Abstract
XRCC3 (X-ray repair complementing defective repair in Chinese hamster cells 3) is a member of the RecA/Rad51-related protein family that participates in homologous recombination, maintaining chromosome stability and participating in DNA repair. Attention has been drawn upon the association of XRCC3 Thr241Met polymorphism with breast cancer risk. The present meta-analysis aims to examine whether XRCC3 Thr241Met polymorphism status is associated with breast cancer risk. Apart from the overall meta-analysis, separate analyses were performed on Chinese and non-Chinese populations, in order to investigate race-specific effects. Eligible articles were identified by a search of MEDLINE bibliographical database for the period up to August 2009. Twenty case–control studies on non-Chinese subjects (19,575 cases and 21,125 controls) and three case–control studies on Chinese subjects (1,216 cases and 1,112 controls) were eligible. Pooled odds ratios (OR) were appropriately derived from fixed-effects or random-effects models. At the overall analysis, the T allele was associated with elevated breast cancer risk mainly following a recessive model (pooled OR = 1.064, 95% CI: 1.007–1.124, fixed effects), given that the effect was more pronounced in homozygous carriers (pooled OR = 1.073, 95% CI: 1.010–1.140, fixed effects). The association seemed confined in non-Chinese populations, once again following a recessive model (pooled OR = 1.072, 95% CI: 1.014–1.133, fixed effects). Concerning Chinese populations, no consistent results were demonstrated. In conclusion, the XRCC3 Thr241Met T allele seems associated with elevated breast cancer risk in non-Chinese subjects. The need for additional studies on Chinese populations seems warranted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
XRCC3 (X-ray repair complementing defective repair in Chinese hamster cells 3) is a member of the RecA/Rad51-related protein family that participates in homologous recombination, maintaining chromosome stability and participating in DNA repair [1]. XRCC3 gene has been found polymorphic in the population; three polymorphisms have been identified: XRCC3 Thr241Met (C to T, rs861539), 5′-UTR A > G (rs1799794), IVS5-14 A > G (rs1799796) [2].
Attention has been mainly drawn at a meta-analytical level upon the association of Thr241Met with breast cancer risk [2–5]; the most recent meta-analysis on the field has reported that the Met allele is associated with elevated breast cancer risk both in Asian and Caucasian populations [3]. Interestingly enough, however, close inspection of the published meta-analyses on Thr241Met reveals the need for a comprehensive approach, as the sizeable studies analyzed by the Breast Cancer Association Consortium [2] have not been included in the most recent meta-analysis by Lee et al. [3].
Under the light of the above, the present meta-analysis aims to examine whether XRCC3 Thr241Met polymorphism status is associated with breast cancer risk. Separate analyses were performed on Chinese and non-Chinese populations, in an attempt to investigate race-specific effects.
Methods
Trial identification
Eligible articles were identified by a search of MEDLINE bibliographical database for the period up to August 2009 (last search: August 31, 2009) using combinations of the following keywords: “XRCC3”, “Thr241Met”, “T241M”, “polymorphism”, “rs861539”, “breast cancer”, “breast”. In addition, we checked all the references of relevant reviews and eligible articles that our search retrieved. Language restrictions were not used and two investigators (KPE and TNS), working independently, searched the literature and extracted data from each eligible case–control study.
Eligible studies and data abstraction
All case–control studies with any sample size examining the association between breast cancer and XRCC3 Thr241Met were considered eligible for this meta-analysis. For each one of the eligible case–control studies, the following data were collected: journal name, year of publication, inclusion and exclusion criteria, demographic characteristics of the population being studied, frequencies of genotypes in cases and controls.
Statistics
Based on the genotype frequencies in cases and controls, crude odds ratios (OR) as well as their standard errors (SE) were calculated. Four different ORs were calculated: (i) CT versus CC (heterozygous carriers), (ii) TT versus CC (homozygous carriers), (iii) T allele carriers (CT and TT grouped together) versus. CC (dominant model) and (iv) TT genotype versus (CT and CC grouped together) (recessive model). Separate analyses were performed in Chinese and non-Chinese populations. In case of zero cells, an appropriate continuity correction (addition of 0.5) was implemented.
