Abstract
The potential for bioaugmentation with aerobic explosive degrading bacteria to remediate hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) contaminated aquifers was demonstrated. Repacked aquifer sediment columns were used to examine the transport and RDX degradation capacity of the known RDX degrading bacterial strains Gordonia sp. KTR9 (modified with a kanamycin resistance gene) Pseudomonas fluorescens I-C, and a kanamycin resistant transconjugate Rhodococcus jostii RHA1 pGKT2:Km+. All three strains were transported through the columns and eluted ahead of the conservative bromide tracer, although the total breakthrough varied by strain. The introduced cells responded to biostimulation with fructose (18 mg L−1, 0.1 mM) by degrading dissolved RDX (0.5 mg L−1, 2.3 µM). The strains retained RDX-degrading activity for at least 6 months following periods of starvation when no fructose was supplied to the column. Post-experiment analysis of the soil indicated that the residual cells were distributed along the length of the column. When the strains were grown to densities relevant for field-scale application, the cells remained viable and able to degrade RDX for at least 3 months when stored at 4 °C. These results indicate that bioaugmentation may be a viable option for treating RDX in large dilute aerobic plumes.
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) is an explosive which is frequently detected in soil and groundwater at military sites, particularly at Department of Defense training and testing ranges (Brannon and Myers 1997; Clausen et al. 2004; Pennington et al. 2001, 2002; Yamamoto et al. 2004). RDX is moderately soluble (e.g., ~40 mg L−1) and has a relatively low octanol–water partitioning coefficient (e.g., log Kow ~0.9), and thus can migrate rapidly from soil to groundwater. Due to its potential toxicity, the U.S. Environmental Protection Agency (EPA) recently established a health advisory level of 2 μg L−1 for RDX in drinking water (http://www.atsdr.cdc.gov/toxprofiles/tp78-c8.pdf).
RDX can be biodegraded by a range of both anaerobic (Adrian and Sutherland 1999; Arnett and Adrian 2009; Fuller et al. 2009; Hawari et al. 2001; Zhang and Hughes 2003; Zhao et al. 2004) and aerobic bacteria (Bernstein et al. 2011; Fournier et al. 2002; Fuller et al. 2010a; Seth-Smith et al. 2002; Thompson et al. 2005). In situ anaerobic bioremediation of RDX (and other explosives) has been demonstrated at several sites (Hatzinger and Lippincott 2012; Michalsen et al. 2013; Newell 2008; Wade et al. 2010). However, creating and maintaining anaerobic conditions across large areas is costly and technically challenging, indicating a need to pursue other potential solutions, especially for large, dilute plumes. Moreover, introduction of carbon substrates required for anaerobic treatment of explosives and other compounds (e.g., volatile organic compounds) often creates a number of secondary groundwater issues including, mobilization of iron, manganese, and arsenic, formation of methane and hydrogen sulfide, and biofouling of wells. Based on these factors, an aerobic bioaugmentation approach may be more effective with less adverse and/or fewer side effects. However, there has only been limited research on developing RDX degrading bioaugmentation cultures (Priestley et al. 2006), and there are no published data providing support for aerobic bioaugmentation of RDX contaminated aquifers.
Aerobic RDX degraders with the xplA/xplB genes have been detected in surface soils (Seth-Smith et al. 2008) and unsaturated deep vadose zone soils and groundwater in Israel (Bernstein et al. 2011; Ronen et al. 2008). However, broader surveys to detect genetic markers have indicated these organisms may not be prevalent in groundwater (Fuller et al. 2010b), so bioaugmentation may be required at many sites for successful aerobic bioremediation of RDX. A case in point is the Umatilla Chemical Depot (UMCD) in Umatilla, OR, where a previous study indicated that aerobic degradation of RDX by indigenous bacteria was not a viable remedial option, presumably due to the absence or low density of organisms with this capability (Michalsen et al. 2013).
Long-term munitions demilitarization operations at the UMCD since 1944 generated explosives-contaminated wastewater that was stored in shallow unlined lagoons. Leachate from the lagoons has resulted in widespread RDX contamination of the underlying aerobic, highly permeable aquifer (average hydraulic conductivity of ~180 m day−1). A pump-and-treat (P&T) system has been operating for several years to remove RDX from groundwater, but treatment efficiency has significantly decreased. RDX concentrations range from 2 to 300 µg L−1 throughout the approximately 80 ha plume. Although anaerobic biostimulation has been successfully demonstrated in situ at the site (Michalsen et al. 2013), quantities of growth substrate required to implement this anaerobic approach at full scale would be substantial and expensive (e.g., millions of US dollars). This provides the impetus for demonstrating aerobic bioaugmentation for remediation of RDX-contaminated groundwater as an innovative, cost-effective approach for UMCD and other sites with large dilute plumes.
One of the critical aspects of successful bioaugmentation for aquifer remediation is adequate transport and distribution of inoculated cells. Based on cell surface charge, cell size, and other characteristics, some microbial cells have been observed to move only a few centimeters in aquifer materials, sometimes leading to clogging of the aquifer or injection wells (Li and Logan 1999; Shaw et al. 1985; Streger et al. 2002). Thus, both cell transport and survival must be evaluated when considering bioaugmentation in the field.
