Abstract
Brown trout of German origin were introduced into Patagonian National Parks in 1905, where they acclimatized and underwent population expansion endangering populations of native species like Galaxiidae. Spawning adults of two populations were sampled in 2004. Their age, length-at-age and migratory behaviour were assessed from scale samples, as well as their variation at the coding LDH-C1* and eight non coding microsatellite loci. Between-population differentiation for life history (spawning time, migratory behaviour, length and weight at age) and reduced genetic variation were revealed. Based on genetic variation, effective population size smaller than 50 individuals has been estimated for the founder stock, and its German origin has been genetically traced. Flexibility in migratory behaviour and spawning time were identified as key factors conferring competitive advantage on those brown trout populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Critical tests in nature of abiotic and biotic factors include transplant experiments at macroscales to decipher the maintenance of genetic diversity (Nevo 2001) and the processes leading to colonisation of new environments and invasions. However, new deliberate macroscale transplants cannot be supported if ecosystem conservation is a concern (Novacek and Cleland 2001; Lodge and Shrader-Frechette 2003; Sanders et al. 2003). Instead, we can learn from transplants performed many years ago and which resulted in successful adaptation of alien species.
The brown trout Salmo trutta L. populations that inhabit the National Parks of the province of Neuquen (Patagonia) since the beginning of the 20th century are famous worldwide, due to landscape values and the trophy quality of catches. Although the region has experienced intense damming during the last 30 years (Mugetti et al. 2004), which is a priori a potential risk for long-term survival of salmonids (Mesquita et al. 2005; Whiteley et al. 2006; Layman et al. 2007), they have successfully adapted to South American ecosystems (Valiente et al. 2007). This species exhibits a consistent ability of adapting to adverse environmental conditions, including damming (Ayllon et al. 2006a). Originally from the northern Hemisphere, it is able to colonise regions as remote as New Zealand (Townsend 2003) and the sub-Antarctic Kerguelen Islands (Ayllon et al. 2006b). It has been declared a potential invader by the IUCN.
Early adaptive evolution in a novel environment can include changes to plastic character states (Parsons and Robinson 2006), sos phenotypic plasticity in life history traits (Pettersson et al. 2001) may help to explain rapid adaptation. Townsend (2003) suggested that hatching time may be one life-history feature involved in brown trout invasiveness, at least in streams where redds of native species like galaxiids are affected by high-flow events. Another factor of invasiveness may be a flexible migratory strategy, which enhances rapid adaptation of naturalised brown trout to new ecosystems (Ayllon et al. 2006b). For example, local adaptation can be enhanced by decreased migratory activity, as it typically occurs when populations subjected to different selective pressures exchange few or no migrants (Garcia de Leaniz et al. 2007).
Based on reduced genetic variability of Patagonian brown trout, Valiente et al. (2007) suggested founder effects and bottlenecks that were not an obstacle for its expansion. The objectives of the present study were two-fold. First, to quantify the bottlenecks overcome by Patagonian brown trout employing their genetic divergence from the donor stock as a basis for estimating effective population sizes. Second, to explore the role of life-history traits in the expansion of this species in Patagonian ecosystems. The final aim of the study was to contribute to understanding what biological features can promote invasive processes, for further application in conservation. Two Salmo trutta populations acclimatized to river-lake-reservoirs ecosystems near the Andean chain were chosen as a case study.
