Abstract
Purpose
Purple acid phosphatases (PAPs) includ the largest classes of non-specific plant acid phosphatases. Most characterized PAPs were found to play physiological functions in phosphorus metabolism. In this study, we investigated the function of AtPAP17 gene encoding an important purple acid phosphatase in Arabidopsis thaliana.
Methods
The full-length cDNA sequence of AtPAP17 gene under the control of CaMV-35S promoter was transferred to the A. thaliana WT plant. The generated homozygote AtPAP17-overexpressed plants were compared by the types of analyses with corresponding homozygote atpap17-mutant plant and WT in both + P (1.2 mM) and − P (0 mM) conditions.
Results
In the + P condition, the highest and the lowest amount of Pi was observed in AtPAP17-overexpressed plants and atpap17-mutant plants by 111% increase and 38% decrease compared with the WT plants, respectively. Furthermore, under the same condition, APase activity of AtPAP17-overexpressed plants increased by 24% compared to the WT. Inversely, atpap17-mutant plant represented a 71% fall compared to WT plants. The comparison of fresh weight and dry weight in the studied plants showed that the highest and the lowest amount of absorbed water belonged to OE plants (with 38 and 12 mg plant−1) and Mu plants (with 22 and 7 mg plant−1) in + P and − P conditions, respectively.
Conclusion
The lack of AtPAP17 gene in the A. thaliana genome led to a remarkable reduction in the development of root biomass. Thus, AtPAP17 could have an important role in the root but not shoot developmental and structural programming. Consequently, this function enables them to absorb more water and eventually associated with more phosphate absorption.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phosphorus (P) is one of the most essential macronutrients for all cells, which plays important roles in macromolecules structure such as nucleic acids, and phospholipids. This element participates in the signal transduction pathways or as a constituent in energy-transferring molecules such as ATP, ADP, or NADPH in the photosynthesis and respiration cycles. Phosphorous makes up about 0.2% of plant dry weight. To ensure functional catabolic reactions, Pi (inorganic phosphate) homeostasis must be conserved between 5 and 20 mM in the plant cytoplasm. Plants acquire P only in the soluble Pi form either as H2PO4− or HPO42−, whereas these forms occur in the soil between 0.1 and 1 μM. Therefore, it is one of the key nutrients for plant growth and development that is considered a challenging agent as well as a major limiting factor in crop yield (Schachtman et al. 1998; Hurley et al. 2010).
Higher plants use many sophisticated responses to overcome nutritional stress by increasing phosphate scavenging and recycling to the maintenance of cellular Pi homeostasis (Yadava et al. 2022). Plants respond to Pi starvation by alteration in root architecture, expression of Pi transporter genes, inducing accommodation and secretion of acid phosphatases (APase), and aggregation of anthocyanin in the leaves (Wang et al. 2011). Up-regulation of APases is a general response to Pi deficiency in higher plants (Kumar et al. 2022). Purple APases (PAPs) are the largest group of plant APases, which have been classified into high and low molecular weight (HMW & LMW) phosphatases (Schenk et al. 2000; Flanagan et al. 2006). The smaller LMW PAPs form monomeric ~ 35 kDa isozymes whereas the larger HMW via ∼ 55–60 kDa forms a homodimeric enzyme. Cysteine residue is required to link subunits of HMW PAPs by a disulfide bridge (Schenk et al. 2013).
All PAPs share heterovalent bimetal‒ligating amino acids with five blocks of conserved amino acid (aa) sequences and seven invariable residues (DXG/GDXXY/GNH(D/E)/VXXH/GHXH; bold letters represent metal-ligating amino acids) at the carboxyl end of PAP members (Schenk et al. 2000; Li et al. 2002; Gahan et al. 2009). They are distinguished from other APases by their special characteristic purple color due to a charge transfer to Fe(III) by tyrosine (Lu et al. 2008; Schenk et al. 2013). The heterovalent bimetallic center is necessary for PAPs to promote hydrolysis activity. One component of this bimetallic center is Fe(III) and the other is Fe(II), Zn(II), or Mn(II). Therefore, these two components usually interact with PAPs in one of the forms of Fe(III)–Fe(II), Fe(III)–Zn(II), or Fe(III)–Mn(II) (Schenk et al. 2000, 2013). Oxidation states are crucial for the function of these isozymes. Accordingly, the coordination of bimetallic ions and seven invariable residues creates highly conserved active sites in all PAPs. Consequently, this feature of the active site makes isozymes catalyze the hydrolysis of a broad range of phosphorylated substrates (Schenk et al. 2013). Despite that, the order and types of amino acids (e.g. polarity, acidic or basic) around the active site play a decisive role in the substrate preference of these isozymes (Feder et al. 2020). In (NtPAP) a PAP from tobacco, an elongated, narrow groove has been located above the catalytic metal center. This groove appears to be suitable for too-bulky substrates such as phytate. Furthermore, the REKA motif is identified close to the active site of NtPAP and probably facilitates strong interactions with the other two phosphate groups from phytate. Accordingly, Glycine max phytase (GmPHY), Oryza sitava phytase (OsPHY1), NtPAP and Arabidopsis thaliana PAP (AtPAP15) act as a phytase (Feder et al. 2020). In AtPAP26, the small circular tunnel is formed above the bimetallic center. This tunnel is probably suitable for smaller substrates such as 2-phosphoenolpyruvate (PEP). Thus, AtPAP26 appears to be a PEPase (Feder et al. 2020). Also, substrate preferences in PAPs are closely related to the structural aspects of isozymes. Nevertheless, the substrate specificity of many PAPs remains unclear. Compatible structural features, the APase activity along with Kinetic parameters assays have demonstrated a large number of plant PAPs which tend to act on a wide range of substrates (Tran et al. 2010a). However, besides the non-specific APase activities of many PAPs, substrate preferences in some of these isozymes, e.g. AtPAP12, AtPAP17 and AtPAP26 prefer PEP (Veljanovski et al. 2006; TRAN et al. 2010b; O’Gallagher et al. 2022), AtPAP10 and red kidney bean (rkbPAP) are an efficient ATPase (Wang et al. 2011; Feder et al. 2020), also, NtPAP, AtPAP15, AtPAP23 are phytate-preferential PAPs (Kaida et al. 2008; Kuang et al. 2009; Wang et al. 2009; Srivastava et al. 2020).
After the genome project was over in organisms, finding true messages and the establishment of PAPs networks can be a novel and exciting field in cell biology. For this purpose, focuses have shifted from structural genomics to functional genomics. Therefore, characterization of the function of Pi-starvation-inducible (PSI) genes and investigation of the appropriate responses of Pi-deficient plants is inevitable. Nowadays, the functions of a few AtPAPs have been almost identified or characterized. AtPAP2 is involved in modulating carbon metabolism (Sun et al. 2012, 2018), and AtPAP5 has a role in the upstream of salicylic acid accumulation. So, being involved with regulating the expression of defense-responsive genes (Ravichandran et al. 2013), AtPAP9 transcript variants highly express in stipule and vascular tissue, particularly in response to fungal infection (Zamani et al. 2014), AtPAP10 is mainly a secreted APase with a key role in plant tolerance to Pi deficiency (Wang et al. 2011, 2014), and AtPAP12 is a secreted PAP with an additional function in Pi recycling from leaked phosphomonoesters (Robinson et al. 2012b), AtPAP15 with phytase activity participate in Ascorbate biosynthesis (Zhang et al. 2008; Wang et al. 2009). Furthermore, AtPAP17 with specific expression profiles, dual-localization to cell wall and cell lytic vacuole, broad substrate catalysis, Pi feedback inhibition, and rapid repression by Pi resupply to − P cells hypothesize to contribute to Pi metabolism during Pi starvation and leaf senescence (O’Gallagher et al. 2022). AtPAP18 is remarkably higher n plant total P content and biomass production (Zamani et al. 2012, 2018), and AtPAP23 is involved in Fe and Mn homeostasis in flower development and metabolism (Zhu et al. 2005). In addition, AtPAP25 has the signaling role during Pi deprivation (Del Vecchio et al. 2014) and finally, AtPAP26 by scavenging Pi from organic phosphate sources and remobilizing phosphate during leaf senescence, is also a secreted PAP (Veljanovski et al. 2006; Hurley et al. 2010; TRAN et al. 2010b; Robinson et al. 2012a; Sabet et al. 2018; Ghahremani et al. 2019).
Insertional mutagenesis is a central method to tag known insertional fragments and construct mutants (Parinov and Sundaresan 2000). These constructed mutant plants have become a huge resource for gene functional analysis in Arabidopsis thaliana (Krysan et al. 1999). Introducing T-DNA fragments into a genome can lead to the loss or the gain of function for an interesting gene (Radhamony et al. 2005). These procedures lead to a wide variety of polymorphism collections in an interesting single gene. Thus, a suitable strategy is inevitable for the selection of desired mutant plants from these collections that almost assure us about the malfunction of the final gene product (O’Malley et al. 2015).
In the bioinformatics era, databases have greatly helped to solve biological structures like DNA, RNA, and proteins. However, for those structures that have not been solved by the database, the alternative approach for the prediction of structures could be homology modeling based on the types of alignment algorithms.