The research presented herein was undertaken to evaluate the potential for bioaugmentation with aerobic bacteria to remediate an RDX contaminated aquifer at UMCD. Specifically, (1) repacked column experiments were performed to determine if the selected strains could be transported effectively through the alluvial aquifer sediments and if the retained cells could survive and maintain RDX degradation capacity over several months, and; (2) cultures were grown in laboratory-scale fermenters to high density and their viability with time was determined to verify that cultures could be prepared and maintained for a large-scale field application. These critical cell transport, degradation, and fermentation characteristics have not been previously evaluated for aerobic RDX degrading bacteria.
Materials and methods
Chemicals and media
RDX was synthesized by Dr. Stephen Fallis at the Naval Air Warfare Center Weapons Division, China Lake, CA. An artificial groundwater (UMAGW) based on the site geochemistry was used for most experiments (see Table S1 in Supplemental Information). Phosphate buffered saline (PBS, pH 7.4) contained (L−1 distilled H2O): 0.24 g K2HPO4, 1.44 g NaH2PO4, 0.20 g KCl, 8.00 g NaCl, and was sterilized by filtration. All other chemicals were reagent grade or purer. Basal salts medium (BSM) was prepared as described in Hareland et al. (1975). R2A agar was purchased from Difco (Becton, Dickinson and Company, Sparks, MD, USA), and Luria broth agar (LB) from Fisher Scientific (Fairlawn, NJ, USA).
Aquifer sediment collection and processing
Sediment samples were collected from the saturated zone by air rotary drilling 38–44 m (125–145 ft) below ground surface, placed on ice, and shipped overnight to CB&I’s laboratory for processing. Sediment was passed through a screen to remove larger material (>3/4″ or 19 mm). The resulting sediment particle size distribution is shown in Table S2, with a mean particle size of 3.4 mm. The sediment organic and inorganic carbon concentrations were on the order of 400 and 5,000 mg kg−1, respectively. Water extractable sulfate and nitrate were below detectable levels (<0.2 mg kg−1). Sieved sediment was stored moist at 15 °C until use.
Bacterial strains
Three RDX degrading bacterial strains were used for this research: the aerobic RDX-degrader Gordonia sp. KTR9 (Thompson et al. 2005; KTR9 hereafter) harboring plasmid pGKT2:Km+, which contained the RDX-degradative genes xplA and xplB (Indest et al. 2010), and an inserted kanamycin resistance marker (Jung et al. 2011); the aerobic RDX-degrader transconjugate Rhodococcus jostii RHA1 pGKT2:Km+ (RHA1 hereafter; Jung et al. 2011); and the anoxic RDX-degrader Pseudomonas fluorescens I-C (Fuller et al. 2009; I-C hereafter). These three strains were selected for their ability to remain viable and active (assessed by RDX degradation activity) for 7 days at 15 °C in UMCD sediment microcosms (see Supplemental Information). The anoxic RDX degrading strain I-C was included because it was possible that spatially variable redox conditions would be created in the columns following substrate additions, (e.g., variable consumption of the carbon source could lead to reductions in the dissolved oxygen (DO) concentrations below what was required by the aerobic RDX degraders). Routine plating was performed using R2A agar, with selective plating for KTR9 and RHA1 on LB agar amended with 50 µg L−1 kanamycin sulfate. All three strains were characterized using a standardized adhesion assay (DeFlaun et al. 1990) using sterilized UMCD sediment.
Repacked column preparation and operation
The repacked columns were similar to those used in a previous study (Schaefer et al. 2007). A schematic of the column setup is presented in Fig. S4. All materials used were stainless steel, Teflon, norprene, glass, or PVC to minimize sorptive losses of RDX. Columns were 30 cm in length with a 4.8 cm inside diameter. Side ports were positioned at 3.25, 7.50, and 15 cm from the influent end of the column, but were not sampled for the results presented here. Moist UMCD sediment was packed into the columns to achieve an approximate bulk density of 1.7 g cm−3. After packing, columns were moved to a walk-in environmental chamber maintained at 15 °C (typical subsurface UMCD temperature) where incubation and sample collection occurred. UMAGW was pumped into the bottom of the column to saturate the sediment at a flow rate of 0.15 mL min−1. Following saturation, a bromide tracer test (tracer concentration 100 mg L−1 Br) was performed to check for packing artifacts (e.g., short circuiting) and to calculate the column pore volume (PV), which was approximately 200 mL. The flow into the column was then switched to UMAGW amended with RDX at a concentration of 0.5 mg L−1 (2.3 µM), and the flow continued until the concentration of RDX in the effluent (C) was roughly equal to the RDX concentration of the influent (C0; C/C0 ≈ 1). Effluent from the column was either directed into a fraction collector, or into a bulk collection bottle. Samples were processed approximately daily during weekdays as described below, and effluent volumes were recorded to monitor actual flow rates.