Materials and methods
Populations analyzed
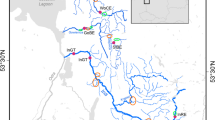
The Patagonian Limay River (Fig. 1) originates from two lakes located in protected natural areas of the Andean chain: Lake Nahuel Huapi (within Nahuel Huapi National Park) and Lake Huechulafquen (within Lanin National Park, near the extinct Lanin Volcano). Brown trout have inhabited the Limay River since their introduction there in 1915, transplanted from Chile where it had been naturalised in 1905. True anadromy (migration to sea) was never possible for these Patagonian trout because access to the Atlantic Ocean is closed by old downstream dams constructed before brown trout introduction. The sampling site on the Limay River (40º 22′S-70º 02′W) was located in the mainstream, near the reservoir Piedra del Aguila. Another sampling site (Currhué Grande River, 39º 52′S-71º 25 W) was located upstream the large river system (Fig. 1). Some habitat characteristics and a list of other species inhabiting these ecosystems are presented in Table 1. In May 2004, these two Argentinean brown trout populations were sampled by electrofishing and anaesthetized, measured and weighted, then released in the sampling area after recovering. The sampling area was similar in both systems, about 2 km2. Scale samples (three-five scales from the anterior dorsal section) were collected from spawning adults of each population for determination of life-history traits (43 and 116 individuals from the Limay and Currhué Grande rivers, respectively) and genetic analysis (50 individuals per population).
Samples from the River Elbe drainage (Lamitz from the mainstream and Gol from a separate tributary) were considered as representative of the German drainage where (via Chile) these Patagonian trout originated (Valiente et al. 2007). Two other populations representative of the two main German drainages were also analysed to confirm the Elbe River origin of the Patagonian trout by eliminating (if pertinent) the two other main German drainages: Dreiherrenbrunnen (Rhine River) and Haarauer Saige (Danube River). Adipose fins of these samples were employed as a source of DNA for genetic analyses (50 samples per population).
Life-history traits
Sex was determined by direct observation, based on secondary sexual traits or, if necessary owing to ambiguous phenotype, by gently stripping the abdomen to obtain gametes. Age and migratory status were determined from scales following Busacker et al. (1990), and size (length) at age was estimated by retrocalculation based on statistical regressions of distances between scale annulus, the scale diameter and fork length. Spawning events can be interpreted in scales as deep striations. Migration to lakes or reservoirs was identified in scales by periods of differential growth, that is, larger spaces between age marks.
Spawning time was directly observed in the Patagonian sampling sites in 2004.
Statistical comparisons between mean length and weight at age of different types of trout (migratory versus resident) were performed with the programme SSPS 8.0. They consisted of two-sided t-tests, after assessing equality of variances by Levene’s test.
Genetic analysis
DNA was extracted following Estoup et al. (1996). Five non-coding microsatellite loci were analyzed: SSOSL85, SOSL311 and SSOSL417 (Slettan et al. 1995), SS4 (Martinez et al. 1999) and BF002 (Sušnik et al. 1997). Microsatellite loci amplification and typing followed the procedures described in Valiente et al. (2007). One protein-coding locus (LDH-C1*) was also analyzed. LDH-C1* genotypes were determined by restriction digestion of the gene DNA sequence (McMeel et al. 2001).
Analysis of population structure
Scoring errors, large allele dropout and null alleles at microsatellite loci were checked with MICROCHECKER (Van Oosterhout et al. 2004). Number of alleles per locus, heterozygosity observed and expected and allele frequencies were determined with the GENETIX (2000) computer package. The programme GENEPOP version 3.4 (Raymond and Rousset 1995) was employed to test conformity to the Hardy Weinberg equilibrium (HWE) by the Markov chain method (10,000 dememorization steps, 1,000 batches and 10,000 iterations per batch).
Evidence of population structuring was obtained employing the programme STRUCTURE (Pritchard et al. 2000). In this programme each individual is considered separately. The programme was designed to identify the K (unknown) populations (genetic clusters) of origin of individuals. For each K value 10 independent runs were run for two different burn-in periods (105 and 106), with 106 iterations. The most probable K value is chosen based on its likelihood and low variance between runs, as well as consistent assignment of individuals to genetic clusters in different runs. The programme ARLEQUIN (Schneider et al. 2000) was employed for estimates of F ST values between pairs of samples and their statistical significance, i.e. population differentiation, with the following settings: 10,000 permutations for significance, 10,000 steps in Markov chain.
