Abstract
Buprofezin is a widely used insecticide that has caused environmental pollution in many areas. However, biodegradation of buprofezin by pure cultures has not been extensively studied, and the transformation pathway of buprofezin remains unclear. In this paper, a buprofezin co-metabolizing strain of DFS35-4 was isolated from a buprofezin-polluted soil in China. Strain DFS35-4 was preliminarily identified as Pseudomonas sp. based on its morphological, physiological, and biochemical properties, as well as 16S rRNA gene analysis. In the presence of 2.0 g l−1 sodium citrate, strain DFS35-4 degraded over 70% of 50 mg l−1 buprofezin in 3 days. Strain DFS35-4 efficiently degraded buprofezin in the pH range of 5.0–10.0 and at temperatures between 20 and 30°C. Three metabolites, 2-imino-5-phenyl-3-(propan-2-yl)-1,3,5-thiadiazinan-4-one, 2-imino-5-phenyl-1,3,5-thiadiazinan-4-one, and methyl(phenyl) carbamic acid, were identified during the degradation of buprofezin using gas chromatography–mass spectrometry (GC–MS) and tandem mass spectrometry (MS/MS). A partial transformation pathway of buprofezin in Pseudomonas sp. DFS35-4 was proposed based on these metabolites.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Buprofezin (2-tert-butylimino-3-isopropyl-5-phenyl-1,3,5-thiadiazinan-4-one), a broad-spectrum insect growth regulator that acts on insects after contact or ingestion by inhibiting chitin biosynthesis and integumentary cuticle deposition (Das et al. 2004; Izawa et al. 1985; Prabhaker and Toscano 2007), was developed by Nihon Nohyaku in 1981. Buprofezin is widely used for its high efficiency in controlling homopteran pests, such as whiteflies (Trialeurodes vaporariorum and Bemisia tabaci), planthoppers (Nilaparvata lugens and Sogatella furcifera), and citrus scales (Aonidiella aurantii and Saissetia oleae) (Atsushi et al. 1996; Hegazy et al. 1990; Yang and Yang 2007).
Because of its high hydrophobicity (log P = 4.31), buprofezin easily adsorbs to soil particles, which may contribute to its persistence in soil (Funayama et al. 1986). Measured half-lives of buprofezin in aerobic soils were on the order of 26–220 days with the rate of degradation related to the soil microbial status. Under aerobic field conditions, the half-lives of buprofezin were on the order of 50–70 days and about 36–104 days under flooded field conditions. The widespread application of buprofezin has resulted in buprofezin residues in many sites. It has been reported that buprofezin is the only significant pesticide residue in tomato, lettuce, and cotton. Buprofezin residues in citrus fruit from trials conducted in Australia, New Zealand, Italy, Spain, and Portugal ranged from 0.05 to 0.69 mg kg−1 for oranges, lemons, and mandarins (Valverde-Garcia et al. 1993). In 2006, when buprofezin was recommended for application on grapes, frequent detection of buprofezin residues was recorded on residue monitoring (Oulkar et al. 2009). In 2008, buprofezin concentrations of 0.181–0.212 mg kg−1 were detected in a Chinese paddy soil in which buprofezin was repeatedly applied.
Even though buprofezin was not highly toxic to mammals (the acute rat oral LD50 was 1,635 mg kg−1 in males), the cleanup of buprofezin residues in the environment is of great concern. Microorganisms play key roles in the detoxification of xenobiotics (Singh et al. 2006), and the use of microorganisms for bioremediation of contaminated sites is considered to be a high-efficient, safe, and cost-effective method. Many studies have been carried out to show that the degradation of buprofezin was mostly due to soil microorganisms. However, to the best of our knowledge, pure cultures capable of degrading buprofezin have not been reported, and the degradation pathway of buprofezin in microorganisms remains unclear. In this study, DFS35-4, a strain of Pseudomonas sp. capable of co-metabolizing buprofezin in the presence of other carbon sources, was isolated from a long-term buprofezin-polluted soil in China. The factors that influence the degradation activities of strain DFS35-4 were evaluated and a buprofezin-transforming pathway was also proposed.