AtPAP17 is among the first LMW AtPAP isozyme to be studied. Among 29 AtPAPs, AtPAP17 is an important LMW PAP that presents actively in Pi deficiency conditions, oxidative stress, and senescing leaves (Del Pozo et al. 1999). This isozyme, which was purified from the soluble intracellular fraction of hydroponic cultures of Pi-deficient Arabidopsis seedlings, exhibits marked induction of AtPAP17 mRNA in shoots and roots during Pi deprivation (Del Pozo et al. 1999). Wang et al. (2014) have demonstrated the effect of mutation of each AtPAP gene on total intracellular APase activity. Their results indicated that the absence of any of the AtPAP12, AtPAP15, and AtPAP26 alone significantly reduced intracellular APase activity. Furthermore, they showed that AtPAP17 knockout did not significantly change this component either in root or shoot. O’Gallagher et al. (2022) purified AtPAP17 from the cell wall of Pi-deficient Arabidopsis cell culture and analyzed its physical, kinetic, and molecular characterization. Their results suggested that AtPAP17 up-regulation occurs in the extracellular matrix and cell vacuole during Pi-deficient, salt-stressed conditions or senescing leaves. Nevertheless, no significant biochemical or phenotypical changes were observed in an atpap17-mutant during mentioned conditions. Furthermore, they concluded that the low peroxidase activity of AtPAP17 probably does not contribute to the detoxification of reactive oxygen species. Unlike that, in another report significant biochemical and phenotypical changes were observed in an atpap17-mutant compared to wild-type (WT) plants under Pi-deficient treatments (Farhadi et al. 2020). According to their report, the root length was longer in the atpap17-mutant. Moreover, they observed higher Apase activity and less seedling fresh weight in atpap17-mutant, as well. They suggested that the loss of the expression of AtPAP17 or AtPAP26 genes was buffered by the up-regulation of the other isozyme. Thus, there is a compensation relationship between AtPAP17 and AtPAP26 genes in Pi homeostasis. A follow-up study suggested the role of both AtPAP17 and AtPAP26 genes in salt tolerance (Abbasi-Vineh et al. 2021). They found that these two phosphatases directly and/or indirectly respond to salt stress by APase activity and/or non-APase activity. Accordingly, atpap17-mutant and AtPAP17-overexpressed plants had the highest and lowest APase activity in no-salinity condition.
The present study aimed to assess AtPAP17 gene function in response to phosphate conditions. To achieve this, A. thaliana was used as a plant model system. An expression binary vector with a native AtPAP17 gene from Arabidopsis under the control of the CaMV-35S promoter was constructed and transformed to wild-type (WT) plants (named OE, for overexpressed) resulting in constitutive expression of this gene. The independent homozygote of T3 generation of OE plants and a homozygote atpap17-mutant plant (Mu) were used to characterize the physiological role of AtPAP17 gene in comparison with WT plants. For investigating AtPAP17 function, we measured APase activity, organic and inorganic content of phosphate, and also morphological effects in three distinct plants under Pi sufficient (+ P) and Pi deficient (− P) conditions.
Materials and methods
Plant materials, seeds culture, and treatments conditions
Arabidopsis thaliana ecotype Columbia (Col-0) was used as a WT plant to compare with atpap17-mutant (Mu) and AtPAP17-overexpressed (OE) homozygote plants. The atpap17-mutant plants have been selected from “The Arabidopsis Information Resource” (TAIR) database on Arabidopsis.org server (Rhee et al. 2003). The seeds of all genotypes were obtained from the National Institute of Genetic Engineering and Biotechnology (NIGEB). For gene transferring, WT plant seeds have grown in soil with 1:1:1 ratio of peat moss:perlite:vermiculite and were fertilized biweekly by Hoagland media (Hoagland and Arnon 1950).
For phosphate treatments, the Arabidopsis seeds of all studied genotypes were soaked in 70% (v/v) ethanol–water, rinsed in distilled water, and treated with 10% sodium hypochlorite. Disinfected seeds were rinsed with sterilized water five times and placed on solid Murashige and Skoog (MS) culture medium (Murashige and Skoog 1962) containing 1.2 mM KH2PO4 supplemented with 1% sucrose and 0.8% plant-agar. Seeds were incubated in dark at 4 °C for 2 days for uniform germination and then transferred to the germinator for 7 days.
The growth conditions were 16-h days and 8-h nights at 25 °C, under 100–150 µmol m−2 s−1 photosynthetically active radiation. The 7-day-old germinated seedlings were aseptically transferred into MS medium with 0 (− P) or 1.2 mM (+ P) KH2PO4 as the sole phosphorus source and grew under 16-h day and 8-h dark at 25 °C for 14 next days. The fresh weight (FW) and dry weight (DW) measurement, APase activity, Pi, and total P content were evaluated with 21-day-old seedlings in − P and + P conditions. Shoots and roots of + P and − P plants were separately harvested and frozen in liquid N2 and stored at − 70 °C until required.
Screening and generation of the homozygote atpap17-mutants
Seeds of atpap17-mutant plant, SALK_097940.47.75.x propagated on the soil with 1:1:1 ratio of peat moss: perlite: vermiculite and grew at a period of 16-h days and 8-h nights at 25 °C. The genomic DNA was isolated according to quick (Young et al. 2001) and CTAB methods (Rogers and Bendich 1985). The homozygous progenies of mutant plants were screened for insertion of T-DNA fragments into the AtPAP17 region using PCR and AtPAP17 gene-specific primers; PAP17mF: 5′-CCACGGCGAGTCTGAGTTTGCTGTTGTGTA-3′ and PAP17mR: 5′-GGAGGCAGTTACTTTCGGGTCGTTCCTTTG-3′ and, a primer anchored in the T-DNA left border; LBb1: 5′-GCGTGGACCGCTTGCTGCAACT-3′.
Isolation and cloning of the AtPAP17 gene
The full-length coding sequence of AtPAP17 (CDS) (Genbank locus No. AT3G17790) was amplified by Expand High Fidelity DNA polymerase and PCR reaction using the cDNA pool of Arabidopsis and gene-specific primers PAP17F: 5′-TACGTTTCCTCGAGCCTACCTAATTCATC-3′ and PAP17R: 5′-TATTAAGGCTTTTTACTCGAGAACAGAGG-3′. The PCR product was purified and integrated into pTZ57R/T (Fermentas, Canada) as a cloning vector. The Escherichia coli competent cells strain DH5α was transformed with pTZ57R/T-AtPAP17 vector and grew on LB selective culture medium (Sezonov et al. 2007) containing ampicillin antibiotic (100 mg L−1). The positive colonies were confirmed by colony PCR and verified by sequencing with M13 universal primers, and enzymatic digestion.
The CDS sequence of the AtPAP17 gene was isolated by digestion with BamHI and XbaI restriction enzymes and finally introduced downstream of the cauliflower mosaic virus CaMV-35S promoter into the corresponding position of pARM2 as a plant expression vector. The E. coli competent cells were transformed with pARM2/AtPAP17 vector. The transformed cells were screened on a solid selective LB culture medium and confirmed using colony PCR and enzymatic digestion. Plasmid extraction was done (Birnboim and Doly 1979) and introduced into Agrobacterium tumefaciens strain GV3101 by freeze and thaw method (Holsters et al. 1978). LB-agar selective medium containing kanamycin and rifampicin antibiotics (50 mg L−1 and 50 mg L−1) was used for screening of the transformed cells.
Plant transformation and detection of overexpressed plants
The flowers of A. thaliana plant (WT) were used for pARM2/AtPAP17 transformation using a vacuum infiltration method supplemented with 2% glucose (Bechtold and Pelletier 1998; Clough and Bent 1998; Zhang et al. 2006). The first transgenic progeny as hemizygous plants (T1 seeds) were screened on MS medium containing 50 mg L−1 kanamycin. The transgenic plants were confirmed by PCR using AtPAP17 gene-specific primers; UniqePAP17F: 5′-CGAGTCTGAGTTTGCTGTTGT-3′ and UniqePAP17R: 5′-ACATAAGAGTTGCGAGATGGAAC-3′. The individual transgenic lines were chosen to eventually produce homozygous third-generation (T3) by self-pollination.
RNA extraction and cDNA synthesis
Phosphate-deficient A. thaliana plants were used for RNA extraction and cDNA synthesis. Total RNA was extracted using a commercial RNA extraction Kit (Sinaclon, Tehran, I.R.) according to the manufacturer’s instructions. The RNA samples were further treated with the DNase I, recombinant, RNase-free (Roche, Germany) according to the supplier’s instruction to eliminate genomic DNA contamination. For cDNA synthesis, 20 µg of each RNA sample was added to a reaction consisting of 1 µL of 100 µM oligo-dT(12–20) primer, 1 µL of 10 mM dNTP mix, and nuclease-free H2O up to a volume of 10 µL. The reaction was heated at 70 °C for 20 min and cooled on ice. 10 µL of the second mixture was added to each above reaction which contained 2 µL 10× RT buffer, 0.1 M dithiothreitol, 50 mM MgCl2, 40 units of RNase inhibitor (Roche, Germany) and 50 units of Superscript II reverse transcriptase (Fermentas, Vilnius, Lithuania). Eventually, the screw cap microcentrifuge tubes were incubated at 42 °C for 90 min.
RT-PCR analysis
Transcriptional levels of AtPAP17 gene were evaluated with one pair of AtPAP17 specific primers Unique PAP17F and Unique PAP17R, on exon 1 and 3, respectively, and α-tubulin gene; Tub + 1: 5′-GCTTTCAACAACTTCTTCAG-3′ and Tub-1: 5′-CATCGTACCACCTTCAGACAC-3′ in semi-quantitative RT-PCR using cDNA as the template. The components of PCR solution were mixed according to the manufacturer’s instruction of a commercial PCR Kit (Sinaclon, I.R.) in a final volume of 20 µL and then placed on a thermocycler (Eppendorf Mastercycler-Gradient, Germany). The PCR reaction was performed by a program with 1 cycle of 4 min denaturation at 94 °C, 35 cycles of 1 min denaturation at 94 °C, 1 min annealing at 60 °C and 1 min extension at 72 °C and a final extension step at 72 °C for 10 min. The specificity of the primers was tested by performing PCR using cDNA. PCR products were analyzed on 1.5% agarose gels. The images of band intensities in ethidium bromide-stained gel were quantified by ImageJ 1.50i software (Schneider et al. 2012) and compared against the level of transcripts of α-tubulin transcripts as the internal control.
