Abstract
Background
The aims of this study were to define the distribution of caveolin 2 (CAV2) in frozen and formalin fixed, paraffin embedded (FFPE) normal breast samples and the significance of CAV2 expression in breast cancer.
Methods
Caveolin 2 distribution in frozen and paraffin-embedded whole tissue sections of normal breast was evaluated using immunohistochemistry and immunofluorescence, in conjunction with antibodies to define luminal epithelial cells (oestrogen receptor and cytokeratin 8/18) and myoepithelial/ basal cells (cytokeratins 14 and 5/6, p63 and smooth muscle actin). CAV2 expression was also immunohistochemically analysed in two independent cohorts of invasive breast carcinomas (n = 245 and n = 418).
Results
In normal breast, CAV2 was expressed in myoepithelial cells, endothelial cells, fibroblasts and adipocytes. Luminal epithelial cells showed no or only negligible staining. CAV2 expression was observed in 9.6% of all breast cancers and was strongly correlated with high histological grade, lack of oestrogen receptor, progesterone receptor and cyclin D1 expression, and positivity for epidermal growth factor receptor, basal markers, p53 expression, and high proliferation index. Furthermore, CAV2 expression was significantly associated with basal-like immunophenotype and proved to be a prognostic factor for breast cancer-specific survival on univariate analysis.
Conclusion
Our results demonstrate that CAV2 is preferentially expressed in basal-like cancers and is associated with poor prognosis. Further in vitro studies are required to determine whether CAV2 has oncogenic properties or is only a surrogate marker of basal-like carcinomas.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The gene CAV2 maps to 7q31.1 and encodes a protein caveolin 2 [1, 2]. Caveolin 2 is one of the components of caveolar membranes, which are specialised invaginated microdomains of the plasma membrane found in the majority of mammalian cells [3, 4]. Caveolae play pivotal roles in cellular transport and also in signal transduction. These microstructures are enriched for growth factor receptors and signalling molecules, which can bind to caveolar proteins and consequently modulate several signalling pathways [3–5]. Therefore the potential role of these proteins in the development of human malignancies has received great attention. The major protein component of caveolae is caveolin 1 and, not surprisingly, most research on the role of caveolar proteins in relation to cancer has been devoted to caveolin 1 and its putative oncogenic [6, 7] and tumour suppressor roles [5, 8].
Caveolin 2 was first isolated from adipocyte-derived [1], caveolin-enriched membranes and shows 38% sequence identity to caveolin 1 [9]. In a way akin to caveolin 1, there are three caveolin 2 isoforms: a long α isoform and the truncated β and γ isoforms [1, 4, 9].
Caveolin 2 is reported to be tightly co-expressed with caveolin 1, with the highest levels of this protein being found in adipocytes, endothelial cells, fibroblasts, smooth-muscle cells, and a variety of epithelial cells [2, 9, 10]. Although caveolin 2 forms complexes with caveolin 1 and is part of the caveolar membranes, caveolin 2 does not seem to be essential for caveolae formation. In fact, Cav2 −/− mice show severe pulmonary dysfunction but have unaltered number of caveolae [4, 10, 11]. Furthermore, caveolin 2 protein sequence does not contain many of the conserved residues that have been implicated in the functionality of caveolin 1 [12, 13].
The role of caveolin 1 in breast cancer has recently received great attention in the literature [6, 7, 14–30]. Some have proposed that caveolin 1 is a tumour suppressor gene frequently inactivated in oestrogen receptor positive cancers [27, 28]. However, we and others have demonstrated that caveolin 1 is preferentially expressed in normal breast myoepithelial cells [6, 14, 17, 18, 20, 29, 30], with luminal cells showing no or only negligible staining, and in tumours with basal-like/myoepithelial phenotype [6, 14, 17, 18, 20, 29, 30]. We have also shown that CAV1 gene amplification is the likeliest mechanism for caveolin 1 expression in 13% of cases with strong caveolin 1 positivity [6]. Furthermore, caveolin 1 has been shown to be expressed in inflammatory breast carcinomas [24, 31], an aggressive type of breast cancer.
The distribution and significance of caveolin 2 expression in normal breast and breast cancer has not received the same attention as that of caveolin 1 [24, 31]. Little information on the expression of this gene has been gathered and most studies have addressed caveolin 2 expression at the mRNA level [23, 31]. However, caveolin 2 is consistently expressed in endothelial cells, therefore studies analysing the prognostic impact of caveolin 2 using mRNA expression assays without microdissection may not provide accurate information. Hence, we have sought to define the distribution of caveolin 2 in normal human breast tissue (by means of immunofluorescence and immunohistochemistry in frozen and formalin-fixed paraffin embedded normal breast samples) and also to evaluate the significance of caveolin 2 expression in breast cancer.