Column biostimulation, bioaugmentation and sampling
Prior to bioaugmentation, biostimulation of the indigenous microbial community was assessed. Three separate additions of fructose were introduced into the column [2 PVs, final influent concentration of 18 mg L−1 (0.1 mM)] at a flow rate of 0.15 mL min−1. Before, during and after the fructose addition, samples of the column influent and effluent were collected and analyzed for RDX concentrations (see below).
After biostimulation, bioaugmentation was performed. The three strains were grown separately in BSM amended with fructose (final concentration 9 g L−1, 50 mM) at 30 °C. Cultures were harvested by centrifugation (3,000×g, 20 min, 5 °C) and washed twice with sterile UMAGW, with final resuspension in sterile UMAGW. The cultures were starved at 15 °C for 48–72 h, then washed twice and resuspended in UMAGW. The optical density (OD) of the starved cultures was measured, and an appropriate dilution was prepared in order to achieve a column inoculum containing approximately 1 × 108 cells mL−1 of each strain in a total volume of 2 PV (400 mL). The inoculum solution also contained RDX (0.5 mg L−1) and Br (100 mg L−1). Samples of the column inoculum were analyzed for initial cell densities (colony forming units, CFUs) by spread plating on general and selective media, and using quantitative polymerase chain reaction (qPCR, see below).
The inoculum was injected into the column at a flow rate of 1.5 mL min−1. After the 2 PV of inoculum solution had entered the column, the influent was switched over to RDX amended UMAGW. Fructose [2 PVs, final influent concentration of 18 mg L−1 (0.1 mM)] was added periodically to assess the ability of the injected cells to degrade RDX upon stimulation with a utilizable carbon source. The flow rate was maintain at 0.15 mL min−1 (which translated to a groundwater velocity of 0.3 m day−1), except during the cell injection when the flow rate was increased to 1.5 mL min−1 (or 3 m day−1), as noted above. These velocities correspond to the natural seepage velocity and the forced gradient velocity when the P&T system is active at UMCD, respectively. Periodic samples of the influent and effluent were collected and analyzed for a range of parameters including RDX, injected strain CFU, anions, alkalinity, pH, and DO (see below). At the end of the experiment, the sediment in the column was removed in discrete intervals and analyzed for residual cell concentrations using qPCR.
Field-scale bioaugmentation culture preparation and characterization
The long term viability and RDX degrading activity of the three strains was also evaluated after they were grown and concentrated to densities required for field-scale application. Starter cultures were initially grown in 3- or 7-L benchtop bioreactors (Applikon Biotechnology B.V., Schiedam, The Netherlands). The bioreactors were continuously mixed, and positive pressure was maintained to minimize foaming. The pH and DO were monitored by specific probes, and residual fructose and ammonium were determined using colorimetric tests. Control of pH was achieved by automatic addition of aqueous solutions of acid (H2SO4) or base (NaOH). Periodic samples were removed for measurement of the cell density (OD600 and CFU).
Once a starter culture had reached a constant OD600, it was used to inoculate a 750-L bioreactor (Abec, Allentown, PA, USA). Growth continued in the 750-L bioreactor until the cell density (as determined by OD600) multiplied by bioreactor volume reached the target number of cells for a hypothetical pilot-scale injection (e.g., 10,000 L at 5 × 107 cells mL−1, or 5 × 1014 total cells). After the required cell density was achieved, the culture was passed through a custom-built cross-flow filtration unit (Kerasep™ tubular ceramic membranes, Novasep, Inc., Boothwyn, PA, USA) to remove the culture media and concentrate the biomass. The culture was further concentrated using a flow-through centrifuge (CEPA Z41, Carl Padberg Zentrigugenbau GmbH, Geroldsecker Vorstadt, Germany; 17,000×g at 21 °C) with final resuspension in sterile UMAGW.
Subsamples (~100 mL) of the concentrated cultures were transferred to duplicate 250 mL sterile glass bottles. One bottle of each culture was placed in a refrigerator at 4 °C (expected shipping and long-term storage temperature during a field injection), and the other was placed in an incubator at 37 °C (anticipated highest temperature the cultures would experience during shipping). Bottles were incubated without shaking. Well-mixed samples (10 mL) were removed from the bottles initially, and after 1, 2, 5, 7, 14 days, and additionally at 30, 60, and 90 days for the bottles incubated at 4 °C. After passing the sample several times through a 25 gauge hypodermic needle using a 20 mL disposable syringe to reduce cell clumping, the optical density (OD550) and viable cell counts were determined (spread plating onto LB + kanamycin and R2A media).
The RDX degradation potential of the cultures over time was assayed on the same schedule (except at the 2 day timepoint) by combining 1 mL of a 1:100 dilution of the sample (in PBS) with 9 mL of sterile UMAGW amended with RDX (10 mg L−1) and fructose (9 mg L−1). The assays with KTR9 and RHA1 were performed in 25 mL serum vials with 15 mL of headspace to maintain aerobic conditions. The assays with I-C were performed in 11 mL serum vials with minimal headspace to create suboxic conditions favorable to RDX degradation by this strain. An uninoculated control was set up with each batch of assays. Vials were incubated with shaking (125 rpm) at room temperature. Subsamples were removed after 24 and 48 h, passed through a 0.45 µm glass microfiber filter, and analyzed for RDX.
