Abstract
Overexpression and alternative splicing of CD44 have been implicated in tumour progression. Here we describe the identification of a high level amplification of human 11p13, encompassing the CD44 gene, in primary breast cancers and cell lines and test whether CD44 acts as the driver of this amplicon. aCGH analysis revealed 11p13 amplification in 3% (3/100) of primary breast carcinomas and in two cell lines. The minimal region of amplification was 34.38–37.62 Mb. Amplification was confirmed by dual-colour FISH in these cell lines and further validated by CISH in an independent tumour cohort. CD44 expression in primary breast cancers was significantly associated with features of basal-like breast cancer. Detection of CD44 expression in breast cancer cell lines confirmed moderate to high expression in basal-like cell lines and minimal expression in luminal cell lines. In both, primary breast cancers and cell lines, 11p13 amplification was associated with high levels of CD44 mRNA expression. CD44 alternative splicing was detected in four of nine cell lines and in tumour samples, irrespective of the amplification status. RNAi mediated knock down of CD44 failed to reveal an increased dependence on CD44 expression for proliferation or survival in amplified cell lines. Given that expression of CD44 is not an absolute requirement for the survival of cells harbouring CD44 gene amplification, CD44 is unlikely to be a driver of the 11p13 amplicon.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human CD44 gene locus maps to 11p13 and contains 20 coding exons [1, 2]. Excessive alternative splicing and posttranslational modifications give rise to a family of transmembrane glycoproteins with diverse structural and functional properties. The standard CD44 isoform (CD44s) contains none of the alternatively spliced exons (variant exons 1–10) and is expressed by nearly all cell types. In contrast, the alternatively spliced, variant isoforms (CD44v) show a highly restricted expression pattern during embryonic development, in pathological backgrounds or in a cell type specific manner, for example, epithelial cells mainly express CD44v8-v10.
CD44 has been implicated in many steps of tumour progression: proliferation, migration, invasion and apoptosis resistance, and at least in part these events are mediated by CD44 binding to its major ligand, the extracellular matrix glycosaminoglycan, hyaluronan [1, 2]. Of particular interest are the numerous reports connecting expression of variant CD44 isoforms and development of metastasis. These in vitro and in vivo studies have lead to a number of investigators examining whether there is a clinical correlation linking CD44 expression with patient prognosis and response. These different studies have all reported that a proportion of breast cancers showed elevated expression of CD44, although there are conflicting reports as to whether this expression correlates with particular subtypes of breast cancer and the pattern of variant CD44 isoform expression [3–7]. With respect to patient outcome, again a consistent trend has failed to emerge. For example, it has been reported that there is a significant correlation between CD44s expression and increased disease-free survival in node-negative patients [8] and that expression of CD44v7-8 correlates with decreased overall survival [6]. Similarly it has been reported both that CD44v6 is and is not a poor prognostic marker [6, 9]. The difficulty in these studies is the complexity of the CD44 variant isoform expression patterns and the lack of reagents to accurately monitor expression of the different splice variants. Together with the lack of evidence that CD44 is an independent prognostic marker, further progress in analysing CD44 expression patterns in breast cancer has been limited.
Recent interest in CD44 has stemmed from reports that high levels of CD44 expression in combination with low level expression of CD24 can be used to prospectively identify a population of breast cancer cells enriched in stem cell-like properties [10]. Similarly, expression of CD44+/CD133+/α2β1+ or CD44+/EpCAM+ have been used to identify a colorectal cancer stem cell population [11, 12]. Subsequently a number of studies have demonstrated that the CD44high/CD24low/negative phenotype in cells cultured from primary cancers or expressed in subpopulations of established breast cancer cell lines, identified a population with stem/progenitor-like properties [13–15]. Of particular interest has been the proposal that these CD44+ve subpopulations are mediators of metastasis and that CD44 may act as a metastasis promoting gene.
Gene amplification is one of the underlying genetic mechanisms leading to overexpression of genes in cancer cells when compared to their normal counterparts. Compared to extensive reported data on the function of CD44 in vitro and its expression in vivo, there have been no systematic study on CD44 gene amplification and the potential implications thereof. Here we describe an amplification on 11p13 spanning the CD44 locus in primary breast cancer samples as well as in established breast cancer cell lines and test a potential role for CD44 as an amplicon driver. Amplification of 11p13 correlates with high expression levels of CD44 but had no clear influence on alternative splicing regulation of this gene. Cell lines harbouring the CD44 amplification are not dependent on CD44 expression, arguing for another gene within the amplicon as amplicon driver.
Materials and methods
Cell lines
All cell lines were cultured in the presence of 100 U/ml penicillin, 10 μg/ml streptomycin and 10% foetal bovine serum (FBS), with the exception of SUM149 cells, which were cultured in 5% FBS. Culture conditions were maintained at 37°C and 5% CO2 in a humidified atmosphere. MDA-MB-468, HS578T, MDA-MB-157, MDA-MB-231 and MCF7 cell lines were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Invitrogen, Paisley, UK). BT474 cells were cultured in the American Tissue Culture Collection (ATCC) Hybricare medium (46-X) supplemented with 1.5 g/l sodium bicarbonate. HCC1500 and HCC1954 cells were cultured in the RPMI-1640 medium (Invitrogen). SUM149 cells were cultured in DMEM/Ham’s F12 medium (Invitrogen) supplemented with 5 μg/ml recombinant human insulin (Sigma, Dorset, UK) and 1 μg/ml hydrocortisone (Sigma). For immunofluorescent staining and Western blot analysis, the pan anti-human CD44 monoclonal antibody E1/2.8 [16] which recognises all CD44 splice variants was used.
Sample collection and characterisation
A consecutive series of 104 grade III invasive ductal carcinomas of no special type (GIII-IDC-NST) cases was retrieved from the University Hospital La Paz, Madrid, Spain and the University of Munster, Germany. This project has been approved by the local Research Ethics Committees of the authors’ institutions. Representative 4 μm thick histological sections of each tumour were cut and stained with antibodies against oestrogen receptor (ER), HER2, epidermal growth factor receptor (EGFR) and cytokeratin (Ck) 5/6 as previously described [17, 18].