Materials and methods
Breast cancer cell lines
Breast carcinoma and epithelial cell lines MDA-MB-468, MCF10A, ZLR-75-1 and T47D were obtained from American Type Culture Collection (Manassas, VA), and maintained in a humidified atmosphere at 5% CO2 and 37°C. MDA-MB-468 cells, which are known to harbour a basal-like phenotype and to express high levels of caveolin 2 mRNA [30], were grown in Dulbecco’s modified Eagle’s medium, with 10% fetal bovine serum, 100 units/ml penicillin and 100 μg/ml streptomycin. ZR-75.1 and T47D cells, which are known to harbour a luminal phenotype and to express very low levels of caveolin 2 mRNA [30], were cultured in RPMI 1640 supplemented with 10% fetal bovine serum, 100 units/ml penicillin and 100 μg/ml streptomycin. MCF10A, an immortalised normal breast epithelial cell line known to have a basal-like/myoepithelial phenotype and to express the highest levels of caveolin 2 mRNA, was grown in DMEM/F12 medium supplemented with 5% horse serum, 10 μg/ml insulin, 20 ng/ml EGF, 10 mM HEPES, 0.1 μg/ml cholera toxin, 0.5 μg/ml hydrocortisone, 100 units/ml penicillin, and 100 μg/ml streptomycin. All cell media and serum were purchased from Invitrogen (Paisley, UK); other supplements were purchased Sigma-Aldrich (Gillingham,Dorset). Cells were grown to 70% confluencey in 175 cm2 flasks (BD FalconTM, Cowley, Oxford).
Western blot
Cells were washed twice in phosphate buffered saline and lysed in ice-cold SDS buffer (2% SDS, 30 mM Tris-HCl pH 6.8, 10% glycerol, 2 mM EDTA). Lysates were sonicated and protein concentration was determined using the bicinchoninic acid (BCA) assay from Perbio Science UK (Cramlington, Northumberland). Prior to loading, 2.5% 2-mercaptoethanol and 0.0125% bromphenol blue were added and lysates were boiled for 5 min. Protein aliquots in the range of 50–150 μg were resolved in 15% SDS gels and transferred to Hybond-C™ nitrocellulose (GE Healthcare; St.Giles, Bucks). Membranes were blocked in 5% non-fat milk and probed with anti-α-tubulin (clone B-5-1-2, 1:30,000, Sigma, Gillingham, Dorset, UK) and anti-caveolin 2 (clone 65, 1:200, BD Transduction Labs, Erembodegem, Belgium) overnight at 4°C. Blots were incubated with HRP-conjugated anti-mouse IgG for 1 h at room temperature (Jackson Immunoresearch, West Grove, PA). Binding was detected using the ECL plus reagent (GE Healthcare, St.Giles, Bucks).
Caveolin 2 expression in normal breast
To define the distribution of caveolin 2 in normal breast samples, 10 μm frozen sections and 4 μm formalin-fixed paraffin-embedded sections and of normal human breast obtained from cosmetic mammoplasties were employed. Frozen sections were cut onto polylysine coated slides (VWR, Leicestershire, UK) and stored at −70°C and formalin-fixed paraffin-embedded sections were cut onto Superfrost Plus slides (VWR, Leicestershire, UK) and immediately used. When the required frozen slides were thawed, the sections were marked using a slide marker pen and fixed in 4% Paraformaldehyde in PBS for 20 min RT. Sections were rinsed twice in PBS then once in PBS + 1% BSA + 2% FCS (IFF). Formalin fixed sections were subjected to heat induced antigen retrieval using Dako target retrieval solution (Dako, Glostrup, Denmark). Primary antibodies used were: Cytokeratins 8/18 (1:100, Novocastra, Newcastle-upon-Tyne, UK), α-smooth muscle actin (α-SMA, 1:5000, Sigma, Gillingham, Dorset, UK), Caveolin 2 (clone 65, 1:100, BD Transduction Labs, Erembodegem, Belgium), Oestrogen Receptor (SP1, 1:400, Neomarkers, Freemont, CA) and p63 (4A4, 1:200, Santa Cruz Biotechnology, Santa Cruz, CA). Antibodies were diluted in IFF and incubated for 40 minutes in a moist chamber followed by 3 × 5 min washes in PBS and then 40 min in conjugates diluted 1:1000 in IFF. The following conjugates were used in various combinations a-Mouse-IgG1-Alexa-488, a-Mouse-IgG2a-Alexa-555, a-Rabbit-IgG-Alexa-488, a-Rabbit-IgG-Alexa-555 (Molecular Probes, Invitrogen, Paisely, UK). Nuclei were counterstained by 3 × 5 min washes in 100 nM 4′,6-diamidino-2-phenylindole (DAPI, Molecular Probes, Invitrogen, Paisely, UK). Sections were mounted in Vectashield (Vector Laboratories, Burlingame, CA) and sealed with nail varnish. Slides were examined and photographed in a Leica TCS-SP2 confocal microscope.