APase activity assay
The seedling of WT plants, homozygous atpap17-mutant, and AtPAP17-overexpressed (T3) plants were ground to a fine powder in liquid nitrogen separately and homogenized in isolation buffer (100 mM NaOAc, pH 5.6). The mixture was centrifuged twice at 18,000×g for 10 min at 4 °C and the separated supernatants were used for enzyme assays. All extraction steps were carried out on the ice (Naseri et al. 2004; Zamani et al. 2012). The APase activities were assayed at 37 °C for 30 min in the same buffer containing 5 mM of p-nitrophenyl phosphate (pNPP) (Heinonen and Lahti 1981). A standard curve was generated using linear regression based on different concentrations of KH2PO4 (0, 5, 10, 20, 40, 60, 80, and 100 mM). The protein concentration was measured according to Bradford’s (1976) method by using bovine serum albumin (BSA) as a reference protein.
Assay of Pi, organic, and total P
Pi and total P were evaluated by a modified Ames (1966) assay. For determination of Pi content, 100 mg fresh 21-day-old shoot tissue samples were ground in liquid nitrogen and centrifuged at 18,000×g for 10 min. 50 µL of the supernatant was diluted with 250 µL distilled water and assayed with 700 µL of reagent containing 10% ascorbic acid and 0.42% ammonium molybdate in 1 N H2SO4 in 1:6 ratio (Zamani et al. 2012). The mixture was incubated at 45 °C for 20 min. Soluble Pi absorbency was measured at 820 nm and its content was expressed as micromoles of soluble Pi per gram fresh weight (µmol g FW−1) using a standard calibration curve of KH2PO4.
For measurement of the total P amount, 50 mg of each fresh shoot tissue was ground in liquid nitrogen and mixed with 10% nitrate magnesium in 95% ethanol up to a volume of 1.5 mL. The material was dried and burned to ash by shaking the tube over a flame until the brown fumes disappear and then allowed the tube to cool down before adding 500 µL of concentrated HCLO4. The final volume was brought to 5 mL by water in the capped tube and incubated in a boiling water bath for 60 min to hydrolyze any pyrophosphate formed in the ash. 50 µL of the homogeneous was assayed as above description (Zamani et al. 2012). Finally, the result of the subtraction of total P content from Pi led to the organic P level which refers to unreachable phosphorus for cells as structural macromolecules containing P.
Measurement of dry and fresh weight
Roots and shoots of 21-day-old plants were separately harvested and assayed by five biological replicates (eight plants per replicate). The plants were dried at 70 °C for 72 h to note dry weights.
Bioinformatic analysis
All primers were designed using Oligo software version 7.56 available at oligo.net. The specificity of the primers was evaluated by Primer-BLAST software at NCBI. SnapGene software from GSL Biotech; available at snapgene.com was utilized for in silico cloning process. The software of Megalign, SeqMan, and Editseq from Lasergene package version 7.1.0 from DNASTAR. Madison, WI.; available at DNAstar.com was applied for sequence alignments.
Amino acid (aa) sequences were obtained from the UniProt database (Pundir et al. 2017) and the three-dimensional structure of solved proteins were acquired from the RCSB protein data bank database (Burley et al. 2019) available at rcsb.org (Berman et al. 2000). Homology modeling was done with different algorithms aimed to build three-dimensional protein structure models using as Swiss Model with different blast algorithms (Camacho et al. 2009; Remmert et al. 2012; Biasini et al. 2014), M4T with multiple templates and multiple mapping method algorithms (Fernandez-Fuentes et al. 2007), and ModBase with psi-blast, impala and based on satisfying spatial restraints (MODELLER) algorithm (Pieper et al. 2014).
The relationship between protein families was investigated through the Pfam database (Finn et al. 2016). Problematic or erroneous parts of predicted protein structure were surveyed according to protein structure analysis (ProSA) local model quality assessment (Wiederstein and Sippl 2007). Modeling according to Threading/Fold Recognition approaches were done through bioinformatics server as I-TASSER (Yang and Zhang 2015), IntFOLD (McGuffin et al. 2015), RaptorX (Källberg et al. 2012; Ma et al. 2012) and Phyre2 (Kelley et al. 2015). The models were analyzed by the sum of validation and quality servers as RAMPAGE with Ramachandran plot analysis (Lovell et al. 2003), QMEAN (Benkert et al. 2008; Studer et al. 2014) from ExPASy server, ERRAT (Colovos and Yeates 1993), Procheck G-factors (Laskowski et al. 1993, 1996) and VERIFY_3D (Bowie et al. 1991; Lüthy et al. 1992) through SAVES v 5.0 server. Loop refining was performed through GalaxyLoop from the GalaxyWEB server (Coutsias et al. 2004; Park et al. 2011, 2014; Park and Seok 2012; Ko et al. 2012). The structure refinement model was gained through the GalaxyRefine server (Heo et al. 2013; Lee et al. 2016). For energy minimization was used DeepView/Swiss-PDBViewer software V4.1.0 (Guex and Peitsch 1997). The resulting model was visualized by PyMOL software V 2.0.7 (Delano 2015).
Statistical analysis
The normality distribution hypothesis of residuals was surveyed for statistical analysis tests. Analysis of variance (ANOVA) was generalized to study the effect of multi-factors based on the completely randomized design (CRD) at least in three replicates. Duncan’s method was used to compare significant differences between means. The whole of statistical analyses such as analysis of variance, chi-square test, t-test, means comparison, and means standard errors were accomplished by using SPSS software V.19.
Results
Identification of suitable templates for AtPAP17 structure prediction
AtPAP17 gene with three exons and two introns is located on the third chromosome of ~ 135 Mbp A. thaliana genome in 6089381 to 6091231 loci. A protein with 338 amino acid sequences and 38.3 kDa molecular mass is encoded by this region. Although the three-dimensional structure of AtPAP17 protein has not been solved in the RCSB protein data bank, the putative structure model could be predicted using the experimentally determined structures of related family members as templates. For the structure prediction, the whole 338 AtPAP17 full-length aa was obtained from the UniProt database. The homology modeling was executed with different algorithms that aim to build three-dimensional protein structure models using as the Swiss Model, M4T, and ModBase databases.
Alignment results from the Swiss model server were matched with six solved templates from the PDB database with maximum sequences identity ~ 30% as follows: 1qhw (purple APase from rat bone), 2bq8 (crystal structure of human purple APase with an inhibitory conformation of the repression loop), 1ute (pig purple APase complexed with phosphate), 3tgh (GAP50 the anchor in the inner membrane complex of Plasmodium), 1war (recombinant human purple APase expressed in Pichia pastoris), and 1qfc (structure of rat purple APase). The QMEAN Z-score for all templates was calculated ǀZǀ > 2. Likewise, a signal peptide with 31 aa lengths elimination for reducing background interference has no significant shift in previous results. Thus, the 338 AtPAP17 full-length aa was introduced to the Pfam database and the metallo-phosphoesterase domain was extracted which contains 210 aa, the positions 47–256. The putative domain was subjected to modeling once again as target sequences. The best hits according to the Swiss model QMEAN Z-score (1 < ǀZǀ < 2) with sequences identity ~ 30% were matched with 1war, 2bq8, and 1ute, respectively (Table 1).
In the ModBase server, the overall estimated models based on 1qfc and 5uq6 (pig purple APase complexed with phosphate in two coordination modes along with a bridging hydroxide ion) templates were considered reliable for AtPAP17 full-length aa according to its evaluation. However, the sequence identity, another quality criterion, was calculated as unreliable with 25% identity for both templates. Afterward, the signal peptide was removed and the main chain was selected as the target sequence. Nevertheless, there are no significant changes in the results. Finally, the amino acid sequence of AtPAP17 metallo-phosphoesterase domain was uploaded in ModBase as input protein sequences. Two templates 2bq8 and 5uq6 with equal sequences identity 32% were considered reliable for all quality criteria parameters (Table 1).
The M4T comparative modeling portal used multiple templates 1qhw-1ute-1war, 1qhw-1ute-1war, and 1ute-3tgh-1qhw for targets sequences as follows AtPAP17 full-length aa, the main chain without a signal peptide and the metallo-phosphoesterase domain, respectively. The first two M4T templates were predicted equivalent. Furthermore, lower problematic or erroneous parts in the two mentioned templates were gained according to the ProSA local model quality assessment (Table 1).
Due to low similarity, the putative structure was predicted with Threading/Fold Recognition approaches through different bioinformatics server as I-TASSER, IntFOLD, RaptorX, and Phyre2.
According to RaptorX server interpretation, the best-matched templates for all three target sequences as AtPAP17 full-length aa, the main chain without signal peptide, and the metallo-phosphoesterase domain predicted 1ute alone. Furthermore, all three mentioned target sequences were assessed separately, and only 15%, 3%, and 0% positions were predicted as disordered, respectively (Table 1).
Based on I-TASSER results, the top three models were calculated based on 1war, 1ute, and 1qhw templates for all three target sequences. The 1war was located in rank 1 absolutely in all cases. The best estimated C-score, TM-score, and RMSD were gained for the main chain without signal peptide with 0.83 confidence score, 0.83 ± 0.08 TM-score, and 4.50 ± 3 Å RMSD followed by metallo-phosphoesterase and then AtPAP17 full-length aa as the targets sequences (Table 1).
In Phyre2, all three targets sequences were ranked with wider templates according to its raw alignment score algorithm, among all of them, the best templates predicted 3tgh with 100% confidence, 97% coverage (97% of sequences have been modeled with 100% confidence) and 8% of disordered for metallo-phosphoesterase target sequences compared with the two other one’s targets. However, when metallo-phosphoesterase domain was introduced as target sequences, after 3tgh the two templates 1ute and 1qhw similarly took the next ranks with 99.97% confidence as well as 98% coverage, respectively (Table 1).
