Abstract
Halogenated organic compounds constitute one of the largest and most diverse groups of chemicals in the environment. Many of these compounds are toxic, persistent and, as a result of their often limited biodegradability, tend to bioaccumulate in the environment. Dibromoneopentyl glycol (DBNPG) and tribromoneopentyl alcohol (TBNPA) are brominated flame retardants commonly used as additives during the manufacture of plastic polymers and as chemical intermediates in the synthesis of other flame retardants. Both are classified as not readily biodegradable. In this paper, we demonstrate the biodegradation of both DBNPG and TBNPA by a common bacterial consortium under aerobic conditions in enrichment cultures containing yeast extract. DBNPG and TBNPA biodegradation is accompanied by a release of bromide into the medium, due to a biological debromination reaction. Molecular analysis of the clone library PCR amplified 16S rRNA gene was used to characterize the bacterial consortium involved in the biodegradation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Halogenated organic compounds constitute one of the largest and most diverse groups of chemicals in the environment. While some of these compounds are generated by naturally occurring biotic and abiotic processes in the oceans and atmosphere (Gribble 2003), over the past 100 years the widespread use of halogen-based chemistry in industrial-scale chemical processes has introduced many additional man-made halogenated organic compounds into the environment (Jain et al. 2005). These compounds are widely used as pharmaceuticals, herbicides, fungicides, insecticides, flame retardants, etc. Additionally, many halogenated organic compounds, such as dioxins and polychlorinated biphenyls, are by-products produced during chemical synthesis (Gribble 2003; Van Pee and Unversucht 2003).
Generally, the chemistry of halogenated organic compounds reflects the physicochemical properties of their halogen substituent(s). The energy of carbon-halogen bonds decrease with the increase in molecular weight of the halogen (i.e., F > Cl > Br > I). Other characteristics, such as electron-withdrawing effects and the physical size and shape of the halogen substituent, impact the chemical reactivity of a compound. The physical size and shape of the halogen substituent can further delay uptake into cells and enzymatic attack during biodegradation. Moreover, the halogen moiety of an organic compound increases lipid solubility and reduces its water solubility. Finally, the halogen substituent and its potential organohalide metabolites may increase the inherent toxicity of a compound (Häggblom and Bossert 2003). The environmental consequences of these characteristic include acute and chronic toxicity, persistence and bioaccumulation in the environment and in animals tissue (Belkin 1992). Thus, the presence of halogenated organic compounds in the environment represents a threat to human and ecological systems and is thus of major concern.
Aerobic and anaerobic biodegradation of halogenated organic compounds, mainly chlorinated organic compounds, have been previously reported. Microorganisms can use halogenated organic compounds in four ways: (1) as a carbon source and an oxidizable substrate (2) as an electron acceptor in the ‘halorespiration’ process (3) in co-metabolic transformation, and (4) in fermentative metabolism, in which a dehalogenated intermediate serves as an electron acceptor (Janssen et al. 2001; Van Pee and Unversucht 2003). The key reaction during biodegradation of halogenated organic compounds is the dehalogenation step, in which the halogen is often replaced by a hydrogen or hydroxyl group. Halogen removal usually reduces resistance to biodegradation, as well as toxicity and the risk of forming toxic intermediates during subsequent metabolic steps (Janssen et al. 2001). There are number of dehalogenation mechanisms, involving a wide range of microorganisms, which are known to occur aerobically and anaerobically (Fetzner 1998; Janssen et al. 2001; Van Pee and Unversucht 2003).
Dibromoneopentyl glycol (DBNPG) and tribromoneopentyl alcohol (TBNPA) are reactive brominated flame retardants that are used as additives during the manufacture of plastic polymers and as chemical intermediates for the synthesis of high molecular-weight flame retardants (Fig. 1). The steric crowding of the halogen group of these bromo-neopentyls provides DBNPG and TBNPA with relatively strong resistance to biodegradation and, indeed, both are classified as not readily biodegradable, having half-lives of more that 100 years (Ezra et al. 2006). DBNPG is believed to be a human carcinogen, based on evidence of carcinogenicity from experimental studies in animals, while TBNPA is considered to cause aquatic environmental damage (Ezra 2005; EPA 2005).