Materials and methods
Chemicals and media
Buprofezin (99% purity) was purchased from Sigma-Aldrich. The antibiotic chloramphenicol was purchased from Amresco. High-performance liquid chromatography (HPLC) grade methanol, acetone, ethyl acetate, and acetonitrile were purchased from the Shanghai Chemical Reagent Co., Ltd, China. All other reagents used in this study were analytical-reagent grade.
Luria–Bertani (LB) medium and mineral salts medium (MM; 1.0 g NH4NO3, 1.6 g K2HPO4, 0.5 g KH2PO4, 0.2 g MgSO4, and 1.0 g NaCl per liter water at pH 7.0) were used in this study. MM supplemented with 50 mg l−1 buprofezin was defined as BMM and MM supplemented with 2.0 g l−1 sodium citrate was defined as SMM. Solid medium plates were prepared by adding 2.0% (w/v) agar.
Strain isolation and characterization
Buprofezin-polluted soil samples were collected from a buprofezin manufacture factory in Jiangsu, China. Soil samples (5.0 g) were added to a 250 ml flask with 100 ml BMM, and incubated at 30°C for 1 week in a rotary shaker at 150 rpm. Five milliliters of the culture, showing degradation of buprofezin, was transferred to 100 ml of fresh BMM (with 1% LB, w/v). This procedure was repeated three times. The final culture which still shows the degradation ability of buprofezin was diluted and plated onto BMM agars (with 1% LB, w/v). The different colonies that formed were isolated, and then tested for their buprofezin-degrading capabilities. One strain designated DFS35-4, which possessed the highest degradation ability, was purified and selected for further investigation.
The isolated strain was identified based on its morphological, physiological, and biochemical properties (with reference to Bergey’s Manual of Determinative Bacteriology), combined with 16S rRNA gene sequence analysis. The DNA of strain DFS35-4 was extracted by high-salt-concentration precipitation (Miller et al. 1988). The 16S rRNA gene was amplified by a previously described PCR method (Li et al. 2010). The nucleotide sequence coding for the 16S rRNA of strain DFS35-4 (1,410 bp) was deposited in the GenBank database under the accession number FJ544978. Alignment of the different 16S rRNA gene sequences from GenBank database was performed using Clustal X 1.8.3 with default settings (Thompson et al. 1997). Phylogeny was analyzed with MEGA version 3.0 software and distance was calculated using the Kimura 2 parameter distance model. An unrooted tree was built by the neighbor joining method (Saitou and Nei 1987). The dataset was bootstrapped 1,000 times.
Degradation of buprofezin by strain DFS35-4 in culture
In our previous study, we found that strain DFS35-4 was resistant to chloramphenicol. In order to prevent the contamination of strain DFS35-4 by other microorganisms, DFS35-4 cells were pre-cultured in LB containing 30 mg l−1 chloramphenicol. Cells were harvested by centrifugation at 6,000×g for 5 min and washed three times with MM. Then, OD 600 nm of cell density was adjusted to 1.0. For all experiments, cells were inoculated at 3% (v/v) into cultures and incubated at 30°C on a shaker at 150 rpm unless otherwise stated. All of the treatments were performed in triplicate.
To investigate the effect of other carbon source on buprofezin biodegradation, 2.0 g l−1 of glucose, sodium citrate, sodium acetate, maltose, galactose, starch, or sucrose was added to BMM. To investigate the effect of the initial pH value on buprofezin biodegradation, the pH value of the medium was adjusted to 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, or 10.0. To investigate the effect of temperature on buprofezin biodegradation at pH 7.0, the culture was incubated at 20, 25, 30, or 37°C. To investigate the effect of the initial concentration of buprofezin on its biodegradation, SMM containing 10, 30, 50, 80, 100, or 200 mg l−1 buprofezin was used.
