Abstract
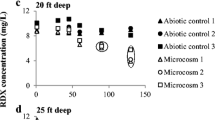
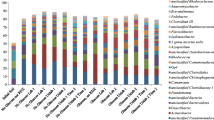
A field-scale experiment to assess biostimulation of uranium reduction is underway at the Natural and Accelerated Bioremediation Research Field Research Center (FRC) in Oak Ridge, Tennessee. To simulate the field experiment, we established replicate batch microcosms containing well-mixed contaminated sediment from a well within the FRC treatment zone, and we added an inoculum from a pilot-scale fluidized bed reactor representing the inoculum in the field experiment. After reduction of nitrate, both sulfate and soluble U(VI) concentration decreased. X-ray absorption near edge structure (XANES) spectroscopy confirmed formation of U(IV) in sediment from biostimulated microcosms, but did not detect reduction of solid-phase Fe(III). Two to three fragments dominated terminal restriction fragment length polymorphism (T-RFLP) profiles of the 16S rDNA gene. Comparison to a clone library indicated these fragments represented denitrifying organisms related to Acidovorax, and Acidovorax isolates from the inoculum were subsequently shown to reduce U(VI). Investigation using the T-RFLP Analysis Program (TAP T-RFLP) and chemical analyses detected the presence and activity of fermenting and sulfate-reducing bacteria after 2 weeks. These organisms likely contributed to uranium reduction. In some microcosms, soluble U(VI) concentration leveled off or rebounded, indicating microbial and/or mineralogical heterogeneity among samples. Sulfate, acetate, and ethanol were depleted only in those microcosms exhibiting a rebound in soluble U(VI). This suggests that rates of U(VI) desorption can exceed rates of U(VI) reduction when sulfate-reducing bacteria become substrate-limited. These observations underscore the importance of effective chemical delivery and the role of serial and parallel processes in uranium reduction.
Article PDF
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
References
Abdelouas A, Lu YM, Lutze W, Nuttall HE (1998) Reduction of U(VI) to U(IV) by indigenous bacteria in contaminated ground waterJ. Contam. Hydrol. 35:217–233
Anderson RT, Vrionis HA, Ortiz-Bernad I, Resch CT, Long PE, Dayvault R, Karp K, Marutzky S, Metzler DR, Peacock A, White DC, Lowe M, Lovley DR (2003) Stimulating the in situ activity of Geobacter species to remove uranium from the groundwater of a uranium-contaminated aquiferAppl. Environ. Microbiol. 69:5884–5891
Arias YM, Tebo BM (2003) Cr(VI) reduction by sulfidogenic and nonsulfidogenic microbial consortia Appl. Environ. Microbiol. 69:1847–1853
Barnett MO, Jardine PM, Brooks SC (2002) U(VI) adsorption to heterogeneous subsurface media: application of a surface complexation modelEnviron. Sci. Technol. 36:937–942
Bertsch PM, Hunter DB, Sutton SR, Bajt S, Rivers ML (1994) In-situ chemical speciation of uranium in soils and sediments by micro X-ray absorption spectroscopyEnviron. Sci. Technol. 28:980–984
Brooks SC, Fredrickson JK, Carroll SL, Kennedy DW, Zachara JM, Plymale AE, Kelly SD, Kemner KM, Fendorf S (2003) Inhibition of bacterial U(VI) reduction by calciumEnviron. Sci. Technol. 37:1850–1858
Coates JD, Chakraborty R, Lack JG, O’Connor SM, Cole KA, Bender KS, Achenbach LA (2001) Anaerobic benzene oxidation coupled to nitrate reduction in pure culture by two strains of Dechloromonas Nature 411:1039–1043
Cole JR, Chai B, Marsh TL, Farris RJ, Wang Q, Kulam SA, Chandra S, McGarrell DM, Schmidt TM, Garrity GM, Tiedje JM (2003) The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomyNucleic Acids Res. 31:442–443
Dunbar J, Ticknor LO, Kuske CR (2001) Phylogenetic specificity and reproducibility and new method for analysis of terminal restriction fragment profiles of 16S rRNA genes from bacterial communitiesAppl. Environ. Microbiol. 67:190–197