Through the IntFOLD server based on its multiple-template modeling approach and applied quality estimates algorithm, for all three target sequences, the top five predicted models were gained only according to 1war as a template. The best model for the main chain without signal peptide as the target sequences was predicted based on the 1war template with p < 0.001 (p-value cut-off) and 0.89 Global model qualities (Table 1).
Prediction of AtPAP17 enzyme structure
To predict, according to the last model of the AtPAP17 enzyme to investigate the role of metal-ligating amino acids in forming active sites, the best model from each of the above-mentioned server was adopted under the criterion of appropriacy. The models were analyzed by the sum of validation and quality servers as RAMPAGE, QMEAN, ERRAT, and VERIFY_3D (Table 1). More confidence and quality were there in the model which was based on the 1ute template through the ExPASy server.
The other aspect of validation was considered based on the available experimental resolution in the PDB database. The resolutions of experimental x-ray diffraction for the seven templates from high to low resolution were solved as follows: 5uq6, 1ute, 3tgh, 1qhw, 2bq8, 1war, 1qfc with 1.18 Å, 1.55 Å, 1.70 Å, 2.20 Å, 2.20 Å, 2.22 Å, 2.70 Å, respectively. The 1ute protein was placed in the high-resolution range. The models created according to the mentioned template provide more confidence in the location of atoms in structures compared to the templates like 1war or 2bq8 with moderate resolution. Furthermore, only in high-resolution ranges can deviations be justified. After the selection steps, the ERRAT plot of the initial model was calculated and regions of the structure that can be rejected at the 99% confidence level were selected and subjected to loop refining. Then, by molecular dynamics simulation, the structure of the model was refined. Eventually, after energy minimization, the resulting model visualized (Fig. 1A, B). The differences between the first and last refined model as shown in Table 2. The small PAPs are closely related to mammalian homologs. Therefore, the last crystal structure of pig PAP (5uq6) with a high resolution, compared with the AtPAP17, predicted a model which is based on 1ute. The superposition of invariable metal-ligating residues between the model of AtPAP17 and 5uq6 indicated full conformity of each residue in the vicinity of the active site (Fig. 1C). The small circular tunnel on the active site of 5uq6 as well as the lack of REKA motif makes 5uq6 like AtPAP17 suitable candidates for smaller substrates (Fig. 1D).
The predicted AtPAP17 protein structure along with 5UQ6 PDB code (Pig purple APase). A The predicted secondary structure of AtPAP17, α-helix, β-sheet and loop are shown by cyan, red and magenta color, respectively. B The predicted 3D structure of AtPAP17. C Superposition of invariable metal-ligating residues between the model of AtPAP17 and 5UQ6 in vicinity of the active site, the residue numbers are according to the 5UQ6 sequence. D The surfaces of the active site of 5UQ6. Invariable metal-ligating amino acids in predicted purple APase 17 enzyme and 5UQ6 are marked by red color. FeM represents the heterovalent bimetallic center which M is Fe(II), Zn(II), or Mn(II)
Selection and verification of AtPAP17 homozygote mutant line
Among all atpap17-mutant polymorphisms recorded in “The Arabidopsis Information Resource (TAIR) database”, a small percentage of atpap17-mutant accessions showed at least a single T-DNA insertion in the coding sequence region. Among those, only a few had insertion either in the neighborhood or inside of the metallo-phosphoesterase region. In this study, the SALK_097940.47.75.x (TAIR Accession; Polymorphism: 1005456219) was selected to determine the function of PAP17 in A. thaliana, and the effect of APase activity on phosphate supply in Pi-sufficient and Pi-starved conditions. The T-DNA fragment has been inserted in only five amino acids far from metallo-phosphoesterase conserved GHDH metal ligating amino acids in this line. Finally, the selected line was preferred by the TAIR database and SALK collection.
The pairwise alignment and assembly of full-length AtPAP17 genomic DNA (GenBank accession: NM_112660) and DNA insertion flanking sequence of polymorphism (TAIR accession: 1005456219) based on the Smith–Waterman algorithm showed a ~ 4478 bp length T-DNA inserted in the third exon of AtPAP17 gene in the reverse direction (Fig. 2A). A multiplex PCR reaction using reverse/forward AtPAP17 gene-specific and T-DNA left border (LBb1) primers verified the position and orientation of inserted T-DNA at the locus of AtPAP17 gene (Fig. 2A). The homozygosity of the inserted T-DNA was confirmed in some atpap17-mutant individual plants by PCR using AtPAP17-specific oligonucleotide primers. Among all cultivated mutants, only the plants represented a single 1031 bp fragment were selected as a homozygous atpap17-mutant line. Other individual plants either showed a single 1500 bp electrophoresis DNA band similar to WT or corresponded to both 1031 and 1500 bp amplicons (Fig. 2B).
Representation of AtPAP17 gene in SALK_097940.47.75.x line. A Schematic diagram of AtPAP17 gene represents the exons (solid boxes), introns (lines) and a T-DNA insertion in exon 3 as reverse form (3′ → 5′). Exon/intron structures were determined by alignment of the AtPAP17 genomic and mRNA sequences. The PAP17m forward/reverse and LBb1 primers are shown by bold arrows. (The length and positions are not drawn to scale). The atg and tga illustrate the putative CDS start and stop codons, respectively. B The multiplex PCR reaction product using three primers PAP17mF, PAP17mR, and LBb1 and DNA samples from the wild-type and three different atpap17-mutant lines (progenies of SALK_097940.47.75.x selfing) on 1% agarose gel: WT represent A. thaliana Col-0 plants as control, lane 1: No T-DNA insertion line similar to the WT, 2: Homozygous atpap17-mutant line, and 3: Heterozygous atpap17-mutant line. M: DNA Ladder mix (Fermentas, Canada) as a molecular weight marker
Generation of transgenic overexpressed plants
The expression vector pARM2/AtPAP17 was transferred into A. thaliana WT genome via Agrobacterium-mediated delivery. The hemizygous transgenic (T1) genotypes were screened using selection on kanamycin-MS culture medium and verified by PCR reaction. For the recognition of the internal AtPAP17 gene from the transgene in the overexpressed plant genomes, Uniqe PAP17F&R primers were designed based on two separated exons (I & III) (Fig. 2A). Based on these primers, the AtPAP17-overexpressed plants showed an extra 544 bp PCR fragment that was shorter than the internal AtPAP17 gene with 737 bp lengths due to the lack of two internal introns. The confirmed transgenic T1 plants were allowed to self-fertilize and produce T2 seeds. Among nineteen distinct T1 transgenic plants producing T2 seeds (Table S1), twelve showed segregation ratios close to 3:1 for kanamycin-resistant: sensitive (p-value < 0.05), and these were selected as single-locus insertion events for further studies. The remaining seven lines (highlighted as dashed line rows in Table S1) represented a deviation from the 3:1 ratio. Among these, three genotypes (11, 15, and 19) matched with 15:1 Mendel’s ratio with the subsequent Chi-square tests. The rest genotypes deviated from Mendelian independent assortment proportions and showed linkage relationships, as cis configuration type (Table S1).
The selected T2 single-locus insertion seedlings were transferred to soil and produced T3 seeds. Subsequently, T3 full green seedling plates on kanamycin-MS culture medium were screened as a homozygote plant. To be more confident about single-locus insertion events, five independent transgenic plants were randomly selected and 100 seeds were collected from each of them. Seeds were mixed and used for further experiments as OE plants.
Gene expression analysis of knockout mutant and overexpressed plants
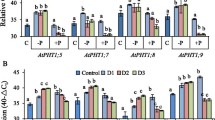
Semi-quantitative RT-PCR reaction was performed to evaluate the transcriptional level of the AtPAP17 gene in the studied plants. The RT-PCR reaction confirmed the lack of transcript either in the root or in the shoot of the atpap17-mutant plant using gene-specific primers (Fig. S1A). Also, the homozygous OE plants were examined for the expression level of the AtPAP17 gene.The AtPAP17 transcripts level was higher in OE versus the WTplants (Fig. S1B). The results indicated a different expression level of AtPAP17 gene in the root than in the shoot compare to the α-tubulin as the reference gene. Although the expression level of the AtPAP17 gene was higher in the root than shoot in the WT plants, the expression level was almost similar in the shoot and root of the OE plants (Fig. S1C).
AtPAP17 significantly contributes to cellular APase activity in both + P and − P conditions
In Pi-sufficient (+ P) condition, the atpap17-mutant plant displayed a 71% decrease in phosphatase activity compared to the WT, whereas an increase of 24% was detected in APase activity of OE plants (Fig. 3). In no phosphate condition (− P), the APase activity was dramatically increased in comparison to a phosphate-sufficient condition in all studied genotypes. The highest and the lowest increment were obtained for mutant and AtPAP17-overexpressed plants with 9.3-folds and 1.6-folds, respectively. Also, the mutant and OE plants represented 61% and 19% more APase activity compared to WT plants in the − P condition. It was also found that the effect of inducible genes was related to phosphate starvation (PSI) conditions at a 66% increase in WT APase activity from + P compared to − P condition. As a result, the AtPAP17 gene knockout showed different effects on APase activity in + P and − P conditions (Fig. 3).
APase activity assay by protein extracted from 21-days-old A. thaliana seedlings (14 days of growth in Pi-fed (+ P; 1.2 mM KH2PO4) and Pi-starved (− P; 0 mM KH2PO4) condition) in the presence of pNPP substrate. Wild-type plant, atpap17-mutant plant and the AtPAP17-overexpressed plants are shown by WT, Mu and OE, respectively. Each histogram is the mean of three assays (50 plants per replication). Vertical bars show standard errors (± SE). Mean values with the same letters have no significant difference as compared by Duncan's method at p < 0.05
Effect of AtPAP17 overexpression on cellular phosphate content
The measurement of organic and inorganic free phosphate proportion in the + P condition showed that the lowest amount of Pi belonging to the atpap17-mutant plant with a 38% decrease compared to the WT. However, the Pi content in OE plants was 111% more than in WT plants in the same condition (Fig. 4A).