Analysis of buprofezin
Buprofezin in the culture was extracted with an equal volume of dichloromethane. The extract was dried over anhydrous Na2SO4 and evaporated using a vacuum rotary evaporator at room temperature. Residual was dissolved in an equal volume of methanol. All samples were analyzed by HPLC (600 controller, Rheodyne 7725i manual injector and 2487 Dual λ Absorbance Detector; Waters Co., Milford, MA). Kromasil 100-5 C18 stationary phase was used in the separation column (4.6 mm internal diameter and 25 cm length). The mobile phase was methanol:water (80:20, v/v), and the flow rate was 0.8 ml min−1. Buprofezin was detected at 245 nm (Xu 2008).
Identification of the metabolites during buprofezin biodegradation
For metabolite identification, DFS35-4 cells were inoculated into 200 ml liquid SMM with 50 mg l−1 buprofezin. A negative control was similarly prepared except that the inoculated cells were heat-killed. Twenty milliliter samples of cultures were collected in 5 days and extracted with an equal volume of dichloromethane. The residual aqueous phase was extracted with an equal volume of ethyl acetate, acidified to pH 2.0, and extracted with an equal volume of ethyl acetate (Li et al. 2010). Extracts from dichloromethane and ethyl acetate were mixed. This mixture was evaporated as described above and re-dissolved in 2 ml of methanol.
Two microliters of the resulting solution was first subjected to gas chromatography–mass spectrometry (GC–MS). GC–MS analysis was performed in electron ionization (EI) mode (70 eV) with a Finnigan GC, equipped with a MS detector. A Finnigan capillary column (15 m length × 0.5 mm inside diameter × 0.25 μm film thickness) was used with the following temperature program: 50°C for 1 min, increased to 240°C at 20°C min−1 and held for 20 min. Helium was used as the carrier gas at a constant flow of 1.0 ml min−1. For MS analysis, the sample was analyzed in split mode (1:20) at an injection temperature of 220°C and an EI source temperature of 250°C, and scanned in the mass range from 30 m/z (mass to charge ratio) to 650 m/z (Nguyen et al. 2010; Paolo et al. 1998).
The metabolites produced during buprofezin biodegradation were further identified by tandem mass spectrometry (MS/MS) using a Finnigan TSQ Quantum Ultra AM instrument (Thermal, USA). The metabolites were confirmed by standard MS using positive electrospray ionization and scanned in the normal mass range from 100 to 400 m/z. Characteristic fragment ions were detected with second-order MS.
Results
Isolation and characterization of buprofezin-degrading strain DFS35-4
Several bacterial strains capable of degrading buprofezin were isolated from the buprofezin-polluted soil samples. Strain DFS35-4, which degraded over 70% of 50 mg l−1 buprofezin in SMM in 3 days, was selected for further study.
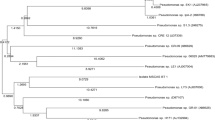
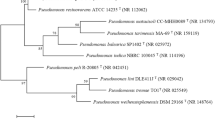
Strain DFS35-4 is a Gram-negative, rod-shaped (0.5–1.0 μm × 1.5–5.0 μm) motile aerobe. It was oxidase, catalase, and arginine dihydrolase positive, and was able to use l-serine, malonate, and glycolate. Strain DFS35-4 was positive for nitrate denitrification. The phylogenetic tree of the 16S rRNA gene sequences is shown in Fig. 1. Stain DFS35-4 was related to the Pseudomonas species lineage and closely clustered with the type strain Pseudomonas citronellolis T, with sequence similarity score of 99.4%. Based on the above phenotypic characteristics and phylogenetic analysis, strain DFS35-4 was primarily identified as Pseudomonas species.