The fixed-effects model (Mantel–Haenszel method), as well as the random effects (DerSimonian Laird) model were used to calculate the pooled OR. Between-study heterogeneity and between-study inconsistency were assessed by using Cochran Q statistic and by estimating I 2, respectively [6]. In case no significant heterogeneity was detected, the fixed effects model was chosen. Evidence of publication bias was determined using Egger’s formal statistical test [7] and by visual inspection of the funnel plot. For the interpretation of Egger’s test, statistical significance was defined as P < 0.1. Meta-analysis was performed using the STATA “metan” command.
In addition, meta-regression was performed to assess whether odds ratio (OR) was associated with publication year. The exponentiated coefficient is provided, since the dependent variable in the meta-regression model is log(OR). Meta-regression was performed with the “metareg” STATA command. Analyses were conducted using STATA 10.0 (STATA Corp. College Station, TX, USA).
Results
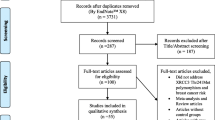
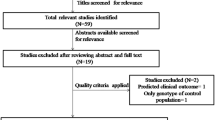
Figure 1 graphically illustrates the trial flow chart. Out of the 55 abstracts retrieved through the search criteria, 25 were irrelevant, four articles [8–11] were excluded because they were conducted on overlapping populations with other eligible studies [2, 3, 5, 12] (these excluded articles represent smaller studies performed on subsets of larger eligible studies), one study [13] was excluded given that it has not included controls in its study design, three articles [4, 14, 15] were reviews/meta-analyses, and three studies [16–18] were excluded due to other reasons (two of them [16, 17] were excluded due to reporting reasons, i.e. no reporting of the relevant genotype frequencies, whereas the other [18] was excluded for examining the association between other XRCC3 polymorphisms and premenopausal breast cancer risk). As a result, 19 case–control articles [2, 3, 5, 12, 19–33] (23 case–control studies, considering that Breast Cancer Association Consortium has more than one studies included) were included in this meta-analysis; 20 case–control studies on non-Chinese subjects (19,575 cases and 21,125 controls) and three case–control studies [3, 24, 29] on Chinese subjects (1,216 cases and 1,112 controls).
Table 1 presents in detail the results of the meta-analysis. At the overall analysis, the T allele was associated with elevated breast cancer risk mainly following a recessive model (pooled OR = 1.064, 95% CI: 1.007–1.124, fixed effects), given that the effect was more pronounced in homozygous carriers (pooled OR = 1.073, 95% CI: 1.010–1.140, fixed effects).
Interestingly enough, stratification by race pointed to discrepancy between non-Chinese and Chinese studies. The association seemed confined in non-Chinese populations, once again following a recessive model (pooled OR = 1.072, 95% CI: 1.014–1.133, fixed effects, Fig. 2). Concerning Chinese populations, no consistent results were found, as the dominant model (pooled OR = 1.102, 95% CI: 0.693–1.949, random effects) and the recessive model (pooled OR = 0.815, 95% CI: 0.580–1.147, fixed effects, Fig. 3) pointed to opposite directions. Interestingly enough, a reverse association, pointing to protective effects of the T allele in homozygous carriers emerged (pooled OR = 0.574, 95% CI: 0.336–0.979, fixed effects); nevertheless, this result should be interpreted with caution given the small number of studies on Chinese populations (n = 3).
