Abstract
Introduced plant pathogens are increasingly recognized as a major threat to biodiversity and ecosystem functioning. One such pathogen, the causal agent of the devastating ash dieback in Europe, Hymenoscyphus fraxineus, was most likely introduced into Europe from eastern Asia in the 1990s. To investigate the genetic population structure of this invasive fungus at the epidemic disease front (Switzerland) and in the post-epidemic phase (Lithuania), a total of 847 H. fraxineus isolates were genotyped at 11 microsatellite loci. Among these isolates, 244 multilocus genotypes were found in five post-epidemic subpopulations (367 isolates) of the fungus and 263 in five epidemic subpopulations (480 isolates). No genetic differentiation was found between isolates recovered from bark lesions and fallen leaf petioles, which suggests that all H. fraxineus genotypes have the potential to induce bark infections on living trees and to survive saprophytically. Moreover, no genetic differentiation and no difference in genetic diversity were detected between the epidemic and post-epidemic populations. The entire genetic diversity present in the original founding populations in north-eastern Europe seems to have been transmitted to the epidemic disease front. Nonetheless, gene flow among post-epidemic subpopulations occurs slightly more random than among epidemic subpopulations. Furthermore, the probability of correct assignment of a particular H. fraxineus genotype to its subpopulation of origin was greater in Switzerland than in Lithuania. These two analyses point to weak founder effects at the disease front.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alien (i.e. exotic, non-native) fungal pathogens have become a major threat to biodiversity and ecosystem functioning worldwide (e.g. Boyd et al. 2013; Santini et al. 2013). Unfortunately, the number of such organisms is expected to increase further in future because of human activity. Global trade breaks down the natural distribution ranges of species by facilitating the movement of pathogens around the globe and their establishment in new territories (Brasier 2008; Westphal et al. 2008; Santini et al. 2013). Once established, exotic pathogens are usually difficult to eradicate in natural ecosystems. Developing successful control strategies requires a thorough knowledge of the biology, epidemiology, and pathways of spread of each target organism (Sakai et al. 2001).
In recent years, the advent of powerful molecular markers has significantly improved our capability to trace the origin of invasive pathogens, to reconstruct their spread in space and time, as well as to investigate population dynamics (e.g. Xhaard et al. 2012; Croucher et al. 2013). Fungi have a wide variety of life cycles, which shape the genetic structure of populations (e.g. Barrett et al. 2008; Billiard et al. 2013). The three main sources of genetic diversity in fungal populations are sexual reproduction, mutation, and the immigration of individuals from other, genetically differentiated populations (e.g. McDonald 1997). In general, a high genetic diversity is considered to indicate a sexually reproducing, native or long-established population (e.g. Burdon and Silk 1997; Dean and Ballard 2004). In contrast, invasive populations and populations at the epidemic front, i.e. populations of recently emerged pathogens that are still expanding their range, are usually characterized by a simple structure and low genetic diversity (e.g. Sakai et al. 2001; Allendorf and Lundquist 2003). For example, native populations of the chestnut blight fungus Cryphonectria parasitica in Asia are genetically more diverse than invasive populations in North America and Europe (Liu and Milgroom 2007). Moreover, European populations of C. parasitica at the epidemic front are characterized by a lower genetic diversity than long-established, post-epidemic populations (Milgroom et al. 2008; Dutech et al. 2010, 2012; Prospero and Rigling 2012).
The loss of genetic diversity observed in new and younger populations is associated with the founding of these populations by a limited number of individuals (the so-called “founder effect”). Range expansions after establishment usually imply a reduction of genetic diversity along the expansion gradient because of repeated founding events (Austerlitz et al. 1997; Fontaine et al. 2013). Such range expansions may occur due to human activities, which unintentionally lead to single or a few individuals being transported into new territories (e.g. by moving infected plant material) or as a consequence of the spontaneous spread of the pathogen (e.g. via spores), and may be driven by climatic changes (e.g. Anderson et al. 2004; Desprez-Loustau et al. 2009). Over time and space, the genetic structure of an expanding population can rapidly change and create complex patterns, with significant differences between the source and the front of the expansion (Excoffier and Ray 2008). For example, alleles that are rare in an established population may become very frequent at the front of the expansion (the so-called “allele surfing”; Klopfstein et al. 2006).
In this study, we focused on the invasive pathogen Hymenoscyphus fraxineus (synonym: H. pseudoalbidus; basionym: Chalara fraxinea), which is causing severe dieback of the common ash (Fraxinus excelsior) in Europe (Pautasso et al. 2013). H. fraxineus is a heterothallic ascomycetous fungus that reproduces sexually on fallen ash-leaf petioles. Abundantly produced ascospores are wind-dispersed and infect ash leaves during summer. The fungus may spread from the infected leaves via petioles into twigs and stems, inducing necrotic bark lesions, which eventually lead to wilting and subsequent dieback of the affected branches (Gross et al. 2014a). As H. fraxineus only rarely sporulates on bark lesions, these represent a dead end for disease epidemiology. In contrast, the petioles of fallen leaves on which the sexual fruiting bodies are formed are the main source of inoculum. Asexual spores serve only as spermatia but not as infective and dispersal propagules (Gross et al. 2014a). This distinguishes H. fraxineus from other major invasive pathogens of forest trees, which are also able to reproduce and spread asexually, such as Cryphonectria parasitica (Prospero and Rigling 2013), Phytophthora ramorum (Grünwald et al. 2012), P. plurivora (Schoebel et al. 2014a), or P. kernoviae (Fichtner et al. 2012).