Breast cancer tissue microarrays (TMAs)
The Breakthrough Breast Cancer TMA (BBCTMA) was constructed with replicate 0.6 mm cores of 245 invasive breast carcinomas (185 invasive ductal carcinomas, 27 invasive lobular carcinomas, 25 invasive mixed carcinomas and 8 invasive breast carcinomas of other special types). All patients in this series were primarily treated with surgery (69 mastectomy and 155 wide local excision), and received adjuvant anthracycline-based chemotherapy at standard doses. Adjuvant endocrine therapy was prescribed for patients with ER positive tumours. Follow-up was available for 244 patients, ranging from 0.5 to 125 months (median––67 months, mean––67 months). Full details of the characterisation of the tissue microarray and the cohort of patients are described elsewhere [6, 32].
The Vancouver TMA was constructed with replicate 0.6 mm cores of 418 breast cancers from women with primary invasive breast cancer who underwent surgery for breast cancer between 1974 and 1995 at Vancouver General Hospital. This TMA comprised a cohort of consecutive cases and the presence of invasive breast carcinoma was the only selection criterion in this study. The patients’ ages ranged from 28 to 93 years with a mean age of 60 years. Patients were primarily treated with curative surgery, either mastectomy or lumpectomy. Axillary lymph node dissection was performed in 370 patients. Histologically confirmed lymph node metastasis was confirmed in 134 patients (36.2%). Outcome data were available for all of the patients, with follow up ranging from 0.03 to 26.7 years (median = 10.8 years). Tumour size information, measured in millimeters, was available for 405 patients; 185 patients had tumors ≤20 mm, and 220 had tumors >20 mm. Adjuvant therapy varied substantially during the period 1974 through 1995, and no information on individual adjuvant treatment was available. The details of this tissue microarrays have been published elsewhere [33, 34].
Tumours were graded according to a modified Bloom-Richardson scoring system [35] and size was categorised according to the TNM staging criteria [36]. This study was approved by the Royal Marsden Hospital Ethics Committee and by the institutional ethical review board of the University of British Columbia, Vancouver, BC.
Caveolin 2 immunohistochemistry
Immunohistochemistry for Caveolin 2 was performed according to a previously described method [24] with a mouse monoclonal antibody (clone 65, BD Transduction Labs, Erembodegem, Belgium) at a dilution of 1:50 and developed with the Envision kit (Dako®, Glostrup, Denmark). Caveolin 2 immunohistochemical distribution on tissue microarray sections was analysed by three of the authors (KS, SMRP & JSR-F) on a multi-headed microscope. Only membranous with or without cytoplasmic reactivity was considered specific. The analysis was performed blinded to the results of other immunohistochemical markers and patients’ outcome. In the BBCTMA cohort, caveolin 2 expression was also correlated with that of the following immunohistochemical markers: oestrogen receptor (ER), progesterone receptor (PgR), Human Epidermal Growth Factor Receptor-2 (HER2), epidermal growth factor receptor (EGFR), cytokeratin (Ck) 14, Ck 5/6, Ck 17, cyclin D1 expression, CCND1 gene amplification and caveolin 1 expression [6, 32]. All cases were classified into luminal, HER2, basal-like and undetermined groups according to the immunohistochemical panel described Nielsen et al. [37]. In the Vancouver TMA cohort, caveolin 2 expression was correlated with that of the following immunohistochemical markers: ER, PgR, HER2, EGFR and Ck 5/6, whose results are described elsewhere [33, 38]. In the Vancouver TMA, caveolin 1 expression was analysed using the mouse monoclonal antibody 2297 (BD Transduction Labs, Erembodegem, Belgium) at 1:150 dilution following heat-induced antigen retrieval [18 min, microwave oven, DAKO antigen retrieval solution (pH 6.0)] as previously described [6]. Slides were analysed by two of the authors (SMRP & JSRF) on a multi-headed microscope using a semi-quantitative scoring system as described by Savage et al. [6]. The presence of CCND1 gene amplification was assessed by means of fluorescent in situ hybridisation (FISH), using a Spectrum Aqua labeled centromeric probe, CEP11 (Vysis), and Spectrum Orange labeled CCND1 (Vysis). FISH signals were enumerated in approximately 40 morphologically intact and non-overlapping nuclei. The average copy number for each probe was calculated and the amplification ratio determined. Amplification was defined as a CCND1: CEP11 ratio ≥1.5, as described by Brown et al. [39].