Phosphorus content in WT, atpap17-mutant line (Mu) and AtPAP17-overexpressed (OE) line of A. thaliana. The seedlings were grown for 14 days on solid MS medium containing 0 mM Pi (− P) or 1.2 mM Pi (+ P). A Pi and B organic P contents. All histograms represent means ± SE of three replicates. Mean values with the same letter have no significant differences as compared by Duncan’s mean comparison method at p < 0.05
The phosphate starvation caused 25% and 50% decreases in total P and free Pi content of WT plants compared to the sufficient conditions, respectively. When plants have grown in Pi-sufficient conditions, total P increased gradually up to 7% in Mu plants with a huge increment of up to 45% in OE plants versus WT (Fig. 4). No significant difference was detected in the organic phosphate of WT and OE plants in + P condition, whereas a significant 38% increase was interestingly observed in the organic phosphate content of atpap17-mutant plants versus WT (Fig. 4B). Consequently, in the + P condition, OE plants showed more increment in Pi content than WT plants, and the Mu plants represented higher organic phosphate than the WT, respectively. Nevertheless, in the − P condition, the statistically insignificant content of organic P and free Pi were observed in OE plants than the WT, whereas the atpap17-mutant plant represented a 42% increase and 73% decrease, respectively (Fig. 4).
Transgenic plants acquired more water in sufficient and deficient phosphate conditions
To show the effect of AtPAP17 expression on Arabidopsis biomass production, we compared the fresh weight (FW) and dry weight (DW) of 21-day-old seedlings after 14 days of growth separately in + P and − P conditions (Fig. 5). Under Pi-sufficient conditions, the OE and Mu plants showed 29% more and 18% less in FW as compared to the WT plants, respectively (Fig. 5A). In the Pi-starved condition, the FW of the atpap17-mutant plant was significantly reduced versus the WT seedling, while no significant changes were detected in OE plants compared to the WT in the same conditions (Fig. 5B).
The growth rate of AtPAP17-overexpressed (OE) lines along with the atpap17-mutant (Mu) and WT of A. thaliana plants. FW and DW were measured by 21-days-old seedlings when grew on MS medium containing A 1.2 mM Pi (+ P) or B no Pi (− P). All values represent means ± SE of five replicates (eight seedlings per replication) in three independent experiments. Mean values with the same letters have no significant difference as compared by Duncan’s method at p < 0.05
Notably, the DW was statistically equal in all studied genotypes both in + P and in − P conditions. The comparison of FW and DW in the studied plants showed that the highest and the lowest amount of absorbed water belonged to OE plants (with 38 and 12 mg plant−1) and Mu plants (with 22 and 7 mg plant−1) in + P and − P conditions, respectively. Separated investigation of root and shoot fresh/dry weights of AtPAP17-overexpressed, atpap17-mutant, and WT seedlings revealed that the shoot FW of OE plants was significantly higher (36%) than in the WT plants at + P conditions, while no notable difference was observed between OE and WT root FW when grown at + P condition (Table 3). Similarly, a significant 31% increase was detected in the shoot FW of OE plants versus WT in − P condition. Inversely, the atpap17-mutant plant demonstrated no significant differences in shoot FW compared to WT plants, whereas a remarkable 93% and 73% reduction was observed in root FW both in + P and − P conditions, respectively (Table 3).
Despite no significant differences between the total dry weight of OE, Mu, and WT plants both in + P and − P conditions, the root DW of the atpap17-mutant plant was 27% and 67% lower than WT plants in + P and − P conditions, respectively. Yet, a significant increase in root biomass was observed in OE root DW under + P condition with a 19% increase and a 26% decrease in − P compare to WT root DW (Fig. 6).
Root growth (dry weight) of A. thaliana plants in 1.2 mM Pi (+ P) or no Pi (− P) conditions. The pooled root obtained from 21-days-old seedlings of AtPAP17-overexpressed (OE), atpap17-mutant (Mu) and WT plants. All values represent means ± SE of five replicates (eight seedlings per replicate) in three independent experiments. Mean values with the same letters have no significant difference as compared by Duncan’s method at p < 0.05
Discussion
Knowledge about gene function, expression pattern, and cellular enzymatic activities has been allowed the identification of novel candidate genes potentially involved in the specific mechanism and activity. So, the prediction of protein structures and their catalytic sites are one of these puzzle pieces. As an evolutionary point, the PAP enzymes belong to a metallo-phosphoesterase family and their members have consisted phospho-protein phosphatases, diadenosine tetraphosphatases, exonucleases, 5-nucleotidases and some of phosphor-monoesterases (Li et al. 2002). Although the protein structure of several metallo-phosphoesterase family members have been experimentally determined until now, the AtPAP17 structure has not been solved on the Protein Data Bank (PDB). Thus, homology modeling has been applied for the prediction of protein structure. To do this, the highly conservative domain was extracted and subjected to comparative modeling. The predicted protein 3D structure demonstrated that seven invariable metal-ligating amino acids have key roles in forming the active site. Therefore, the best mutant plant for functional assessments could be destructed genomes in these sites. So, the SALK_097940.47.75.x line was preferred as the T-DNA insertion in this line has located only five amino acids further than GHDH metal-ligating conserved sequences. Semi-quantitative RT-PCR revealed no AtPAP17 transcript in the mentioned line (Fig. S1A). This result ensured that the AtPAP17 transcript would not participate in the translational and post-translational events. Accordingly, the AtPAP17 function is disrupted.
In the A. thaliana genome, the AtPAP17 gene is regulated under a native internal inducible promoter, while in OE plants, the expression of AtPAP17 is also under the control of CaMV-35S promoter, in addition to the internal promoter. So, the WT plants were fortified with AtPAP17 cDNA under the control of CaMV-35S promoter causing a high level of gene expression. Later semi-quantitative RT-PCR results confirmed this postulation (Fig. S1). Hence, the fluctuation of APase activity and phosphate metabolism in three distinct genotypes as WT, verified atpap17-mutant, and created AtPAP17-overexpressed plants in both phosphates sufficient and deficient conditions could be evidence of AtPAP17 function in A. thaliana plant (Wang et al. 2014).
Roots development is a typical symptom of plants when grown in a circumstance where Pi availability is the only limitation to growth. Our results indicated that in Pi-sufficient conditions, the AtPAP17 gene knockout led to a significant decrease in root dry weight, while the transgenic plants with an excess of the AtPAP17 gene were able to develop their roots to get more phosphate from the medium culture (Fig. 6). In this condition, Pi was the highest and the lowest in OE and Mu plants, respectively (Fig. 4). These results are consistent with Abbasi-Vineh et al. (2021). WT plants with internal AtPAP17 could spread their roots and subsequently acquire more phosphate. However, a significant shift in root development was observed when WT plants grew in a Pi-starved condition (Fig. S2). The observed difference in 21-day-old roots may refer to the first 7 days where the whole samples have grown in sufficient phosphate. Since the transgenic plants have a more surplus copy of the AtPAP17 gene in addition to the internal one, this advantage empowers them to achieve more Pi than WT plants during the first 7 days (Fig. 4A). Therefore, the extension of the root was unnecessary for OE plants, while the WT plants expand their roots to attain more Pi at Pi-starved conditions. Also, the atpap17 knockout plants should expand their roots, whereas the AtPAP17 locus has been impeded by T-DNA insertion, and the lack of PAP17 enzyme reduced root dry biomass and consequently the Pi content. Thus, as a major result, AtPAP17 has an important role in root morphology and structure, but not shoot developmental and structural programming (Fig. S2).