In a previous study, we demonstrated the complete bacterial biodegradation of DBNPG and the resulting accumulation of bromides in the tested solution, indicating debromination (Segev et al. 2007). DBNPG was found to be biodegraded by a bacterial consortium under aerobic conditions in an enrichment culture containing sterile soil in the medium (the source of the bacterial consortium was contaminated soil from the Ramat Hovav Industrial Park, located in the Negev desert, Israel. The same soil was used in the medium). This sterile soil was found to be crucial for the biodegradation. In the current study, we report biodegradation of both DBNPG and TBNPA under aerobic conditions without external soil-supporting medium.
Materials and methods
Bacterial consortium
After 16 sequential transfers of the bacterial consortium, capable of biodegrading DBNPG to a soil-dependent enrichment culture (Segev et al. 2007), we established a stable bacterial consortium, capable of biodegrading DBNPG without the presence of sterile soil in the medium. DBNPG and TBNPA biodegradation experiments with the bacterial consortium were conducted on this soil-free enrichment culture medium.
Medium and growth
The bacterial consortium was grown at 30°C under aerobic conditions in an enrichment culture containing a sterile (autoclave; 20 min at 121°C) mineral salt medium, following the Zahn-Wellens/EMPA test (OECD 1992), with addition of 1,000 mg L−1 yeast extract and 1,000 mg L−1 DBNPG (Bromine Compounds, Israel), or 100 mg L−1 TBNPA (Sigma Aldrich) as the sole carbon source (hereinafter referred to as DBNPG-YE medium and TBNPA-YE medium, respectively).
DBNPG, TBNPA and bromide analyses
DBNPG concentration was determined by Agilent 1,100 High Performance Liquid Chromatography (HPLC), performed with a Diode Array Detector 1 (DAD) G1315B and a Lichrocart® 250-4 HPLC-Cartridge Lichrosphere® 100 rp-18 column. HPLC samples were prepared by mixing DBNPG enrichment culture samples with HPLC-grade methanol (1:1). After centrifugation, the samples were filtered through a 0.45 μm membrane filter (Schleicher & Schuell MicroScience, Germany). TBNPA concentration was determined with an Agilent 19091S-433 Gas Chromatography-Mass Spectrometry (GC-MS), with a 5,973 Network mass selective detector and an HP-5MS (0.25 mm × 30 m × 0.25 μm) column. GC-MS samples were prepared using StartaTM–X 33 μ polymeric sorbent (Phenomenex®, USA). Bromide concentration was determined by an ISE25Br-9 Ion Selective Electrode (Radiometer Analytical, USA). Samples for bromide analysis were prepared by diluting enrichment culture samples with distilled water. Bromide concentration in the samples was calculated from a bromide calibration curve. Initial concentrations were measured 3 h after inoculation of the bacterial consortium. All experiments relied upon three replicates.
DNA extraction and PCR amplification
Total genomic DNA was extracted using a MoBio power soil DNA isolation kit (MoBio Laboratories, Solana Beach, CA) according to the manufacturer’s protocol, with one modification: The purified DNA was eluted in 40 μl of C6 solution (MoBio Laboratories) and stored at −20°C. DNA concentration was determined by an ND-1,000 UV-Vis spectrophotometer (NanoDrop Technologies, Wilmington, DE). Total DNA was amplified by PCR with Mastercycler gradient thermocycler (Eppendorf, Westbury, NY) using general 16S rRNA primers for bacteria (Sigma-Genosys); forward primer, 8F, GGATCCAGACTTTGATYMTGGCTCAG (as described by Felske et al. 1997) and the reverse primer, 907R, CCGTCAATTCMTTTGAGTTT (as described by Lane et al. 1985). The PCR reaction mixtures included 12.5 μl ReddyMix (PCR master mix containing 1.5 m M MgCl2 and a 0.2 mM concentration of each deoxynucleoside triphosphate) (ABgene, Surrey, UK), 1 μl each of the forward and reverse primers, and 2 μl of the sample preparation. Double-distilled water was added to bring the total volume to 25 μl. The PCR protocol included an initial step of 4 min at 95°C, followed by 34 cycles of the following incubation pattern: 94°C for 30 s, 54°C for 40 s, and 72°C for 70 s. A final extension at 72°C for 20 min concluded the reaction. PCR samples were stored at −20°C until further processed.