Genetic identification of the original German donor population
Self-assignment tests (Bayesian method) were done using the programme GENECLASS 1.0.02 (Cornuet et al. 1999). This is a likelihood-based method in which individuals are assigned to the population in which the likelihood of their genotype is highest. The following settings were used: 10,000 simulated individuals, threshold = 0.0100 and Bayesian estimation of frequencies.
Estimates of effective population size
Two different methods were employed to estimate effective population sizes. One was based on temporal changes in allele frequencies between the ancestral Elbe River and the Patagonian populations, using a likelihood-based estimation (Berthier et al. 2002). Temporal stability of the Elbe River population was assumed, as well as isolation of the Patagonian systems.
The other method, more direct, was based on the multilocus F ST value under complete isolation as a function of time (generations), the effective population size Ne, the number of subpopulations and the heterozygosity averaged across subpopulations, under the infinite alleles model (IAM), following Jin and Chakraborty (1995) modified by Raeymaekers et al. (2005). Although an IAM mutational model is unrealistic for microsatellites, it is simple and robust and has been used for microsatellites in other fish species (Raeymaekers et al. 2005; Palkovacs et al. 2008).
Results
Brown trout coexist in both studied ecosystems with South American native species, including Galaxiids like Galaxias maculatus and G. platei (Table 1). They also share the habitat with one (mainstream Limay) or more (Currhué Grande River) other exotic salmonids.
In 2004 there was a clear difference in spawning time between both sites. Spawning occurred earlier in Currhué Grande (June–July; Table 1) than downstream (mainstream Limay), where it happened between August and September. This trend has been consistent at least for the last 10 years. Statistics are unnecessary given the observational nature of these data.
The age at spawning was not equal for the two Patagonian populations. In Currhué Grande River the brood stock was composed of 11, 26, 42, 22 and 5 individuals aged 3–7 years old, respectively, whereas the mature adults sampled from the mainstream Limay were 12, 15, 10 and 6 individuals 2, 3, 4 and 5 years old, respectively (Table 2). This gives an age at spawning of 3.5 and 2.7 for Currhué Grande and Limay Rivers, respectively. Sex-ratio (females:males) was 1.23 and 0.72 for the Currhué and Limay Rivers, respectively. In the Limay River the difference between migratory and sedentary trout for sex-ratio was not significant (60 and 32.1% females for migratory and sedentary trout, respectively, Table 2); contingency Chi-square of 3.06, 1 df, P > 0.05.
With respect to migratory behaviour, the two Patagonian populations exhibited different life histories (Fig. 2). Trout inhabiting the Currhué Grande River were all migratory whereas 65% of trout sampled from the mainstream Limay were sedentary. After measuring distances between scale annulus, scale diameter and trout length, length at age was individually estimated for the spawning adults analyzed in both populations. The graphs obtained for length at age in Currhué Grande and Limay migratory trout (Fig. 3) indicate that the trout grew faster in the mainstream Limay. This was confirmed by statistical comparison of weight between trout of the same age inhabiting the two ecosystems (Table 3). Values of t (P-value) were 3.76 (0.002), 2.47 (0.018) and 3.52 (0.002) for 3, 4 and 5 year old migratory trout, respectively (equal variances confirmed by Levene’s tests in each case). Comparison between migratory and resident trout was possible only for the Limay River population, as resident trout did not appear in Currhué Grande River. Migratory brown trout exhibited higher growth, were significantly longer (Fig. 3; P < 0.001 at all ages except at 1 year old for which P = 0.042) and heavier (Table 3) than sedentary trout (t = 3.11 with P = 0.011, 6.31 with P = 0.0001, 4.54 with P = 0.002 and 2.91 with P = 0.044 for 2, 3, 4 and 5 years old trout, respectively). Actually, sedentary trout grew even less than the Currhué Grande all-migratory individuals, at least at 3 and 4 years old (t = 3.96 with P = 0.001 and t = 3.89 with P = 0.0001, respectively).