Analytical
The concentrations of RDX and its nitroso-containing metabolites hexahydro-1-nitroso-3,5-dinitro-1,3,5-triazine, hexahydro-1,3-dinitroso-5-nitro-triazine, and hexahydro-1,3,5-trinitroso-1,3,5-triazine were monitored using high performance liquid chromatography (HPLC) according to a modified EPA Method 8330 (www.epa.gov/epawaste/hazard/testmethods/sw846/pdfs/8330a.pdf) using a Dionex 3000 Ultimate HPLC with a Agilent Zorbax Bonus-RP column (4.6 × 75 mm, 3.5 µm particle diameter), variable wavelength detector (254 nm), and a photodiode array detector collecting peak spectral data. The mobile phase was 50:50 methanol:0.2 % (v:v) trifluoroacetic acid in water at a flow rate of 1 mL min−1. The column temperature was 33 °C. The practical quantitation limit was approximately 10 µg L−1. The RDX metabolites 4-nitro-2,4-diazabutanal and methylenedinitramine were not measured during these column experiments.
Anion concentrations in 0.2 µm filtered samples were measured by ion chromatography (EPA Method 300.0). Alkalinity (as CaCO3) was measured according to EPA Method 310.1. DO was measured using visual colorimetric CHEMetric tubes (Midland, VA, USA), and pH was measured using a standard laboratory probe.
qPCR analysis
qPCR to enumerate the three RDX degrading strains was performed on DNA extracted from column effluent samples (2.5 mL) using the Qiagen DNeasy Tissue Kit (Qiagen, Valencia, CA) according to the manufacturer’s instructions for gram positive bacteria. DNA was extracted from sediment samples using the MO BIO PowerSoil® DNA Extraction Kit (MO BIO Laboratories, Carlsbad, CA) according to the manufacturer’s instructions using bead beating to lyse cells. The DNA extracts from three replicate sediment samples (0.35–0.93 g dry weight) per location were pooled by ethanol precipitation. All qPCR amplification reactions were performed using Applied Biosystems 7900HT Fast Real‐time PCR system (Foster City, CA). qPCR was performed in 20 μL reaction volumes in 384‐well optically clear plates. Analysis of the xenB gene used a SYBR Green PCR Master Mix, 300 nM of the respective primers (Table S4), and 1 μL of template DNA. Thermal cycler conditions were 95 °C for 10 min; then 40 cycles of 95 °C for 15 s; 60 °C for 60 s; followed by a final dissociation stage. Analysis of the 16S rRNA and xplA genes used a Quantitect PCR Probe Mix, 300 nM of the respective primers (Table S4), 200 nM of the respective TaqMan probe and 1 μL of template DNA. Thermal cycler conditions were 95 °C for 12 min; then 40 cycles of 95 °C for 30 s; 50 °C for 60 s; and 72 °C for 20 s. Standard curves for each qPCR assay were obtained from serial dilutions of genomic DNA isolated from strain I-C (16S rRNA and xenB), or an xplA containing plasmid (pET11a, Celtek Genes, Franklin, TN).
Data analysis
Zero-order RDX degradation rates during different phases of the column experiment were estimated using CXTFIT (Toride et al. 1995), which accounts for advection and dispersion within the column (based on the bromide curve data). Rates were also calculated by simple linear curve fitting of the effluent RDX concentration versus time data. In addition, the mass of RDX degraded per unit of added fructose was calculated by integrating the area of sustained decreased effluent RDX concentrations after each addition of fructose. RDX was assumed to be degrading in response to fructose addition during any period in which the effluent C/C0 was 0.05 lower than the C/C0 measured before the fructose addition.
Results and discussion
Column transport experiments
During the column experiment, the influent DO concentration remained at 7 mg L−1, the effluent DO concentration averaged 3.4 ± 0.8 mg L−1 (n = 111), and only minor decreases in effluent DO concentration (0.5–1 mg L−1) were observed upon fructose addition. No significant changes in influent or effluent nitrate, sulfate, or alkalinity were observed. Based on these data, the column system remained aerobic during the entire experiment. The influent and effluent pH averaged 8.1 ± 0.1 and 7.9 ± 0.1 SU, respectively (n = 77).
An overview of the RDX concentrations in the effluent of the column (C), relative to the influent concentration (C0) over the duration of the entire experiment is presented in Fig. 1. No inherent RDX degradation was observed in the absence of carbon amendment (Phase 1). The amendment of the column with fructose three separate times before bioaugmentation (Phase 2) did not result in any significant RDX degradation (Fig. S5). The apparent zero-order RDX degradation rates based on linear fits to the change in effluent RDX concentration versus time during the Phase 2 fructose additions was 0.008 ± 0.004 day−1 (n = 3). Zero-order rates estimated using CXTFIT were essentially identical to the simple linear curve fits due to the relatively short column residence time and small dispersivity of the packed sediment in the column, so only the linear fitting results are reported. RDX removal per unit of fructose added during Phase 2 yielded values of 0.07, 0.11, and 0.02 mg RDX mg fructose−1 for the first, second, and third fructose additions, respectively (average 0.07 ± 0.04). These data indicate that the indigenous microbial community in the aquifer sediment prior to bioaugmentation had very low densities of bacterial strains able to aerobically degrade RDX even when supplied with a labile carbon source, and are in agreement with previous testing conducted at the UMCD site (Michalsen et al. 2013).