Clone library
The PCR products were purified by electrophoresis on a 0.8% agarose gel (Sigma), stained with ethidium bromide, and visualized with a UV transilluminator. The approximately 0.9-kbp heterologous 16S rRNA products were excised from the gel and the DNA products were purified from the gel slice, using a Wizard PCR prep kit (Promega, Madison, Wisconsin). The gel-purified PCR products were cloned into the pCRII-TOPO-TA cloning vector, as specified by Invitrogen (Carlsbad, CA) and transformed into calcium chloride-competent Escherichia coli DH5α cells, according to the manufacturer’s protocol and standard techniques. Clone colonies were scanned and plasmid DNA was amplified with M13-F and M13-R primers annealed to the plasmid. The clones, with the correct plasmid insert, were then used for sequencing.
Sequencing
Clones were sequenced at McLab DNA Sequencing Services (South San Francisco, CA). A phylogenetic tree was constructed by the neighbor-joining method with Molecular Evolutionary Genetics Analysis, version 4 (MEGA 4) (Kumar et al. 2004).
Results
Using HPLC analysis and an ion selective electrode, we were able to demonstrate complete DBNPG biodegradation under aerobic condition in DBNPG-YE medium after 21 days. As shown in Fig. 2, the DBNPG concentration in the medium decreased and the bromide concentration increased accordingly, while the level of their concentrations in the sterile control remaining constant.
In order to determine the effect of yeast extract on DBNPG biodegradation, various concentrations of this ingredient were tested. The bromide concentration in the medium was used as an indicator for biodegradation. Figure 3 shows that addition of yeast extract supports and accelerates DBNPG biodegradation. Furthermore, without the addition of yeast extract to the medium, biodegradation does not occur.
Since the DBNPG and TBNPA only differ by an additional bromide in place of hydrogen, we examined the bacterial consortium’s ability to biodegrade TBNPA. We found that TBNPA was degraded by the DBNPG biodegrading consortium, with a similar trend of bromide accumulation in the medium (Fig. 4). TBNPA concentration in the sterile control remained constant throughout the experiment (data not shown in figure), with no bromide increase being detected.
Our previous attempts to isolate debrominating bacteria on different agar plates were not successful; bacterial isolates grown on agar plates did not show biodegradation activity (Segev et al. 2007). In order to characterize the bacterial consortium involved in the observed biodegradation, a clone library of PCR-amplified 16S rRNA genes was applied to the consortium grown on the DBNPG-YE medium. Sequencing of 60 bacterial clones and their classification revealed three groups of bacterial families: Sphingomonadaceae (52% of the clones), Burkholderiaceae (41% of the clones) and Rhizobiaceae (7% of the clones). At the level of bacterial species, these clones are closely related to Sphingomonas desiccabilis (99% similarity, 1 clone), Sphingopyxis macrogoltabida (98% similarity, 30 clones) Ralstonia sp. (99% similarity, 25 clones) and Sinorhizobium meliloti (99% similarity, 4 clones) (see Fig. 5). Alignment of these sequences with previously published consortium sequences (Segev et al. 2007) reveals that there is one mutually related species. Clone DBNPG 64 and the excised DGGE band RH Soil 227 are closely related to Sinorhizobium sp. Alignment of the two sequences presents 99% similarity.
Phylogenetic tree of PCR amplified 16S rRNA gene—DBNPG enrichment culture clones (▲) and their related species compared to the previously reported consortium species (Segev et al. 2007): excised denaturing gradient gel electrophoresis (DGGE) band (■); LB Agar isolates (□). Accession numbers are given in parentheses. The clones are representative clones
Discussion
In this paper, we describe the bacterial biodegradation of DBNPG and TBNPA under aerobic conditions, by a bacterial consortium in mineral medium with an addition of yeast extract. During the biodegradation process, there is a release of bromide into the medium, while the sterile controls remain constant. Thus, it is clearly demonstrated that the biodegradation and debromination reactions are due to bacterial activity. Since it was suggested that removal of the halogen substituent makes a compound more susceptible to complete mineralization (Ronen and Abeliovich 2000), we assume that the debromination reaction is the first step in DBNPG and TBNPA biodegradation and that the resulting intermediates then undergo complete mineralization. To this point, no intermediates were detected, however, further research is required. In addition, since biodegradation of contaminants is most often controlled by environmental conditions and/or the availability of compound (Bouwer and Zehnder 1993), and since different studies have demonstrated that intermediates and degradation products can be even more toxic and harmful than the original compound(s) (Cámara et al. 2004; Ronen and Abeliovich 2000; Salkinoja-Salonen et al. 1995), it is of utmost importance to identify DBNPG- and TBNPA- derived intermediates.