Phylogenetic analysis of strain DFS35-4 and related species by the neighbor joining method. Bootstrap values (%) are indicated at the nodes, the scale bars represent 0.005 substitutions per nucleotide position. The GenBank accession number for each microorganism is shown in parentheses after the species name
Degradation of buprofezin by strain DFS35-4
In SMM, strain DFS35-4 degraded over 70% of 50 mg l−1 buprofezin in 3 days, and degraded over 85% buprofezin in 5 days. However, in MM, only 10% of the total buprofezin was degraded in 3 days (Fig. 2). These results showed that the degradation of buprofezin by strain DFS35-4 was greatly affected by other carbon sources. When strain DFS35-4 grew with sodium citrate as the carbon source, it simultaneously co-metabolized buprofezin. It was found that other carbon sources, such as starch, sodium acetate, galactose, maltose, and sucrose could also significantly promote the degradation of buprofezin (Fig. 2).
Strain DFS35-4 degraded buprofezin at a wide range of concentrations, including high concentrations. For instance, over 60% of the total buprofezin at a concentration of 200 mg l−1 was degraded in 3 days. Strain DFS35-4 efficiently degraded buprofezin at temperatures between 20 and 30°C and in pH from 5.0 to 10.0. The optimum degradation temperature and pH were 30°C and 9.0, respectively.
Identification of the metabolites during buprofezin biodegradation
GC–MS was first applied to identify the metabolites produced during buprofezin biodegradation. Four compounds were detected in the total ion chromatography (TIC) profiles (Fig. 3a). Compounds with retention times of 10.27, 3.75, and 2.65 min were designated as compounds A, B, and C, respectively. It was found a compound with a retention time of 4.79 min did not match any compound in the National Institute of Standards and Technology (NIST) library, and was designated as an unknown compound (Fig. 3a).
According to its mass spectra and the NIST library identification program, compound A was found to buprofezin (Fig. 3b). The molecular ion (M+) peak of compound A was 305 m/z with characteristic ions at 276.99 m/z (M+–2CH2), 247.94 m/z (M+–C(CH3)3), 215.94 m/z (M+–C(CH3)3–S), 189.94 m/z (M+–C(CH3)3–S–CN), 171.95 m/z (M+–C(CH3)3–S–CN–H2O), 130.94 m/z (M+–C(CH3)3–S–CN–H2O–CH(CH2)2), 104.94 m/z (M+–C(CH3)3–S–CN–H2O–CH(CH2)2–C–N), 118.80 m/z (M+ –N(CH(CH3)2)–CN (C(CH3)3)–CH2S), and 42.92 m/z (M+–N(CH(CH3)2)–CN(C(CH3)3)–CH2S–C6H4). Compound B was identified by its M+ peak of 207 m/z. The characteristic fragment ion peaks at 192.81 m/z (M+–N) and 132.87 m/z (M+–N–S–C–NH2) (Fig. 3c) allowed compound B to be identified as 2-imino-5-phenyl-1,3,5-thiadiazinan-4-one. Compound C was identified as methyl(phenyl) carbamic acid. The (M+) peak of compound C was 151 m/z with characteristic ions at 132.86 m/z (M+–H2O) and 118.86 m/z (M+–H2O–CH2) (Fig. 3d). All the products along with their characteristic ions of the mass spectra and retention times are summarized in Table 1.
The metabolites produced were further identified using MS/MS. In the first MS, prominent protonated molecular ions [M + H]+ occurred at 306, 250, 234, and 201 m/z (Fig. 4a). The peak at 306 m/z [M + H]+, corresponding to a (M+) of 305 m/z, was identified as buprofezin. This was in agreement with the GC/MS identification. The identity of the compounds corresponding to [M + H]+ peaks at 234 and 201 m/z were unknown. The compound corresponding to the [M + H]+ peak at 250 m/z was designated as compound D. The [M + H]+ peak for compound D at 250 m/z enabled the assignment of the (M+) at 249 m/z. The characteristic second-order MS fragment ion peaks at 207.93 m/z [M + H]+, 193.95 m/z [M + H]+, 133.97 m/z [M + H]+, 106.01 m/z [M + H]+, 86.03 m/z [M + K]+, and 43.20 m/z [M + H]+ are shown in Fig. 4b. These fragments were used to identify compound D as 2-imino-5-phenyl-3-(propan-2-yl)-1,3,5-thiadiazinan-4-one.