Statistical analysis
The Statview software package was used for all calculations. Correlations between categorical variables were performed using the chi-square test and Fisher’s exact test. Correlations between continuous and categorical variables were performed with analysis of variance (ANOVA). Breast cancer specific survival (BCSS) was expressed as the number of months from diagnosis to the occurrence of an event (disease-related death). Cumulative survival probabilities were calculated using the Kaplan-Meier method. Differences between survival rates were tested with the log-rank test. All tests were two-tailed, with a confidence interval of 95%.
Multivariate analysis was performed using the Cox multiple hazards model. A P value of 0.05 in the univariate survival analysis was adopted as the limit for inclusion in the multivariate model. Cases with missing values were excluded in the multivariate analysis model.
Results
Validation of the specificity of caveolin 2 monoclonal antibody
To validate the anti-caveolin 2 monoclonal antibody (clone 65, BD Transduction Labs, Erembodegem, Belgium), cell lines known to express the caveolin 2 mRNA at high and exceedingly low levels were grown according to standard protocols. As expected, the basal-like cell lines MDA-MB-468 and MC10A, known to express high levels of caveolin 2 mRNA, showed discrete 20 kDa bands, confirming the expression of caveolin 2 at protein level. The luminal cell lines T47D and ZR-75.1, which are reported to express low levels of cavolin 2 expression, showed no or very low levels of caveolin 2 protein expression, respectively (Fig. 1).
Validation of caveolin 2 antibody by Western blot. Proteins were resolved in 15% SDS gels, under reducing conditions and probed with anti-α-tubulin and anti-caveolin 2 monoclonal antibodies, followed by HRP-conjugated anti-mouse IgG. The blot shows expression of caveolin 2 in the immortalized breast epithelial cell line MCF10A and to a lesser extent in MDA-MB-468, basal-like carcinoma. In contrast, T47D and ZR-75.1 (right panel) express no detectable caveolin 2
Expression of caveolin 2 in normal breast
The distribution of caveolin 2 in both frozen and formalin-fixed, paraffin-embedded normal breast tissues was identical. Caveolin 2 was strongly expressed in adipocytes, endothelial cells, perineurial cells, a subset of intra and inter-lobular fibroblasts. Within normal breast ducts and lobules, caveolin 2 was restricted to myoepithelial cells. Luminal epithelial cells showed no or only negligible staining (Fig. 2 and 3).
Analysis of caveolin 1 and 2 expression in normal breast by immunofluorescence. Normal breast ducts (A and B) and terminal duct-lobular units (C and D) display strong expression of caveolin 1 (A and C, green) and 2 (B and D, green) in myoepithelial cells, which also co-express p63 (A, B, C and D, red). Note that within breast ducts and lobules, caveolins 1 and 2 are expressed in similar cell populations (i.e., myoepithelial cells). In addition, caveolin are expressed in fibroblasts and endothelial cells. In E and F, please note the co-expression of caveolin 2 (E and F, green) and α-smooth muscle actin (E and F, red) in normal breast myoepithelial cells. In vessels, α-smooth muscle actin is expressed in pericytes (E and F, red, arrow), whereas caveolin 2 is expressed in endothelial cells (E and F, green, arrowheads)
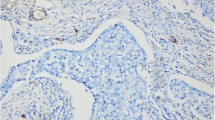
Expression of caveolin 2 in a formalin-fixed paraffin-embedded section of normal breast (A and B) and in basal-like breast cancers (C and D). In normal breast, note the similar distribution of caveolin 2 when compared to that seen with immunofluorescence. Caveolin 2 expression in basal-like breast cancer was found in the membranous and cytoplasmic subcellular compartments
Caveolin 2 expression in invasive breast cancer
In breast cancers, caveolin 2 expression was seen in 9.6% of breast cancers (i.e., 13 out of 210 informative cases (6.2%) of the BBCTMA and in 37 out of 309 (11.9%) informative cases of the Vancouver TMA). In both cohorts, caveolin 2 expression was significantly associated with high histological grade, lack of ER and PgR, and positivity for EGFR and basal keratins (Tables 1 and 2 and Fig. 3). Caveolin 2 positive tumours more frequently displayed p53 nuclear immunostaining and were significantly associated with high proliferation rates (Table 1). In addition, in the BBCTMA cohort, caveolin 2 expression was inversely correlated with cyclin D1 expression. Given the above correlations it is not surprisingly that caveolin 2 showed a strong association with basal-like immunophenotype as defined by Nielsen et al.’s criteria [37].