Variability found at the microsatellite loci analyzed in the Patagonian populations and the German stocks is depicted in Table 4. The Limay River population did not differ significantly from HWE after a sequential Bonferroni correction for multiple tests (Chi-square = 19.8, P = 0.0328), whereas the Currhue population significantly deviated from it (Chi-square = Infinity, Highly Significant) due to an excess of different homozygotes. Allelic richness values were significantly lower (P = 0.04) for the two Patagonian populations than for the Elbe River stocks (Fig. 4), but not significantly different from those obtained for the other two German populations analyzed.
Using a Bayesian self-assignment test on microsatellite datasets (Table 5), only 14 (28%) and 12 (24%) individuals from the Limay and Currhué Grande River populations, respectively, were still assigned to German stocks, mostly to the Elbe River, confirming the expected origin of Patagonian brown trout from a local Elbe River hatchery. However some contribution from the Rhine drainage cannot be excluded, as some individuals from the Currhué Grande River were significantly assigned to the Rhine and the Elbe systems simultaneously.
Differentiation between the two Patagonian samples, measured as F ST = 0.0711 (Table 6), was highly significant after strict sequential Bonferroni correction, but lower than their differentiation with respect to the ancestral Elbe River populations. The global value of F ST between Patagonian and Elbe River samples was 0.1436. The programme STRUCTURE allowed us to identify each Patagonian population as a separate genetic unit, also separated from the ancestral German populations. When the two Patagonian populations (Limay, Currhue Grande) were run in the programme STRUCTURE together with the populations inhabiting two Elbe River tributaries (Lamitz, Gol), the number of separate population units estimated was four. For K = 4 all runs were consistent, with a mean variance of likelihood of 512, alpha value 0.035 stable along the runs. More than 90% of Patagonian individuals were strongly assigned (>0.9 likelihood) to Limay or Currhue; similarly, more than 90% Elbe River individuals were strongly assigned to Lamitz or Gol. These results support strong differentiation between the two Patagonian populations, as well as between the two Elbe River populations.
The results obtained for the coding locus LDH-C1* did not reveal significant differentiation between the two Patagonian populations (Table 4). The Patagonian samples were variable at this locus, exhibiting the two alleles *90 and *100. Their allele frequencies (0.20 and 0.15 for Currhue Grande and Limay samples, respectively) were not statistically different. Significantly higher *100 allele frequency was found for the Patagonian trout samples than for their ancestors from the Elbe River (Chi-Square = 4.28, P < 0.05 pooling together the two samples from each region).
The Patagonian National Parks were colonised by brown trout populations likely smaller than 50 individuals as effective population size. From the variance of allele frequencies between the donor Elbe River and the Patagonian populations (Berthier et al. 2002), the estimated effective population sizes were 39.6 and 49.2 for the Limay and Currhué Grande populations, respectively, assuming 28 generations since 1905 (mean age at first reproduction in this area, 3.63 ± 0.93 years). When the method based on multilocus F ST value was employed, effective population size for Patagonian brown trout populations ranged between 17.18 and 35.6, for Ho averaged across populations of 0.590 (SD 0.182, range 0.408–0.772).
Discussion
Although based on modest sample sizes due to inherent sampling difficulties, this study provides some insights into the invasive capacity of brown trout. Flexibility in life-history traits is probably one key factor for invasiveness. Migratory behaviour is a good example. It is characteristic of northwest European brown trout populations, including those established in Germany such as the Elbe River stocks (Elliott 1994). However, the Patagonian populations evolved—or just adopted—different life history patterns. Although the Currhué Grande population is still all-migratory, an important portion of the Limay River population, 65%, is sedentary at the present. Such a drastic change in migratory behaviour may explain why Salmo trutta can become an invader. Even at low densities, brown trout achieve high biomass and this ability has been signalled as one of the main reasons for their impact on native fish communities (Townsend 2003), which are less able to achieving high biomass (Huryn 1998). In this study we have shown that migratory trout achieve higher biomass than sedentary morphs. In the mainstream Limay, where growth is fast likely due to abundant food and space in the vicinity of reservoirs, brown trout are surely less pressed for abandoning the river habitat for migrating to reservoirs and may choose staying at native places. Upstream near the glacial Nahuel Huapi lake, in Currhué Grande, migratory behaviour enables faster growth and exploitation of non-local resources. Olsson et al. (2006) have demonstrated that high density and scarcity of food determine migratory behaviour in this species, so this trait seems to be more environmentally than genetically determined. Between-population differences found for Patagonian trout doubtless reflect plasticity in life-history tactics known for this species (e.g. Pettersson et al. 2001).