Discussion
The wide use of buprofezin has resulted in residues of buprofezin in the environment. Microbial degradation of buprofezin is a promising way to clean up the buprofezin-contaminated sites. However, microbial biodegradation of buprofezin by pure cultures has not been reported. In this paper, a buprofezin-degrading strain Pseudomonas sp. DFS35-4, which could degrade buprofezin in the presence of some other carbon sources, was isolated from the long term buprofezin-polluted soil. In SMM, the strain DFS35-4 could degrade over 70% of total buprofezin at the concentration of 50 mg l−1 in 3 days, and efficiently degrade buprofezin in a wide range of pH values and temperatures. These results illuminate that strain DFS35-4 has a great potential in the bioremediation of buprofezin-contaminated sites.
The proposed route for hydrolysis of buprofezin in water (pH 5) involves opening of the thiadiazinane ring to form thiobiuret followed by amide cleavage to produce isopropylphenylurea. Alternatively, the sulfur in thiobiuret may be replaced with oxygen to form biuret, followed by amide cleavage to produce isopropylphenylurea. However, no metabolites were identified during buprofezin degradation by a pure culture in previous studies and the degradation pathway in microorganisms remained unclear. In this study, three metabolites, 2-imino-5-phenyl-3-(propan-2-yl)-1,3,5-thiadiazinan-4-one, 2-imino-5-phenyl-1,3,5-thiadiazinan-4-one, and methyl(phenyl) carbamic acid, were detected and identified by GC–MS and MS/MS. Based on the structure of these metabolites, we proposed a partial transformation pathway of buprofezin in Pseudomonas sp. DFS35-4 (Fig. 5). The first step in the transformation pathway of buprofezin was the sequential loss of N-tert-butyl and N-isopropyl to form 2-imino-5-phenyl-3-(propan-2-yl)-1,3,5-thiadiazinan-4-one and 2-imino-5-phenyl-1,3,5-thiadiazinan-4-one. N-dealkylation is believed to result from a hydroxylation of the C–N carbon (α position), followed by decomposition of the unstable intermediate into the N-dealkylated product and the alkyl fragment as an aldehyde or ketone (Okazaki and Guengerich 1993; Nagy et al. 1995). We had tried to identify the extract structures of the leaving alkyl groups. It was found that acetone was not detected in our study and could not be used by our strain. However, it was found that both isopropanol and tert-butanol could be used by our strain for growth, indicating that the leaving alkyls might be isopropanol and tert-butanol, even though isopropanol and tert-butanol were not detected. The heterocyclic ring of the product was opened by hydrolysis and the phenyl urea methyl (phenyl) carbamic acid was formed from the presumed metabolite [(carbamimidoylsulfanyl) methyl] phenylcarbamic acid. This study offers useful information about the biodegradation of buprofezin and presents the transformation mechanism of buprofezin in microorganisms.