An independent series of 245 invasive breast carcinomas was arrayed in a tissue microarray (TMA) constructed with replicate 0.6 mm cores. All patients were diagnosed and managed at the Royal Marsden Hospital (RMH) between 1994 and 2000. Patients were selected on the basis of being eligible for therapeutic surgery, being followed up at the RMH, having representative histological blocks in the RMH pathology files, and receiving standard anthracycline-based adjuvant chemotherapy. Adjuvant endocrine therapy was prescribed for patients with ER positive tumours (tamoxifen alone in 96.4% of the patients for the available follow-up period). Complete follow-up was available for 244 patients, ranging from 0.5 to 125 months (median = 67 months, mean = 67 months). Tumours were graded according to a modified Bloom–Richardson scoring system [19] and size was categorised according to the TNM staging criteria [20]. The study was approved by the Royal Marsden Hospital Ethics Committee. 3 μm sections from the TMAs were mounted on polylysine-coated slides and immunohistochemistry was performed with antibodies raised against the following markers: ER, progesterone receptor (PR), HER2, EGFR, Ck 5/6, Ck 14, Ck 17, Cyclin D1, KI67, p53, topoisomerase II α, caveolin 1 (CAV1), caveolin 2 (CAV2) and FOXA1 as reported elsewhere [21]. TMA sections were also subjected to chromogenic in situ hybridisation (CISH) with SpotLight probes for CCND1, MYC, HER2, TOP2A, and chromosome 8 centromere as previously reported [21–23].
Microarray CGH (aCGH)
Five consecutive 8 μm thick frozen sections of GIII-IDC-NST tumours were microdissected as previously described [24, 25] to obtain a percentage of tumour cells in the remaining tissue greater than 90%. The DNA concentration in tumours and cell lines was measured with Picogreen® according to the manufacturer’s instructions (Invitrogen, Paisley, UK) and optimal DNA quality was objectively assessed using a previously described multiplex quality control PCR [26]. Microarray-based comparative genomic hybridisation (aCGH) was successfully performed in 100 consecutive fresh/frozen samples (four cases were excluded due to insufficient DNA quantity) and nine breast cancer cell lines using the 32 K BAC re-array collection (CHORI) tiling path aCGH platform [24]. This type of BAC array platform has been shown to be as robust as and to have comparable resolution with high density oligonucleotide arrays [27–29]. DNA labelling, hybridisations, washes, slide scanning and data acquisition were performed as previously described [24], and data analysed as described in detail elsewhere [18]. Thresholds were applied smoothed Log2 ratios, which were converted into categorical data as previously described [18], with low level gain defined as a smoothed Log2 ratio of between 0.08 and 0.4, corresponding to approximately 3–4 copies of the locus, whilst gene amplification was defined as having a Log2 ratio >0.4, corresponding to more than 5 copies [18, 24, 25]. A detailed analysis of the profiles of the IDC-NSTs is described elsewhere [18]. Tumours were classified into simplex, complex ‘sawtooth’ and complex ‘firestorm’ as previously described [18, 25, 30].
CD44 immunohistochemical, chromogenic in situ hybridisation and fluorescence in situ hybridisation analysis
Immunohistochemistry for CD44 was performed on the TMA cohort using the pan anti-human CD44 monoclonal antibody clone DF1485 (Dako, Cambridgeshire, UK) at a dilution of 1:20. Detection was carried out using the Dako EnVision system with 3,3′-diaminobenzidine as chromogen. Only membranous with or without cytoplasmic reactivity was considered specific. Expression of CD44 was determined using the Allred semi-quantitative scoring system: Intensity of staining was scored as 0 (no staining at high magnification), 1 (weak staining only visible at high magnification), 2 (moderate staining readily visible at low magnification), 3 (strong staining visible at low magnification). The proportion of tumours showing positive staining was scored as follows: 0 (0%), 1 (<1%), 2 (1–10%), 3 (11–33%), 4 (34–66%), 5 (67–100%). Intensity and proportion of staining were added and cases classified as negative (0–2), low expression (3–5) and high expression (6–8). CD44 immunohistochemical analysis was performed by two observers (CM & JRF) blinded to the results of other immunohistochemical markers, CD44 CISH and patients’ outcome.
CISH for CD44 was performed on the TMA cohort utilising in house, biotin labelled probes as previously described [31]. These probes were generated using the BAC clones RP11-30D23 and RP11-790K21, which encompass the region between 34.99 and 35.25 Mb on chromosome 11. The specificity of the probe and its genomic position were confirmed by SP6/T7 end pair sequencing and fluorescence in situ hybridisation (FISH) mapping using normal lymphocyte metaphase spreads as previously described [31]. For analysis, signals were counted in at least 60 non-overlapping nuclei of morphologically unequivocal cells. Cases were considered amplified by CISH if >50% of neoplastic cells harboured large signal clusters or >5 signals/nucleus or large gene clusters, as previously described [22, 32]. CISH analysis was performed by two observers (CM & JRF) blinded to the results of CD44 and other immunohistochemical markers, and patients’ outcome.
FISH analysis was performed on interphase and metaphase chromosomes from breast carcinoma cell lines. A CD44 locus specific FISH probe BAC RP11-79OK21 was labelled with digoxigenin as previously described [31] and detected with anti-digoxigenin-fluorescein (Roche). The biotinylated chromosome 11 centromere was labelled with Cy3-streptavidin (Sigma). Nuclei were counterstained with DAPI (4,6-diamino-2-phenylindole). Metaphase images were captured using a Zeiss Axioplan2 microscope, equipped with Applied Imaging CCD camera and Cytovision software, version 2.81 (Applied Imaging).
RT-PCR and splice variant analysis
Total RNA of cell lines was isolated using the RNeasy Kit (Qiagen, Crawley, UK), and reverse transcribed with the Omniscript RT Kit using oligo dT primers (Qiagen). For identification of CD44 isoforms, variant sequences were amplified with CD44p1 and CD44p2 primers, which are located outside the variant region, as previously described [33]. Amplified sequences were subcloned using the TOPO TA cloning Kit (Invitrogen) and identified by sequencing. RNA from tumour samples was isolated using Trizol according to manufactures instructions (Invitrogen, UK) reverse transcribed and amplified with SuperScript III One step RT-PCR System (Invitrogen) using CD44p1 and CD44p2 primers. Amplified products were subcloned and sequenced as described above.