Changes in other life-history traits, like spawning time, may be explained by environmental factors (the Currhué Grande sampling site is located 500 m altitude over the mainstream Limay and nearer the Andean mountains, so climate conditions are likely different), but also by the accompanying species. Competition with rainbow trout (Oncorhynchus mykiss) for relatively reduced spawning area (Table 1) could contribute to explain delayed spawning in the mainstream Limay site. Rainbow trout spawning, occurring in the spring, can destroy migratory brown trout spawning redds produced during the winter (Landergren 1999). Delayed spawning time (August–September) of sedentary brown trout would prevent redd (nest) destruction at the time of fry emergence and early dispersal due to rainbow trout spawning activity. Late hatching has been suggested as a factor promoting less vulnerability of brown trout redds to environmental disturbances like high-flow events (Townsend 2003). Our observations reinforce this idea. On the other hand, differences in spawning time may promote population structuring in brown trout. Population isolation by spawning time has been described for fish species from eel (Anguilla anguilla) (Maes et al. 2006) to herring (Clupea harengus) (Ruzzante et al. 2006) to sockeye salmon (Oncorhynchus nerka) (Ramstad et al. 2004). Genetic determination of spawning time has been recently found in rainbow trout through QTL analysis (Leder et al. 2006). If subjacent genetic components of this life history trait also exist in brown trout, we can speculate that change in spawning time may produce biological reproductive barriers between Patagonian brown trout populations that could lead to incipient speciation. Changes in life-history traits are not necessarily a true case of evolution—change at genetic level. However, environmentally initiated novelties may have greater evolutionary potential than mutationally induced ones. Thus, genes are probably more often followers than leaders in evolutionary change (West-Eberhard 2005). Further population evolution associated with this change in migratory and spawning behaviour can be expected.
Significant genetic differentiation between the Patagonian populations was detected at neutral microsatellite loci, at a level as high as the differentiation existing between populations inhabiting different European drainages. Brown trout populations are highly structured and significant genetic differences often exist between closely neighbouring subpopulations (Ferguson 1989; Altukhov et al. 2000), thus this result can be expected. However, significant changes in allele frequencies at the coding locus LDH-C1* have not arisen between the two Patagonian populations. Slight but significant increase of the *100 allele in both Patagonian stocks with respect to the Elbe River populations may suggest some selection. Lactate dehydrogenase protein variation is selectively constrained by temperature (Colosimo et al. 2003). The variation detected in this study affects the active site of the enzyme (McMeel et al. 2001), which may have a direct role in catalysis (Henry and Ferguson 1985). The ancestral allele *100, characteristic of southern European regions (Hamilton et al. 1989) may not be neutral in the new environmental conditions. Rapid population evolution due to combined differential gene flow and selection has been reported for introduced brown trout in other regions (Ayllon et al. 2006b). The results presented in this study reinforce the strong role of these factors for shaping population genetic structure in this species.
Based on F ST values between Patagonian and their donor Elbe River stock, as well as in changes in microsatellite allele frequencies, our results suggest that colonisation of Argentine Patagonian national parks was probably achieved by less than 50 trout founders—or maybe the Patagonian populations have overcome bottlenecks reducing effective population size to those low levels. Reduced effective population size has not been an obstacle for adaptation and proliferation of brown trout in Patagonia (Valiente et al. 2007), as it is not for other invaders (Tsutsui et al. 2000; Sax et al. 2005). Plasticity in life-history traits, as suggested by Lande (1988), seems to be more important than genetics for successful adaptation and expansion of populations. A few individuals can colonise entire drainages or ecosystems.