Elias DA, Senko JM, Krumholz LR (2003) A procedure for quantitation of total oxidized uranium for bioremediation studiesJ. Microbiol. Methods 53:343–353
Finneran K, Housewright M, Lovley D (2002a) Multiple influences of nitrate on uranium solubility during bioremediation of uranium-contaminated subsurface sedimentsEnviron. Microbiol. 4:510–516
Finneran KT, Anderson RT, Nevin KP, Lovley DR (2002b) Potential for bioremediation of uranium-contaminated aquifers with microbial U(VI) reductionSoil Sediment Contam. 11:339–357
Francis AJ, Dodge CJ, Lu FL, Halada GP, Clayton CR (1994) XPS and XANES studies of uranium reduction by Clostridium sp Environ. Sci. Technol. 28:636–639
Gentile M, Yan TF, Tiquia S, Fields M, Nyman J, Zhou J & Criddle C (2005) Stability in a denitrifying fluidized bed reactor. Microbiol Ecology (forthcoming)
Gorby Y, Lovley D (1992) Enzymatic uranium precipitationEnviron. Sci. Technol. 26:205–207
Istok JD, Senko JM, Krumholz LR, Watson D, Bogle MA, Peacock A, Chang YJ, White DC (2004) In situ bioreduction of technetium and uranium in a nitrate-contaminated aquiferEnviron. Sci. Technol. 38:468–475
Kaplan CW, Kitts CL (2003) Variation between observed and true Terminal Restriction Fragment length is dependent on true TRF length and purine contentJ. Microbiol. Methods 54:121–125
Kitts CL (2001) Terminal Restriction Fragment patterns: a tool for comparing microbial communities and assessing community dynamicsCurr. Issues Intest. Microbiol. 2:17–25
Kemner KM, Kelly SD, Orlandini KA, Tsapin AI, Goldfeld MG, Perfiliev YD, Nealson KH (2001) XAS investigations of Fe(VI)J. Synchrotron Radiat. 8:949–951
Kniemeyer O, Probian C, Rossello-Mora R, Harder J (1999) Anaerobic mineralization of quaternary carbon atoms: isolation of denitrifying bacteria on dimethylmalonateAppl. Environ. Microbiol. 65:3319–3324
Lane DJ, (1991) 16S/23S rRNA sequencing In: Stackebrandt E, Goodfellow M (eds), Nucleic Acid Techniques in Bacterial Systematics. John Wiley & Sons Ltd., Chichester, England, pp 115–175
Lovley DR, Phillips EJP, Gorby YA, Landa ER (1991) Microbial reduction of uraniumNature 350:413–416
Lovley DR, Roden EE, Phillips EJP, Woodward JC (1993) Enzymatic iron and uranium reduction by sulfate-reducing bacteriaMar. Geol. 113:41–53
Lytle FW, Greego RB, Sandston DR, Marques EC, Wong J, Spiro CL, Huffman GP, Huggins FE (1984) Measurements of soft X-ray absorption spectra with a fluorescent ion chamberNucl. Instrum. Methods Phys. Res. Sect. A 226:542–548
Marsh TL, Saxman P, Cole J, Tiedje J (2000) Terminal restriction fragment length polymorphism analysis program, a web-based research tool for microbial community analysisAppl. Environ. Microbiol. 66:3616–3620
Myers CR, Nealson KH (1988) Bacterial manganese reduction and growth with manganese oxide as the sole electron acceptorScience 240: 1319–1321
Nevin KP, Finneran KT, Lovley DR (2003) Microorganisms associated with uranium bioremediation in a high-salinity subsurface sedimentAppl. Environ. Microbiol. 69:3672–3675
Newville M (2001) IFEFFIT: interactive XAFS analysis and FEFF fittingJ. Synchrotron Radiat. 8:322–324
Nyman JL, Williams SM & Criddle CS (2005) Bioengineering for the in-situ remediation of metals. In: Grassian VH (Ed) Environmental Catalysis. (pp 493–520) Taylor & Francis Boca Raton, FL
Patterson RR, Fendorf S, Fendorf M (1997) Reduction of hexavalent chromium by amorphous iron sulfideEnviron. Sci. Technol. 31:2039–2044
Peterson ML, White AF, Brown GE, Parks GA (1997) Surface passivation of magnetite by reaction with aqueous Cr(VI): XAFS and TEM resultsEnviron. Sci. Technol. 31:1573–1576