The pathogen was most probably introduced into Europe from eastern Asia by two individuals (Gross et al. 2014b). Although ash dieback was first observed in Poland in the early 1990s, H. fraxineus was only later identified as the causal agent of the disease (Kowalski and Holdenrieder 2009). Currently, a large part of the native distribution range of F. excelsior is affected (Timmermann et al. 2011; Pautasso et al. 2013; McKinney et al. 2014). Different phases of the epidemic have been observed, due to the gradual spread of H. fraxineus from the putative source population in Poland towards the periphery of the continent. In areas where the pathogen has been present for a long time (e.g. Poland, Baltic States), ash dieback is already in its chronic, post-epidemic phase (Kraj et al. 2012; Lygis et al. 2014). In other areas (e.g. Great Britain, south-western France), the epidemic front is still expanding (Husson et al. 2011; DEFRA 2013).
In Lithuania, ash dieback was first noticed in the mid 1990s. To date, virtually all stands of F. excelsior have been affected by the disease (Lygis et al. 2014). In Switzerland, however, ash dieback was first observed in 2008 (Engesser et al. 2009), and the front of the disease is still expanding, with H. fraxineus colonizing new areas (e.g. Southern Switzerland). In this study, we aimed to genetically characterize epidemic (Swiss) and post-epidemic (Lithuanian) subpopulations of H. fraxineus with the aid of microsatellite markers. Specifically, we (1) tested whether genotypes originating from necrotic lesions (parasitic phase of H. fraxineus) are genetically differentiated from genotypes isolated from fallen ash leaf petioles (saprophytic phase of H. fraxineus); (2) determined the extent of population differentiation at the disease front and in the post-epidemic region; and (3) tested the hypothesis that epidemic subpopulations of the pathogen are less diverse than post-epidemic subpopulations.
Materials and methods
Sampling and isolation of H. fraxineus
To isolate H. fraxineus, symptomatic tissues of Fraxinus excelsior (i.e. bark lesions on shoots, branches or stems, and fallen leaf petioles) were sampled in five Lithuanian and five Swiss forest stands (Table 1). In both countries, the sampled stands were at least 40 km apart, at distances ranging between 60–260 km in Lithuania and 40–250 km in Switzerland. The sampled stands in the two countries were 1250–1660 km apart. In each stand, the samples were collected at 60 sampling points, which consisted of a circular area approx. 2 m in diameter and included at least one F. excelsior tree showing typical dieback symptoms (Kowalski and Holdenrieder 2009). The minimum distance between two sampling points in a stand was at least 5 m. At each sampling point, 2–4 approx. 15–20-cm long pieces from shoots, branches or stems with fresh necrotic bark lesions were collected from 1–3 ash trees (1–20-years-old). In addition, fallen ash leaf petioles with pseudosclerotial plates were collected from the forest floor at each sampling point. H. fraxineus from leaf petioles was isolated as described by Kirisits et al. (2013), and from necrotic bark lesions as described by Schoebel et al. (2014b). Not more than one isolate obtained from lesions and not more than one isolate obtained from petioles were picked for further analyses from each sampling point, i.e. not more than two tissue samples (one lesion and one petiole) per sampling point. For further analyses, country-level H. fraxineus populations (N = 2) were considered as “populations”, while stand-level populations (N = 10) as “subpopulations”.
DNA extraction and species identification
For DNA extraction from H. fraxineus cultures, an agar plug of growing mycelium was transferred to new plates containing diamalt agar covered with a cellophane layer (Cellolaire, Switzerland) to separate the mycelium from the agar. After 4–5 weeks of incubation in the dark at 20 °C, mycelia were harvested from the cellophane and DNA was extracted as described by Schoebel et al. (2014b) using the Norgen Total RNA Purification Kit (Norgen Biotek Corp., Canada) following the manufacturer’s protocol but omitting the DNase treatment step (in this way the kit yields amplifiable DNA). As genotyping was performed using H. fraxineus-specific microsatellite primers (Gross et al. 2012), no additional species identification was necessary.
Microsatellite genotyping
Eleven polymorphic microsatellite loci (Gross et al. 2012; Online Resource 1) were selected for the population genetic analyses. PCR was performed in two newly developed multiplex reactions (MP1 and MP2, Online Resource 1) using 1 µl of 10 × diluted DNA and the Type-it Microsatellite PCR Kit (Qiagen, Valencia, CA, USA), following the manufacturer’s protocol, except that only 3 µl of the Master Mix were used per reaction. PCR conditions were set as follows: initial denaturation for 5 min at 95 °C, followed by 28 cycles consisting of 30 s at 95 °C, 90 s at 58 °C and 30 s at 72 °C, with a final extension for 30 min at 60 °C. PCR amplifications were performed on the Veriti® Thermal Cyclers (Applied Biosystems, Foster City, CA, USA). Subsequently, the PCR products obtained were run on an ABI 3730xl DNA Analyser with the GeneScan™ 500 LIZ® Size Standard for fragment analysis. All fragments were scored using the GeneMapper software, version 3.7 (all Applied Biosystems).