Forest plot for the overall association between XRCC3 Thr241Met polymorphism and breast cancer risk for non-Chinese subjects following a recessive model. Each study is shown by the point estimate of the Odds Ratio (OR) (the size of the square is proportional to the weight of each study) and 95% confidence interval for the OR (extending lines); the pooled OR and 95% confidence interval have been appropriately derived from fixed effects model
Meta-regression with publication year did not point to any major modifying effects of publication year (for heterozygous carriers: exponentiated coefficient = 1.037, 95% CI: 0.987–1.089, P = 0.138; for homozygous carriers: exponentiated coefficient = 1.048, 95% CI: 0.986–1.114, P = 0.127; for the recessive model: exponentiated coefficient = 1.041, 95% CI: 0.988–1.096, P = 0.128), apart from a trend of borderline significance in the dominant model (exponentiated coefficient = 1.041, 95% CI: 0.995–1.089, P = 0.080).
No significant publication bias was detected (P = 0.957 for heterozygous carriers, P = 0.116 for homozygous carriers, P = 0.690 for the dominant model, P = 0.334 for the recessive model).
Discussion
The principal message of this meta-analysis is that the T allele of the XRCC3 Thr241Met polymorphism is associated with elevated breast cancer risk in non-Chinese populations; the results are principally compatible with a recessive model. Although controversy exists at a functional level, with studies supporting [34] or not [35] the functioning properties of the Thr241Met polymorphism, this meta-analysis points to significant effects in terms of breast cancer risk.
Having analyzed an almost twofold larger number of studies than the previous meta-analysis [3], our results seem to confirm and establish the positive trend in non-Chinese populations that the data by Lee et al. had indicated. Specifically, under the light of the larger number of studies in the present, formal statistical significance has been reached in the subanalysis of non-Chinese studies; in the most recent meta-analysis [3] solely a trend of borderline significance had emerged. In other words, the accumulation of data has led to adequate power.
More importantly, however, the results of the present meta-analysis are not in accordance with those reported by Lee et al. [3] concerning Chinese populations. Although the latter stated that the (positive association) “trend was shown little stronger in Asian than in Caucasian”, the present meta-analysis does not confirm this observation. According to our data, the results in Chinese populations were inconsistent, with dominant and recessive models pointing to opposite directions; worthy of note, a reverse association in homozygous carriers reached significance. At any case, the association between Thr241Met and breast cancer risk essentially remains an open field in Chinese populations, as the number of studies (n = 3) is considerably smaller than that needed for the achievement of robust conclusions [36].
In conclusion, the XRCC3 Thr241Met T allele seems associated with elevated breast cancer risk in non-Chinese subjects. The need for additional studies on Chinese populations seems warranted, as the results remain inconclusive.
References
Brenneman MA, Weiss AE, Nickoloff JA, Chen DJ (2000) XRCC3 is required for efficient repair of chromosome breaks by homologous recombination. Mutat Res 459:89–97
Breast Cancer Association Consortium (2006) Commonly studied single-nucleotide polymorphisms and breast cancer: results from the Breast Cancer Association Consortium. J Natl Cancer Inst 98:1382–1396
