification by qRT-PCR and demonstrated PRLr expression in 29/40 (73%) cases of lobular neoplasia by IHC. Amplification of PRLr was neither detected in 10 cases of DCIS nor in 5 areas of normal breast tissue by qRT-PCR and only 14/40 (35%) cases of DCIS showed PRLr expression by IHC (P = 0.0008). Our study suggests the prolactin receptor gene is a molecular target that may be important in the pathogenesis and progression of lobular neoplasia. Investigation of the status of this gene in cases of DCIS has indicated that it may not be as important in the progression of this type of breast cancer, supporting the view that lobular and ductal carcinomas may evolve along separate pathways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pre-invasive breast cancer exists in two forms: ductal carcinoma in situ (DCIS), and lobular carcinoma in situ (LCIS). Both confer an increased risk of future invasive breast cancer. The treatment and management of women diagnosed with LCIS is a subject of ongoing debate. Epidemiological evidence supports that LCIS is a marker of increased risk of developing breast cancer. Invasive carcinomas, subsequent to an initial diagnosis of LCIS, can occur many years later. Although the risk of malignancy was shown to apply to both breasts as noted in earlier studies, the risk appears to be greater in the ipsilateral than the contralateral breast as noted in more recent studies [1, 2]. Furthermore, while the subsequent invasive carcinomas are more likely to be ductal, no special type, than lobular, the proportion of lobular (ILC) is much greater than the rate of ILC in the general population [3, 4]. Cumulative molecular evidence suggests that LCIS may be a premalignant lesion with the capacity to progress toward invasive carcinoma. The strongest indication comes from the studies of E-cadherin (CDH1), a gene often inactivated in invasive lobular carcinoma (ILC) and LCIS, as shown both by immunohistochemical studies [5] and mutation analysis of the gene [6–8]. This loss of E-cadherin is often accompanied by abnormalities in the E-cadherin/catenin complex [9]. In addition, studies using comparative genomic hybridization (CGH) have led to the discovery of a number of changes in lobular carcinomas (reviewed in [10]). However, CGH data from these studies have shown little consensus (with the exception of recurrent loss of 16q and frequent gain of 1q) over the regions of amplification and deletions in these tumors [11–16]. Interestingly, it is of note that the classic changes of lobular lesions (loss of 16q and gain of 1p), and in this regard, the overall numbers of changes in these lesions, resemble those found in the low grade ductal lesions [17].
More recently, array CGH studies have suggested that the relationship between LCIS and synchronous ILC might be clonal. Hwang et al. [13], studying matched pairs of LCIS and ILC, demonstrated similar genomic alterations in a majority of paired specimens. Similar results obtained by other groups [18, 19] further support the hypothesis of a progression pathway. Additional data are, however, required to determine if all the examples of LCIS are precursors to invasive carcinoma and if particular genomic alterations can predict which cases are most likely to invade.
In order to delineate the genetic basis of LCIS, we have conducted a genomewide scan using array CGH on microdissected formalin-fixed, paraffin-embedded (FFPE) tissue samples. We have chosen to study LCIS adjacent to ILC because it is plausible that LCIS adjacent to ILC may have given rise to the invasive carcinoma. This analysis provides detailed profiles of genomic amplifications in this study group and draws attention to a novel observation of the amplification of the prolactin receptor gene (PRLr) on chromosome 5p.
Materials and methods
Sample collection
Eight cases of LCIS with corresponding ILC were identified from the case files of the Department of Pathology at Mount Sinai Hospital (Toronto, Canada). Owing to the small size of the lesions, the amount of tissue available from this study group was limited for our microarray study. Ten cases of DCIS with invasive ductal carcinoma (IDC) and five cases of normal breast from reduction mammoplasty specimens were also selected for PCR validation. A new and independent series of 40 cases of lobular neoplasia (LN) and 40 cases of DCIS was identified from the files of the Department of Pathology at the University Health Network (Toronto, Canada) for immunohistochemical studies. In this series, lobular neoplasia cases were classified either as ALH (atypical lobular hyperplasia), LCIS, or pleomorphic LCIS (PLCIS) according to the proposed criteria [20, 21]. A case was considered as PLCIS if it demonstrated an in situ lesion with high grade cytology ± central necrosis and negative E-cadherin staining [21]. In addition, synchronous lesions when present were also noted from the pathology report (ALH/LCIS with/without ILC, DCIS or IDC). DCIS cases contained ductal lesions only (DCIS ± atypical ductal hyperplasia (ADH), with or without invasive carcinoma or lobular neoplasia) and were graded using the World Health Organization scale [22]. Institutional research ethics board approval was obtained for this study.
DNA preparation
From the FFPE samples, 5-μm sections were cut and prepared for microdissection. LCIS and invasive components were identified by S.J.D. and microdissected using laser capture microdissection on the PixCell II (Arcturus, Mountain View, CA). Normal lymph nodes (also FFPE) were microdissected to be used as reference DNA. Captured cells were incubated in 50 μl of Proteinase K extraction buffer (50 mm Tris, 1 mM EDTA, 0.5% Tween-20) for 72 h at 55°C. Proteinase K was inactivated by incubation at 95°C for 10 min prior to use for PCR.