RDX concentrations (squares) as C/C0 in the effluent of the repacked column. Phases are designated with numbers above the plot and correspond to: Phase 1 before biostimulation, Phase 2 biostimulation with fructose before bioaugmentation, Phase 3 bioaugmentation, and Phase 4 biostimulation with fructose after bioaugmentation. Start and end of the fructose additions are indicated by the dashed vertical lines. Start and end of bioaugmentation is indicated by the solid vertical lines. 1 PV is equivalent to 1 day, except during the Phase 3 cell injection, when 1 PV = 0.1 days
The bioaugmentation of the column occurred during Phase 3. Beginning with the cell injection, the RDX concentration in the effluent decreased to about 45 % of the influent concentration, and then slowly increased to 90 % of the influent concentration over a period of about 16 days (Fig. 2a). The apparent zero-order RDX degradation rate during the injection period was 0.14 day−1. Although the cells were washed and no fructose was added with the injected cells, appreciable RDX degradation occurred during the injection. It is possible that this RDX degradation was supported by either carbon coming from dead cells or extracellular polymers in the inoculum. This phenomena was also observed in a parallel column experiment (Fig. S6).
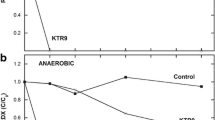
a Relative breakthrough curves of RDX (squares), cells (as OD600, diamonds), and conservative tracer (solid lines) during bioaugmentation (Phase 3) of the repacked column. b Absolute cell concentrations of the RDX degrading cells (KTR9, diamonds; RHA1, triangles; I-C, circles). c Analysis of the cell breakthrough curve during column bioaugmentation based on qPCR of target genes (16S gene, diamonds; xplA, triangles; xenB, circles). Start and end of the bioaugmentation is indicated by the dashed vertical lines. 1 PV is equivalent to 1 day, except during the Phase 3 cell injection, when 1 PV = 0.1 days
The percent of the injected cells observed in the effluent during the bioaugmentation was small (1.2, 0.01, and 8.1 % for KTR9, RHA1, and I-C, respectively). This was not totally unexpected, given that 99+ % of KTR9 and RHA1 cells, and 96 % of I-C cells, adhered to UMCD sediment in a standardized adhesion assay (data not shown). The peak cell concentrations detected in the effluent averaged 106–107 CFU mL−1 (Fig. 2b), indicating that some cells were transported through the aquifer column. The peak cell concentrations, as indicated by the OD600 measurements of the effluent, also eluted before the peak concentration of the conservative bromide tracer, indicating pore exclusion effects were occurring (i.e., cells excluded from some small pores accessible to Br−; Dong et al. 2002; Ginn 2002) (Fig. 2a).
Periodic analysis of effluent samples showed that KTR9 and I-C cultures were present at 102–103 CFU mL−1 for approximately the next 60 days after the peak cell concentration eluted, and then fell below the detection limit of 10 CFU mL−1, while concentrations of RHA1 fell to below the detection limit within 1 PV of the main cell peak (data not shown). Analysis of the effluent using qPCR yielded similar breakthrough curves to the plate counts for the three strains (Fig. 2c).
There is only one previous report examining transport of aerobic RDX degraders through saturated porous media, in which Rhodococcus DN22 was pumped through very small columns packed with clean sand (Priestley et al. 2006). However, the experimental design of that work does not allow a meaningful comparison with the present results. Overall, the extent of transport and injected cell recoveries are not inconsistent with previous column transport results with other types of bacteria or bioaugmentation cultures (e.g., Streger et al. 2002; Stumpp et al. 2011). In the coarse-grained UMCD aquifer, it is anticipated that some fraction of any bioaugmentation culture will be transported a reasonable distance.
In response to the addition of fructose (Phase 4), effluent RDX concentrations quickly decreased to approximately 35 % of the influent concentration, then increased back to a C/C0 of 0.9 over the course of 12 days after fructose addition ceased (Fig. 3). The apparent zero-order RDX degradation rate in response to the fructose addition was 0.08 day−1, or approximately 10-fold higher than the RDX degradation rate in response to fructose before bioaugmentation (Phase 2). The RDX removal per unit of fructose during Phase 4 was calculated to be 1.82 mg RDX mg fructose−1, which was approximately 26-fold higher than before bioaugmentation (Phase 2).