Previously, we reported the complete biodegradation of DBNPG in a soil-dependent enrichment culture (Segev et al. 2007). After 16 sequential transfers, we established a stable bacterial consortium capable of biodegrading DBNPG without the presence of sterile soil in the medium but with an addition of yeast extract. Furthermore, we found that the consortium is also capable of biodegrading TBNPA under the same conditions in the presence of yeast extract. Yeast extract provides substrate and inorganic nutrients for stimulating microbial activity and supports aerobic and anaerobic dehalogenation (Erable et al. 2005; Lee et al. 1998; Olaniran et al. 2004). The ability to replace the sterile Ramat Hovav soil with yeast extract suggests that the latter provides a factor or co-metabolite that exists in the Ramat Hovav soil which is necessary for the biodegradation process and debromination reaction. Aerobic dehalogenation, as a result of co-metabolism, is a known mechanism in which halogenated organic compounds compete with the growth substrate in the enzyme active site (Fetzner 1998). The replacement of sterile soil with yeast extract and consequently the establishment of soil-free enrichment culture are important to the understanding of the biodegradation processes of DBNPG and TBNPA and for future bioremediation, since there is no use of an external specific soil in the medium, which is defined.
Although the DBNPG concentration in the medium is greater than that of TBNPA concentration, the DBNPG biodegradation rate is faster. We believe that the number and arrangement of halogen substituents have important effects on the biodegradation and further study should be conducted. In general, the greater the number of halogens per organic compound, the more difficult it is to biodegrade (Slater et al. 1995).
At present, our attempts to isolate debrominating bacteria were not successful (Segev et al. 2007). Various studies have shown that the degradation activity of microorganism consortia is much more efficient than the activity of single strains (Bastos et al. 2002; Olaniran et al. 2001). Thus, it is possible that the debromination of DBNPG and TBNPA is more active in a consortium. Since culture-independent technique allows for microorganism characterization without isolation, we applied the clone library of PCR-amplified 16S rRNA gene technique to characterize the bacterial consortium. Sequencing of 60 bacterial clones revealed that most of the clones are closely related to Sphingopyxis sp., which has a potential to carry out debromination reactions (Aranda et al. 2004). Nonetheless, it is important to note that specific populations involved in the dehalogenating reactions do not always dominate their consortia (Smidt and De Vos 2004). Actually, all the bacterial clone sequences that are closely related species to S. meliloti, S. desiccabilis and Ralstonia sp. have a potential for performing the debromination reaction (Finan et al. 2001; Schenzle et al. 1999; Song et al. 2000; Wilkes et al. 1996). Additionally, a previously published bacterial Denaturing Gradient Gel Electrophoresis (DGGE) band sequence, which is closely related to Paracoccus sp., also has debromination potential (Segev et al. 2007).
It is important to notice that the previously published bacterial DGGE band sequences (Segev et al. 2007) and the clone library sequences share only one sequence, closely related species of S. meliloti, in common. The reason for this can be that although the same bacterial consortium was characterized, the two techniques are applied on different cultures, namely a soil-dependent (DGGE) enrichment culture and a soil-free (clone library) enrichment culture. Moreover, the DGGE and the clone library are two different molecular techniques, in which different set of primers are used and thus, different fractions of the population are revealed (Ben-Dov et al. 2006; Díez et al. 2001). The fact that the two techniques share one sequence in common suggests that the bacterial consortium population has low diversity. S. meliloti is known to possess genes that are involved in dehalogenation reactions (Finan et al. 2001). This might suggest that a Sinorhizobium sp. bacterium is the abundant debrominating microorganism, although further research is still needed to confirm this hypothesis.
We believe that understanding the biodegradation process of these bromo-neopentyls and the characterization of the dehalogenating microorganisms will be an important milestone in developing practical biological treatment processes for wastewater and sediments contaminated with compounds like DBNPG and TBNPA and, perhaps, other halogenated organic compounds.