DOP-PCR
Degenerate oligonucleotide-primed (DOP) PCR was performed according to the protocol of Kuukasjarvi et al. [23] with modifications as proposed by Huang et al. [24]. In brief, 10–100 ng of DNA was amplified by Thermo Sequenase DNA Polymerase (Amersham Biosciences, Canada) in a low stringency pre-amplification step (5 cycles) followed by regular PCR amplification in less stringent conditions. Several DOP-PCR reactions were pooled from each sample (to reduce the effect of bias on each sample) and cleaned by phenol:chloroform extraction before quantification and labeling. The resulting DNA was quantified using a DynaQuant 200 Fluorometer (Amersham Biosciences, Canada).
Array CGH
Array CGH using cDNA microarrays was performed according to the protocol of Pollack et al. [25] with some modifications. In brief, 2–3 μg of pooled DOP-PCR products from each sample as well as an equal amount of reference DNA were labeled by random priming (Bioprime DNA labelling Kit, Invitrogen, Canada) in three separate reactions with either Cy3 or Cy5. (The reference sample used was DNA extracted from the pooled normal lymph nodes that were also FFPE and treated in the same manner as the samples of interest. For example, if the test DNA was DOP-amplified, then the control DNA was also DOP-amplified.) Dye swap experiments were performed on each sample. Labeled products were mixed in appropriate combinations in DIG Easy Hyb (Roche, Canada) hybridization buffer and hybridized for 16–24 h at 37°C to the human 19K arrays from the University Health Network Microarray Centre (Toronto, Canada) which contains 19,008 characterized and unknown human genes and/or ESTs [26, 27]. Slides were rinsed in 0.1× SSC to remove the cover slips and then washed three times in 0.1× SSC/0.1% SDS for 10 min at room temperature. Slides were rinsed once in 1× SSC and twice in 0.1× SSC and centrifuged to dryness. Arrays were scanned using the GenePix 4000A scanner (Axon Instruments, USA). The PMT gain for each laser was adjusted to give an average ratio of Cy3–Cy5 of 1.0 and to minimize the number of saturated pixels. Images were then analyzed using the GenePix Pro 3.0 software (Axon Instruments, USA). The resulting data were normalized using the Bioconductor genomic analysis software package available on line at: http://www.bioconductor.org/. The data were normalized within each subarray as well as for the entire array. The values for the foreground and background for the duplicate spots were first averaged. Then the gene amplification level was calculated as the log base 2 of the ratio of the test to normal values of the differences between foreground and background. The gene amplification level for each “fluor-flip” experiment was averaged. Log2 ratios higher than 0.5 were considered amplified. The differential analysis was performed using Significance Analysis of Microarrays (SAM), which is a software developed at Stanford University Labs by Tusher et al. [28].
Quantitative PCR
Primers were designed using Primer Express Version 1.05 (PE Applied Biosystems, Canada) for the prolactin receptor gene and a control gene (gamma-interferon, γ-IFN). Primer sequences were as follows: for PRLr, forward 5′-GGTCTGGCTGAACTGCAGAAA and reverse 5′-AAGGTGCAAGCAATGAGTGCT; for γ-IFN, forward 5′-CTGGCGACAGTTCAGCCAT and reverse 5′-CCAACGCAAAGCAATACATGA. PCR was performed according to the ABI7700 (PE Applied Biosystems, Canada) protocols using the QuantiTectTM SYBR® Green PCR System (Qiagen, Canada). All the PCR reactions were performed in duplicate, and each duplicate set of reactions was repeated. The comparative threshold cycle (CT) method was used for the calculation of copy number as specified by the manufacturer. Groups were analyzed using Pearson’s chi-square test for proportions and Student’s t-test to compare levels of amplification.