Lee SA, Lee KM, Park SK, Choi JY, Kim B, Nam J, Yoo KY, Noh DY, Ahn SH, Kang D (2007) Genetic polymorphism of XRCC3 Thr241Met and breast cancer risk: case–control study in Korean women and meta-analysis of 12 studies. Breast Cancer Res Treat 103:71–76
Manuguerra M, Saletta F, Karagas MR, Berwick M, Veglia F, Vineis P, Matullo G (2006) XRCC3 and XPD/ERCC2 single nucleotide polymorphisms and the risk of cancer: a HuGE review. Am J Epidemiol 164:297–302
Han J, Hankinson SE, Ranu H, De Vivo I, Hunter DJ (2004) Polymorphisms in DNA double-strand break repair genes and breast cancer risk in the Nurses’ Health Study. Carcinogenesis 25:189–195
Higgins JP, Thompson SG, Deeks JJ, Altman DG (2003) Measuring inconsistency in meta-analyses. BMJ 327:557–560
Egger M, Davey Smith G, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. BMJ 315:629–634
Smith TR, Levine EA, Perrier ND, Miller MS, Freimanis RI, Lohman K, Case LD, Xu J, Mohrenweiser HW, Hu JJ (2003) DNA-repair genetic polymorphisms and breast cancer risk. Cancer Epidemiol Biomarkers Prev 12:1200–1204
Kuschel B, Auranen A, McBride S, Novik KL, Antoniou A, Lipscombe JM, Day NE, Easton DF, Ponder BA, Pharoah PD et al (2002) Variants in DNA double-strand break repair genes and breast cancer susceptibility. Hum Mol Genet 11:1399–1407
Kang D (2003) Genetic polymorphisms and cancer susceptibility of breast cancer in Korean women. J Biochem Mol Biol 36:28–34
Han J, Hankinson SE, Zhang SM, De Vivo I, Hunter DJ (2004) Interaction between genetic variations in DNA repair genes and plasma folate on breast cancer risk. Cancer Epidemiol Biomarkers Prev 13:520–524
Smith TR, Levine EA, Freimanis RI, Akman SA, Allen GO, Hoang KN, Liu-Mares W, Hu JJ (2008) Polygenic model of DNA repair genetic polymorphisms in human breast cancer risk. Carcinogenesis 29:2132–2138
Bewick MA, Conlon MS, Lafrenie RM (2006) Polymorphisms in XRCC1, XRCC3, and CCND1 and survival after treatment for metastatic breast cancer. J Clin Oncol 24:5645–5651
Han S, Zhang HT, Wang Z, Xie Y, Tang R, Mao Y, Li Y (2006) DNA repair gene XRCC3 polymorphisms and cancer risk: a meta-analysis of 48 case-control studies. Eur J Hum Genet 14:1136–1144
Goode EL, Ulrich CM, Potter JD (2002) Polymorphisms in DNA repair genes and associations with cancer risk. Cancer Epidemiol Biomarkers Prev 11:1513–1530
Andreassen CN, Alsner J, Overgaard J, Herskind C, Haviland J, Owen R, Homewood J, Bliss J, Yarnold J (2005) TGFB1 polymorphisms are associated with risk of late normal tissue complications in the breast after radiotherapy for early breast cancer. Radiother Oncol 75:18–21
Synowiec E, Stefanska J, Morawiec Z, Blasiak J, Wozniak K (2008) Association between DNA damage, DNA repair genes variability and clinical characteristics in breast cancer patients. Mutat Res 648:65–72
Han J, Haiman C, Niu T, Guo Q, Cox DG, Willett WC, Hankinson SE, Hunter DJ (2009) Genetic variation in DNA repair pathway genes and premenopausal breast cancer risk. Breast Cancer Res Treat 115:613–622
Jacobsen NR, Nexo BA, Olsen A, Overvad K, Wallin H, Tjonneland A, Vogel U (2003) No association between the DNA repair gene XRCC3 T241M polymorphism and risk of skin cancer and breast cancer. Cancer Epidemiol Biomarkers Prev 12:584–585
Smith TR, Miller MS, Lohman K, Lange EM, Case LD, Mohrenweiser HW, Hu JJ (2003) Polymorphisms of XRCC1 and XRCC3 genes and susceptibility to breast cancer. Cancer Lett 190:183–190
Figueiredo JC, Knight JA, Briollais L, Andrulis IL, Ozcelik H (2004) Polymorphisms XRCC1-R399Q and XRCC3-T241M and the risk of breast cancer at the Ontario site of the Breast Cancer Family Registry. Cancer Epidemiol Biomarkers Prev 13:583–591