Quantitative RT-PCR
Reverse transcription of 75 ng total RNA was performed as described above. qRT-PCR was performed with Taqman Gene Expression Assays (Applied Biosystems, Drive Foster City, CA, USA) using human CD44 (Hs00153304_m1), human glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (4310884E) and β-actin (4310881E) probes on an ABI Prism 7900HT sequence detection system (Applied Biosystems, Warrington, UK). Data were collected for each cell line in triplicates for at least two independent RNA isolations. Data were analysed with the SDS 2.2.1 software. Fold change is represented by relative quantification (RQ) units, error bars indicate calculated RQmin and RQmax values, based on a confidence level of 95%.
siRNA treatment, proliferation and survival assays
A single CD44 siRNA 5-GUAUGACACAUAUUGCUUCdT-3 and 5-GAAGCAAUAUGUGUCAUACdT-3 [34] (Dharmacon, Cramlington, UK) was used for transfection of all cell lines with Oligofectamine Reagent (Invitrogen) according to the manufacturers’ instructions at a final concentration of 100 nM. A non-targeting siRNA (On-Target plus control siRNA; Dharmacon) and mock transfections were used as controls. To assess proliferation, cells were seeded 24 h after siRNA transfection into 96 well plates at 2 × 103 (HCC1500: 1 × 104) cells per well in medium containing 0.5 or 10% FCS in triplicates. At different time points, cell number was quantified by CellTiter-Blue (Promega, Southampton, UK). Proliferation was estimated relative to time 0 (48 h after transfection). For survival assays, cells were seeded 24 h after siRNA transfection into 96 well plates at 1 × 104 (HCC 1500: 5 × 104) cells per well in triplicate. 48 h after transfection the medium was aspirated and replaced with medium containing doxorubicin (Sigma). Survival of treated cells relative to untreated cells was quantified using CellTiter-Blue (Promega) 72 h after drug treatment.
Statistical analysis
The Statview version 5 statistical package was used for all statistical analysis of the TMA series. Correlations between categorical variables were performed using the Chi-square test and Fisher’s exact test. Correlations between continuous and categorical variables were performed with the Mann–Whitney U test. Disease-free survival (DFS) was expressed as the number of months from diagnosis to the occurrence of local recurrence in the form of ductal carcinoma in situ and/or invasive breast cancer. Metastasis-free survival (MFS) was expressed as the number of months from diagnosis to the occurrence of distant relapse. Breast cancer specific survival (BCSS) was expressed as the number of months from diagnosis to the occurrence of an event (disease-related death). Cumulative survival probabilities were calculated using the Kaplan–Meier method. Differences between survival rates were tested with the Log-Rank test. Correlations between CD44 mRNA expression levels, 11p13 amplification and molecular subtype were performed using two tailed unpaired t-test. All tests were two-tailed, with a confidence interval of 95%.
Results
Amplification of CD44 in breast cancers
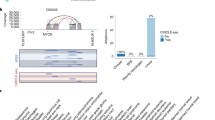
Out of the 100 GIII-IDC-NSTs analysed with aCGH, 25 were classified as basal-like, 25 as HER2, 45 as luminal and 5 of indeterminate phenotype using a previously validated surrogate for the breast cancer molecular subgroups [35]. The detailed analysis of the genetic profiles of these tumours is reported elsewhere [18]. aCGH analysis revealed amplifications of 11p13-p12 (34.38–37.62 Mb) in 3/100 (3%) tumours (Fig. 1a), with the minimal region of amplification mapping to 34.38–37.62 Mb. According to Ensembl release 51 (http://www.ensembl.org), this region incorporated 19 genes ELF5, CAT, EHF, APIP, PDHX, CD44, SLC1A2, AC090625.1, AC090625.5, FJX1, TRIM44, AC090692.9, C11orf55, TRAF6, COMMD9, RAG1, AC009656.11, RAG2 and C17ORF74 (Fig. 1b).
11p12-p13 amplification in primary grade III breast cancers. a Representative chromosome 11 plots of primary grade III invasive ductal carcinomas of no special type with 11p12-p13 amplification (tumours 1064, 1063, 1023). Log2 ratios are plotted on the Y axis against each clone according to genomic location on the X axis. AWS-smoothed ratios are co-plotted (blue). The box highlights the common region of amplification at 11p13. b Detail of the genomic position and genes mapping to the smallest region of amplification of 11p12-p13 obtained from http://www.ensembl.org
In this cohort, amplification at this locus was seen in 3/29 triple negative tumours (ER−, PgR−, HER2−), and 0/71 tumours of non-triple negative phenotype (P = 0.0226, Fishers-exact test). One amplified tumour also displayed positivity for basal marker expression and all had a ‘sawtooth’ pattern of genomic alterations [30]. No other significant associations between amplification of 11p13 and other clinicopathological or immunohistochemical parameters were found. No material from the cohort subjected to aCGH was available for CD44 immunohistochemical assessment.
To determine the prevalence of 11p13 amplification in an independent cohort of breast cancers, we generated CISH probes mapping to the smallest region of amplification, specifically to CD44, and applied these to a TMA containing 245 primary breast cancers. CD44 gene amplification was observed in 3 out of 125 (2.4%) interpretable samples (Fig. 2). All cases harbouring CD44 amplification were of histological grade III; two were invasive ductal carcinomas and one was a pleomorphic lobular carcinoma. In this cohort, we could not confirm the association between triple negative phenotype and CD44 amplification; of the three amplified cases, two were of luminal phenotype (ER+, PR−, HER2−) and one was of basal-like phenotype (ER−, PR−, HER2−, and positive for EGFR and basal keratins). CD44 amplification was not significantly correlated with any clinico-pathological features of the tumours after Bonferroni correction for multiple comparisons. No correlations with disease-free, metastasis-free or overall survival were found in the whole series or when exploratory analysis was performed in ER+, ER−, lymph node positive and lymph node negative subgroups (data not shown). It should be noted, however, that owing to the small sample size and the low prevalence of CD44 amplification, type II or β errors cannot be ruled out.