The data demonstrated that the cells remained viable and active (e.g., able to degrade RDX in response to fructose addition) for at least 3 months after injection. Similar sustained RDX degradation capacity was observed in a parallel column experiment, in which rapid RDX degradation was observed in response to fructose even after a 4-month period without any fructose addition (Fig. S6). Previous observations with KTR9 have demonstrated that labile nitrogen (e.g., \({\text{NO}}_{2}^{ - } ,\;{\text{NO}}_{3}^{ - } ,\;{\text{NH}}_{4}^{ + }\)) inhibits RDX degradation (Indest et al. 2010), and further, that nitrogen starvation conditions induce the expression of the RDX degradative gene xplA and leads to rapid RDX degradation (Indest et al. 2010, 2013; Jung et al. 2011). These previous observations with KTR9 support the sustained RDX degradation capacity of the bioaugmentation culture during the column experiments, in which the AGW contained no \({\text{NH}}_{4}^{ + }\) or \({\text{NO}}_{2}^{ - } .\) It also appears that the presence of approximately 40 mg L−1 \({\text{NO}}_{3}^{ - }\) in the AGW did not completely inhibit RDX degradation. The presence of relatively high nitrate concentrations may have resulted in lower RDX degradation rates than if it had been absent. As these conditions are typical not only at UMCD, but at many other contaminated sites, the column experiment results indicate that the aerobic RDX degrading strains added during bioaugmentation will possess sustained RDX degradation activity in situ.
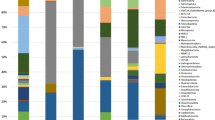
The distribution of retained cells in the repacked column is shown in Fig. 4. The cell density was greater at the influent end of the column than the effluent end of the column (as might be expected as the result of straining and attachment of the cells), but the concentration of KTR9 + RHA1 reached ~105 g−1 dry soil 30 cm into the column based on gene copy numbers for the xplA gene. Concentrations of I-C were much lower throughout the column based on gene copy numbers for xenB. It is likely that the addition of fructose to the column increased the levels of indigenous microorganisms, especially near the influent end of the column, given the trends in total bacteria based on the 16S rRNA gene copy values (background 16S gene copy number was ~104–105 g−1 dry sediment). The relative retained cell distribution throughout the column was comparable to previous column cell transport experiments using both intact sediment cores (e.g., Dong et al. 2002; Fuller et al. 2000) and repacked sediment columns (e.g., Streger et al. 2002). Overall, the data suggest that KTR9 and RHA1 were distributed and survived in the aquifer matrix, whereas distribution and/or survival of I-C was relatively poor by comparison.
Field-scale bioaugmentation culture evaluation
As part of the evaluation of bioaugmentation for RDX remediation, it is critical to confirm that large volumes of degradative strains can be produced, stored, and deployed to the field without loss of cell viability or activity. Similar studies have been performed for dechlorination consortia, which are now widely used for anaerobic bioaugmentation for chlorinated solvent remediation (Steffan and Vainberg 2013; Vainberg et al. 2009). After pilot-scale fermentation and concentration, the cell density (relative to the initial OD550) of all three of the bioaugmentation cultures remained constant for at least 90 days at 4 °C for KTR9 and I-C, and 60 days for RHA1 (Fig. S7). The data for culturable cells (as CFU mL−1) was similar to the OD550 data (Fig. 5). Some decrease in cultivable cells was observed for all three strains incubated at 4 °C during the first 14 days, followed by a period of stable CFU counts for up to 90 days for KTR9 and I-C, and 60 days for RHA1 (Fig. 5). CFU counts on R2A agar were the same as on LB + kanamycin agar for KTR9 and RHA1 (data not shown). OD-normalized RDX degradation potential (defined as percent of initial RDX degraded in 24 h divided by the relative OD550) remained relatively stable for KTR9, while some decrease was observed for RHA1 after 30 days (Fig. 6). This is in agreement with nitrogen starvation inducing the RDX degrading genes in these strains. The RDX degradation assay was not performed under suboxic conditions for strain I-C, but some activity was observed by this culture during the experiment (data not shown). For RHA1 and I-C, incubation at 37 °C resulted in rapid decreases in OD550, loss of viability, and reduced RDX degradation potential. KTR9 incubated at 37 °C showed similar patterns in viability and RDX degradation to the other two strains, but less of a reduction in OD550, possibly indicating that KTR9 cells were dying, but were not lysing. These results clearly indicate that large volumes of high density cultures could be produced in advance of a field application and stored at 4 °C for at least 2 months without significant loss of cell density, and more importantly, RDX degradation activity.
Conclusions
The results presented herein are the first demonstrating transport of aerobic RDX degraders in repacked saturated site sediments. The selected strains were transported through repacked UMCD sediment columns and effectively colonized the porous media, retaining RDX degradation activity for at least 3 months (and for at least 6 months in a parallel experiment), as evidenced by rapid RDX degradation upon the addition of fructose. Sustained RDX-degrading activity following periods of starvation in situ is encouraging for future field-scale applications of this remediation approach because reduced substrate injection quantities and durations translate to reduced materials and field labor costs and a greater likelihood of maintaining bulk aerobic conditions in the aquifer, which is desirable for these RDX-degrading strains. Furthermore, the fact that nitrogen starvation strongly induces xplA gene expression, but that the presence of some labile nitrogen is not completely inhibitory to RDX degradation, indicates that many contaminated aquifers should be amenable to bioaugmentation. High degradation efficiency could translate into inoculation with fewer cells or reduced treatment time schedules. Additionally, this is the first reported production of aerobic RDX degraders at a scale that is relevant for field application. Further efforts are currently underway to evaluate the transport, longevity, and in situ RDX degradation activity of these strains in the field at UMCD.