Population diversity analyses
In each subpopulation, the number of multilocus genotypes (MLGs) obtained only from petioles, only from lesions, and from both petioles and lesions was calculated using the R package POPPR v. 1.1.2 (Kamvar et al. 2014; R Development Core Team 2014). To assess how much power we had to discriminate between unique MLGs using the selected 11 microsatellite loci, we randomly sampled (1000 times) from one to 10 loci and plotted the number of MLGs detected against the number of loci, as implemented in the R package POPPR (Kamvar et al. 2014; R Development Core Team 2014). The genotypic diversity of the subpopulations (i.e. total, only within samples from lesions, only within samples from petioles) was characterized by the Shannon-Wiener diversity index H = −Σ p i × ln p i (where p i is the frequency of the ith genotype) and the number of expected genotypes (i.e. genotypic richness, eMLG). Both indices were calculated by using the R package VEGAN (Dixon 2003; R Development Core Team 2014) for the smallest sample size. Differences in eMLG between the Swiss and Lithuanian populations and between the lesion and petiole isolates were tested for significance by conducting a t test in SPSS Statistics v. 17.0 (SPSS Inc., Chicago).
For each locus, the total number of alleles (AM), as well as the allele frequencies, were calculated using the software GENEPOP (Rousset 2008). Subpopulation-specific (i.e. total, only within samples from lesions, only within samples from petioles) allelic richness (AR) and incidence of private alleles (APR) were calculated using a generalized rarefaction approach as implemented in the program ADZE (Szpiech et al. 2008) and tested for significance by conducting a t test in SPSS Statistics v. 17.0.
The overall genetic differentiation (FST value with a 95 % confidence interval) among Swiss and Lithuanian subpopulations was estimated using FSTAT 2.9.3.2. (Goudet 2001). Pairwise FST values (Weir and Cockerham 1984) between isolates from petioles and lesions within each subpopulation, as well as between all subpopulation pairs (petiole and lesion isolates combined), were calculated by using GENEPOP for full data set (847 individuals). The existence of a pattern of isolation-by-distance between subpopulations (i.e. all subpopulations, Swiss subpopulations, Lithuanian subpopulations) was determined by analysing the regression of FST/(1 − FST) on the natural logarithm of geographic distance (Rousset 1997). The significance of the regression was determined by performing Mantel tests using the program GENALEX v. 6.41 (Peakall and Smouse 2006).
H. fraxineus is considered to be a strictly outcrossing fungus (Gross et al. 2014a). However, we verified that all MLGs detected resulted from random sexual reproduction rather than from clonal spread by estimating the probability of obtaining a specific MLG under the assumption of random mating (P Gen). Moreover, for repeated MLGs, i.e. MLGs observed more than once in a subpopulation (see Results), we also estimated the probability of a second encounter, assuming that the isolates with the same MLG were derived from different, independent sexual reproductive events (P Sex). Calculations were performed using GENCLONE v. 2.0 (Arnaud-Haond and Belkhir 2007). The hypothesis of random mating within subpopulations was tested using index of association statistics (Agapow and Burt 2001). Specifically, the index rBarD, which corresponds to the index of association IA, but is independent of the number of loci considered, was calculated using the R package POPPR (Kamvar et al. 2014). rBarD is expected to be zero if subpopulations freely recombine and significantly greater than zero if association between alleles is non-random (clonality). This was again calculated using the full data set (847 individuals).
Haplotype clustering analyses
The genetic structure of the combined H. fraxineus dataset and of the Swiss and Lithuanian subpopulations was analysed by using two different approaches. First, the probability of MLGs being distributed into K number of divergent clusters was calculated using a Bayesian clustering method implemented in STRUCTURE v. 2.3.3 (Pritchard et al. 2000). For all MLGs in the clone-corrected data set, i.e. where each MLG was considered only once per subpopulation, the membership coefficient for each cluster was calculated. An admixture model without prior population information and with allele frequencies correlated across subpopulations was considered (Falush et al. 2003). Each model was simulated 10 times for K = 1 − 20, with a burn-in of 105 iterations and a run length of 106 iterations after the burn-in. The optimal number of clusters is generally set at the threshold at which the mean logarithm of the probability of the data [ln P(X|K)] reaches a plateau, i.e. no additional information can be obtained by increasing the number of clusters any further (Pritchard et al. 2000; Falush et al. 2003).
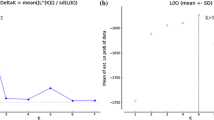
Second, we performed a Discriminant Analysis of Principal Components (DAPC, Jombart et al. 2010), as implemented in the R package ADEGENET (Jombart 2008), to genetically cluster similar individuals according to their MLG. DAPC is a multivariate method and consists of a two-step procedure. First, DAPC transforms the data using Principal Component Analysis (PCA) before performing a Linear Discriminant Analysis (LDA) on the retained principal components. Discriminant functions are expressed as linear combinations of alleles, which allows the allele contributions to be computed. Unlike STRUCTURE, this method does not use any underlying population genetic model and focuses on the genetic differentiations among groups of MLGs. We performed DAPC analysis using the population of origin as a prior and ran four different sets of analyses: (1) for two populations (Switzerland and Lithuania), (2) for all 10 subpopulations, (3) for only Swiss subpopulations, and (4) for only Lithuanian subpopulations. Further, re-assignment of all individuals to their (sub-) population of origin was performed for all data sets. In order to measure the trade-off between over-fitting and the power of discrimination, an a-score optimisation, indicating the optimal number of PCs to be retained (Jombart 2010), was performed for each of the datasets. Analyses were conducted using clone-corrected data sets.