References
Atsushi K, Rikio Y, Takamichi K (1996) Effect of buprofezin on oviposition of brown planthopper, Nilaparvata lugens, at sub-lethal dose. J Pestic Sci 21:153–157
Das C, Roy S, Pal R, Kole RK, Chowdhury A (2004) Effect of pH on the persistence behavior of the insecticide buprofezin in water under laboratory conditions. Environ Contam Toxicol 72:307–311
Funayama S, Uchida M, Kanno H, Ysuchiya K (1986) Degradation of buprofezin in flooded and upland soils under laboratory conditions. J Pestic Sci 11:605–610
Hegazy G, Cock AD, Degheele D (1990) Ultrastructural changes in the cuticle of the greenhouse whitefly Trialeurodes vaporariorum, induced by the insect growth inhibitor, buprofezin. Entomol Exp Appl 57:299–302
Izawa Y, Uchida M, Sugimoto T, Asai T (1985) Inhibition of chitin synthesis by buprofezin analogs in relation to their activity controlling Nilaparvata lugens. Pestic Biochem Physiol 24:343–347
Li R, Zheng JW, Wang R, Song Y, Chen QM, Yang XJ, Li SP, Jiang JD (2010) Biochemical degradation pathway of dimethoate by Paracoccus sp. Lgjj-3 isolated from treatment wastewater. Int Biodeter Biodegrad 64:51–57
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acid Res 16:1215
Nagy I, Compernolle F, Ghys K, Vanderleyden J, Mot RD (1995) A single cytochrome P-450 system is involved in degradation of the herbicides EPTC (S-Ethyl Dipropylthiocarbamate) and atrazine by Rhodococcus sp. strain NI86/21. Appl Environ Microbiol 61:2056–2060
Nguyen TD, Lee KJ, Lee MH, Lee GH (2010) A multiresidue method for the determination 234 pesticides in Korean herbs using gas chromatography mass spectrometry. Microchem J 95:43–49
Okazaki O, Guengerich FP (1993) Evidence for specific base catalysis in N-dealkylation reactions catalyzed by cytochrome P450 and chloroperoxidase. J Biol Chem 268:1546–1552
Oulkar DP, Banerjee K, Patil SH, Upadhyay AK, Taware PB, Deshmukh MB, Adsule PG (2009) Degradation kinetics and safety evaluation of buprofezin residues in grape (Vitis vinifera L.) and three different soils of India. Pest Manag Sci 65:183–188
Paolo C, Alberto A, Vincenzo LG, Marinella M, Filippo MP, Franco C, Fabrizio D, Sandro N (1998) Determination of buprofezin, pyridaben, and tebufenpyrad residues by gas chromatography-mass-selective detection in clementine citrus. J Agric Food Chem 46:4255–4259
Prabhaker N, Toscano NC (2007) Toxicity of the insect growth regulators, buprofezin and pyriproxyfen, to the glassy-winged sharpshooter, Homalodisca coagulata say (Homoptera:Cicadellidae). Crop Prot 26:495–502
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Singh BK, Walker A, Wright DJ (2006) Bioremedial potential of fenamiphos and chlorpyrifos degrading isolates: influence of different environmental conditions. Soil Biol Biochem 38:2682–2693
Thompson JD, Gibson TJ, Plewniak F, Jeamougin F, Higgins DG (1997) The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Valverde-Garcia E, Gonzalez-Pradas R, Del Aguilera A (1993) Analysis of buprofezin residues in vegetables. Application to the degradation study of eggplant grown in a greenhouse. J Agric Food Chem 41:2319–2323
Xu Z (2008) Analysis of mixture of buprofezin and methidathion by HPLC. Pestic Sci Adm 29:14–16
Yang YT, Yang LF (2007) Control effects of chlorpyrifos and buprofezin against brown planthopper. Pestic Sci Adm 28:24–27
Acknowledgments
We gratefully acknowledge Dr. Weiyou Zhou of Nanjing Science and Technology University for excellent assistance in GC–MS analysis. This work was supported by grants from the Chinese National Natural Science Foundation (31070100), the Major Projects on Control and Rectification of Water Body Pollution (2009ZX07103-002), and the Key Technology R&D Program of Jiangsu Province (BE2009670).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, K., Liu, XM., Li, R. et al. Isolation of a buprofezin co-metabolizing strain of Pseudomonas sp. DFS35-4 and identification of the buprofezin transformation pathway. Biodegradation 22, 1135–1142 (2011). https://doi.org/10.1007/s10532-011-9469-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-011-9469-x