CD44 gene copy number and protein expression in invasive breast cancer. a, b Invasive ductal carcinoma displaying CD44 gene amplification as defined by chromogenic in situ hybridisation (a), and strong, membranous and cytoplasmic immunohistochemical expression (b). Note the presence of small (a) and large (a—inset) CD44 gene clusters. c, d Invasive ductal carcinoma with one to two CD44 gene copies per nucleus of neoplastic cells (c) displaying low levels of CD44 membranous protein expression (d)
Immunohistochemical analysis demonstrated that all cases with CD44 amplification as determined by CISH displayed high levels of CD44 expression (Table 1; Fig. 2). Within the whole cohort, CD44 expression was significantly associated with lack of ER and PR, lack or low levels of cyclin D1 expression and with expression of EGFR, Ck 5/6, Ck 14 and Ck 17, high proliferation rates as defined by Ki67 (MIB1) and topoisomerase IIα, and p53 nuclear expression. Furthermore, high levels of CD44 expression were preferentially found in tumours of basal-like and triple negative phenotype (Table 1). All cases with CD44 gene amplification expressed high levels of CD44 protein expression. Univariate survival analysis failed to reveal any association between CD44 expression and disease-free, metastasis-free or overall survival (data not shown).
Amplification of CD44 in breast cancer cell lines
Re-analysis of publicly available aCGH data from 50 breast cancer cell lines using a 1 Mb BAC array, revealed the presence of an 11p13 amplification in four basal-like breast cancer cell lines (SUM149, MDA-MB-468, HCC1954 and HCC1500) [36]. By performing aCGH of these cell lines on our 32 K tiling path BAC array platform (resolution of 50 Kb), amplification of this genomic locus was confirmed in SUM149 and MDA-MB-468 cells. HCC1954 cells were found not to be amplified at the CD44 locus but harboured a majority of tetraploid cells. HCC1500 harboured amplification of 11p13 but this amplified region lay proximal (centromeric) to the CD44 gene. These data were further verified by dual colour metaphase FISH analysis (Fig. 3). Multiple copies of CD44 were detected in SUM149 and MDA-MB-468 cells, in the form of clusters of signals indicative of HSR’s (homologously staining regions), whereas no copy number gains or amplification of 11p13 were detected in control basal-like breast cancer cell lines HCC1500 or HCC1954 (Fig. 3) or other breast cancer cells lines examined (HS578T, MCF7, BT474, SKBR3, MDA-MB-231, ZR75-1; data not shown; range of the 11p13:CEP11 ratios 0.81-1).
11p12-p13 amplification in breast cancer cell lines. Representative chromosome 11 plots and metaphase spreads of breast cancer cell lines lacking (HCC1500 and HCC1954) or harbouring (MDA-MB-468 and SUM149) 11p12-p13 amplification. Log2 ratios are plotted on the X axis against each clone according to genomic location on the Y axis. AWS-smoothed ratios are co-plotted (blue). The box highlights the common region of amplification at 11p13. In the metaphase spreads, fluorescence in situ hybridisation (FISH) was performed with probes for chromosome 11 centromere (red) and CD44 (green). A perfect agreement between array CGH and FISH results was observed. Note the presence of multiple CD44 signals in MDA-MB-468 and SUM149 cells
CD44 amplification is associated with high levels of CD44 mRNA expression
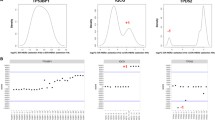
CD44 mRNA levels were assessed by real-time quantitative PCR (qRT-PCR) in a panel of human breast cancer cell lines (Fig. 4a) and correlated with 11p13 gene amplification and the molecular phenotype. Cell lines with 11p13 amplification displayed significantly higher levels of CD44 mRNA expression (3.1480 ± 0.6536) than cell lines lacking amplification of this region (1.0018 ± 0.2978, t-test two-tailed P value = 0.013). CD44 mRNA levels were significantly higher in basal-like (2.0124 ± 0.4475) than in luminal breast cancer cell lines (0.0921 ± 0.0021, t-test two-tailed P value = 0.008). Given the association between CD44 expression and basal-like phenotype, we tested whether the expression of CD44 mRNA would be higher in basal-like cell lines with 11p13 amplification than in cell lines lacking amplification of this region: basal-like cell lines with 11p13 amplification displayed expression levels significantly higher (3.1480 ± 0.6536) than the remaining basal-like cell lines (1.3656 ± 0.2653, t-test two-tailed P value = 0.025). Equivalent results were obtained when CD44 mRNA levels were normalised to GAPDH (data not shown).
CD44 expression in amplified and non-amplified cell lines. a Real time quantitative RT-PCR analysis of CD44 mRNA levels in breast cancer cell lines. Fold increase is shown relative to HCC1954 cells. β-actin served as endogenous control. Black bars, amplified basal-like cell lines; grey bars, non-amplified basal-like cell lines, white bars, non-amplified luminal cell lines. b Cells were fixed and stained for CD44 with E1/2 antibody followed by Alexa488 goat anti-mouse Ig. Nuclei were counterstained with DAPI. Scale bar, 200 μm. c Cells were lysed and subject to Western blot analysis with the E1/2 anti-CD44 antibody which recognises all CD44 isoforms. Blotting for α-tubulin served as loading control. d End point RT-PCR of amplified cell lines and primary breast cancer samples. Isoforms indicated by arrows were identified by sequencing
CD44 protein levels were assessed by Western blotting and immunofluorescence (Fig. 4b–c). As previously reported [14] and in agreement with the mRNA levels, negligible CD44 protein was detected in the luminal breast cancer cell lines, MCF7 and BT474. High levels of CD44 protein were observed in the two CD44 amplified basal-like cell lines, MDA-MB-468 and SUM149, but these levels were not notably higher than in the non-amplified basal-like cell lines HCC1500, MDA-MB-157, MDA-MB-231 and HS578T. Expression of alternatively spliced variant CD44 isoforms was observed in the two amplified cell lines, MDA-MB-468 and SUM149, but also in the non-amplified cells lines, HCC1500 and HCC1954. All other cell lines expressed only the standard CD44 isoform, CD44s, which contains none of the alternatively splice exons. All cell lines expressing variant CD44 isoforms showed decreased expression of the standard CD44s isoform (Fig. 4c).