Univariate survival analysis revealed that expression of caveolin 2 was significantly associated with a shorter breast cancer-specific survival in both BBCTMA and Vancouver TMA cohorts (Fig. 4). Caveolin 1 expression was also associated with a shorter breast cancer-specific survival in both BBCTMA and Vancouver TMA cohorts on univariate survival analysis ([6] and Huntsman D and Reis-Filho JS, data not shown). On a multivariate model including tumour size, histological grade, lymph node metastasis and caveolin 2 expression proved to be an independent predictor of outcome for BCSS in the BBCTMA cohort, but not in the Vancouver TMA cohort (Table 3). Furthermore, in a model including all parameters associated with BCSS on univariate analysis, caveolin 2 failed to provide additional prognostic information in both cohorts (Table 3). Given the strong association between caveolin 2 expression and basal-like phenotype, it is likely that its prognostic significance stems from their strong direct correlation (i.e., in the BBCTMA, 39.3% of all basal-like cancers harboured caveolin 2 expression whereas 0.6% of non-basal-like cancers showed positivity for caveolin, P < 0.0001; and in the Vancouver TMA, 45.6% of all basal-like cancers harboured caveolin 2 expression whereas 4.1% of non-basal-like cancers showed positivity for caveolin, P < 0.0001).
Discussion
In this study we demonstrate that caveolin 2 is preferentially expressed in myoepithelial cells of normal breast, whereas luminal epithelial cells showed negligible staining both in frozen and formalin-fixed, paraffin-embedded tissue sections. We have also demonstrated that caveolin 2 is co-expressed with caveolin 1 in normal breast myoepithelial cells.
In previous expression profiling studies, caveolin 2 was one of the genes featuring in the intrinsic gene list used to classify breast cancer into molecular subgroups described by described by the Stamford group [40, 41]. In those studies, caveolins 1 and 2 were shown to be expressed at the highest levels in normal breast-like cancers. Interestingly, unlike caveolin 1, the levels of caveolin 2 mRNA were also high in the majority of basal-like cancers [40]. These findings suggest that although caveolin 1 and 2 map to the same chromosomal region (i.e., 7q31.1) and are known to form heterodimers, distinct mechanisms regulate the expression of these two components of the caveolar membranes in breast cancers. Interestingly, when we carried out a detailed analysis of the publicly available data from two microarray studies [42, 43] (http://www.oncomine.org), caveolin 2 was shown to be expressed at significantly higher levels (P < 0.001, t-test) in tumours that frequently show a basal-like phenotype [44], namely those arising in BRCA1 mutation carriers [43] and in TP53-mutated tumours [42]. However, results of expression profile studies should be interpreted with caution, given that the high levels of caveolins 1 and 2 in normal breast-like samples may be derived from the expression of caveolins in angiogenic endothelial cells and fibroblasts of the tumour stroma [6].
This is the first study where expression of caveolin 2 at the protein level was shown to be strongly correlated with that of basal/ myoepithelial markers. We have also demonstrated that although caveolin 1 and caveolin 2 are frequently co-expressed and significantly correlated, 43.5% (20 out of 46) of tumours expressing caveolin 2 failed to show caveolin 1 protein expression and that 35% (14 out 40) of tumours expressing caveolin 1 lacked caveolin 2 expression. Interactions between caveolins 1 and 2 have been shown to be complex and context dependent [4, 10]. Their role and significance in the formation of caveolae in tumours remains to be determined. Although there are arguable data to suggest that caveolin 1 may negatively regulate the activity of EGFR [12, 13, 45], it has recently been demonstrated that EGFR and caveolin 1 are co-expressed in several tumour types [6, 18, 29, 30, 46] but do not necessarily co-localise to the same subcellular compartment [45, 47]: whilst caveolin 1 localises to the caveolae, EGFR seems to be preferentially expressed in flat lipid rafts [45, 47]. In this study, 42–58% of EGFR positive breast cancers co-expressed caveolin 2. However, caveolin 2 lacks the motifs reported to be required for the interaction between caveolins and EGFR [13]; therefore the significance of this finding remains to be determined. Given that myoepithelial/basal cells express caveolins 1 and 2 and EGFR [6, 14, 18, 26, 29, 30], one might speculate that expression of these markers in basal-like breast cancers could either be a reflection of the cell of origin of basal-like/myoepithelial breast cancers or part of a transcriptomic programme that leads to a basal-like/myoepithelial phenotype.