Induction of APases genes in Pi limitation stress is a general response of higher plants to Pi scarcity (Mukatira et al. 2001; Wu et al. 2003; Lin et al. 2009). Since most known APases have an inducible promoter, there is expected to observe a significant shift in their expression pattern when faced with phosphate deficiency. According to that, transcriptional level mainly controlled AtPAP17 up-regulation during Pi deprivation. Thus, PHR1 as a transcription factor for the binding site in the AtPAP17 promoter has a key role in Pi-Starvation-Response (PSR) (Bustos et al. 2010). In such a situation, it seems to be impossible to relate an appropriate role for a specific phosphatase. Unlike that, Pi sufficient condition could be a better option when expressed APase genes profile associated with phosphate deficiency stress has less interference with the contribution of the AtPAP17 gene. Gallagher et al. found non-denaturing PAGE followed by in-gel APase activity staining of the soluble intracellular proteome from Arabidopsis seedling extracts. Also, the AtPAP17 isozyme was greatly downregulated in phosphate-sufficient conditions, whereas it is highly up-regulated when faced with phosphate deficient condition. Furthermore, Pi resupply with 2 mM NaPi for 48 h to − P cells or seedlings of Arabidopsis caused a significant downregulate of AtPAP17 isozyme. Interestingly, PAP17 was the most downregulated among all proteins detected in the + P cells, relative to − P controls (Mehta et al. 2021). Thus, it is difficult to understand how eliminating AtPAP17 in atpap17 knockout plants could have any impact on growth or APase activities in Pi-sufficient conditions. Nevertheless, the creation of overexpressing lines of 35S:AtPAP10, 35S:AtPAP12 and 35S:AtPAP26 indicated significant increases in their APase activities in all + P and – P conditions (Wang et al. 2014). The results indicated that in the Pi-starved condition, the mutant plants displayed a further 61% increment in APase activity than WT plants. This surplus alteration or adaptation may be acquired due to a difference in the expression level of the complementary genes to compensate for the lack of AtPAP17 expression level in atpap17-mutant plant. In seedlings of the atpap17-mutant a significant up-regulation is observed of HRP9, AtPAP8, and AtPAP26 genes (Farhadi et al. 2020). However, there is a contradiction with O’Gallagher et al. (2022). This discrepancy may be due to various terms of the treatments or experiments such as periods of growth length and/or Pi deficiency. So, 21-day-old seedlings in this work as well as 14-day-old seedlings in previous studies (Wang et al. 2014; O’Gallagher et al. 2022) were used for APase activity measurement. Indeed, in this study, an additional 7-day period of Pi deficient stress was considered for the seedlings as compared to previous studies (Wang et al. 2014; O’Gallagher et al. 2022). Therefore, according to (Farhadi et al. 2020) the longer period of Pi deficient stress likely induced more APase genes and finally may result in more APase activity in atpap17-mutant lines. The other outcome indicated that less increment in APase activity in OE plants with a 60% increase from + P to − P compared to Mu plants with a 9.3-folds increase from + P to − P arose from the extra copy number of AtPAP17 gene under the control of CaMV-35S promoter. Therefore, the presence of this gene may lead to fewer efficacies or less perception of Pi deficiency stress. In 14-day-old AtPAP10-overexpressed Arabidopsis seedlings grown on both + P and − P medium, root-associated APase activity was 150–200% greater than WT. In contrast, intracellular APase activity of AtPAP12 and AtPAP26-overexpressed plants was 150–300% greater than WT (Wang et al. 2014). Also, APase activity increased due to the heterologous expression of AtPAP26 as overexpressed in tobacco plants (Zamani et al. 2012, 2018; Sabet et al. 2018). Inversely, the lack of the AtPAP17 gene in atpap17-mutant plant resulted in the APase activity being greatly reduced when cultivated in sufficient phosphate conditions. These results are identically consistent with Farhadi et al. (2020). Also in another study, the two independent atpap15 mutants had 15–20% more phytate than WT plants. These results demonstrated a significant reduction in their APase activity because AtPAP15 acts as phytase (Zhang et al. 2008). Nevertheless, O’Gallagher et al. (2022) reported no obvious differences in APase activity for all conditions which may be due to various terms of the treatments or experiments as mentioned earlier. Comparing phosphatase activity between WT and atpap17-mutant plants in the phosphate-free medium may imply that the degradation of AtPAP17 gene encourages a portion of complementary Pi hemostasis genes to change their expression to tolerate the AtPAP17 gene deficiency (Farhadi et al. 2020). Hence, in the phosphate-free condition, a huge increase in phosphatase activity of the atpap17-mutant plant can prove that the AtPAP17 enzyme has a key role in phosphate homeostasis in A. thaliana plants. Several studies have shown that in the absence of phosphate, the AtPAP17 gene expression level significantly increased in WT Arabidopsis plants, so AtPAP17 is a highly up-regulated purple APases in the phosphate-deficient condition (Del Pozo et al. 1999; Veljanovski et al. 2006; Hurley et al. 2010). Wang et al. (2009) demonstrated that overexpressed AtPAP15 plants containing CaMV-35S promoter exhibited significantly improved P efficiency and enriched plant P content as compared to WT when planted on phytate as the lone P source. Also, the only reason for reducing the amount of Pi in the atpap17-mutant plant arose from the lack of AtPAP17 activity due to absorbing more Pi in both conditions. Hurley et al. (2010) has revealed that the atpap26-mutant plant shows a 35% reduction in Pi concentration. On the other hand, the AtPAP17-overexpressed plants were the most Pi absorption compared to WT in Pi-sufficient conditions. Non-denaturing PAGE followed by in-gel APase activity staining clearly showed that the AtPAP17 isozyme existed in both intracellular proteome and secretome (O’Gallagher et al. 2022). Although AtPAP17 is up-regulated in Pi-deficient conditions, the kinetic studies revealed that AtPAP17 catalytic efficiency is much less than AtPAP12 and AtPAP26. Therefore, AtPAP17 cannot be the dominant Purple Apase in Arabidopsis (TRAN et al. 2010b; O’Gallagher et al. 2022). Nevertheless, AtPAP17 displayed maximal APase activity between pH 5.5 and 7.5. This broad pH profile which is much wider than the majority of characterized AtPAPs with optimal activity between pH 5.5 and 6.0 plays a specialized role in P metabolism during salinity or drought stress or leaf senescence. During these stresses the extracellular matrix and/or cytoplasm pH shifts to alkalinization. In these situations, AtPAP17 could facilitate the remobilization and recycling of Pi from intracellular P monoesters and anhydrides of older tissues and/or scavenge Pi from external organophosphate compounds (Wang et al. 2014; O’Gallagher et al. 2022). Many studies show that Pi uptake from the environment is regulated by changes in water absorption in plants (Greenway and Klepper 1968). Thus, the increased fresh weight of OE plants compared with WT cause them to absorb more water to absorb more phosphate. Therefore, greater fresh weight in OE plants is related to more water absorption, but, the dry weight of these OE plants as compared with WT has not revealed any remarkable difference in similar conditions. Semi-quantitative RT-PCR revealed that the AtPAP17 gene expression in the root tissue was more than in the shoot tissue in WT. So, AtPAP17 gene knockout may leave more extensive damage to the root tissue (Fig. S2). Also, the results indicated a significant reduction in the fresh and dry weight of atpap17-mutant roots compared with the roots of WT both in + P and − P conditions. After the perception of stress, the phosphate-free plants alter their root structure (Schachtman et al. 1998; Plaxton 2004; Czarnecki et al. 2013). In this vein, earlier studies (Farhadi et al. 2020) have also reported that the root length of atpap17-mutant was less and more than WT in + P and − P conditions, respectively whereas OE plants represented more root length in both + P and − P conditions. Furthermore, lateral root numbers in all three WT, atpap17-mutant, and OE plants indicated no differences in − P while atpap17-mutant and OE plants have more and less lateral root numbers in + P condition. In the − P condition the root-to-shoot ratio was observed the most and the least in WT and atpap17-mutant plant respectively for both fresh and dry weight. Our results clearly show that the WT plants perceived phosphate deficiency stress and expanded their roots to maintain nutrient uptake from the environment when grown in phosphate-free conditions. As a result, the atpap17-mutant plant despite sensing phosphate deficiency stresses cannot develop roots, due to the lack of the AtPAP17 gene, and eventually, it led to the least root biomass production (Fig. S2).
Conclusion
This study integrates structural modeling and genetic tools to assess the function of the purple acid phosphatase AtPAP17 isozyme in Arabidopsis Pi metabolism. It would be of significant interest to those concerned with the role of PAPs in plant Pi acquisition and use efficiency. For this, homology modeling was applied for the prediction of the AtPAP17 isozyme structure. For more confidence, the best mutant plant for functional assessments was selected based on T-DNA insertion around invariable metal-ligating amino acids which have key roles in forming the isozyme active site. In the following, OE plants generated and accompanied by WT were subjected to biochemical studies.
It was found in this study that the AtPAP17 isozyme plays a direct role in Apase activity and interacts with other phosphatases to maintain phosphorus homeostasis, and it has a special role in root growth and development. Wang et al. (2014) suggested that APases may have other roles besides improving Pi recycling or Pi scavenging for plants so that the lack of AtPAP17 isozyme leads to a significant change in root biomass. Therefore, the study of interactions between PAP genes provides a better understanding of cellular P homeostasis regulatory processes. Finally, these processes may lead to facilitating the bioengineering of P-efficient crop varieties, urgently needed to reduce or eliminate the use of inefficient, non-renewable, and polluting Pi-containing fertilizers in agriculture.
Data availability
All data generated or analyzed during this study are included in the article/supplementary material, further inquiries can be directed to the corresponding author/s.