Results
Population diversity
A total of 847 out of the 849 isolates recovered from the collected tissue samples were successfully genotyped at 11 H. fraxineus-specific microsatellite loci. Amongst those, 480 came from Swiss subpopulations and 367 from Lithuanian subpopulations (Table 2). About half of the genotyped isolates (49.8 %, N = 425) originated from lesions and half (50.2 %, N = 422) from petioles. Altogether, 390 different H. fraxineus MLGs were identified, 263 in the Swiss subpopulations and 244 in the Lithuanian (Table 2).
Multilocus genotype accumulation curves based on a re-sampling procedure for Switzerland and Lithuania indicated that, with 11 microsatellite loci, we still did not reach the plateau of the number of MLGs expected to be present in both populations (Fig. 1). One-hundred-seventeen MLGs (30 %) were shared between the two countries. Single MLGs comprised a maximum of 12 isolates, and 213 of them (54.6 %) were only represented by one isolate (Fig. 2). All MLGs including more than eight isolates were found in both countries in isolates recovered from both petioles and lesions (data not shown) and in relatively large proportions (40–90 %) of the subpopulations sampled in each country. The number of MLGs identified per subpopulation ranged between 71 (BIE) and 90 (SAR) in Switzerland, and between 49 (VIL) and 81 (BIR) in Lithuania (Table 2). Calculations based on a rarefaction procedure indicated that the number of eMLGs in a H. fraxineus subpopulation of N = 56, i.e. the smallest size of the subpopulations analysed, varied between 47.2 ± 0.88 (KED) and 53.4 ± 1.18 (BIR) (Table 3). Few differences among subpopulations were observed in the Shannon-Wiener index (H), which ranged between 3.69 (KUE) and 3.98 (UKM) (Table 3). About one-third (122 out of 390) of the MLGs identified were shared between petioles and lesions. In both the epidemic and post-epidemic populations, no significant differences (at P < 0.05) were observed between the number of eMLGs recovered from bark lesions and the number from fallen leaf petioles.
Multilocus genotype accumulation curves for H. fraxineus using nuclear microsatellite data of 719 H. fraxineus isolates from Switzerland (A) and Lithuania (B) obtained by randomly sampling one to 10 loci 1000 times (threshold 90 %). Each multilocus genotype was considered only once per subpopulation
Frequency distribution of all multilocus genotypes (MLGs) identified in the whole H. fraxineus population (i.e. Lithuanian and Swiss subpopulations combined) with respect to the number of isolates per MLG (in total, 847 isolates grouped into 390 MLGs). No. of MLGs, number of multilocus genotypes; No. of isolates per MLG, number (count) of isolates per multilocus genotype
Combining both the Swiss and Lithuanian H. fraxineus subpopulations, a total of 29 alleles were detected across the 11 microsatellite loci. All loci were polymorphic and harboured from two to five different alleles (Fig. 3). The mean number of alleles per locus (AM) ranged from 1.9 (KED) to 2.4 (BIE) (Table 3). Allelic richness (AR) and the number of private alleles (APR) calculated for the smallest sample size analysed (N = 56) were the highest in subpopulations BIR (AR = 2.08 ± 0.02; APR = 0.09 ± 0.01) and BIE (AR = 2.06 ± 0.02; APR = 0.07 ± 0.00), while the lowest values were observed in KED (AR = 1.86 ± 0.01; APR = 0.00 ± 0.00) and AIG (AR = 1.87 ± 0.02; APR = 0.00 ± 0.00) (Table 3). AR and APR in isolates obtained from lesions and from petioles were not significantly different (P < 0.05) in either subpopulation. Private alleles were present in one Swiss MLG (from subpopulation BIE, isolated from a petiole) at loci 53 (293 bp) and 69 (256 bp), and in three Lithuanian MLGs at loci 15 (one individual from subpopulation KRE, isolated from a lesion; 197 bp), 38 (one individual from subpopulation BIR, isolated from a lesion; 234 bp), and 69 (one individual from subpopulation UKM, isolated from a petiole; 268 bp) (Fig. 3). On excluding the five private alleles, no significant differences (P < 0.05) were detected between the allele frequencies in the Swiss and Lithuanian populations.
Allele frequencies at the 11 nuclear microsatellite loci genotyped in the 10 H. fraxineus subpopulations studied in A Switzerland (480 isolates) and B Lithuania (367 isolates). Numbers (upper line below the x axis) indicate the allele sizes in base pairs. See Table 1 for subpopulation name abbreviations, and Online Resource 1 for locus name abbreviations (lower line below the x-axis). Private alleles for the specific population are indicated in bold
None of the rBarD values were significantly different from zero, indicating that all subpopulations investigated were freely recombining, i.e. there were virtually no associations between alleles (no clonality) (Table 3). Calculations conducted with GENCLONE indicated that all MLGs detected in the Swiss and Lithuanian subpopulations had a probability of P Gen < 0.05 being obtained by chance through a sexual event. In contrast, the probability of the same MLG occurring more than once because of random sexual recombination (P Sex) was less than 0.05 only for a few MLGs detected at least twice (Table 3).