We and others have previously shown that ER expression is inversely correlated with the expression of caveolin 1 in normal breast and breast cancer [6, 14, 17, 18, 30]. In the present study, we demonstrate that ER-positive normal breast luminal epithelial cells express negligible levels of caveolin 2 and that, in breast cancer, caveolin 2 expression is inversely correlated with the expression of ER. Our findings support the idea that the ER signalling inhibits the expression of caveolin 1 and 2 in normal smooth muscle cells and breast cancer cells [48–50]. The study by Razandi et al. [48] demonstrated, in cell line experiments, that ER associates with caveolin proteins in the plasma membrane and that oestrogen can modulate this association depending on the cellular context. These authors also suggested that CAV1 expression facilitated ER translocation to the cell membrane and inhibited oestrogen induced MAPK activation, suggesting that ER signalling and the caveolar proteins modulate each other [48]. In addition, given that normal breast luminal epithelial cells have negligible levels of caveolin 1 [6, 17] and 2 expression, our findings call into question the significance of the apparent ‘loss’ of caveolins expression, the arguable dominant negative CAV1 gene mutations in ER-positive breast cancers [15, 21, 27, 28] and the model recently put forward by Li et al. [28]. Taken together, the above findings add to the increasing evidence to suggest that caveolins may not have tumour suppressive properties in the basal-like subgroup of breast cancer [6, 14, 18, 25, 26, 29, 30].
Caveolin 2 has been shown to be expressed in highly aggressive, inflammatory breast carcinomas [24, 31]. Interestingly, it has been shown that inflammatory breast cancers preferentially display either a basal-like or a HER2 phenotype by gene expression analysis [51]. Although no inflammatory breast cancers were included in the present study due to the TMA design, on univariate analysis, we demonstrated that caveolin 2 was associated with a poor breast cancer specific survival in two cohorts of patients. In addition, its impact was independent of size, grade and lymph node metastasis in a cohort of patients uniformly treated with surgery followed by anthracycline-based adjuvant chemotherapy. However, when basal-like status was included in the multivariate model, caveolin 2 expression lost its significance, suggesting that it is unlikely to add any independent prognostic information to that offered by other basal/myoepithelial markers. Nevertheless, both caveolin 1 and 2 seem to play a pivotal role in the biology of basal-like breast cancer, principally because they seem to be involved in the sensitivity of this subgroup of breast cancers to crosslinking agents [52] and tyrosine kinase inhibitors [53]. Huang et al. [53] have recently demonstrated that caveolins 1 and 2 are significantly downregulated when breast cancer cell lines were subjected to treatment with dasatinib, a multitargeted kinase inhibitor. Interestingly, breast cancer cell lines sensitive to this drug are preferentially of basal-like phenotype and express high levels of caveolins 1 and 2 [53]. Furthermore, these caveolins have been shown to be substrates for SRC family kinases and functionally involved in their signalling pathways [53, 54]. In addition, caveolins 1 and 2 have been included in a six gene predictive signature for response to this tyrosine kinase inhibitor [53].
In conclusion, this study provides another line of evidence to suggest that caveolin 2 is unlikely to play a tumour suppressive role in basal-like breast carcinomas and that it may be used, in conjunction with other markers (i.e., ER, HER2, ‘basal’ keratins, EGFR and P-cadherin) [6, 14, 17, 18, 26, 29, 30, 32, 37, 44], to identify tumours with a basal-like phenotype. Further in vitro and in vivo studies are required to determine whether caveolin 2 has oncogenic properties or is only a surrogate marker for basal-like cancers.
References
Engelman JA, Zhang XL, Lisanti MP (1998) Genes encoding human caveolin-1 and -2 are co-localized to the D7S522 locus (7q31.1), a known fragile site (FRA7G) that is frequently deleted in human cancers. FEBS Lett 436:403–410
Scherer PE, Okamoto T, Chun M et al (1996) Identification, sequence, and expression of caveolin-2 defines a caveolin gene family. Proc Natl Acad Sci U S A 93:131–135
Hnasko R, Lisanti MP (2003) The biology of caveolae: lessons from caveolin knockout mice and implications for human disease. Mol Interv 3:445–464
Stan RV (2005) Structure of caveolae. Biochim Biophys Acta 1746:334–348