It is not easy to extract from this idea practical lessons to be applied in conservation, except the general precautionary approach of totally preventing introduction of foreign species into new environments. Total restriction of stock and species movements should be strictly respected if conservation of wild ecosystems is an issue for scientists and managers. Removal of brown trout from Patagonian ecosystems, although it is known that native species like galaxiidae are threatened by this invader (Huryn 1998; Baigun and Ferriz 2003), is currently impossible precisely due to its extraordinary ability to adapt to different ecosystems. Besides, the species sustains local economies attracting tourism and is highly valued, in Argentina and in many other places, so its eradication is probably not desirable from the economic point of view because the functioning of native ecosystems is not equally valued (Townsend 2003). Development of international directives and their implementation in real life, together with environmental education emphasising the risk of invasions, are suggested as methods for preventing future threats to biodiversity.
References
Altukhov YP, Salmenkova EA, Omelchenko VT (2000) Salmonid fishes. Population biology, genetics and management. Fish Biology and Aquatic Resources series. Blackwell Science Ltd., Oxford, 368 pp
Ayllon F, Moran P, Garcia-Vazquez E (2006a) Maintenance of a small anadromous subpopulation of brown trout (Salmo trutta L.) by straying. Freshw Biol 51:351–358. doi:10.1111/j.1365-2427.2005.01486.x
Ayllon F, Davaine P, Beall E et al (2006b) Dispersal and rapid evolution in brown trout colonizing virgin Subantarctic ecosystems. J Evol Biol 19:1352–1358. doi:10.1111/j.1420-9101.2005.01075.x
Baigun C, Ferriz R (2003) Distribution patterns of native freshwater fishes in Patagonia (Argentina). Org Divers Evol 3:151–159. doi:10.1078/1439-6092-00075
Berthier P, Beaumont MA, Cornuet JM et al (2002) Likelihood based estimation of the effective population size using temporal changes in allele frequencies: a genealogical approach. Genetics 160:741–751
Busacker GP, Adelman IR, Goolish EM (1990) Growth. In: Schreck CB, Moyle PB (eds) Methods for fish biology. American Fisheries Society, Bethesda, pp 363–387
Colosimo A, Giuliani A, Maranghi F et al (2003) Physiological and genetical adaptation to temperature in fish populations. Cont Shelf Res 23:1919–1928. doi:10.1016/j.csr.2003.06.012
Cornuet JM, Piry S, Luikart G et al (1999) New methods employing multilocus genotypes to select or exclude populations as origins of individuals. Genetics 153:1989–2000
Elliott JM (1994) Quantitative ecology of the Brown Trout. Oxford University Press, London
Estoup A, Largiader CR, Perrot E et al (1996) Rapid one-tube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Mol Mar Biol Biotechnol 5:295–298
Ferguson A (1989) Genetic differences among brown trout, Salmo trutta, stocks and their importance for the conservation and management of the species. Freshw Biol 21:35–46. doi:10.1111/j.1365-2427.1989.tb01346.x
Garcia de Leaniz C, Fleming IA, Einum S et al (2007) A critical review of adaptive genetic variation in Atlantic salmon: implications for conservation. Biol Rev Camb Philos Soc 82:173–211. doi:10.1111/j.1469-185X.2006.00004.x
Hamilton KE, Ferguson A, Taggart JB et al (1989) Post-glacial colonization of brown trout, Salmo trutta L. Ldh-5 as a phylogeographic marker locus. J Fish Biol 35:651–664. doi:10.1111/j.1095-8649.1989.tb03017.x