References
Abbasi-Vineh MA, Sabet MS, Karimzadeh G (2021) Identification and functional analysis of two purple acid phosphatases AtPAP17 and AtPAP26 involved in salt tolerance in Arabidopsis thaliana plant. Front Plant Sci 11:618716. https://doi.org/10.3389/fpls.2020.618716
Ames BN (1966) Assay of inorganic phosphate, total phosphate and phosphatases. In: Neufeld EF, Ginsburg V (eds) Methods in enzymology, vol 8. Academic Press, New York, pp 115–118. https://doi.org/10.1016/0076-6879(66)08014-5
Bechtold N, Pelletier G (1998) In planta Agrobacterium mediated transformation of adult Arabidopsis thaliana plants by vacuum Infiltration. In: Martinez-Zapater JM, Salinas J (eds) Arabidopsis protocols. Humana Press, Totowa, pp 259–266
Benkert P, Tosatto SCE, Schombu D (2008) QMEAN: a comprehensive scoring function for model quality assessment. Proteins 71:261–277. https://doi.org/10.1002/prot.21715
Berman HM, Westbrook J, Feng Z et al (2000) The protein data bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Biasini M, Bienert S, Waterhouse A et al (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42:W252–W258. https://doi.org/10.1093/nar/gku340
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1523. https://doi.org/10.1093/nar/7.6.1513
Bowie JU, Lüthy R, Eisenberg D (1991) A method to identify protein sequences that fold into a known three-dimensional structure. Science 253:164–170. https://doi.org/10.1126/science.1853201
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Burley SK, Berman HM, Bhikadiya C et al (2019) RCSB protein data bank: biological macromolecular structures enabling research and education in fundamental biology, biomedicine, biotechnology and energy. Nucleic Acids Res 47:D464–D474. https://doi.org/10.1093/nar/gky1004
Bustos R, Castrillo G, Linhares F et al (2010) A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet 6:e1001102. https://doi.org/10.1371/journal.pgen.1001102
Camacho C, Coulouris G, Avagyan V et al (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. https://doi.org/10.1186/1471-2105-10-421
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2:1511–1519. https://doi.org/10.1002/pro.5560020916
Coutsias EA, Seok C, Jacobson MP, Dill KA (2004) A kinematic view of loop closure. J Comput Chem 25:510–528. https://doi.org/10.1002/jcc.10416
Czarnecki O, Yang J, Weston DJ et al (2013) A dual role of strigolactones in phosphate acquisition and utilization in plants. Int J Mol Sci 14:7681–7701. https://doi.org/10.3390/ijms14047681
Del Pozo JC, Allona I, Rubio V et al (1999) A type 5 acid phosphatase gene from Arabidopsis thaliana is induced by phosphate starvation and by some other types of phosphate mobilizing/oxidative stress conditions. Plant J 19:579–589. https://doi.org/10.1046/j.1365-313X.1999.00562.x
Del Vecchio HA, Ying S, Park J et al (2014) The cell wall-targeted purple acid phosphatase AtPAP25 is critical for acclimation of Arabidopsis thaliana to nutritional phosphorus deprivation. Plant J 80:569–581. https://doi.org/10.1111/tpj.12663
Delano W (2015) The PyMOL molecular graphics system. Schrodinger, V 2.0.7. http://www.pymol.sourceforge.net
Farhadi S, Sabet MS, Malboobi MA, Moieni A (2020) The critical role of AtPAP17 and AtPAP26 genes in Arabidopsis phosphate compensation network. Front Plant Sci 11:565865. https://doi.org/10.3389/fpls.2020.565865
Feder D, McGeary RP, Mitić N et al (2020) Structural elements that modulate the substrate specificity of plant purple acid phosphatases: avenues for improved phosphorus acquisition in crops. Plant Sci 294:110445. https://doi.org/10.1016/j.plantsci.2020.110445
Fernandez-Fuentes N, Madrid-Aliste CJ, Rai BK et al (2007) M4T: a comparative protein structure modeling server. Nucleic Acids Res 35:W363–W368. https://doi.org/10.1093/nar/gkm341
Finn RD, Coggill P, Eberhardt RY et al (2016) The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res 44:D279–D285. https://doi.org/10.1093/nar/gkv1344
Flanagan JU, Cassady AI, Schenk G et al (2006) Identification and molecular modeling of a novel, plant-like, human purple acid phosphatase. Gene 377:12–20. https://doi.org/10.1016/j.gene.2006.02.031
Gahan LR, Smith SJ, Neves A, Schenk G (2009) Phosphate ester hydrolysis: metal complexes as purple acid phosphatase and phosphotriesterase analogues. Eur J Inorg Chem 2009:2745–2758. https://doi.org/10.1002/ejic.200900231
Ghahremani M, Tran H, Biglou SG et al (2019) A glycoform of the secreted purple acid phosphatase AtPAP26 co-purifies with a mannose-binding lectin (AtGAL1) up-regulated by phosphate-starved Arabidopsis. Plant Cell Environ 42:1139–1157. https://doi.org/10.1111/pce.13432
Greenway H, Klepper B (1968) Phosphorus transport to the xylem and its regulation by water flow. Planta 83:119–136. https://doi.org/10.1007/BF00385017
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-Pdb Viewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723. https://doi.org/10.1002/elps.1150181505
Heinonen JK, Lahti RJ (1981) A new and convenient colorimetric determination of inorganic orthophosphate and its application to the assay of inorganic pyrophosphatase. Anal Biochem 113:313–317. https://doi.org/10.1016/0003-2697(81)90082-8
Heo L, Park H, Seok C (2013) GalaxyRefine: protein structure refinement driven by side-chain repacking. Nucleic Acids Res 41:W384–W388. https://doi.org/10.1093/nar/gkt458
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil, 2nd edn. California Agricultural Experiment Station, Berkeley, California, pp 1–32
Holsters M, de Waele D, Depicker A et al (1978) Transfection and transformation of Agrobacterium tumefaciens. Mol Gen Genet 163:181–187. https://doi.org/10.1007/BF00267408
Hurley BA, Tran HT, Marty NJ et al (2010) The dual-targeted purple acid phosphatase isozyme AtPAP26 is essential for efficient acclimation of Arabidopsis to nutritional phosphate deprivation. Plant Physiol 153:1112–1122. https://doi.org/10.1104/pp.110.153270
Kaida R, Hayashi T, Kaneko TS (2008) Purple acid phosphatase in the walls of tobacco cells. Phytochemistry 69:2546–2551. https://doi.org/10.1016/j.phytochem.2008.07.008
Källberg M, Wang H, Wang S et al (2012) Template-based protein structure modeling using the RaptorX web server. Nat Protoc 7:1511–1522. https://doi.org/10.1038/nprot.2012.085
Kelley LA, Mezulis S, Yates CM et al (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845–858. https://doi.org/10.1038/nprot.2015.053
Ko J, Park H, Heo L, Seok C (2012) GalaxyWEB server for protein structure prediction and refinement. Nucleic Acids Res 40:W294–W297. https://doi.org/10.1093/nar/gks493
Krysan PJ, Young JC, Sussman MR (1999) T-DNA as an insertional mutagen in Arabidopsis. Plant Cell 11:2283–2290. https://doi.org/10.1105/tpc.11.12.2283
Kuang R, Chan KH, Yeung E, Lim BL (2009) Molecular and biochemical characterization of AtPAP15, a purple acid phosphatase with phytase activity, in Arabidopsis. Plant Physiol 151:199–209. https://doi.org/10.1104/pp.109.143180
Kumar K, Yadava P, Gupta M et al (2022) Narrowing down molecular targets for improving phosphorus-use efficiency in maize (Zea mays L.). Mol Biol Rep 49:12091–12107. https://doi.org/10.1007/s11033-022-07679-5
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291. https://doi.org/10.1107/S0021889892009944
Laskowski RA, Rullmann JAC, MacArthur MW et al (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8:477–486. https://doi.org/10.1007/BF00228148
Lee GR, Heo L, Seok C (2016) Effective protein model structure refinement by loop modeling and overall relaxation. Proteins 84:293–301. https://doi.org/10.1002/prot.24858
Li D, Zhu H, Liu K et al (2002) Purple acid phosphatases of Arabidopsis thaliana. Comparative analysis and differential regulation by phosphate deprivation. J Biol Chem 277:27772–27781. https://doi.org/10.1074/jbc.M204183200
Lin W-Y, Lin S-I, Chiou T-J (2009) Molecular regulators of phosphate homeostasis in plants. J Exp Bot 60:1427–1438. https://doi.org/10.1093/jxb/ern303
Lovell SC, Davis IW, Arendall WB III et al (2003) Structure validation by Cα geometry: ϕ, ψ and Cβ deviation. Proteins 50:437–450. https://doi.org/10.1002/prot.10286
Lu K, Li J, Zhong W et al (2008) Isolation, characterization and phosphate-starvation inducible expression of potential Brassica napus purple acid phosphatase 17 (BnPAP17) gene family. Bot Stud 17:199–213
Lüthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three-dimensional profiles. Nature 356:83–85. https://doi.org/10.1038/356083a0
Ma J, Peng J, Wang S, Xu J (2012) A conditional neural fields model for protein threading. Bioinformatics 28:i59–i66. https://doi.org/10.1093/bioinformatics/bts213
McGuffin LJ, Atkins JD, Salehe BR et al (2015) IntFOLD: an integrated server for modelling protein structures and functions from amino acid sequences. Nucleic Acids Res 43:W169–W173. https://doi.org/10.1093/nar/gkv236
Mehta D, Ghahremani M, Pérez-Fernández M et al (2021) Phosphate and phosphite have a differential impact on the proteome and phosphoproteome of Arabidopsis suspension cell cultures. Plant J 105:924–941. https://doi.org/10.1111/tpj.15078
Mukatira UT, Liu C, Varadarajan DK, Raghothama KG (2001) Negative regulation of phosphate starvation-induced genes. Plant Physiol 127:1854–1862. https://doi.org/10.1104/pp.010876
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Naseri JI, Truong NT, Hörentrup J et al (2004) Porcine purple acid phosphatase: heterologous expression, characterization, and proteolytic analysis. Arch Biochem Biophys 432:25–36. https://doi.org/10.1016/j.abb.2004.08.008
O’Gallagher B, Ghahremani M, Stigter K et al (2022) Arabidopsis PAP17 is a dual-localized purple acid phosphatase up-regulated during phosphate deprivation, senescence, and oxidative stress. J Exp Bot 73:382–399. https://doi.org/10.1093/jxb/erab409
O’Malley RC, Barragan CC, Ecker JR (2015) A user’s guide to the Arabidopsis T-DNA insertion mutant collections. In: Alonso JM, Stepanova AN (eds) Plant functional genomics: methods and protocols. Springer, New York, pp 323–342
Parinov S, Sundaresan V (2000) Functional genomics in Arabidopsis: large-scale insertional mutagenesis complements the genome sequencing project. Curr Opin Biotechnol 11:157–161. https://doi.org/10.1016/S0958-1669(00)00075-6