Immunohistochemistry
Antibodies to the prolactin receptor (clone B6.2, Thermo Scientific, Fremont, CA), prolactin (Dako Inc, Canada), signal transducer and activator of transcription-5a (STAT5a) (Invitrogen, Canada), gross cystic disease fluid protein-15 (GCDFP-15) (clone 23A3, Thermo Scientific), E-cadherin (Vector Laboratories, Canada), p120 catenin (clone 98, BD Biosciences, Canada), estrogen receptor (ER) (clone SP1, Thermo Scientific), and HER-2 (clone 4B5, Ventana, Tucson, AZ). were used for the immunohistochemical study. In brief, 5 μm-sections were deparaffinized in xylene and rehydrated in descending alcohols. Endogenous peroxidase activity was blocked by incubation in 3% hydrogen peroxide for 10 min. Antigen retrieval was performed by microwave treatment of sections in citrate buffer at pH 6 (STAT5a, GCDFP-15, E-cadherin, ER, HER-2, prolactin and p120 catenin) or Tris–EDTA buffer at pH 9 (PRLr) for 2 min. Primary antibodies were incubated overnight at the following dilutions: PRLr (1/100), prolactin (1/200), STAT5a (1/50), GCDFP-15 (1/60), ER (1/200), HER-2 (1/50), E-cadherin (1/200), p120 catenin (1/100). The slides were then incubated with the appropriate biotinylated secondary antibody for 45 min, and a detection kit (NovaRed Substrate Kit, Vector Laboratories, Canada) was used for demonstration of the antibodies. Counterstaining was done with hematoxylin. Positive controls were included in each staining series (placental tissue for PRLr [29], normal breast for STAT5a [30] and salivary gland for GCDFP-15 [31]). Immunohistochemistry slides were evaluated independently by two pathologists (D.T.T. and A.D.). For PRLr, prolactin, GCDFP-15, and STAT5a, both the percentage and intensity (scale from 0 to 3+) of tumor cells staining were scored. Cases were considered positive if there is staining of >10% of tumor cells with an intensity of 2+ or more [30, 32]. E-cadherin and p120 catenin scorings were performed as previously described [33]. ER and HER-2 stainings were evaluated according to published criteria [34, 35]. Groups were compared using Pearson’s chi-square test or Fisher’s exact test.
Results
Analysis of microarray data
The CGH microarray analysis was successfully carried out on four of the paired samples using DOP-PCR amplification. In the other cases, insufficient material was obtained for labeling. The data from the microarray experiments in this study have been deposited in the NCBI Gene Expression Omnibus database (GEO; http://www.ncbi.nlm.nih.gov/geo/) and are accessible through GEO Series accession number GSE20474. Although a myriad of genomic alterations identified by CGH have been reported by multiple studies, these reports all agree that lobular carcinomas have a recurrent loss of 16q and frequent gain of 1q [11–14]. The genomic profiles obtained from our array comparative genomic hybridization (aCGH) studies are in line with previously published studies on genomic alterations in lobular carcinoma. Our profiled cases show similar changes with deletion of genes at the 16q location present in 7/8 (88%) samples and amplification of genes at the 1q location in 7/8 (88%) samples. In addition, when we look at the 50 genes with the highest SAM scores in our differential analysis (amplification or deletion), 74% of them were found in regions previously reported to be altered in lobular carcinoma by CGH (see electronic supplementary material—Online Resource 1). We investigated the top 10 genes with the highest SAM scores in the LCIS samples which represented the genes most likely to be amplified in this data set (see electronic supplementary material—Online Resource 2). We were the most interested in the SAM scores for the LCIS data because they would be a subset of genes that may undergo amplification early in the development of lobular neoplasia. These results indicate that not all the genes that are amplified in LCIS are also amplified in the adjacent ILC, with a higher number of significant amplified or deleted genes in the pre-invasive lesions compared to the adjacent invasive carcinoma (see electronic supplementary material—Online Resource 3). Owing to the limited amount of genetic material available for validation by quantitative PCR analysis, we used the following criteria to select a gene for validation: (a) the spot on the array corresponded to a known and/or somewhat characterized gene; (b) the gene showed high levels of amplification in several samples or had an interesting pattern of amplification (i.e., in LCIS only); (c) the gene was located at a chromosomal region known to be amplified in breast cancer and in particular in lobular carcinoma; and (d) the gene had the possibility of being biologically relevant to breast cancer or the progression to invasive carcinoma. Of the top 10 genes that we found to be amplified to different levels in LCIS and ILC, we selected one candidate gene, the prolactin receptor gene (PRLr) for further analysis. Prolactin has been implicated in the progression of breast cancer [36], and thus PRLr appeared a promising candidate for further validation and study.
Quantitative real time PCR validation
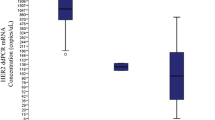
Quantitative real time PCR (qRT-PCR) was performed on the PRLr gene. As qRT-PCR requires a smaller amount of starting material, it was also performed on the remaining cases of LCIS and ILC in the study for which we were not able to obtain microarray data. At least, two independently microdissected samples were used for validation. In addition, qRT-PCR was also performed for PRLr on 10 cases of DCIS with corresponding invasive ductal carcinoma (IDC) and five cases containing normal breast ducts from reduction mammoplasty specimens. As the upper limit of the 95% confidence interval for copy number for all the qRT-PCR results is 1.36, we used this as our threshold for amplification (Table 1). We found the amplification status using qRT-PCR correlated with the data obtained from microarray analysis. Four of the eight samples of LCIS and 4/8 samples of ILC showed amplification of PRLr (all the 6 samples with aCGH amplification of PRLr showed similar results using RT-PCR). In comparison, 0/10 samples of DCIS and 3/10 samples of IDC showed amplification of the gene. When LCIS and ILC are looked at together, there is overall more amplification of PRLr when compared to the DCIS and IDC group (P = 0.01). Interestingly, the levels of amplification between the two groups, lobular and ductal, also differed in the range of copy number values where the ductal group is much lower (0.78–1.58, n = 20) than the lobular group (0.92–3.68, n = 16) (P = 0.03). None of the normal breast specimens showed amplification of PRLr.