Population structure
The global FST value across epidemic H. fraxineus subpopulations was 0.025 (95 % confidence interval: 0.001–0.004), and across post-epidemic subpopulations −0.001 (95 % confidence interval: −0.008 to 0.001). Pairwise subpopulation differentiation was only significant (P < 0.001) between the Swiss subpopulation AIG and all other subpopulations (Online Resource 5). Within subpopulations, the FST values between isolates from petioles and lesions were also low (−0.002 to 0.029; Table 2). No significant signal of isolation-by-distance could be detected in any of the three H. fraxineus populations, i.e. not in the Swiss (P = 0.206), nor the Lithuanian (P = 0.648), nor the two combined (P = 0.270).
STRUCTURE analysis for the entire H. fraxineus population (i.e. Switzerland and Lithuania combined) showed that ln P(X\K) decreased more-or-less constantly until K = 5, increased again until K = 10, and then reached a clear plateau (Online Resource 2A). Therefore, to detect the optimal number of clusters to represent our data best, we considered the change in ΔK, i.e. the ad hoc quantity related to the second-order rate of change of the log probability of data, with respect to the number of clusters (Evanno et al. 2005). The value of ΔK decreased until K = 3, then increased until K = 6, before decreasing and increasing again (Online Resource 2B). As ΔK never remained constant, we decided that our data could be best explained with one genetic cluster (K = 1). Qualitatively similar results were obtained when performing STRUCTURE analysis with the Lithuanian and the Swiss population separately (data not shown).
No discreet population structure could be determined by DAPC for any of the datasets analysed (Online Resources 3 and 4). The first discriminant function for the comparison between the Swiss and Lithuanian populations (Fig. 4) does not indicate any significant divergence in the two groups. Nonetheless, when reassigning the MLGs of each population to their population of origin (Fig. 5), about 80 % of all MLGs from the Swiss site-subpopulations could be reassigned to the Swiss population, while <40 % of the MLGs from the Lithuanian site-subpopulations could be re-assigned to the Lithuanian population (Fig. 5A). In the Swiss site-subpopulation AIG, the majority (58 %) of the MLGs found could be successfully reassigned to this subpopulation (Fig. 5B). Among the Lithuanian subpopulations, the highest reassignment rate was observed for BIR, but was still well below 30 %.
Reassignment probability of the H. fraxineus (sub-) populations investigated using nuclear microsatellite data of 719 H. fraxineus isolates. Each multilocus genotype was considered only once per subpopulation. A Individuals grouped by epidemic (Switzerland) and post-epidemic population (Lithuania). B All 10 subpopulations shown separately. For the subpopulation name abbreviations, see Table 1
Discussion
In the present study we aimed to genetically characterize epidemic and post-epidemic subpopulations of H. fraxineus, the causal agent of ash dieback. In both stages of the epidemic, no significant differences were found between the genotypic diversity of the H. fraxineus isolates derived from bark lesions and from the fallen leaf petioles. Moreover, the fungal multilocus genotypes (MLGs) obtained from bark lesions were not genetically differentiated from those recovered from petioles. These results suggest that all H. fraxineus MLGs identified in the present study may be able to induce bark lesions on trees and to saprophytically survive on fallen leaf petioles. The fungus usually first infects a living ash tree through the leaves (Gross et al. 2014a). Subsequently, H. fraxineus spreads along the leaf veins and may penetrate into the bark of twigs through leaf petioles. The bark infections represent an epidemiological dead-end, only the leaf infections are necessary for the reproduction of the pathogen. Thus, one could hypothesize that MLGs may differ in their ability to cause bark lesions, i.e. only a fraction of the MLGs infecting the leaves are also able to infect the bark. Similarly, not all MLGs colonizing living leaves may survive on the fallen leaves and petioles. Our study, however, seems to exclude niche partitioning of H. fraxineus MLGs between bark lesions on living trees and dead leaves and petioles in the litter.
The genetic diversity of H. fraxineus from sites in Switzerland, presumed to represent the epidemic disease phase and those from sites in Lithuania, presumed to represent post-epidemic populations, was unexpectedly very similar. By revealing high genotypic, but low allelic, diversity within the 10 subpopulations analysed, our results confirm previous findings (Bengtsson et al. 2012; Kraj et al. 2012; Gross et al. 2014b). The index of association statistics clearly points to the absence of clonal reproduction in either the epidemic or the post-epidemic subpopulations. This suggests that most genotypic diversity arises through random re-assorting of alleles during sexual reproduction. However, the GENCLONE analyses indicated that almost none of the repeated H. fraxineus MLGs, i.e. genotypes occurring more than once, seems to have appeared because of random mating within the subpopulations investigated. This could lead to the conclusion that clonal reproduction occurs in H. fraxineus, but given that this fungus is considered to be a strictly outcrossing organism (Gross et al. 2014a), such an assumption is not very realistic. On the other hand, the repeated MLGs may be due to the limited discriminatory power of the microsatellite markers used in this study, i.e. additional loci would be necessary to distinguish all MLGs present in our H. fraxineus populations. A simulation exercise showed that there is still no saturation in the number of genotypes detected when utilizing 11 loci.