Forsti A, Angelini S, Festa F, Sanyal S, Zhang Z, Grzybowska E, Pamula J, Pekala W, Zientek H, Hemminki K et al (2004) Single nucleotide polymorphisms in breast cancer. Oncol Rep 11:917–922
Dufloth RM, Costa S, Schmitt F, Zeferino LC (2005) DNA repair gene polymorphisms and susceptibility to familial breast cancer in a group of patients from Campinas, Brazil. Genet Mol Res 4:771–782
Zhang L, Zhang Z, Yan W (2005) Single nucleotide polymorphisms for DNA repair genes in breast cancer patients. Clin Chim Acta 359:150–155
Millikan RC, Player JS, Decotret AR, Tse CK, Keku T (2005) Polymorphisms in DNA repair genes, medical exposure to ionizing radiation, and breast cancer risk. Cancer Epidemiol Biomarkers Prev 14:2326–2334
Webb PM, Hopper JL, Newman B, Chen X, Kelemen L, Giles GG, Southey MC, Chenevix-Trench G, Spurdle AB (2005) Double-strand break repair gene polymorphisms and risk of breast or ovarian cancer. Cancer Epidemiol Biomarkers Prev 14:319–323
Garcia-Closas M, Egan KM, Newcomb PA, Brinton LA, Titus-Ernstoff L, Chanock S, Welch R, Lissowska J, Peplonska B, Szeszenia-Dabrowska N et al (2006) Polymorphisms in DNA double-strand break repair genes and risk of breast cancer: two population-based studies in USA and Poland, and meta-analyses. Hum Genet 119:376–388
Thyagarajan B, Anderson KE, Folsom AR, Jacobs DR Jr, Lynch CF, Bargaje A, Khaliq W, Gross MD (2006) No association between XRCC1 and XRCC3 gene polymorphisms and breast cancer risk: Iowa Women’s Health Study. Cancer Detect Prev 30:313–321
Sangrajrang S, Schmezer P, Burkholder I, Boffetta P, Brennan P, Woelfelschneider A, Bartsch H, Wiangnon S, Cheisilpa A, Popanda O (2007) The XRCC3 Thr241Met polymorphism and breast cancer risk: a case-control study in a Thai population. Biomarkers 12:523–532
Costa S, Pinto D, Pereira D, Rodrigues H, Cameselle-Teijeiro J, Medeiros R, Schmitt F (2007) DNA repair polymorphisms might contribute differentially on familial and sporadic breast cancer susceptibility: a study on a Portuguese population. Breast Cancer Res Treat 103:209–217
Brooks J, Shore RE, Zeleniuch-Jacquotte A, Currie D, Afanasyeva Y, Koenig KL, Arslan AA, Toniolo P, Wirgin I (2008) Polymorphisms in RAD51, XRCC2, and XRCC3 are not related to breast cancer risk. Cancer Epidemiol Biomarkers Prev 17:1016–1019
Loizidou MA, Michael T, Neuhausen SL, Newbold RF, Marcou Y, Kakouri E, Daniel M, Papadopoulos P, Malas S, Kyriacou K et al (2008) Genetic polymorphisms in the DNA repair genes XRCC1, XRCC2 and XRCC3 and risk of breast cancer in Cyprus. Breast Cancer Res Treat 112:575–579
Krupa R, Synowiec E, Pawlowska E, Morawiec Z, Sobczuk A, Zadrozny M, Wozniak K, Blasiak J (2009) Polymorphism of the homologous recombination repair genes RAD51 and XRCC3 in breast cancer. Exp Mol Pathol 87:32–35
Au WW, Salama SA, Sierra-Torres CH (2003) Functional characterization of polymorphisms in DNA repair genes using cytogenetic challenge assays. Environ Health Perspect 111:1843–1850
Araujo FD, Pierce AJ, Stark JM, Jasin M (2002) Variant XRCC3 implicated in cancer is functional in homology-directed repair of double-strand breaks. Oncogene 21:4176–4180
Higgins JPT, Green S (eds) (2008) Cochrane handbook for systematic reviews of interventions version 5.0.1. The Cochrane collaboration, 2008. Available at www.cochrane-handbook.org. Accessed 29 Aug 2009
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Economopoulos, K.P., Sergentanis, T.N. XRCC3 Thr241Met polymorphism and breast cancer risk: a meta-analysis. Breast Cancer Res Treat 121, 439–443 (2010). https://doi.org/10.1007/s10549-009-0562-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-009-0562-3