Bouras T, Lisanti MP, Pestell RG (2004) Caveolin-1 in breast cancer. Cancer Biol Ther 3:931–941
Savage K, Lambros MB, Robertson D et al (2007) Caveolin 1 is overexpressed and amplified in a subset of basal-like and metaplastic breast carcinomas: a morphologic, ultrastructural, immunohistochemical, and in situ hybridization analysis. Clin Cancer Res 13:90–101
Williams TM, Lisanti MP (2005) Caveolin-1 in oncogenic transformation, cancer, and metastasis. Am J Physiol Cell Physiol 288:C494–C506
Razani B, Schlegel A, Liu J et al (2001) Caveolin-1, a putative tumour suppressor gene. Biochem Soc Trans 29:494–499
Scherer PE, Lewis RY, Volonte D et al (1997) Cell-type and tissue-specific expression of caveolin-2. Caveolins 1 and 2 co-localize and form a stable hetero-oligomeric complex in vivo. J Biol Chem 272:29337–29346
Le Lay S, Kurzchalia TV (2005) Getting rid of caveolins: phenotypes of caveolin-deficient animals. Biochim Biophys Acta 1746:322–333
Razani B, Wang XB, Engelman JA et al (2002) Caveolin-2-deficient mice show evidence of severe pulmonary dysfunction without disruption of caveolae. Mol Cell Biol 22:2329–2344
Couet J, Li S, Okamoto T et al (1997) Identification of peptide and protein ligands for the caveolin-scaffolding domain. Implications for the interaction of caveolin with caveolae-associated proteins. J Biol Chem 272:6525–6533
Couet J, Sargiacomo M, Lisanti MP (1997) Interaction of a receptor tyrosine kinase, EGF-R, with caveolins. Caveolin binding negatively regulates tyrosine and serine/threonine kinase activities. J Biol Chem 272:30429–30438
Charafe-Jauffret E, Ginestier C, Monville F et al (2005) Gene expression profiling of breast cell lines identifies potential new basal markers. Oncogene 25:2273–2284
Chen ST, Lin SY, Yeh KT et al (2004) Mutational, epigenetic and expressional analyses of caveolin-1 gene in breast cancers. Int J Mol Med 14:577–582
Engelman JA, Zhang XL, Lisanti MP (1999) Sequence and detailed organization of the human caveolin-1 and -2 genes located near the D7S522 locus (7q31.1). Methylation of a CpG island in the 5′ promoter region of the caveolin-1 gene in human breast cancer cell lines. FEBS Lett 448:221–230
Hurlstone AF, Reid G, Reeves JR et al (1999) Analysis of the CAVEOLIN-1 gene at human chromosome 7q31.1 in primary tumours and tumour-derived cell lines. Oncogene 18:1881–1890
Pinilla SM, Honrado E, Hardisson D et al (2006) Caveolin-1 expression is associated with a basal-like phenotype in sporadic and hereditary breast cancer. Breast Cancer Res Treat 99:85–90
Hayashi K, Matsuda S, Machida K et al (2001) Invasion activating caveolin-1 mutation in human scirrhous breast cancers. Cancer Res 61:2361–2364
Jones C, Mackay A, Grigoriadis A et al (2004) Expression profiling of purified normal human luminal and myoepithelial breast cells: identification of novel prognostic markers for breast cancer. Cancer Res 64:3037–3045
Lee H, Park DS, Razani B et al (2002) Caveolin-1 mutations (P132L and null) and the pathogenesis of breast cancer: caveolin-1 (P132L) behaves in a dominant-negative manner and caveolin-1 (−/−) null mice show mammary epithelial cell hyperplasia. Am J Pathol 161:1357–1369
Park SS, Kim JE, Kim YA et al (2005) Caveolin-1 is down-regulated and inversely correlated with HER2 and EGFR expression status in invasive ductal carcinoma of the breast. Histopathology 47:625–630
Sagara Y, Mimori K, Yoshinaga K et al (2004) Clinical significance of Caveolin-1, Caveolin-2 and HER2/neu mRNA expression in human breast cancer. Br J Cancer 91:959–965
Van den Eynden GG, Van Laere SJ, Van der Auwera I et al (2006) Overexpression of caveolin-1 and -2 in cell lines and in human samples of inflammatory breast cancer. Breast Cancer Res Treat 95:219–228
Yang G, Truong LD, Timme TL et al (1998) Elevated expression of caveolin is associated with prostate and breast cancer. Clin Cancer Res 4:1873–1880
Jacquemier J, Padovani L, Rabayrol L et al (2005) Typical medullary breast carcinomas have a basal/myoepithelial phenotype. J Pathol 207:260–268
Sotgia F, Rui H, Bonuccelli G et al (2006) Caveolin-1, mammary stem cells, and estrogen-dependent breast cancers. Cancer Res 66:10647–10651
Li T, Sotgia F, Vuolo MA et al (2006) Caveolin-1 mutations in human breast cancer: functional association with estrogen receptor {alpha}-positive status. Am J Pathol 168:1998–2013
Chin K, DeVries S, Fridlyand J et al (2006) Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10:529–541
Neve RM, Chin K, Fridlyand J et al (2006) A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10:515–527