Immunohistochemistry validation
An immunohistochemical study was performed on an independent series of 40 cases of LCIS/ALH and 40 cases of DCIS to validate the microarray and qRT-PCR data. The median age at diagnosis for the LCIS and DCIS cases was 54 years (range 35–83 years) and 56 years (range 32–85), respectively. Thirty-nine out of 40 (98%) lobular neoplasia cases were estrogen-receptor (ER)-positive compared to 34/40 (85%) cases of DCIS (P > 0.05) (Table 2). For HER-2, one (3%) case of LCIS was positive compared to five (13%) cases of DCIS (P > 0.05) (Table 2). All the cases of lobular neoplasia showed negative staining for E-cadherin (absence of membrane staining) and diffuse cytoplasmic staining for p120 catenin, while all the DCIS cases demonstrated linear membranous staining for E-cadherin (Fig. 1) and p120 catenin (data not shown).
Histological sections of normal terminal duct-lobular unit, atypical lobular hyperplasia (ALH), lobular carcinoma in situ (LCIS) and ductal carcinoma in situ (DCIS). Cases were stained with hematoxylin and eosin (H&E), or antibodies to E-cadherin, prolactin receptor (PRLr) or signal transducer and activator of transcription-5a (STAT5a), with hematoxylin counterstaining. Representative sections are shown (total magnification at ×200)
The PRLr antibody used for this study (clone B6.2) has been widely employed and validated in multiple reports that studied the expression of PRLr in breast carcinomas and other cancers [29, 37, 38]. Our results demonstrate that 73% (29/40) of lobular lesions express PRLr compared to only 35% (14/40) of ductal lesions (P = 0.0008) (Fig. 1; Tables 2, 3). PRLr expression was mainly localized to the cytoplasm of positive cells, similar to previous reports [32, 39]. Analysis of PRLr staining in adjacent normal ducts/lobules shows at most faint staining that would not meet our criteria for positive staining in the majority of cases, whether ductal or lobular. On occasion, the luminal border of epithelial cells from ducts with columnar cell change and apocrine metaplasia do show some stronger PRLr staining, as previously reported [29, 36]. Expression of PRLr was comparable between the different lobular neoplasia histologic subtypes (ALH, LCIS or pleomorphic LCIS) (Table 3). Similarly, when DCIS lesions were subdivided into low grade and high grade, the difference with lobular lesions was also preserved (Table 3). When we stratified lobular cases according the presence or the absence of synchronous lesions (ILC, DCIS, or IDC), no difference in the rate of PRLr positivity was observed (P = 0.944, Table 4). Of note, cases containing both LCIS/ALH and DCIS on the same section demonstrated positive staining for PRLr in the lobular lesion and negative staining in DCIS in three (out of four) cases, and positive staining in both lobular and ductal lesions in one (out of four) case. Staining for prolactin showed only low positivity rates in both lobular and ductal lesions (3% in both groups, data not shown).
STAT5a is a downstream transcription factor of PRLr activation by prolactin [40]. In all the cases, normal ducts/lobules were positively stained with STAT5a, as reported previously [30]. However, similar to PRLr, STAT5a was expressed in a higher numbers of cases in the lobular group (33/40, 83%) compared to the ductal group (11/40, 28%) (P = 0.0001) (Table 3). GCDFP-15, also known as prolactin-inducible protein (PIP), is a marker, which has been widely used to differentiate breast primary tumors from adenocarcinomas of other organs [41]. Its expression is thought to be regulated by many hormones, including androgen and prolactin [42]. GCDFP-15 staining was observed in 30/40 (75%) of lobular cases compared to 22/40 (55%) cases of DCIS (P = 0.06). Of note, 26/40 (65%) lobular cases showed consensus staining among all the three markers (PRLr, STAT5a, and GCDFP-15, all positive or all negative), while for DCIS cases, there was a 28% (11/40) consensus rate. Neither STAT5a nor GCDFP-15 showed amplification/deletion in our aCGH data.
Although some of our study cases harbor invasive carcinomas in addition to in situ lesions, the number is low (eight and seven cases with evaluable invasive carcinomas in the lobular and ductal groups, respectively). We have scored their staining patterns in each case out of interest (6/8 invasive carcinomas positive for PRLr in the lobular group, 4/7 positive in the ductal group). As the initial aim of our study was to evaluate pre-invasive lobular lesions and as it was not possible to achieve any statistical significance with the number of invasive cases at hand, no attempt at formulating any reliable conclusion was made with these results.
Discussion
In this study, we used array CGH to evaluate copy number changes in LCIS and adjacent ILC in archival tissues. We used SAM analysis to look for amplification patterns in LCIS in an attempt to determine genes that may be involved in the pathogenesis of pre-invasive lobular lesions. Of the top 10 amplified genes in our analysis, we further characterized the prolactin receptor gene (PRLr) based on previous studies in breast cancer. To our knowledge, our results show for the first time the preferential expression of PRLr in pre-invasive lobular lesions compared to pre-invasive ductal lesions of the breast.