References
Aranda C, Godoy F, Becerra J, Barra R, Martínez M (2004) Aerobic secondary utilization of a non-growth and inhibitory substrate 2,4,6-trichlorophenol by Sphingopyxis chilensis S37 and Sphingopyxis-like strain S32. Biodegradation 14:265–274. doi:10.1023/A:1024752605059
Bastos F, Bessa J, Pacheco CC, De Marco P, Castro PML, Silva M, Jorge RF (2002) Enrichment of microbial culture able to degrade 1, 3-dichloro-2-propanol: a comparison between batch and continuous methods. Biodegradation 13:211–220. doi:10.1023/A:1020834603785
Belkin S (1992) Biodegradation of haloalkanes. Biodegradation 3:299–313. doi:10.1007/BF00129090
Ben-Dov E, Shapiro OH, Siboni N, Kushmaro A (2006) Advantage of using Inosine at the 3′ termini of 16S rRNA gene universal primers for the study of microbial diversity. Appl Environ Microbiol 72:6902–6906. doi:10.1128/AEM.00849-06
Bouwer EJ, Zehnder AJB (1993) Bioremediation of organic compound–putting microbial metabolism to work. Bioremediat J 11:360–367
Cámara B, Herrera C, González M, Couve E, Hofer B, Seeger M (2004) From PCBs to highly toxic metabolites by the biphenyl pathway. Environ Microbiol 6:842–850. doi:10.1111/j.1462-2920.2004.00630.x
Díez B, Pedrós-Alió C, Marsh TL, Massana R (2001) Application of denaturing gradient gel electrophoresis (DGGE) to study the diversity of marine picoeukaryotic assemblages and comparison of DGGE with other molecular techniques. Appl Environ Microbiol 67:2942–2951. doi:10.1128/AEM.67.7.2942-2951.2001
Environmental Protection Agency (EPA) (2005) Furniture flame retardancy partnership: environmental profiles of chemical flame-retardant alternatives for low-density polyurethane foam. Chem Hazard Rev 2:2.1–2.26
Erable B, Goubet I, Lamare S, Seltana A, Dominique M, Maugard T (2005) Non conventional hydrolytic dehalogenation of 1-chlorobutane by dehydrated bacteria. Biotechnol Bioeng 91:304–313. doi:10.1002/bit.20437
Ezra S (2005) The fate of brominated neopentyl alcohols in a fractured chalk aquifer. PhD thesis. Ben-Gurion University of the Negev
Ezra S, Feinstein S, Yakirevich A, Adar E, Bilkis I (2006) Retadation of organo-bromides in a fractured chalk aquitard. J Contam Hydrol 86:195–214. doi:10.1016/j.jconhyd.2006.02.016
Felske A, Rheims H, Wolterink A, Stackebrandt E, Akkermans DL (1997) Ribosome analysis reveals prominent activity of an uncultured member of the class Actinobacteria in grassland soil. Microbiology 143:2983–2989
Fetzner S (1998) Bacterial dehalogenation. Appl Microbiol Biotechnol 50:633–657. doi:10.1007/s002530051346
Finan TM, Weidner S, Wong K, Buhrmester J, Chain P, Vorhölter FJ, Hernandez-Lucas I, Becker A, Cowie A, Gouzy J, Golding B, Pühler A (2001) The complete sequence of the 1,683-kb pSymB megaplasmid from the N2-fixing endosymbiont Sinorhizobium meliloti. Proc Natl Acad Sci USA 98:9889–9894. doi:10.1073/pnas.161294698
Gribble GW (2003) The diversity of naturally produced organohalogens. Chemosphere 52:289–297. doi:10.1016/S0045-6535(03)00207-8
Häggblom MM, Bossert ID (2003) Dehalogenation–microbial processes and environmental applications, 1st edn. Kluwer Academic Publishers, Norwell Massachusetts
Jain RK, Kapur M, Labana S, Lal B, Sarma PM, Bhattacharya D, Thakur S (2005) Microbial diversity: application of microorganisms for the biodegradation of xenobiotics. Curr Sci 89:101–112
Janssen BD, Oppentocht JE, Poelarends GJ (2001) Microbial dehalogenation. Curr Opin Biotechnol 12:254–258. doi:10.1016/S0958-1669(00)00208-1
Kumar S, Tomura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163. doi:10.1093/bib/5.2.150