Van Laere S, Van der Auwera I, Van den Eynden GG et al (2005) Distinct molecular signature of inflammatory breast cancer by cDNA microarray analysis. Breast Cancer Res Treat 93:237–246
Reis-Filho JS, Savage K, Lambros MB et al (2006) Cyclin D1 protein overexpression and CCND1 amplification in breast carcinomas: an immunohistochemical and chromogenic in situ hybridisation analysis. Mod Pathol 19:999–1009
Makretsov NA, Huntsman DG, Nielsen TO et al (2004) Hierarchical clustering analysis of tissue microarray immunostaining data identifies prognostically significant groups of breast carcinoma. Clin Cancer Res 10:6143–6151
Prentice LM, Shadeo A, Lestou VS et al (2005) NRG1 gene rearrangements in clinical breast cancer: identification of an adjacent novel amplicon associated with poor prognosis. Oncogene 24:7281–7289
Elston CW, Ellis IO (1991) Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 19:403–410
Singletary SE, Connolly JL (2006) Breast cancer staging: working with the sixth edition of the AJCC Cancer Staging Manual. CA Cancer J Clin 56:37–47; quiz 50–1
Nielsen TO, Hsu FD, Jensen K et al (2004) Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res 10:5367–5374
Tischkowitz M, Brunet JS, Begin LR et al (2007) Use of immunohistochemical markers can refine prognosis in triple negative breast cancer. BMC Cancer 7(1):134
Brown LA, Irving J, Parker R et al (2006) Amplification of EMSY, a novel oncogene on 11q13, in high grade ovarian surface epithelial carcinomas. Gynecol Oncol 100:264–270
Perou CM, Sorlie T, Eisen MB et al (2000) Molecular portraits of human breast tumours. Nature 406:747–752
Sorlie T, Perou CM, Tibshirani R et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A 98:10869–10874
Miller LD, Smeds J, George J et al (2005) An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. Proc Natl Acad Sci U S A 102:13550–13555
Pawitan Y, Bjohle J, Amler L et al (2005) Gene expression profiling spares early breast cancer patients from adjuvant therapy: derived and validated in two population-based cohorts. Breast Cancer Res 7:R953–R964
Turner NC, Reis-Filho JS (2006) Basal-like breast cancer and the BRCA1 phenotype. Oncogene 25:5846–5853
Pike LJ (2005) Growth factor receptors, lipid rafts and caveolae: an evolving story. Biochim Biophys Acta 1746:260–273
Abulrob A, Giuseppin S, Andrade MF et al (2004) Interactions of EGFR and caveolin-1 in human glioblastoma cells: evidence that tyrosine phosphorylation regulates EGFR association with caveolae. Oncogene 23:6967–6979
Ringerike T, Blystad FD, Levy FO et al (2002) Cholesterol is important in control of EGF receptor kinase activity but EGF receptors are not concentrated in caveolae. J Cell Sci 115:1331–1340
Razandi M, Oh P, Pedram A et al (2002) ERs associate with and regulate the production of caveolin: implications for signaling and cellular actions. Mol Endocrinol 16:100–115
Kiss AL, Turi A, Mullner N et al (2005) Oestrogen-mediated tyrosine phosphorylation of caveolin-1 and its effect on the oestrogen receptor localisation: an in vivo study. Mol Cell Endocrinol 245:128–137
Yang FL, He H, Liu XX et al (2006) Effects of raloxifene on caveolin-1 mRNA and protein expressions in vascular smooth muscle cells. Acta Biochim Biophys Sin (Shanghai) 38:747–752
Van Laere SJ, Van den Eynden GG, Van der Auwera I et al (2006) Identification of cell-of-origin breast tumor subtypes in inflammatory breast cancer by gene expression profiling. Breast Cancer Res Treat 95:243–255
Vekris A, Meynard D, Haaz MC et al (2004) Molecular determinants of the cytotoxicity of platinum compounds: the contribution of in silico research. Cancer Res 64:356–362
Huang F, Reeves K, Han X et al (2007) Identification of candidate molecular markers predicting sensitivity in solid tumors to dasatinib: rationale for patient selection. Cancer Res 67:2226–2238
Brown MT, Cooper JA (1996) Regulation, substrates and functions of src. Biochim Biophys Acta 1287:121–149
Acknowledgments
This study was funded in part by Breakthrough Breast Cancer. The Genetic Pathology Evaluation Centre is funded in part through an unrestricted educational grant from Sanofi-Aventis. We thank the Queen Victoria Hospital NHS Trust, East Grinstead, for provision of normal breast reduction mammoplasties.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Savage, K., Leung, S., Todd, S.K. et al. Distribution and significance of caveolin 2 expression in normal breast and invasive breast cancer: an immunofluorescence and immunohistochemical analysis. Breast Cancer Res Treat 110, 245–256 (2008). https://doi.org/10.1007/s10549-007-9718-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-007-9718-1