The most significant challenge when working with small pre-invasive lesions such as LCIS is obtaining enough material to perform large-scale analysis such as array CGH. The additional problem of only being able to observe LCIS lesions in FFPE material further limits such studies. It is only after fixation that the cellular architecture of these lesions can be seen and diagnosed. This limits the study of these types of lesions to the use of FFPE material that often suffers from sub-optimal quality and quantity of genomic DNA. Further confounding the study of these lesions is their size. Many consist of as few as 50 cells making microdissection necessary to obtain pure populations of cells for analysis. In addition, after the initial identification, the lesion may not be present in all the additional sequential sections cut for microdissection. In previous validation studies (data not shown); test DNA mixed with as little as 25% normal DNA showed a drastic reduction in the ability to detect a known gene amplification. This finding highlights the importance of employing microdissection in the analysis of small pre-invasive breast lesions. Unexpectedly, our analysis suggests that LCIS lesions appear to have more genetic alterations than invasive lobular carcinoma. There are several possible explanations for this phenomenon. It could be that since the LCIS lesions are composed of larger groupings of cells they are easier to microdissect and consequently the genetic profile is not masked by contaminating stromal cells, lymphocytes, etc. Alternatively, pre-invasive lesions may also represent a more heterogeneous population of cells that have acquired different genomic alterations and that only some of these cells may be capable of progressing to invasive disease. A larger scale analysis of pre-invasive lesions compared to their invasive counterparts is needed to address this issue more thoroughly. Interestingly, a previously reported study from Mastracci et al. [12] seemed to demonstrate a similar phenomenon where more genetic aberrations (as determined by CGH) were found in less advanced lesions, namely ALH, compared to the more advanced LCIS lesions.
Among the genes we identified by aCGH, we chose to further characterize the prolactin receptor gene (PRLr) based on previous studies in breast cancer. PRLr maps to 5p13-12 [43] which is a region that has been found to be amplified in lobular carcinoma in previous studies [13, 44, 45] and has been identified as a recurrent high-level amplification region in breast cancer cell lines [46, 47]. Although its role in the development of breast cancer is unclear, evidence implicating prolactin and its receptor in mammary carcinogenesis is mounting (reviewed in [36]). The main signaling pathways activated by binding of prolactin to PRLr include the Jak-STAT, Ras-MAPK and PI3K-Akt pathways [48]. Through activation of Jak-STAT signaling, prolactin can cause induction of genes involved in cell proliferation, such as cyclin D1. Similarly, prolactin-PRLr activation of the Ras–MAPK pathway results in cell proliferation and survival, as has been nicely demonstrated in mammary tumor cell lines [49, 50]. In transgenic mouse models, loss of PRLr delays tumor formation [51] while aberrant induction of prolactin causes mammary tumors [52]. Epidemiologic studies also link elevated levels of serum prolactin and risk of breast cancer [53]. Finally, the prolactin receptor is upregulated in several breast cancer cell lines [47] and higher levels of PRLr mRNA [39] and protein [32] have been found in breast tumors. Interestingly, none of these latter studies has looked at variations in expression between different breast cancer subtypes, and although previous studies have shown high expression of PRLr (up to 95%) in invasive breast carcinomas [32, 39], they have not included a significant number of lobular lesions in their series to permit observation of the differential expression we found in our study.
To our knowledge, we have demonstrated in this study, for the first time, a differential expression of PRLr in pre-invasive lesions of the breast. The gene was found to be amplified by qRT-PCR in 50% (4/8) of LCIS, and protein expression was demonstrated in 73% (29/40) of lobular neoplasia cases by IHC. In DCIS, the prolactin receptor gene does not appear to be amplified (0%, 0/10 cases, P = 0.01) nor highly expressed (35%, 14/40 cases, P = 0.0008). Interestingly, a downstream transcription factor of PRLr activation, STAT5a, is also more highly expressed in lobular lesions compared to ductal lesions (83% vs. 28%, P = 0.001). Similarly, Bratthauer et al. [30], studying STAT5a expression in a variety of breast lesions demonstrated a higher frequency of expression in LCIS compared to DCIS (LCIS 32% (8/25 cases) positive vs. DCIS 4% (1/25 cases) positive). GCDFP-15, also known as prolactin inducible protein (PIP), has been shown, in early in vitro studies, to be regulated by a number of hormones, including prolactin [42]. It is thought that this regulation is mediated via STAT5a [54, 55]. In our series, there was a higher expression rate of GCDFP-15 in lobular compared to ductal in situ lesions (75% vs. 55%), although our numbers did not reach statistical significance. There are only few reports on the rate of GCDFP-15 expression in in situ breast lesions. Expression rates in pre-invasive lobular lesions (including ALH, LCIS, and PLCIS) vary between 71 and 100% [21, 56–58], while those for DCIS, are slightly lower lying between 42 and 70% [56, 59, 60]. Of interest, an early report on GCDFP-15 expression in different breast lesions demonstrated a higher positivity rate in in situ lobular compared to ductal lesions of the breast (80% (8/10) vs. 50% (12/22)) [56].