The lack of significant differences in genetic diversity of H. fraxineus in epidemic and post-epidemic populations may be explained as follows. First, most of the genetic diversity that developed in the area of first introduction to Europe (i.e. Poland, Kowalski 2006) through sexual recombination between the initial MLGs has already been spread by range expansion. This contrasts with other invasive plant pathogens, for example the chestnut blight fungus Cryphonectria parasitica (Dutech et al. 2010; Prospero and Rigling 2012), whose range expansion in Europe mostly occurred with only a fraction of the genetic diversity present in the areas where the pathogen was first introduced. For H. fraxineus, the scenario suggests a natural windborne spread of the pathogen via clouds of ascospores rather than a human-mediated dispersal of single genotypes. Alternatively, it is possible that only a few divergent MLGs have spread across Europe. They may, after becoming established in healthy ash stands, have rapidly undergone sexual reproduction and thereby produced new MLGs.
Despite the large number of isolates genotyped, we only detected five novel (i.e. not previously reported) alleles. The longer disease history in Lithuania is probably the reason for the slightly higher number of mutations in the post-epidemic population there. Three private alleles were found in three MLGs (all from different isolates and subpopulations in Lithuania) compared to one MLG with two private alleles in the epidemic population in Switzerland. However, any differences in the private allelic richness (APR) of the epidemic and post-epidemic H. fraxineus populations were negligible. This indicates that 10–15 years, i.e. the approximate time lag between the appearance of ash dieback in Lithuania and in Switzerland, are not enough for a significant number of mutations to occur and become established. Moreover, as the allele frequencies in Switzerland (epidemic) were nearly the same as those in Lithuania (post-epidemic), we can conclude that allele surfing (Klopfstein et al. 2006) is still not occurring. In future, it would be interesting to check whether the frequency of the particular alleles may change in epidemic and/or post-epidemic populations, leading to a clear population differentiation.
Clustering analyses with STRUCTURE and DAPC revealed virtually no distinct groups of MLGs. Hence, we assume a single genetic cluster of H. fraxineus MLGs is present comprising all epidemic and post-epidemic subpopulations. This finding is in agreement with the results of previous studies (Bengtsson et al. 2012; Gross et al. 2014b). Strong gene flow both in Lithuania and Switzerland is further suggested by the lack of significant differentiation among the herein investigated subpopulations. It is worth noting that the gene flow among post-epidemic (Lithuanian) subpopulations seems to occur slightly more random than among epidemic (Swiss) subpopulations. In fact, the chances of correctly assigning a specific H. fraxineus MLG to its own subpopulation of origin were higher in Switzerland, especially for the pre-alpine subpopulations AIG (Aigle) and SAR (Sargans). This may reflect different founder events at the disease front, whose effects have not yet been completely wiped out by random mating and genetic exchange among the subpopulations. The gene flow in the expanding range of H. fraxineus in Switzerland may also be affected by physical barriers, such as the high Alpine mountain ranges.
Our study shows that there is only little geographic or ecological differentiation of the European H. fraxineus populations either at the disease front or in the post-epidemic area. The Swiss and Lithuanian subpopulations are genetically very similar despite the geographic distance between them and the very different dates of the first disease occurrence. Thus, the range expansion of H. fraxineus in Europe seems to have occurred through pushed colonization waves, i.e. with all of the genetic diversity of the population conserved at the disease front (Roques et al. 2012). Such an expansion pattern is consistent with the special annual life cycle of H. fraxineus, with its strictly outbreeding reproductive mode and heterothallic mating system, and the absence of a clonal spread (Gross et al. 2014a). Sexually produced ascospores are wind-dispersed and lead to a continuous genetic exchange between the epidemic and post-epidemic populations. This counteracts the effects of the founder events at the disease front. The emergence of new alleles via mutations seems to be rare, as does the increase in frequency of those already occurring. This rather special genetic population structure may make it difficult to develop an effective biological control of H. fraxineus. For example, several mycoviruses that reduce the virulence of their fungal host have been found in plant pathogens and may have the potential to be used as biological control agents (Ghabrial and Suzuki 2009). Recently, Schoebel et al. (2014b) detected a novel mitovirus (HfMV1) in European H. fraxineus isolates. The high genetic diversity of the fungal host population may, however, hinder the spread of such viruses that could potentially induce hypovirulence in their host. In fungi, the genetic diversity assessed by microsatellite markers typically correlates with the diversity of vegetative compatibility (vc) types (Liu et al. 1996; Milgroom et al. 2008). Consistent with these findings, the diversity of vc types in H. fraxineus invading Great Britain was recently reported to be high (Brasier and Webber 2013). As horizontal transmission of mycoviruses is restricted between fungal individuals of different vc types, the effectiveness of a biological control of H. fraxineus using mycoviruses will also depend on whether these can efficiently spread within H. fraxineus populations.