Henry T, Ferguson A (1985) Kinetic studies on the lactate dehydrogenase (LDH-5) isozymes of brown trout, Salmo trutta L. Comp Biochem Physiol 82B:95–98
Huryn AD (1998) Ecosystem level evidence for top–down and bottom–up control of production in a grassland stream system. Oecologia 115:173–183. doi:10.1007/s004420050505
Jin L, Chakraborty R (1995) Population structure, stepwise mutations, heterozygote deficiency and their implications in DNA forensics. J Hered 74:274–285. doi:10.1038/hdy.1995.41
Lande R (1988) Genetics and demography in biological conservation. Science 241:1455–1460. doi:10.1126/science.3420403
Landergren P (1999) Spawning of anadromous rainbow trout, Oncorhynchus mykiss (Walbaum): a threat to sea trout, Salmo trutta L, populations? Fish Res 40:55–63
Layman CA, Quattrochi JP, Peyer CM et al (2007) Niche width collapse in a resilient top predator following ecosystem fragmentation. Ecol Lett 10:937–944. doi:10.1111/j.1461-0248.2007.01087.x
Leder EH, Danzmann RG, Ferguson MM (2006) The candidate gene, clock, localizes to a strong spawning time quantitative trait locus region in rainbow trout. J Hered 97:74–80. doi:10.1093/jhered/esj004
Lodge DM, Shrader-Frechette K (2003) Nonindigenous species: ecological explanation, environmental ethics, and public policy. Conserv Biol 17:31–37. doi:10.1046/j.1523-1739.2003.02366.x
Maes GE, Pujolar JM, Hellemans B et al (2006) Evidence for isolation by time in the European eel (Anguilla anguilla L.). Mol Ecol 15:2095–2107. doi:10.1111/j.1365-294X.2006.02925.x
Martinez JL, Moran P, Garcia-Vazquez E (1999) Dinucleotide repeat polymorphism at the SS4, SS6 and SS11 loci in Atlantic salmon (Salmo salar). Anim Genet 30:462–478. doi:10.1046/j.1365-2052.1999.00498-3.x
McMeel OM, Hoey EM, Ferguson A (2001) Partial nucleotide sequences, and routine typing by polymerase chain reaction–restriction fragment length polymorphism, of the brown trout (Salmo trutta) lactate dehydrogenase, LDH-C1*90 and *100 alleles. Mol Ecol 10:29–34. doi:10.1046/j.1365-294X.2001.01166.x
Mesquita N, Hänfling B, Carvalho GR et al (2005) Phylogeography of the cyprinid Squalius aradensis and implications for conservation of the endemic freshwater fauna of southern Portugal. Mol Ecol 14:1939–1954. doi:10.1111/j.1365-294X.2005.02569.x
Mugetti AC, Calcagno AT, Brieva CA et al (2004) Aquatic habitat modifications in La Plata River basin, Patagonia and associated marine areas. Ambio 33:78–87. doi:10.1639/0044-7447(2004)033[0078:AHMILP]2.0.CO;2
Nevo E (2001) Evolution of genome-phenome diversity under environmental stress. Proc Natl Acad Sci USA 98:6233–6240. doi:10.1073/pnas.101109298
Novacek MJ, Cleland EE (2001) The current biodiversity extinction events: scenarios for mitigation and recovery. Proc Natl Acad Sci USA 98:5466–5470. doi:10.1073/pnas.091093698
Olsson IC, Greenberg LA, Bergman E et al (2006) Environmentally induced migration: the importance of food. Ecol Lett 9:645–651. doi:10.1111/j.1461-0248.2006.00909.x
Palkovacs EP, Dion CB, Post DM et al (2008) Independent evolutionary origins of landlocked alewife populations and rapid parallel evolution of phenotypic traits. Mol Ecol 17:582–597
Parsons KJ, Robinson BW (2006) Replicated evolution of integrated plastic responses during early adaptive divergence. Evol Int J Org Evol 60:801–813