The grounds behind the preferential expression of PRLr in lobular compared to ductal lesions still remains, however, unresolved. Recently, Simpson et al. [17] proposed the concept of a family of related low nuclear grade lesions, which include lesions such as lobular neoplasia, flat epithelial atypia (FEA), ADH and low-grade DCIS, as precursors of low grade invasive breast carcinomas. Secondary specific alterations during tumorigenesis, such as mutations to E-cadherin (CDH1), would then be required for the development and differentiation of a lobular phenotype. A recent study has in fact shown the amplification of the fibroblast growth factor receptor 1 (FGFR1) gene as another supposedly specific and distinctive molecular change seen in lobular carcinomas [45]. It is, therefore, tempting to speculate that PRLr might also act in a similar fashion, and thereby support the model that the progression of LCIS and DCIS proceeds along different molecular pathways.
With respect to prolactin and the prolactin receptor as a therapeutic agent, a prolactin antagonist, G129R, has been shown to inhibit breast cancer cell proliferation in vitro [61] and slow the growth rate of tumors in mice [62]. More recently, a study has demonstrated the therapeutic efficacy of a fusion protein containing the PRLr antagonist with the angiogenesis inhibitor endostatin [63], and another study has shown an additive effect of G129R to trastuzumab in xenograft models of breast cancer [64].
Developing a molecular profile of breast cancer and its many subtypes is crucial to the development of potential diagnostic, prognostic, and therapeutic targets. Array CGH is a powerful tool, which can vastly increase the amount of information obtained from histologically identified, microdissected lesions. We have utilized this technology in the hope of identifying chromosomal/genetic events that would result in the development of pre-invasive and invasive mammary lobular neoplasia and found evidence for amplification of the prolactin receptor gene in this process.
References
Marshall LM, Hunter DJ, Connolly JL et al (1997) Risk of breast cancer associated with atypical hyperplasia of lobular and ductal types. Cancer Epidemiol Biomarkers Prev 6:297–301
Page DL, Schuyler PA, Dupont WD et al (2003) Atypical lobular hyperplasia as a unilateral predictor of breast cancer risk: a retrospective cohort study. Lancet 361:125–129
Rosen PP, Kosloff C, Lieberman PH et al (1978) Lobular carcinoma in situ of the breast. Detailed analysis of 99 patients with average follow-up of 24 years. Am J Surg Pathol 2:225–251
Garreau JR, Nelson J, Look R et al (2005) Risk counseling and management in patients with lobular carcinoma in situ. Am J Surg 189:610–614; discussion 614–615
De Leeuw WJ, Berx G, Vos CB et al (1997) Simultaneous loss of E-cadherin and catenins in invasive lobular breast cancer and lobular carcinoma in situ. J Pathol 183:404–411
Berx G, Cleton-Jansen AM, Nollet F et al (1995) E-cadherin is a tumour/invasion suppressor gene mutated in human lobular breast cancers. EMBO J 14:6107–6115
Mastracci TL, Tjan S, Bane AL et al (2005) E-cadherin alterations in atypical lobular hyperplasia and lobular carcinoma in situ of the breast. Mod Pathol 18:741–751
Vos CB, Cleton-Jansen AM, Berx G et al (1997) E-cadherin inactivation in lobular carcinoma in situ of the breast: an early event in tumorigenesis. Br J Cancer 76:1131–1133
Dabbs DJ, Kaplai M, Chivukula M et al (2007) The spectrum of morphomolecular abnormalities of the E-cadherin/catenin complex in pleomorphic lobular carcinoma of the breast. Appl Immunohistochem Mol Morphol 15:260–266
Mastracci TL, Boulos FI, Andrulis IL et al (2007) Genomics and premalignant breast lesions: clues to the development and progression of lobular breast cancer. Breast Cancer Res 9:215
Lu YJ, Osin P, Lakhani SR et al (1998) Comparative genomic hybridization analysis of lobular carcinoma in situ and atypical lobular hyperplasia and potential roles for gains and losses of genetic material in breast neoplasia. Cancer Res 58:4721–4727
Mastracci TL, Shadeo A, Colby SM et al (2006) Genomic alterations in lobular neoplasia: a microarray comparative genomic hybridization signature for early neoplastic proliferationin the breast. Genes Chromosom Cancer 45:1007–1017
Hwang ES, Nyante SJ, Yi Chen Y et al (2004) Clonality of lobular carcinoma in situ and synchronous invasive lobular carcinoma. Cancer 100:2562–2572
Etzell JE, Devries S, Chew K et al (2001) Loss of chromosome 16q in lobular carcinoma in situ. Hum Pathol 32:292–296