To characterise the CD44 isoform expression, mRNA from the cell lines was amplified by RT-PCR using primers located outside the variant regions (Fig. 4d). Consistent with the Western blot data (Fig. 4c), sequencing of the RT-PCR products demonstrated that in addition to the standard CD44s isoform, the CD44v8-10 isoform was prominently expressed by the HCC1954, HCC1500, MDA-MB-468 and SUM149 cell lines, with the SUM149 cells also showing expressing of the CD44v3-10 and CD44v6-10 isoforms. As MDA-MB-468 and SUM149 cells, the two cell lines carrying the amplification, showed alternative splicing, we tested the possibility that amplification deregulates the splicing control of CD44. The three primary breast tumours identified by aCGH to harbour amplification of CD44 (Fig. 1) and five control tumours were analysed for expression of splice variants by RT-PCR. All analysed samples showed expression of CD44s and/or CD44v8-10 with no clear differences between tumours carrying the CD44 amplification and those with a non-amplified locus (Fig. 4d).
Cell lines with 11p13 amplification are not dependent on CD44 expression for proliferation or survival
Given that primary tumours and cell lines harbouring 11p13 amplification consistently expressed high levels of CD44 and that genes overexpressed when amplified are potential amplicon drivers, we hypothesised that CD44 would be the driver of 11p13 amplification. To test this hypothesis, we performed siRNA-mediated downregulation of CD44 in a panel of breast cancer cell lines and analysed the proliferative and survival potential. Transfection with CD44 siRNA resulted in a substantial downregulation of CD44 expression as revealed by western blotting and immunohistochemistry (Fig. 5a). Of note, these siRNAs were directed against sequences in the non-variant CD44 backbone and efficiently resulted in downregulation of both standard and alternatively spliced isoforms. Effective downregulation was achieved 2 days after transfection and was stable for 4 days (data not shown). As shown in Fig. 5b, proliferation of MDA-MB-468, SUM149, HCC1954 and MDA-MB-231 cells was unaffected by CD44 downregulation both in high and low serum conditions. Survival upon drug treatment was analysed after 3 days of doxorubicin treatment in the same cell lines. Again, no CD44 dependent differences in susceptibility were observed in the cell lines tested (Fig. 6). Similar results were obtained for all the other cell lines examined (data not shown). Taken together, these results provide strong circumstantial evidence to suggest that CD44 is not the driver of the 11p13 amplicon in breast cancers.
Downregulation of CD44 expression has no differential impact on the proliferation of cell lines with or without the CD44 containing amplicon. Cells were transfected with CD44 siRNA oligonucleotides (CD44si) or non-targeting control siRNA oligonucleotides (CONsi). a Western blot analysis demonstrating efficient downregulation of all CD44 isoforms in all cell lines tested. α-tubulin served as loading control. b Proliferation was analysed for 6 days under high (10%) or low (0.5%) serum conditions and is presented as relative fold increase. The two basal-like cell lines carrying the amplicon (MDA-MB-468, SUM149) and two non-amplified basal-like cell lines (HCC1954, MDA-MB-231) are shown
Downregulation of CD44 expression has no differential impact on the sensitivity to drug treatment of cell lines with or without the CD44 containing amplicon. Cells were transfected with CD44 siRNA (CD44si) or a non-targeting control siRNA (CONsi) oligonucleotides. Survival was monitored after 3 days of doxorubicin treatment at stated concentrations. The two basal-like cell lines carrying the amplicon (MDA-MB-468, SUM149) and two non-amplified basal-like cell lines (HCC1954, MDA-MB-231) are shown
Discussion
The development of high-resolution aCGH platforms have allowed the rapid identification in both tumour tissue and cancer cell lines of regions of chromosomal amplification, harbouring potential oncogenes, and chromosomal deletions, harbouring potential tumour suppressor genes [37]. Here we describe the identification of a subset of breast cancers and cell lines harbouring amplification at 11p13 in which the minimal region of amplification encompasses 19 genes including CD44 (Fig. 1b). Of these the presence of CD44 was intriguing given the wealth of information on a role for CD44 in tumour progression and metastasis in a variety of cancer types [1, 2] and more recently in the identification of CD44 as a marker of cancer stem/progenitor cells [38]. Amplification of 11p12-p13 has been identified in head and neck squamous cell carcinoma and gastric carcinoma cell lines [39, 40] and the presence of this amplification was shown to be associated with significantly increased levels of CD44 expression. Consistent with these observations, here we provide the first direct evidence that CD44 is consistently expressed at high levels in primary breast cancers and breast cancer cell lines harbouring 11p13 amplification. It should be noted, however, that in a way akin to EGFR, CAV1, MYC, CCND1 [32, 36, 41–43], high level expression of CD44 was more pervasive than gene amplification. In agreement with previous studies [14, 44], we detected high levels of CD44 expression, as determined by qRT-PCR and Western blotting, in all basal-like cell lines examined and low or negligible levels of expression in the luminal cell lines. Moreover, by staining a large cohort of invasive breast cancers we found a significant correlation between expression of CD44 and basal-like breast cancers (Table 1). Despite the association with basal-like phenotype, CD44 expression did not show a correlation with the outcome of breast cancer patients.
The mechanisms that regulate CD44 splicing are not fully understood but it is clear that in normal cells, splicing is regulated by tissue specific factors. Cells of mesenchymal origin predominantly express the standard CD44 isoform, CD44s, in which all of the variant exons are spliced out whereas normal epithelial cells frequently show expression of the variant CD44 isoforms, in particular CD44v8-10, CD44v3-10 and CD44v3,v8-10 [2]. It is also well established that the pattern of CD44 splice variant expression is frequently altered in tumour cells, with expression of variant exons v6 and v7 being commonly observed. Consequently we explored in this study whether amplification of the CD44 locus was associated with a different pattern of CD44 isoform expression. It was notable that the two amplified cell lines did indeed express high levels of variant CD44 isoforms as detected by sequencing and Western blotting, however, again this variant expression was not restricted to the amplified cell lines. Similarly, no differences in CD44 splicing were detected between amplified and non-amplified primary breast cancers. It was of interest to note, that the MDA-MB-157, MDA-MB-231 and HS578T cell lines predominantly expressed CD44s and these cell lines are all placed into the mesenchymal group based on expression profiling [44].