Ressler T, Wong J, Roos J, Smith IL (2000) Quantitative speciation of Mn-bearing particulates emitted from autos burning (methylcyclopentadienyl)manganese tricarbonyl-added gasolines using XANES spectroscopyEnviron. Sci. Technol. 34:950–958
Riley RG, Zachara JM & Wobber FJ (1992) Chemical Contaminants on DOE Lands and Selection of Contaminant Mixtures for Subsurface Science Research. Report No. DOE/ER-0547T, Office of Energy Research, US Department of Energy, Washington, DC
Sani RK, Peyton BM, Amonette JE, Geesey GG (2004) Reduction of uranium(VI) under sulfate-reducing conditions in the presence of Fe(III)-(hydr)oxides Geochim. Cosmochim. Acta 68:2639–2648
Schulze R, Spring S, Amann R, Huber I, Ludwig W, Schleifer KH, Kampfer P (1999) Genotypic diversity of Acidovorax strains isolated from activated sludge and description of Acidovorax defluvii sp. novSyst. Appl. Microbiol. 22:205–214
Senko J, Istok J, Suflita J, Krumholz L (2002) In-situ evidence for uranium immobilization and remobilizationEnviron. Sci. Technol. 36:1491–1496
Shelobolina ES, O’Neill K, Finneran KT, Hayes LA, Lovley DR (2003) Potential for in situ bioremediation of a low-pH, high-nitrate uranium-contaminated groundwaterSoil Sediment Contam. 12:865–884
Suzuki Y, Kelly SD, Kemner KA, Banfield JF (2003) Microbial populations stimulated for hexavalent uranium reduction in uranium mine sedimentAppl. Environ. Microbiol. 69:1337–1346
Suzuki Y, Kelly SD, Kemner KM, Banfield JF (2004) Enzymatic U(VI) reduction by Desulfosporosinus speciesRadiochim. Acta 92:11–16
Swenson W, Arendt J, Wilson DS (2000) Artificial selection of microbial ecosystems for 3-chloroaniline biodegradationEnviron. Microbiol. 2:564–571
Waychunas GA, Apted MJ, Brown GE (1983) X-ray K-edge absorption spectra of Fe minerals and model compounds: near-edge structurePhys. Chem. Miner. 10:1–9
Widdel F, Pfennig N (1984) Dissimilatory sulfate- or sulfur-reducing bacteria In: Krieg NR, Holt JG (eds), Bergey’s Manual of Systematic Bacteriology 1. Williams and Wilkins, Baltimore, MD, pp. 663–679
Acknowledgements
This work was funded from the National Science Foundation graduate fellowship program and a United States Department of Energy (DOE) Natural and Accelerated
Bioremediation Research (NABIR) Biological and Environmental Research (BER) grant (#DE-F603-00ER63046). The authors would like to thank David Watson of the Oak Ridge Field Research Center for providing sediment samples, and Dr. Wei-Min Wu for providing denitrified synthetic groundwater. Portions of this research were carried out at the Stanford Synchrotron Radiation Laboratory, a national user facility operated by Stanford University on behalf of the United States Department of Energy, Office of Basic Energy Sciences. The SSRL Structural Molecular Biology Program is supported by the Department of Energy, Office of Biological and Environmental Research, and by the National Institutes of Health, National Center for Research Resources, Biomedical Technology Program. Other portions were performed at GeoSoilEnviroCARS (Sector 13), Advanced Photon Source (APS), Argonne National Laboratory. GeoSoilEnviroCARS is supported by the National Science Foundation – Earth Sciences (EAR-0217473), Department of Energy – Geosciences (DE-FG02-94ER14466) and the State of Illinois. Use of the APS was supported by the US Department of Energy, Basic Energy Sciences, Office of Energy Research, under Contract No. W-31-109-Eng-38.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nyman, J.L., Marsh, T.L., Ginder-Vogel, M.A. et al. Heterogeneous response to biostimulation for U(VI) reduction in replicated sediment microcosms. Biodegradation 17, 303–316 (2006). https://doi.org/10.1007/s10532-005-9000-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-005-9000-3