References
Adrian N, Sutherland K (1999) RDX biodegradation by a methanogenic enrichment culture obtained from an explosives manufacturing wastewater treatment plant. U.S. Army Corps of Engineers, Construction Engineering Research Laboratories. Report# 99/15
Arnett CM, Adrian NR (2009) Cosubstrate independent mineralization of hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) by a Desulfovibrio species under anaerobic conditions. Biodegradation 20:15–26
Bernstein A, Adar E, Nejidat A, Ronen Z (2011) Isolation and characterization of RDX-degrading Rhodococcus species from a contaminated aquifer. Biodegradation 22:997–1005
Brannon J, Myers T (1997) Review of fate and transport process of explosives. Army Corps of Engineers, Waterways Experiment Station. Report# IRRP-97-2
Clausen J, Robb J, Curry D, Korte N (2004) A case study of contamination on military ranges: Camp Edwards, Massachusetts, USA. Environ Pollut 129:13–21
DeFlaun M, Tanzer A, McAteer A, Marshall B, Levy S (1990) Development of an adhesion assay and characterization of an adhesion-deficient mutant of Pseudomonas fluorescens. Appl Environ Microbiol 56:112–119
Dong H, Rothmel RK, Onstott TC, Fuller ME, DeFlaun MF, Dunlap R, Fletcher M (2002) Simultaneous transport of two bacterial strains in intact cores from Oyster, Virginia: biological effects and numerical modeling. Appl Environ Microbiol 68:2120–2132
Fournier D, Halasz A, Spain J, Fiurasek P, Hawari J (2002) Determination of key metabolites during biodegradation of hexahydro-1,3,5-trinitro-1,3,5-triazine with Rhodococcus sp. strain DN22. Appl Environ Microbiol 68:166–172
Fuller ME, Dong H, Mailloux BJ, Onstott TC, DeFlaun MF (2000) Examining bacterial transport in intact cores from Oyster, Virginia: effect of sedimentary facies type on bacterial breakthrough and retention. Water Resour Res 36:2417–2431
Fuller ME, McClay K, Hawari J, Paquet L, Malone TE, Fox BG, Steffan RJ (2009) Transformation of RDX and other energetic compounds by xenobiotic reductases XenA and XenB. Appl Microbiol Biotechnol 84:535–544
Fuller ME, Hawari J, Perreault N (2010a) Microaerophilic degradation of hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) by three Rhodococcus strains. Lett Appl Microbiol 51:313–318
Fuller ME, McClay K, Higham M, Hatzinger PB, Steffan RJ (2010b) Hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) bioremediation in groundwater: are known RDX-degrading bacteria the dominant players? Bioremediat J 14:121–134
Ginn TR (2002) A travel time approach to exclusion on transport in porous media. Water Resour Res 38:12-1–12-11. doi:10.1029/2001WR000865
Hareland WA, Crawford RL, Chapman PJ, Dagley S (1975) Metabolic function and properties of 4-hydroxyphenylacetic acid 1-hydroxylase from Pseudomonas acidovorans. J Bacteriol 121:272–285
Hatzinger PB, Lippincott D (2012) In situ bioremediation of energetic compounds in groundwater. Environmental Security Technology Certification Program (ESTCP). Report# ER-200425
Hawari J, Halasz A, Beaudet S, Paquet L, Ampleman G, Thiboutot S (2001) Biotransformation routes of octahydro-1,3,5,7-tetranitro-1,3,5,7-tetrazocine by municipal anaerobic sludge. Environ Sci Technol 35:70–75
Indest KJ, Jung CM, Chen H-P, Hancock D, Florizone C, Eltis LD, Crocker FH (2010) Functional characterization of pGKT2, a 182-kilobase plasmid containing the xplAB genes, which are involved in the degradation of hexahydro-1,3,5-trinitro-1,3,5-triazine by Gordonia sp. strain KTR9. Appl Environ Microbiol 76:6329–6337. doi:10.1128/aem.01217-10
Indest KJ, Hancock DE, Jung CM, Eberly JO, Mohn WW, Eltis LD, Crocker FH (2013) Role of nitrogen limitation in transformation of RDX (hexahydro-1,3,5-trinitro-1,3,5-triazine) by Gordonia sp. strain KTR9. Appl Environ Microbiol 79:1746–1750. doi:10.1128/aem.03905-12
Jung CM, Crocker FH, Eberly JO, Indest KJ (2011) Horizontal gene transfer (HGT) as a mechanism of disseminating RDX-degrading activity among Actinomycete bacteria. J Appl Microbiol 110:1449–1459
Li Q, Logan BE (1999) Enhancing bacterial transport for bioaugmentation of aquifers using low ionic strength solutions and surfactants. Water Res 33:1090–1100. doi:10.1016/S0043-1354(98)00291-7
Michalsen MM, Weiss R, King A, Gent D, Medina VF, Istok JD (2013) Push–pull tests for estimating RDX and TNT degradation rates in groundwater. Groundw Monit Remediat 33:61–68. doi:10.1111/gwmr.12016
Newell C (2008) Treatment of RDX & HMX plumes using mulch biowalls. Environmental Security Technology Certification Program (ESTCP). Report# ER-0426