Weber-Mangal S, Sinn HP, Popp S et al (2003) Breast cancer in young women (< or = 35 years): genomic aberrations detected by comparative genomic hybridization. Int J Cancer 107:583–592
Mohsin SK, O’Connell P, Allred DC et al (2005) Biomarker profile and genetic abnormalities in lobular carcinoma in situ. Breast Cancer Res Treat 90:249–256
Simpson PT, Reis-Filho JS, Gale T et al (2005) Molecular evolution of breast cancer. J Pathol 205:248–254
Morandi L, Marucci G, Foschini MP et al (2006) Genetic similarities and differences between lobular in situ neoplasia (LN) and invasive lobular carcinoma of the breast. Virchows Arch 449:14–23
Aulmann S, Penzel R, Longerich T et al (2008) Clonality of lobular carcinoma in situ (LCIS) and metachronous invasive breast cancer. Breast Cancer Res Treat 107:331–335
Page DL, Dupont WD, Rogers LW et al (1985) Atypical hyperplastic lesions of the female breast. A long-term follow-up study. Cancer 55:2698–2708
Sneige N, Wang J, Baker BA et al (2002) Clinical, histopathologic, and biologic features of pleomorphic lobular (ductal-lobular) carcinoma in situ of the breast: a report of 24 cases. Mod Pathol 15:1044–1050
Tavassoli FA, Devilee P (2003) Intraductal proliferative lesions. In: Anonymous pathology and genetics. Tumours of the breast and female genital organs. IARC Press, Lyon
Kuukasjarvi T, Tanner M, Pennanen S et al (1997) Optimizing DOP-PCR for universal amplification of small DNA samples in comparative genomic hybridization. Genes Chromosom Cancer 18:94–101
Huang Q, Schantz SP, Rao PH et al (2000) Improving degenerate oligonucleotide primed PCR-comparative genomic hybridization for analysis of DNA copy number changes in tumors. Genes Chromosom Cancer 28:395–403
Pollack JR, Perou CM, Alizadeh AA et al (1999) Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nat Genet 23:41–46
Iakovlev VV, Arneson NC, Wong V et al (2008) Genomic differences between pure ductal carcinoma in situ of the breast and that associated with invasive disease: a calibrated aCGH study. Clin Cancer Res 14:4446–4454
Wang C, Iakovlev VV, Wong V et al (2009) Genomic alterations in primary breast cancers compared with their sentinel and more distal lymph node metastases: an aCGH study. Genes Chromosom Cancer 48:1091–1101
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 98:5116–5121
Galsgaard ED, Rasmussen BB, Folkesson CG et al (2009) Re-evaluation of the prolactin receptor expression in human breast cancer. J Endocrinol 201:115–128
Bratthauer GL, Strauss BL, Tavassoli FA (2006) STAT 5a expression in various lesions of the breast. Virchows Arch 448:165–171
Satoh F, Umemura S, Osamura RY (2000) Immunohistochemical analysis of GCDFP-15 and GCDFP-24 in mammary and non-mammary tissue. Breast Cancer 7:49–55
Gill S, Peston D, Vonderhaar BK et al (2001) Expression of prolactin receptors in normal, benign, and malignant breast tissue: an immunohistological study. J Clin Pathol 54:956–960
Dabbs DJ, Bhargava R, Chivukula M (2007) Lobular versus ductal breast neoplasms: the diagnostic utility of p120 catenin. Am J Surg Pathol 31:427–437
Wolff AC, Hammond ME, Schwartz JN et al (2007) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 25:118–145
Hammond ME, Hayes DF, Dowsett M et al (2010) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol 28:2784–2795
Clevenger CV, Furth PA, Hankinson SE et al (2003) The role of prolactin in mammary carcinoma. Endocr Rev 24:1–27
Ferreira M, Mesquita M, Quaresma M et al (2008) Prolactin receptor expression in gynaecomastia and male breast carcinoma. Histopathology 53:56–61
Mertani HC, Garcia-Caballero T, Lambert A et al (1998) Cellular expression of growth hormone and prolactin receptors in human breast disorders. Int J Cancer 79:202–211
Reynolds C, Montone KT, Powell CM et al (1997) Expression of prolactin and its receptor in human breast carcinoma. Endocrinology 138:5555–5560
Nevalainen MT, Xie J, Bubendorf L et al (2002) Basal activation of transcription factor signal transducer and activator of transcription (Stat5) in nonpregnant mouse and human breast epithelium. Mol Endocrinol 16:1108–1124
Bhargava R, Beriwal S, Dabbs DJ (2007) Mammaglobin vs GCDFP-15: an immunohistologic validation survey for sensitivity and specificity. Am J Clin Pathol 127:103–113
Myal Y, Robinson DB, Iwasiow B et al (1991) The prolactin-inducible protein (PIP/GCDFP-15) gene: cloning, structure and regulation. Mol Cell Endocrinol 80:165–175
Arden KC, Boutin JM, Djiane J et al (1990) The receptors for prolactin and growth hormone are localized in the same region of human chromosome 5. Cytogenet Cell Genet 53:161–165