A key aspect of genomic analysis is to identify regions of amplification which contain oncogenic drivers and which are therefore potential therapeutic targets. In breast cancer, this is best exemplified by the HER2 (ErbB2) amplification found in 18–20% of breast cancer patients and the clinical use of the HER2 targeted treatments, trastuzumab and lapatinib [45]. However, the identification of a region of genomic amplification poses two major problems. First, it is necessary to identify which of the co-amplified genes is the oncogenic driver. Second, owing to the inherent genomic instability of cancer cells, cancers may also contain somatically acquired genomic amplifications that do not necessarily confer a selective advantage. We and others have demonstrated that drivers of specific amplicons are consistently expressed at high levels in the presence of amplification of their loci and that amplicons often harbour more than one driver [18, 46, 47]. For instance, RNA interference studies have demonstrated that cell lines harbouring 17q12 amplification, are not only dependent on HER2 expression, but also on the expression of GRB7 and STARD3 [47]. Given the longstanding interest in the role of CD44 in tumorigenesis and as a potential cancer stem/progenitor cell marker, together with our observation that CD44 was consistently expressed at high levels when amplified, we examined whether cells harbouring CD44 gene amplification were dependent on CD44 expression for their proliferation and survival. We found no evidence that CD44 expression was required for the proliferation or survival of cells with 11p13 amplification, strongly suggesting that CD44 is not the driver of this amplicon. Further studies will be required to establish whether cell lines and tumours harbouring this region of amplification are dependent on the expression of any of the other 18 genes.
Conclusions
A subset of breast cancers and breast cancer cell lines harbour an amplification at 11p13 encompassing the CD44 gene. CD44 is consistently expressed at high levels in primary cancers and cell lines with CD44 amplification. Unlike many other genes consistently overexpressed when amplified (e.g. HER2, EGFR, PPM1D) [18], CD44 expression is not required for the proliferation or survival of cells with 11p13 amplification. Together these data provide strong circumstantial evidence to suggest that CD44 is unlikely to be the driver of 11p13 amplification.
Abbreviations
- aCGH:
-
Array chromosomal genomic hybridisation
- CISH:
-
Chromogenic in situ hybridisation
- Ck:
-
Cytokeratin
- ER:
-
Oestrogen receptor
- EGFR:
-
Epidermal growth factor receptor
- FISH:
-
Fluorescent in situ hybridisation
- PR:
-
Progesterone receptor
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
References
Naor D, Sionov RV, Ish-Shalom D (1997) CD44: structure, function, and association with the malignant process. Adv Cancer Res 71:241–319. doi:10.1016/S0065-230X(08)60101-3
Ponta H, Sherman L, Herrlich PA (2003) CD44: from adhesion molecules to signalling regulators. Nat Rev Mol Cell Biol 4:33–45. doi:10.1038/nrm1004
Joensuu H, Klemi PJ, Toikkanen S, Jalkanen S (1993) Glycoprotein CD44 expression and its association with survival in breast cancer. Am J Pathol 143:867–874
Auvinen P, Tammi R, Tammi M, Johansson R, Kosma VM (2005) Expression of CD 44s, CD 44 v 3 and CD 44 v 6 in benign and malignant breast lesions: correlation and colocalization with hyaluronan. Histopathology 47:420–428. doi:10.1111/j.1365-2559.2005.02220.x
Berner HS, Nesland JM (2001) Expression of CD44 isoforms in infiltrating lobular carcinoma of the breast. Breast Cancer Res Treat 65:23–29. doi:10.1023/A:1006417412046
Kinoshita J, Haga S, Shimizu T, Imamura H, Watanabe O, Kajiwara T (1999) The expression of variant exon v7–v8 CD44 antigen in relation to lymphatic metastasis of human breast cancer. Breast Cancer Res Treat 53:177–183. doi:10.1023/A:1006130601575
Rys J, Kruczak A, Lackowska B, Jaszcz-Gruchala A, Brandys A, Stelmach A, Reinfuss M (2003) The role of CD44v3 expression in female breast carcinomas. Pol J Pathol 54:243–247
Diaz LK, Zhou X, Wright ET, Cristofanilli M, Smith T, Yang Y, Sneige N, Sahin A, Gilcrease MZ (2005) CD44 expression is associated with increased survival in node-negative invasive breast carcinoma. Clin Cancer Res 11:3309–3314. doi:10.1158/1078-0432.CCR-04-2184
Ma W, Deng Y, Zhou L (2005) The prognostic value of adhesion molecule CD44v6 in women with primary breast carcinoma: a clinicopathologic study. Clin Oncol (R Coll Radiol) 17:258–263. doi:10.1016/j.clon.2005.02.007
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF (2003) Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 100:3983–3988. doi:10.1073/pnas.0530291100
Dalerba P, Dylla SJ, Park IK, Liu R, Wang X, Cho RW, Hoey T, Gurney A, Huang EH, Simeone DM, Shelton AA, Parmiani G, Castelli C, Clarke MF (2007) Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci USA 104:10158–10163. doi:10.1073/pnas.0703478104
Collins AT, Berry PA, Hyde C, Stower MJ, Maitland NJ (2005) Prospective identification of tumorigenic prostate cancer stem cells. Cancer Res 65:10946–10951. doi:10.1158/0008-5472.CAN-05-2018
Grimshaw MJ, Cooper L, Papazisis K, Coleman JA, Bohnenkamp HR, Chiapero-Stanke L, Taylor-Papadimitriou J, Burchell JM (2008) Mammosphere culture of metastatic breast cancer cells enriches for tumorigenic breast cancer cells. Breast Cancer Res 10:R52. doi:10.1186/bcr2106