Park H, Seok C (2012) Refinement of unreliable local regions in template-based protein models. Proteins 80:1974–1986. https://doi.org/10.1002/prot.24086
Park H, Ko J, Joo K et al (2011) Refinement of protein termini in template-based modeling using conformational space annealing. Proteins 79:2725–2734. https://doi.org/10.1002/prot.23101
Park H, Lee GR, Heo L, Seok C (2014) Protein loop modeling using a new hybrid energy function and its application to modeling in inaccurate structural environments. PLoS ONE 9:e113811. https://doi.org/10.1371/journal.pone.0113811
Pieper U, Webb BM, Dong GQ et al (2014) ModBase, a database of annotated comparative protein structure models and associated resources. Nucleic Acids Res 42:D336–D346. https://doi.org/10.1093/nar/gkt1144
Plaxton WC (2004) Plant response to stress: biochemical adaptations to phosphate deficiency. In: Goodman RM (ed) Encyclopedia of plant and crop science. Marcel Dekker, New York, pp 976–980. https://doi.org/10.1081/E-EPCS-120010648
Pundir S, Martin MJ, O’Donovan C (2017) UniProt protein knowledgebase. In: Wu CH, Arighi CN, Ross KE (eds) Protein bioinformatics: from protein modifications and networks to proteomics. Springer, New York, pp 41–55
Radhamony RN, Mohan Prasad A, Srinivasan R (2005) T-DNA insertional mutagenesis in Arabidopsis: a tool for functional genomics. Electron J Biotechnol 8:82–106
Ravichandran S, Stone SL, Benkel B, Prithiviraj B (2013) Purple acid phosphatase 5 is required for maintaining basal resistance against Pseudomonas syringae in Arabidopsis. BMC Plant Biol 13:107. https://doi.org/10.1186/1471-2229-13-107
Remmert M, Biegert A, Hauser A, Söding J (2012) HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat Methods 9:173–175. https://doi.org/10.1038/nmeth.1818
Rhee SY, Beavis W, Berardini TZ et al (2003) The Arabidopsis information resource (TAIR): a model organism database providing a centralized, curated gateway to Arabidopsis biology, research materials and community. Nucleic Acids Res 31:224–228. https://doi.org/10.1093/nar/gkg076
Robinson WD, Carson I, Ying S et al (2012a) Eliminating the purple acid phosphatase AtPAP26 in Arabidopsis thaliana delays leaf senescence and impairs phosphorus remobilization. New Phytol 196:1024–1029. https://doi.org/10.1111/nph.12006
Robinson WD, Park J, Tran HT et al (2012b) The secreted purple acid phosphatase isozymes AtPAP12 and AtPAP26 play a pivotal role in extracellular phosphate-scavenging by Arabidopsis thaliana. J Exp Bot 63:6531–6542. https://doi.org/10.1093/jxb/ers309
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76. https://doi.org/10.1007/BF00020088
Sabet MS, Zamani K, Lohrasebi T et al (2018) Functional assessment of an overexpressed Arabidopsis purple acid phosphatase gene (AtPAP26) in tobacco plants. Iran J Biotechnol 16(1):31–41. https://doi.org/10.21859/ijb.2024
Schachtman DP, Reid RJ, Ayling SM (1998) Phosphorus uptake by plants: from soil to cell. Plant Physiol 116:447–453. https://doi.org/10.1104/pp.116.2.447
Schenk G, Guddat LW, Ge Y et al (2000) Identification of mammalian-like purple acid phosphatases in a wide range of plants. Gene 250:117–125. https://doi.org/10.1016/S0378-1119(00)00186-4
Schenk G, Mitić N, Hanson GR, Comba P (2013) Purple acid phosphatase: a journey into the function and mechanism of a colorful enzyme. Coord Chem Rev 257:473–482. https://doi.org/10.1016/j.ccr.2012.03.020
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. https://doi.org/10.1038/nmeth.2089
Sezonov G, Joseleau-Petit D, D’Ari R (2007) Escherichia coli physiology in luria-bertani broth. J Bacteriol 189:8746–8749. https://doi.org/10.1128/JB.01368-07
Srivastava R, Akash PAP et al (2020) Identification, structure analysis, and transcript profiling of purple acid phosphatases under Pi deficiency in tomato (Solanum lycopersicum L.) and its wild relatives. Int J Biol Macromol 165:2253–2266. https://doi.org/10.1016/j.ijbiomac.2020.10.080
Studer G, Biasini M, Schwede T (2014) Assessing the local structural quality of transmembrane protein models using statistical potentials (QMEANBrane). Bioinformatics 30:i505–i511. https://doi.org/10.1093/bioinformatics/btu457
Sun F, Suen PK, Zhang Y et al (2012) A dual-targeted purple acid phosphatase in Arabidopsis thaliana moderates carbon metabolism and its overexpression leads to faster plant growth and higher seed yield. New Phytol 194:206–219. https://doi.org/10.1111/j.1469-8137.2011.04026.x
Sun Q, Li J, Cheng W et al (2018) AtPAP2, a unique member of the PAP family, functions in the plasma membrane. Genes 9:257. https://doi.org/10.3390/genes9050257
Tran HT, Hurley BA, Plaxton WC (2010a) Feeding hungry plants: the role of purple acid phosphatases in phosphate nutrition. Plant Sci 179:14–27. https://doi.org/10.1016/j.plantsci.2010.04.005
Tran HT, Qian W, Hurley BA et al (2010b) Biochemical and molecular characterization of AtPAP12 and AtPAP26: the predominant purple acid phosphatase isozymes secreted by phosphate-starved Arabidopsis thaliana. Plant Cell Environ 33:1789–1803. https://doi.org/10.1111/j.1365-3040.2010.02184.x
Veljanovski V, Vanderbeld B, Knowles VL et al (2006) Biochemical and molecular characterization of AtPAP26, a vacuolar purple acid phosphatase up-regulated in phosphate-deprived Arabidopsis suspension cells and seedlings. Plant Physiol 142:1282–1293. https://doi.org/10.1104/pp.106.087171
Wang X, Wang Y, Tian J et al (2009) Overexpressing AtPAP15 enhances phosphorus efficiency in soybean. Plant Physiol 151:233–240. https://doi.org/10.1104/pp.109.138891
Wang L, Li Z, Qian W et al (2011) The Arabidopsis purple acid phosphatase AtPAP10 is predominantly associated with the root surface and plays an important role in plant tolerance to phosphate limitation. Plant Physiol 157:1283–1299. https://doi.org/10.1104/pp.111.183723
Wang L, Lu S, Zhang Y et al (2014) Comparative genetic analysis of Arabidopsis purple acid phosphatases AtPAP10, AtPAP12, and AtPAP26 provides new insights into their roles in plant adaptation to phosphate deprivation. J Integr Plant Biol 56:299–314. https://doi.org/10.1111/jipb.12184
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:W407–W410. https://doi.org/10.1093/nar/gkm290
Wu P, Ma L, Hou X et al (2003) Phosphate starvation triggers distinct alterations of genome expression in Arabidopsis roots and leaves. Plant Physiol 132:1260–1271. https://doi.org/10.1104/pp.103.021022
Yadava P, Dayaman V, Agarwal A et al (2022) Fine-tuning the transcriptional regulatory model of adaptation response to phosphate stress in maize (Zea mays L.). Physiol Mol Biol Plants 28:885–898. https://doi.org/10.1007/s12298-022-01155-x
Yang J, Zhang Y (2015) I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res 43:W174–W181. https://doi.org/10.1093/nar/gkv342
Young JC, Krysan PJ, Sussman MR (2001) Efficient screening of Arabidopsis T-DNA insertion lines using degenerate primers. Plant Physiol 125:513–518. https://doi.org/10.1104/pp.125.2.513
Zamani K, Sabet MS, Lohrasebi T et al (2012) Improved phosphate metabolism and biomass production by overexpression of AtPAP18 in tobacco. Biologia 67:713–720. https://doi.org/10.2478/s11756-012-0072-3
Zamani K, Lohrasebi T, Sabet MS et al (2014) Expression pattern and subcellular localization of Arabidopsis purple acid phosphatase AtPAP9. Gene Expr Patterns 14:9–18. https://doi.org/10.1016/j.gep.2013.08.001
Zamani K, Sabet MS, Lohrasebi T, Malboobi MA (2018) Functional analysis of AtPAP18 gene encoding a purple acid phosphatase in Arabidopsis thaliana. Mod Genet J 13:321–332
Zhang X, Henriques R, Lin S-S et al (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1:641–646. https://doi.org/10.1038/nprot.2006.97
Zhang W, Gruszewski HA, Chevone BI, Nessler CL (2008) An Arabidopsis purple acid phosphatase with phytase activity increases foliar ascorbate. Plant Physiol 146:431–440. https://doi.org/10.1104/pp.107.109934
Zhu H, Qian W, Lu X et al (2005) Expression patterns of purple acid phosphatase genes in Arabidopsis organs and functional analysis of AtPAP23 predominantly transcribed in flower. Plant Mol Biol 59:581–594. https://doi.org/10.1007/s11103-005-0183-0
Acknowledgements
This research was supported by a grant from the Tarbiat Modares University of I.R. Iran. We gratefully acknowledge the National Institute of Genetic Engineering and Biotechnology (NIGEB) for kindly supporting us in preparing of AtPAP17 construct. In the end, we should appreciate their valuable technical assistance.
Supplementary information
Table S1—Chi-square test for genotypes conformity from Mendelian 3:1 ratio (kanamycin resistant: sensitive).
Figure S1—Semi-quantitative RT-PCR test. The comparison of AtPAP17 transcript levels in (A) root or shoot of Arabidopsis WT against homozygous atpap17-mutant (Mu) line, or (B) some of distinct homozygous T3 overexpressed (OE) plants versus WT (samples prepared from whole of plant), or (C) root or shoot of WT and homozygous OE plant. RT-PCR reactions were done for 35 cycles for all experiments with the specific primers for AtPAP17 (Unique PAP17 F, R). The α-Tubulin transcript level was selected as an internal control (bottom panel). Samples were obtained by the whole of plant or shoot/root pool of WT plants, Mu line and OE plants.
Figure S2—Phenotypically comparison of 21-day-old seedlings. The AtPAP17-overexpressed (OE), atpap17-mutant (Mu) and WT plants have grown for 14 days on MS medium culture containing (A) 1.2 mM Pi (+ P) or (B) no Pi (− P). Unit scale is 1 cm.
Funding
This study was funded by a grant for a Master’s thesis from the Tarbiat Modares University of I.R. Iran (Grant No. 9085326).
Author information
Authors and Affiliations
Contributions
All the experiments, data analysis, and manuscript writing were done by AJL. MSS supervised the project and provided editorial input into the writing. MJ-J, KZ, TL, and MAM advised the project. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jamali Langeroudi, A., Sabet, M.S., Jalali-Javaran, M. et al. Functional assessment of AtPAP17; encoding a purple acid phosphatase involved in phosphate metabolism in Arabidopsis thaliana. Biotechnol Lett 45, 719–739 (2023). https://doi.org/10.1007/s10529-023-03375-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-023-03375-x