Pennington JC et al (2001) Distribution and fate of energetics on DoD test and training ranges: Interim Report 1. U.S. Army Engineer Research and Development Center. Report# ERDC TR-01-13
Pennington JC et al (2002) Distribution and fate of energetics on DoD test and training ranges: Interim Report 2. U.S. Army Engineer Research and Development Center. Report# ERDC TR-02-8
Priestley JT, Coleman NV, Duxbury T (2006) Growth rate and nutrient limitation affect the transport of Rhodococcus sp. strain DN22 through sand. Biodegradation 17:571–576
Ronen Z, Yanovich Y, Goldin R, Adar E (2008) Metabolism of the explosive hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) in a contaminated vadose zone. Chemosphere 73:1492–1498
Schaefer CE, Fuller ME, Condee CW, Lowey JM, Hatzinger PB (2007) Comparison of biotic and abiotic treatment approaches for co-mingled perchlorate, nitrate, and nitramine explosives in groundwater. J Contam Hydrol 89:231–250
Seth-Smith HMB, Rosser SJ, Basran A, Travis ER, Dabbs ER, Nicklin S, Bruce NC (2002) Cloning, sequencing, and characterization of the hexahydro-1,3,5-trinitro-1,3,5-triazine degradation gene cluster from Rhodococcus rhodochrous. Appl Environ Microbiol 68:4764–4771
Seth-Smith HMB, Edwards J, Rosser SJ, Rathbone DA, Bruce NC (2008) The explosive-degrading cytochrome P450 system is highly conserved among strains of Rhodococcus spp. Appl Environ Microbiol 74:4550–4552. doi:10.1128/aem.00391-08
Shaw JC, Bramhill B, Wardlaw NC, Costerton JW (1985) Bacterial fouling in a model core system. Appl Environ Microbiol 49:693–701
Steffan RJ, Vainberg S (2013) Production and handling of Dehalococcoides bioaugmentation cultures. In: Stroo HF, Leeson A, Ward CH (eds) Bioaugmentation for groundwater remediation. SERDP ESTCP Environmental Remediation Technology. Springer, New York, pp 89–115. doi:10.1007/978-1-4614-4115-1_3
Streger SH, Vainberg S, Dong H, Hatzinger PB (2002) Enhancing transport of Hydrogenophaga flava ENV735 for bioaugmentation of aquifers contaminated with methyl tert-butyl ether. Appl Environ Microbiol 68:5571–5579. doi:10.1128/aem.68.11.5571-5579.2002
Stumpp C, Lawrence JR, Hendry MJ, Maloszewski P (2011) Transport and bacterial interactions of three bacterial strains in saturated column experiments. Environ Sci Technol 45:2116–2123. doi:10.1021/es103569u
Thompson KT, Crocker FH, Fredrickson HL (2005) Mineralization of the cyclic nitramine explosive hexahydro-1,3,5-trinitro-1,3,5-triazine by Gordonia and Williamsia spp. Appl Environ Microbiol 71:8265–8272
Toride N, Leij FJ, van Genuchten MT (1995) The CXTFIT code for estimating transport parameters from laboratory or field tracer experiments, Version 2.0. U.S. Salinity Laboratory, USDA, ARS, Riverside
Vainberg S, Condee CW, Steffan RJ (2009) Large-scale production of bacterial consortia for remediation of chlorinated solvent-contaminated groundwater. J Ind Microbiol Biotechnol 36:1189–1197. doi:10.1007/s10295-009-0600-5
Wade R, Davis JL, Wani AH, Felt D (2010) Biologically active zone enhancement (BAZE) for in situ RDX degradation in ground water. Environmental Security Technology Certification Program (ESTCP). Report# ER-0110
Yamamoto H, Morley MC, Speitel GE Jr, Clausen J (2004) Fate and transport of high explosives in a sandy soil: adsorption and desorption. Soil Sediment Contam 13:459–477
Zhang C, Hughes JB (2003) Biodegradation pathways of hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) by Clostridium acetobutylicum cell-free extract. Chemosphere 50:665–671
Zhao J-S, Paquet L, Halasz A, Manno D, Hawari J (2004) Metabolism of octahydro-1,3,5,7-tetranitro-1,3,5,7-tetrazocine by Clostridium bifermentans strain HAW-1 and several other H2-producing fermentative anaerobic bacteria. FEMS Microbiol Lett 237:65–72
Acknowledgments
This project was supported by the Environmental Security Technology Certification Program (ESTCP) under contract W912DW-12-C-0029. Views, opinions, and/or findings contained in this report are those of the author(s) and should not be construed as an official Department of Defense position or decision unless so designated by other official documentation.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fuller, M.E., Hatzinger, P.B., Condee, C.W. et al. Laboratory evaluation of bioaugmentation for aerobic treatment of RDX in groundwater. Biodegradation 26, 77–89 (2015). https://doi.org/10.1007/s10532-014-9717-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-014-9717-y