Pettersson JCE, Hansen MM, Bohlin T (2001) Does dispersal from landlocked trout explain the coexistence of resident and migratory trout females in a small stream? J Fish Biol 58:487–495. doi:10.1111/j.1095-8649.2001.tb02267.x
Pritchard JK, Stephens M, Donnelly PJ (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raeymaekers JAM, Maes GE, Audenaert E et al (2005) Detecting Holocene divergence in the anadromous-freshswater three-spined stickleback (Gasterosteus aculeatus) system. Mol Ecol 14:1001–1014. doi:10.1111/j.1365-294X.2005.02456.x
Ramstad KM, Woody CA, Sage GK et al (2004) Founding events influence genetic population structure of sockeye salmon (Oncorhynchus nerka) in Lake Clark, Alaska. Mol Ecol 13:277–290. doi:10.1046/j.1365-294X.2003.2062.x
Raymond M, Rousset F (1995) GENEPOP (version 1.2): a population genetics software for exact test and ecumenicism. J Hered 86:248–249
Ruzzante DE, Mariani S, Bekkevold D et al (2006) Biocomplexity in a highly migratory pelagic marine fish, Atlantic herring. Proc R Soc B Biol Sci 273:1459–1464. doi:10.1098/rspb.2005.3463
Sanders NJ, Gotelli NJ, Heller NE et al (2003) Community disassembly by an invasive species. Proc Natl Acad Sci USA 100:2474–2477. doi:10.1073/pnas.0437913100
Sax DF, Stachowicz JJ, Gaines SD (eds) (2005) Species invasions: insights into ecology, evolution, and biogeography. Sinauer Associates, Sunderland
Schneider S, Roessli D, Excoffier L (2000) Arlequin ver. 2000: a software for population genetics data analysis. Genetics and Biometry Laboratory, University of Geneva, Switzerland
Slettan A, Olsaker I, Lie Ø (1995) Atlantic salmon, Salmo salar, microsatellites at the SSOSL25, SSOSL85, SSOSL311, SSOSL417 loci. Anim Genet 26:281–282
Sušnik S, Snoj A, Pohar J et al (1997) The microsatellite marker (BRFO 002) characteristic for different geographically remote brown trout, Salmo trutta L., populations. Anim Genet 28:370–383
Townsend CR (2003) Individual, population, community and ecosystem consequences of a fish invader in New Zealand streams. Conserv Biol 17:38–47. doi:10.1046/j.1523-1739.2003.02017.x
Tsutsui ND, Suarez AV, Holway DA et al (2000) Reduced genetic variation and the success of an invasive species. Proc Natl Acad Sci USA 97:5948–5953. doi:10.1073/pnas.100110397
Valiente AG, Juanes F, Nuñez P et al (2007) Is genetic variability so important? Non-native salmonids in South America. J Fish Biol 71:1–12
Van Oosterhout C, William F, Hutchinson DP et al (2004) Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538. doi:10.1111/j.1471-8286.2004.00684.x
West-Eberhard MJ (2005) Developmental plasticity and the origin of species differences. Proc Natl Acad Sci USA 102:6543–6549. doi:10.1073/pnas.0501844102
Whiteley AR, Spruell P, Allendorf FW (2006) Can common species provide valuable information for conservation? Mol Ecol 15:2767–2786
Acknowledgments
We thank Ivan G. Pola (University of Oviedo) for technical assistance and Dr Ulrich Schliewen (Zoologische Staatssammlung München, Germany) for German samples. Alexandros Triantafyllidis (University of Thessaloniki, Greece) kindly revised the manuscript. Spanish Grants MCYT REN2003-00303 and PR2004-0084 (to EGV) and a Hatch Grant (to FJ) provided funding.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Valiente, A.G., Juanes, F., Nuñez, P. et al. Brown trout (Salmo trutta) invasiveness: plasticity in life-history is more important than genetic variability. Biol Invasions 12, 451–462 (2010). https://doi.org/10.1007/s10530-009-9450-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-009-9450-3