References
Agapow P-M, Burt A (2001) Indices of multilocus linkage disequilibrium. Mol Ecol Notes 1:101–102
Allendorf FW, Lundquist LL (2003) Introduction: population biology, evolution, and control of invasive species. Conserv Biol 17(1):24–30
Anderson PK, Cunningham AA, Patel NG, Morales FJ, Epstein PR, Daszak P (2004) Emerging infectious diseases of plants: pathogen pollution, climate change and agrotechnology drivers. Trends Ecol Evol 19(10):535–544
Arnaud-Haond S, Belkhir K (2007) GENCLONE: a computer program to analyse genotypic data, test for clonality and describe spatial clonal organization. Mol Ecol Notes 7:15–17
Austerlitz F, Jung-Muller B, Godelle B, Gouyon P-H (1997) Evolution of coalescence times, genetic diversity and structure during colonization. Theor Popul Biol 51:148–164
Barrett LG, Thrall PH, Burdon JJ, Linde CC (2008) Life history determines genetic structure and evolutionary potential of host-parasite interactions. Trends Ecol Evol 23(12):678–685
Bengtsson SBK, Vasaitis R, Kirisits T, Solheim H, Stenlid J (2012) Population structure of Hymenoscyphus pseudoalbidus and its genetic relationship to Hymenoscyphus albidus. Fungal Ecol 5(2):147–153
Billiard S, Lopez-Villavicencio M, Hood ME, Giraud T (2013) Sex, outcrossing and mating types? Unsolved questions in fungi and beyond. J Evol Biol 25:1020–1038
Boyd IL, Freer-Smith PH, Gilligan CA, Godfray HCJ (2013) The consequence of tree pests and diseases for ecosystem services. Science 342(6160):1235773
Brasier CM (2008) The biosecurity threat to the UK and global environment from international trade in plants. Plant Pathol 57:792–808
Brasier C, Webber J (2013) Vegetative incompatibility in the ash dieback pathogen Hymenoscyphus pseudoalbidus and its ecological implications. Fungal Ecol 6(6):501–512
Burdon JJ, Silk J (1997) Sources and patterns of diversity in plant pathogenic fungi. Phytopathology 87:664–669
Croucher PP, Mascheretti S, Garbelotto M (2013) Combining field epidemiological information and genetic data to comprehensively reconstruct the invasion history and the microevolution of the sudden oak death agent Phytophthora ramorum (Stramenopila: Oomycetes) in California. Biol Invasions 15:2281–2297
Dean MD, Ballard JWO (2004) Linking phylogenetics with population genetics to reconstruct the geographic origin of a species. Mol Phylogenet Evol 32:998–1009
DEFRA (2013) Chalara Management Plan. Department for Environment Food and Rural Affairs, London. https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/221051/pb13936-chalara-management-plan-201303.pdf. Accessed 10 June 2014
Desprez-Loustau M-L, Courtecuisse R, Robin C et al (2009) Species diversity and drivers of spread of alien fungi (sensu lato) in Europe with a particular focus on France. Biol Invasions 12:157–172
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14:927–930
Dutech C, Fabreguettes O, Capdevielle X, Robin C (2010) Multiple introductions of divergent genetic lineages in an invasive fungal pathogen, Cryphonectria parasitica in France. Heredity 105(2):220–228
Dutech C, Barrès B, Bridier J, Robin C, Milgroom MG, Ravigné V (2012) The chestnut blight fungus world tour: successive introduction events from diverse origins in an invasive plant fungal pathogen. Mol Ecol 21(16):3931–3946
Engesser R, Queloz V, Meier F, Kowalski T, Holdenrieder O (2009) Das Triebsterben der Esche in der Schweiz. [Ash dieback in Switzerland]. Wald und Holz 6:24–27 (in German)
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Ray N (2008) Surfing during population expansions promotes genetic revolutions and structuration. Trends Ecol Evol 23(7):347–351
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fichtner EJ, Rizzo DM, Kirk SA, Webber JF (2012) Infectivity and sporulation potential of Phytophthora kernoviae to select North American native plants. Plant Pathol 61(2):224–233
Fontaine M, Austerlitz F, Giraud T et al (2013) Genetic signature of a range expansion and leap-frog event after the recent invasion of Europe by the grapevine downy mildew pathogen Plasmopara viticola. Mol Ecol 22:2771–2786
Ghabrial SA, Suzuki N (2009) Viruses of plant pathogenic fungi. Annu Rev Phytopathol 47:353–384
Goudet J (2001) FSTAT, A Program to Estimate and Test Gene Diversities and Fixation Indices. Version 2.9.3, Department of Ecology and Evolution, University of Lausanne, Switzerland. http://www2.unil.ch/popgen/softwares/
Gross A, Grünig CR, Queloz V, Holdenrieder O (2012) A molecular toolkit for population genetic investigations of the ash dieback pathogen Hymenoscyphus pseudoalbidus. Forest Pathol 42:252–264
Gross A, Holdenrieder O, Pautasso M, Queloz V, Sieber TN (2014a) Hymenoscyphus pseudoalbidus, the causal agent of European ash dieback. Mol Plant Pathol 15(1):5–21
Gross A, Hosoya T, Queloz V (2014b) Population structure of the invasive forest pathogen Hymenoscyphus pseudoalbidus. Mol Ecol 23:2943–2960
Grünwald NJ, Garbelotto M, Goss EM, Heungens K, Prospero S (2012) Emergence of the sudden oak death pathogen Phytophthora ramorum. Trends Microbiol 20(3):131–138
Husson C, Scala B, Caёl O, Frey P, Feau N, Ioos R, Marçais B (2011) Chalara fraxinea is an invasive pathogen in France. Eur J Plant Pathol 130:311–324
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11(1):94
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281. http://dx.doi.org/10.7717/peerj.281. http://cran.r-project.org/web/packages/poppr/