Loveday RL, Greenman J, Simcox DL et al (2000) Genetic changes in breast cancer detected by comparative genomic hybridisation. Int J Cancer 86:494–500
Reis-Filho JS, Simpson PT, Turner NC et al (2006) FGFR1 emerges as a potential therapeutic target for lobular breast carcinomas. Clin Cancer Res 12:6652–6662
Forozan F, Mahlamaki EH, Monni O et al (2000) Comparative genomic hybridization analysis of 38 breast cancer cell lines: a basis for interpreting complementary DNA microarray data. Cancer Res 60:4519–4525
Peirce SK, Chen WY, Chen WY (2001) Quantification of prolactin receptor mRNA in multiple human tissues and cancer cell lines by real time RT-PCR. J Endocrinol 171:R1–R4
Swaminathan G, Varghese B, Fuchs SY (2008) Regulation of prolactin receptor levels and activity in breast cancer. J Mammary Gland Biol Neoplasia 13:81–91
Das R, Vonderhaar BK (1996) Activation of raf-1, MEK, and MAP kinase in prolactin responsive mammary cells. Breast Cancer Res Treat 40:141–149
Perks CM, Keith AJ, Goodhew KL et al (2004) Prolactin acts as a potent survival factor for human breast cancer cell lines. Br J Cancer 91:305–311
Oakes SR, Robertson FG, Kench JG et al (2007) Loss of mammary epithelial prolactin receptor delays tumor formation by reducing cell proliferation in low-grade preinvasive lesions. Oncogene 26:543–553
Rose-Hellekant TA, Arendt LM, Schroeder MD et al (2003) Prolactin induces ERalpha-positive and ERalpha-negative mammary cancer in transgenic mice. Oncogene 22:4664–4674
Tworoger SS, Eliassen AH, Rosner B et al (2004) Plasma prolactin concentrations and risk of postmenopausal breast cancer. Cancer Res 64:6814–6819
Carsol JL, Gingras S, Simard J (2002) Synergistic action of prolactin (PRL) and androgen on PRL-inducible protein gene expression in human breast cancer cells: a unique model for functional cooperation between signal transducer and activator of transcription-5 and androgen receptor. Mol Endocrinol 16:1696–1710
Debily MA, Marhomy SE, Boulanger V et al (2009) A functional and regulatory network associated with PIP expression in human breast cancer. PLoS One 4:e4696
Le Doussal V, Zangerle PF, Collette J et al (1985) Immunohistochemistry of a component protein of the breast cystic disease fluid with mol. wt 15,000. Eur J Cancer Clin Oncol 21:715–725
Middleton LP, Palacios DM, Bryant BR et al (2000) Pleomorphic lobular carcinoma: morphology, immunohistochemistry, and molecular analysis. Am J Surg Pathol 24:1650–1656
Radhi JM (2000) Immunohistochemical analysis of pleomorphic lobular carcinoma: higher expression of p53 and chromogranin and lower expression of ER and PgR. Histopathology 36:156–160
Selim AA, El-Ayat G, Wells CA (2001) Immunohistochemical localization of gross cystic disease fluid protein-15, -24 and -44 in ductal carcinoma in situ of the breast: relationship to the degree of differentiation. Histopathology 39:198–202
Mazoujian G, Bodian C, Haagensen DE Jr et al (1989) Expression of GCDFP-15 in breast carcinomas. Relationship to pathologic and clinical factors. Cancer 63:2156–2161
Chen WY, Ramamoorthy P, Chen N et al (1999) A human prolactin antagonist, hPRL-G129R, inhibits breast cancer cell proliferation through induction of apoptosis. Clin Cancer Res 5:3583–3593
Chen NY, Holle L, Li W et al (2002) In vivo studies of the anti-tumor effects of a human prolactin antagonist, hPRL-G129R. Int J Oncol 20:813–818
Beck MT, Chen NY, Franek KJ et al (2003) Prolactin antagonist-endostatin fusion protein as a targeted dual-functional therapeutic agent for breast cancer. Cancer Res 63:3598–3604
Scotti ML, Langenheim JF, Tomblyn S et al (2008) Additive effects of a prolactin receptor antagonist, G129R, and herceptin on inhibition of HER2-overexpressing breast cancer cells. Breast Cancer Res Treat 111:241–250
Acknowledgments
We would like to thank Dr Frances O’Malley (Mount Sinai Hospital, Toronto) for providing us cases for the microarray analysis. This study was supported by grants from the Dean’s Fund, Faculty of Medicine, University of Toronto and the Streams of Excellence Program of the Canadian Breast Cancer Research Alliance to S.J.D., the Canadian Institutes of Health Research (CIHR) Molecular Pathology Fellowship (STP-53912) and the Ontario Institute for Cancer Research (OICR) grant TSAO-OICR-RF-007 to D.T.T.
Conflict of interest statement
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tran-Thanh, D., Arneson, N.C., Pintilie, M. et al. Amplification of the prolactin receptor gene in mammary lobular neoplasia. Breast Cancer Res Treat 128, 31–40 (2011). https://doi.org/10.1007/s10549-010-1025-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-010-1025-6