Lane DJ, Pace B, Olsen GJ, Stahlt DA, Sogint ML, Pace NR (1985) Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci USA 82:6955–6959. doi:10.1073/pnas.82.20.6955
Lee MD, Odom JM, Buchanan RJ Jr (1998) Microbial dehalogenation of chlorinated solvents: insight from the field. Annu Rev Microbiol 52:423–452. doi:10.1146/annurev.micro.52.1.423
Olaniran AO, Babalola GO, Okoh AI (2001) Aerobic dehalogenation potentials of four bacterial species isolated from soil and sewage sludge. Chemosphere 45:45–50. doi:10.1016/S0045-6535(01)00075-3
Olaniran AO, Pillay D, Pillay B (2004) Haloalkane and haloacid dehalogenases from aerobic bacterial isolates indigenous to contaminated sites in Africa demonstrate diverse substrate specificities. Chemosphere 55:27–33. doi:10.1016/j.chemosphere.2003.10.067
Organisation for Economic Cooperation and Development (OECD). (1992) OECD guideline for testing of chemicals—Zahn-Wellens/EMPA test (302B)
Ronen Z, Abeliovich A (2000) Anaerobic-aerobic process for microbial degradation of tetrabromobisphenol A. Appl Environ Microbiol 66:2372–2377. doi:10.1128/AEM.66.6.2372-2377.2000
Salkinoja-Salonen M, Uotila J, Jokela ML, Saski E (1995) Organic halogens in the environment: studies of environmental biodegradability and human exposure. Environ Health Perspect 103:63–69. doi:10.2307/3432482
Schenzle A, Lenke H, Spain JCS, Knackmuss HJ (1999) Chemoselective nitro group reduction and reductive dechlorination initiate degradation of 2-chloro-5-nitrophenol by Ralstonia eutropha JMP134. Appl Environ Microbiol 65:2317–2323
Segev O, Abeliovich A, Kushmaro A (2007) Biodegradation of dibromonropentyl glycol by bacterial consortium. Chemosphere 68:958–964. doi:10.1016/j.chemosphere.2007.01.014
Slater JH, Bull AT, Hardman DJ (1995) Microbial dehalogenation. Biodegradation 6:181–189. doi:10.1007/BF00700456
Smidt H, De Vos WM (2004) Anaerobic microbial dehalogenation. Annu Rev Microbiol 58:43–73. doi:10.1146/annurev.micro.58.030603.123600
Song B, Palleroni NJ, Haggblom MM (2000) Isolation and characterization of diverse halobenzoate degrading denitrifying bacteria from soils and sediments. Appl Environ Microbiol 66:3446–3453. doi:10.1128/AEM.66.8.3446-3453.2000
Van Pee KH, Unversucht S (2003) Biological dehalogenation and halogenation reactions. Chemosphere 52:299–312. doi:10.1016/S0045-6535(03)00204-2
Wilkes H, Wittich RM, Timmis KN, Fortnagel P, Francke W (1996) Degradation of chlorinated dibenzofurans and dibenzo-p-dioxins by Sphingomonas sp. strain RW1. Appl Environ Microbiol 62:367–371
Acknowledgments
The work was supported by a grant from BMBF-MOST Cooperation in Water Technologies (Grant WT-501) and a grant from the Ramat Hovav Council, Israel. We thank the Rieger Foundation and the Israel Commercial Industrial club for O. Segev’s generous fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Segev, O., Meusel, W., Friedenberger, M. et al. Aerobic biodegradation of the brominated flame retardants, dibromoneopentyl glycol and tribromoneopentyl alcohol. Biodegradation 20, 621–627 (2009). https://doi.org/10.1007/s10532-009-9249-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-009-9249-z



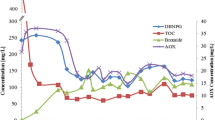



 ); sterile control (
); sterile control ( ). Bromide concentrations: bacterial consortium (
). Bromide concentrations: bacterial consortium ( ); sterile control (
); sterile control ( )
)
 ); Bacterial Consortium after 30 days (□); Sterile control after 30 days (■). Tested compound in all experiments was DBNPG
); Bacterial Consortium after 30 days (□); Sterile control after 30 days (■). Tested compound in all experiments was DBNPG