Sheridan C, Kishimoto H, Fuchs RK, Mehrotra S, Bhat-Nakshatri P, Turner CH, Goulet R Jr, Badve S, Nakshatri H (2006) CD44+/CD24-breast cancer cells exhibit enhanced invasive properties: an early step necessary for metastasis. Breast Cancer Res 8:R59. doi:10.1186/bcr1610
Ponti D, Costa A, Zaffaroni N, Pratesi G, Petrangolini G, Coradini D, Pilotti S, Pierotti MA, Daidone MG (2005) Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties. Cancer Res 65:5506–5511. doi:10.1158/0008-5472.CAN-05-0626
Isacke CM, Sauvage CA, Hyman R, Lesley J, Schulte R, Trowbridge IS (1986) Identification and characterization of the human Pgp-1 glycoprotein. Immunogenetics 23:326–332. doi:10.1007/BF00398797
Korsching E, Packeisen J, Agelopoulos K, Eisenacher M, Voss R, Isola J, van Diest PJ, Brandt B, Boecker W, Buerger H (2002) Cytogenetic alterations and cytokeratin expression patterns in breast cancer: integrating a new model of breast differentiation into cytogenetic pathways of breast carcinogenesis. Lab Invest 82:1525–1533
Natrajan R, Lambros MB, Rodriguez-Pinilla SM, Moreno-Bueno G, Tan DSP, Marchio C, Vatcheva R, Rayter S, Mahler-Araujo B, Fulford LG, Hungermann D, Mackay A, Grigoriadis A, Fenwick K, Tamber N, Hardisson D, Tutt A, Palacios J, Lord CJ, Buerger H, Ashworth A, Reis-Filho JS (2009) Tiling path genomic profiling of grade III invasive ductal breast cancers. Clin Cancer Res 15(8). doi: 10.1158/1078-0432.CCR-08-1878
Ellis IO, Galea M, Broughton N, Locker A, Blamey RW, Elston CW (1992) Pathological prognostic factors in breast cancer. II. Histological type. Relationship with survival in a large study with long-term follow-up. Histopathology 20:479–489
Singletary SE, Connolly JL (2006) Breast cancer staging: working with the sixth edition of the AJCC cancer staging manual. CA Cancer J Clin 56:37–47. doi:10.3322/canjclin.56.1.37 quiz 50-31
Tan DS, Marchio C, Jones RL, Savage K, Smith IE, Dowsett M, Reis-Filho JS (2008) Triple negative breast cancer: molecular profiling and prognostic impact in adjuvant anthracycline-treated patients. Breast Cancer Res Treat 111:27–44. doi:10.1007/s10549-007-9756-8
Bhargava R, Gerald WL, Li AR, Pan Q, Lal P, Ladanyi M, Chen B (2005) EGFR gene amplification in breast cancer: correlation with epidermal growth factor receptor mRNA and protein expression and HER-2 status and absence of EGFR-activating mutations. Mod Pathol 18:1027–1033. doi:10.1038/modpathol.3800438
Reis-Filho JS, Milanezi F, Carvalho S, Simpson PT, Steele D, Savage K, Lambros MB, Pereira EM, Nesland JM, Lakhani SR, Schmitt FC (2005) Metaplastic breast carcinomas exhibit EGFR, but not HER2, gene amplification and overexpression: immunohistochemical and chromogenic in situ hybridization analysis. Breast Cancer Res 7:R1028–R1035. doi:10.1186/bcr1341
Marchio C, Iravani M, Natrajan R, Lambros MB, Savage K, Tamber N, Fenwick K, Mackay A, Senetta R, Di Palma S, Schmitt FC, Bussolati G, Ellis LO, Ashworth A, Sapino A, Reis-Filho JS (2008) Genomic and immunophenotypical characterization of pure micropapillary carcinomas of the breast. J Pathol 215:398–410. doi:10.1002/path.2368
Marchio C, Natrajan R, Shiu KK, Lambros MB, Rodriguez-Pinilla SM, Tan DS, Lord CJ, Hungermann D, Fenwick K, Tamber N, Mackay A, Palacios J, Sapino A, Buerger H, Ashworth A, Reis-Filho JS (2008) The genomic profile of HER2-amplified breast cancers: the influence of ER status. J Pathol 216:399–407. doi:10.1002/path.2423
Arriola E, Marchio C, Tan DS, Drury SC, Lambros MB, Natrajan R, Rodriguez-Pinilla SM, Mackay A, Tamber N, Fenwick K, Jones C, Dowsett M, Ashworth A, Reis-Filho JS (2008) Genomic analysis of the HER2/TOP2A amplicon in breast cancer and breast cancer cell lines. Lab Invest 88:491–503. doi:10.1038/labinvest.2008.19
Coe BP, Ylstra B, Carvalho B, Meijer GA, Macaulay C, Lam WL (2007) Resolving the resolution of array CGH. Genomics 89:647–653. doi:10.1016/j.ygeno.2006.12.012
Tan DS, Lambros MB, Natrajan R, Reis-Filho JS (2007) Getting it right: designing microarray (and not ‘microawry’) comparative genomic hybridization studies for cancer research. Lab Invest 87:737–754. doi:10.1038/labinvest.3700593
Gunnarsson R, Staaf J, Jansson M, Ottesen AM, Goransson H, Liljedahl U, Ralfkiaer U, Mansouri M, Buhl AM, Smedby KE, Hjalgrim H, Syvanen AC, Borg A, Isaksson A, Jurlander J, Juliusson G, Rosenquist R (2008) Screening for copy-number alterations and loss of heterozygosity in chronic lymphocytic leukemia—a comparative study of four differently designed, high resolution microarray platforms. Genes Chromosom Cancer 47:697–711. doi:10.1002/gcc.20575
Hicks J, Krasnitz A, Lakshmi B, Navin NE, Riggs M, Leibu E, Esposito D, Alexander J, Troge J, Grubor V, Yoon S, Wigler M, Ye K, Borresen-Dale AL, Naume B, Schlicting E, Norton L, Hagerstrom T, Skoog L, Auer G, Maner S, Lundin P, Zetterberg A (2006) Novel patterns of genome rearrangement and their association with survival in breast cancer. Genome Res 16:1465–1479. doi:10.1101/gr.5460106
Lambros MB, Simpson PT, Jones C, Natrajan R, Westbury C, Steele D, Savage K, Mackay A, Schmitt FC, Ashworth A, Reis-Filho JS (2006) Unlocking pathology archives for molecular genetic studies: a reliable method to generate probes for chromogenic and fluorescent in situ hybridization. Lab Invest 86:398–408. doi:10.1038/labinvest.3700390