Kirisits T, Dämpfle L, Kräutler K (2013) Hymenoscyphus albidus is not associated with an anamorphic stage and displays slower growth than Hymenoscyphus pseudoalbidus on agar media. Forest Pathol 43(5):386–389
Klopfstein S, Currat M, Excoffier L (2006) The fate of mutations surfing on the wave of a range expansion. Mol Biol Evol 23(3):482–490
Kowalski T (2006) Chalara fraxinea sp. nov. associated with dieback of ash (Fraxinus excelsior) in Poland. Forest Pathol 36(4):264–270
Kowalski T, Holdenrieder O (2009) Pathogenicity of Chalara fraxinea. Forest Pathol 39:1–7
Kraj W, Zarek M, Kowalski T (2012) Genetic variability of Chalara fraxinea, dieback cause of European ash (Fraxinus excelsior L.). Mycol Prog 11:37–45
Liu YC, Milgroom MG (2007) High diversity of vegetative compatibility types in Cryphonectria parasitica in Japan and China. Mycologia 99(2):279–284
Liu YC, Cortesi P, Double ML, MacDonald WL, Milgroom MG (1996) Diversity and multilocus genetic structure in populations of Cryphonectria parasitica. Phytopathology 86(12):1344–1351
Lygis V, Bakys R, Gustiene A, Burokiene D, Matelis A, Vasaitis R (2014) Forest self-regeneration following clear-felling of dieback-affected Fraxinus excelsior: focus on ash. Eur J Forest Res 133:501–510
McDonald BA (1997) The population genetics of fungi: tools and techniques. Phytopathology 87:448–453
McKinney LV, Nielsen LR, Collinge DB, Thomsen IM, Hansen JK, Kjær ED (2014) The ash dieback crisis: genetic variation in resistance can prove a long-term solution. Plant Pathol 63(3):485–499
Milgroom MG, Sotirovski K, Spica D et al (2008) Clonal population structure of the chestnut blight fungus in expanding ranges in Southeastern Europe. Mol Ecol 17(20):4446–4458
Pautasso M, Aas G, Queloz V, Holdenrieder O (2013) European ash (Fraxinus excelsior) dieback—a conservation biology challenge. Biol Conserv 158:37–49
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Prospero S, Rigling D (2012) Invasion genetics of the chestnut blight fungus Cryphonectria parasitica in Switzerland. Phytopathology 102:73–82
Prospero S, Rigling D (2013) Chestnut blight. In: Nicolotti G, Gonthier P (eds) Infectious forest diseases. CAB International, Wallingford, pp 318–339
R Development Core Team (2014) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/. Accessed 6 November 2014
Roques L, Garnier J, Hamel F, Klein EK (2012) Allee effect promotes diversity in traveling waves of colonization. PNAS 109(23):8828–8833
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rousset F (2008) Genepop’007: a complete reimplementation of the Genepop software for Windows and Linux. Mol Ecol Resources 8:103–106
Sakai AK, Allendorf FW, Holt JS et al (2001) The population biology of invasive species. Annu Rev Ecol Syst 32:305–332
Santini A, Ghelardini L, De Pace C et al (2013) Biogeographical patterns and determinants of invasion by forest pathogens in Europe. New Phytol 197:238–250
Schoebel CN, Stewart J, Gruenwald NJ, Rigling D, Prospero S (2014a) Population history and pathways of spread of the plant pathogen Phytophthora plurivora. PLoS ONE 9(1):e85368
Schoebel CN, Zoller S, Rigling D (2014b) Detection and genetic characterization of a novel mycovirus in Hymenoscyphus fraxineus, the causal agent of ash dieback. Infect Gen Evol 28:78–86
Szpiech ZA, Jakobsson M, Rosenberg NA (2008) ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24:2498–2504
Timmermann V, Børja I, Hietala AM, Kirisits T, Solheim H (2011) Ash dieback: pathogen spread and diurnal patterns of ascospore dispersal, with special emphasis on Norway. EPPO Bull 41:14–20
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population-structure. Evolution 38:1358–1370
Westphal MI, Browne M, MacKinnon K, Noble I (2008) The link between international trade and the global distribution of invasive alien species. Biol Invasions 10:391–398
Xhaard C, Barrès B, Andrieux A, Bousset L, Halkett F, Frey P (2012) Disentangling the genetic origins of a plant pathogen during disease spread using an original molecular epidemiology approach. Mol Ecol 21:2383–2398
Acknowledgments
We would like to thank the foresters in both countries for their help in finding diseased ash stands, as well as Hélène Blauenstein, Romina Sigrist and Donata Kinčiūtė for their help with field and laboratory work. We are also grateful to the Genetic Diversity Center of ETH Zurich for their support, and to the European Cooperation in the Field of Scientific and Technical Research (COST) Programme Action No. FP1103 ‘FRAXBACK’ for granting two Short-Term Scientific Missions (STSM’s). This study was financially supported by the Lithuanian-Swiss cooperation programme to reduce economic and social disparities within the enlarged European Union (project grant agreement no. CH-3-ŠMM-01/12). Finally, we are grateful to Silvia Dingwall for English revision of the manuscript and to the two anonymous reviewers for valuable comments on the manuscript.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Burokiene, D., Prospero, S., Jung, E. et al. Genetic population structure of the invasive ash dieback pathogen Hymenoscyphus fraxineus in its expanding range. Biol Invasions 17, 2743–2756 (2015). https://doi.org/10.1007/s10530-015-0911-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-015-0911-6