Reis-Filho JS, Pinheiro C, Lambros MB, Milanezi F, Carvalho S, Savage K, Simpson PT, Jones C, Swift S, Mackay A, Reis RM, Hornick JL, Pereira EM, Baltazar F, Fletcher CD, Ashworth A, Lakhani SR, Schmitt FC (2006) EGFR amplification and lack of activating mutations in metaplastic breast carcinomas. J Pathol 209:445–453. doi:10.1002/path.2004
Hudson DL, Sleeman J, Watt FM (1995) CD44 is the major peanut lectin-binding glycoprotein of human epidermal keratinocytes and plays a role in intercellular adhesion. J Cell Sci 108(Pt 5):1959–1970
Roscic-Mrkic B, Fischer M, Leemann C, Manrique A, Gordon CJ, Moore JP, Proudfoot AE, Trkola A (2003) RANTES (CCL5) uses the proteoglycan CD44 as an auxiliary receptor to mediate cellular activation signals and HIV-1 enhancement. Blood 102:1169–1177. doi:10.1182/blood-2003-02-0488
Nielsen TO, Hsu FD, Jensen K, Cheang M, Karaca G, Hu Z, Hernandez-Boussard T, Livasy C, Cowan D, Dressler L, Akslen LA, Ragaz J, Gown AM, Gilks CB, van de Rijn M, Perou CM (2004) Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res 10:5367–5374. doi:10.1158/1078-0432.CCR-04-0220
Neve RM, Chin K, Fridlyand J, Yeh J, Baehner FL, Fevr T, Clark L, Bayani N, Coppe JP, Tong F, Speed T, Spellman PT, DeVries S, Lapuk A, Wang NJ, Kuo WL, Stilwell JL, Pinkel D, Albertson DG, Waldman FM, McCormick F, Dickson RB, Johnson MD, Lippman M, Ethier S, Gazdar A, Gray JW (2006) A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10:515–527. doi:10.1016/j.ccr.2006.10.008
Chin L, Gray JW (2008) Translating insights from the cancer genome into clinical practice. Nature 452:553–563. doi:10.1038/nature06914
Woodward WA, Sulman EP (2008) Cancer stem cells: markers or biomarkers? Cancer Metastasis Rev 27:459–470. doi:10.1007/s10555-008-9130-2
Jarvinen AK, Autio R, Kilpinen S, Saarela M, Leivo I, Grenman R, Makitie AA, Monni O (2008) High-resolution copy number and gene expression microarray analyses of head and neck squamous cell carcinoma cell lines of tongue and larynx. Genes Chromosom Cancer 47:500–509. doi:10.1002/gcc.20551
Fukuda Y, Kurihara N, Imoto I, Yasui K, Yoshida M, Yanagihara K, Park JG, Nakamura Y, Inazawa J (2000) CD44 is a potential target of amplification within the 11p13 amplicon detected in gastric cancer cell lines. Genes Chromosom Cancer 29:315–324. doi:10.1002/1098-2264(2000)9999:9999<::AID-GCC1047>3.0.CO;2-E
Savage K, Lambros MB, Robertson D, Jones RL, Jones C, Mackay A, James M, Hornick JL, Pereira EM, Milanezi F, Fletcher CD, Schmitt FC, Ashworth A, Reis-Filho JS (2007) Caveolin 1 is overexpressed and amplified in a subset of basal-like and metaplastic breast carcinomas: a morphologic, ultrastructural, immunohistochemical, and in situ hybridization analysis. Clin Cancer Res 13:90–101. doi:10.1158/1078-0432.CCR-06-1371
Chin K, DeVries S, Fridlyand J, Spellman PT, Roydasgupta R, Kuo WL, Lapuk A, Neve RM, Qian Z, Ryder T, Chen F, Feiler H, Tokuyasu T, Kingsley C, Dairkee S, Meng Z, Chew K, Pinkel D, Jain A, Ljung BM, Esserman L, Albertson DG, Waldman FM, Gray JW (2006) Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10:529–541. doi:10.1016/j.ccr.2006.10.009
Andre F, Job B, Dessen P, Tordai A, Michiels S, Liedtke C, Richon C, Yan K, Wang B, Vassal G, Delaloge S, Hortobagyi GN, Symmans WF, Lazar V, Pusztai L (2009) Molecular characterization of breast cancer with high-resolution oligonucleotide comparative genomic hybridization array. Clin Cancer Res 15:441–451. doi:10.1158/1078-0432.CCR-08-1791
Charafe-Jauffret E, Ginestier C, Monville F, Finetti P, Adelaide J, Cervera N, Fekairi S, Xerri L, Jacquemier J, Birnbaum D, Bertucci F (2006) Gene expression profiling of breast cell lines identifies potential new basal markers. Oncogene 25:2273–2284. doi:10.1038/sj.onc.1209254
Pegram M, Slamon D (2000) Biological rationale for HER2/neu (c-erbB2) as a target for monoclonal antibody therapy. Semin Oncol 27:13–19
Guan Y, Kuo WL, Stilwell JL, Takano H, Lapuk AV, Fridlyand J, Mao JH, Yu M, Miller MA, Santos JL, Kalloger SE, Carlson JW, Ginzinger DG, Celniker SE, Mills GB, Huntsman DG, Gray JW (2007) Amplification of PVT1 contributes to the pathophysiology of ovarian and breast cancer. Clin Cancer Res 13:5745–5755. doi:10.1158/1078-0432.CCR-06-2882
Kao J, Pollack JR (2006) RNA interference-based functional dissection of the 17q12 amplicon in breast cancer reveals contribution of coamplified genes. Genes Chromosom Cancer 45:761–769. doi:10.1002/gcc.20339
Acknowledgments
This work was funded by Breakthrough Breast Cancer grants to JSR-F and CMI. We acknowledge NHS funding to the NIHR Biomedical Research Centre. We thank Kay Savage and Suzanne Parry (Breakthrough Histopathology Laboratory, The Institute of Cancer Research) who performed the TMA immunohistochemical staining.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Pamela Klingbeil and Rachael Natrajan have contributed equally to this work. RN and RV performed aCGH and data analysis and RV performed FISH and CISH. CM and JSR-F interpreted the immunohistochemical and in situ hybridisation experiments. JP and HB provided the breast cancer samples for aCGH. PK and GE performed all the cell based assays. CMI and JSR-F designed the experiments. PK, RN, JSR-F and CMI wrote the manuscript.
Rights and permissions
About this article
Cite this article
Klingbeil, P., Natrajan, R., Everitt, G. et al. CD44 is overexpressed in basal-like breast cancers but is not a driver of 11p13 amplification. Breast Cancer Res Treat 120, 95–109 (2010). https://doi.org/10.1007/s10549-009-0380-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-009-0380-7