Abstract
Objectives
Metal-ion independent non-specific nucleases are of high potential for applications in EDTA-containing bioprocessing workflows.
Results
A novel extracellular non-specific nuclease EcNuc from the enterobacterium Escherichia coli has been identified. The recombinant gene was expressed and the protein was purified. Maximum activity of the enzyme was detected at 41.7 °C and at an acidic pH of 5.8. EcNuc tolerates EDTA in the reaction buffer at concentrations of up to 20 mM and the activity is not impaired by high concentrations of mono- and divalent metal ions in the absence of EDTA. The viscosity of crude protein extracts after cell lysis in EDTA-containing buffers is reduced when supplemented with EcNuc.
Conclusion
Proof-of-concept has been demonstrated that a metal-ion independent non-specific nuclease can be applied for removal of nucleic acids in EDTA-containing buffers for the subsequent purification of proteins from crude extracts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nucleases are ubiquitous in pro- and eukaryotic organisms. Their roles comprise the utilization of DNA as nutrients, they are involved in biofilm degradation or they act as a barrier to transformation of (host) cells by nucleic acids (Brown et al. 2012; Rangarajan and Shankar 2001). Due to their catalytic activity, nucleases are highly demanded in biotechnology and molecular biology approaches. Specific nucleases are widely applied as restriction enzymes, while non-specific nucleases are mainly utilized for the degradation of nucleic acids in crude protein extracts to decrease viscosity and to remove contaminating host cell nucleic acids during protein purification (Rangarajan and Shankar 2001).
Nucleases are classified based on primary sequences, sequence specificities, reaction mechanisms, substrates and reaction products (Yang 2011). Most non-specific nucleases are metal-ion dependent enzymes with NucA from Serratia marcescens being the best characterized isozyme. In this nuclease, a magnesium ion is obligatory and mainly coordinated by Asn119 acting as the principle ligand, while His89 is a general base that activates a water molecule for a nucleophilic attack on the phosphorous ester in the nucleic acid backbone (Benedik and Strych 1998). NucA from S. marcescens is commercially sold (trademark “Benzonase® Nuclease”) to be used in downstream processing, mainly for the elimination of nucleic acid contaminations in protein purification processes (Rangarajan and Shankar 2001).
There is only a limited number of nucleases known that do not require divalent metal ions for their catalytic activity including the restriction enzyme LlaKI from Lactococcus lactis KLDS4, GBSV1-NSN from the thermophilic bacteriophage GBSV1 and R.PabI, which is encoded by transposable elements in Archaea (Belkebir and Azeddoug 2012; Song and Zhang 2008; Wang et al. 2016). The non-specific DNase II-like proteins isolated from parasitic nematodes and the BfiI-type restriction enzymes from members of the genus Bacillus can be assigned to the phospholipase D family (Bao et al. 2008; Liao et al. 2014; Yang 2011). This superfamily of proteins is defined by a common catalytic domain represented by a typical HxK(x)4D(x)6GSxN motif that has been structurally characterized for the endonuclease Nuc from Salmonella enterica subsp. enterica serovar Typhimurium (Stuckey and Dixon 1999). Bacterial nucleases exhibit a single motif, while related proteins from eukaryotes are characterized by two copies of the consensus sequence. This group of enzymes is not only restricted to nucleases, but also contains phosphatidylserine and cardiolipin synthases and a Yersinia murine toxin (Rudolph et al. 1999; Stuckey and Dixon 1999).
In this study, a distantly related homologue of Nuc was identified in Escherichia coli. The gene was cloned and expressed and the recombinant protein was purified. Biochemical properties are reported and the potential application of EcNuc in downstream processing with a focus on EDTA-containing buffers was investigated.
Methods
Computational analysis
BLASTP analyses were used to identify uncharacterized nucleases in bacterial genomes. A putative endonuclease from Escherichia coli, which is deposited under the accession no. KZO82453.1, was identified and the SignalP 4.1 server (http://www.cbs.dtu.dk/services/SignalP/) was used to predict a hydrophobic signal peptide sequence. GraphPad was used for curve fitting purposes (Motulsky 2016).
Gene cloning, expression and purification
Culture conditions, plasmid propagation and maintenance as well as transformation of bacterial cells were done using standard molecular biology techniques. Plasmid pET-24d(+) in combination with Escherichia coli Veggie BL21 (DE3) Singles (both Merck, Darmstadt, Germany) was used for gene expression purposes of a signal peptide free and codon optimized gene variant (ATUM, California, USA) (Supplementary Figure 1). The recombinant protein was produced as a double 6xHIS-tag fusion enzyme (Fig. 1a) and purified in a three-step purification process: (1) affinity chromatography using Ni2+-NTA agarose (Qiagen, Hilden, Germany), (2) ion exchange chromatography using SP-Sepharose (GE Healthcare, Solingen, Germany), (3) proteolytic cleavage followed by another affinity chromatography using Ni2+-NTA agarose (Fig. 1b, c). The following buffers were used for purification approaches: (1) 50 mM NaPO4, 50 mM NaCl, pH 7.0 (equilibration buffer), plus 150 mM imidazole (washing buffer), or plus 500 mM imidazole (elution buffer), (2) 25 mM NaPO4, pH 6.0 (equilibration buffer), plus 300 mM NaCl (washing buffer), or plus 1000 mM NaCl (elution buffer), (3) flow-through containing non-tagged nuclease was sampled. Finally, buffer exchange was done using PD-10 columns (GE Healthcare, Solingen, Germany) and the protein was stored in 25 mM NaPO4, 25 mM NaCl, pH 7.0. A semi-dry Western blotting system (Biometra, Göttingen, Germany) was used to transfer proteins onto Roti®-PVDF membrane (Carl Roth GmbH + Co. KG, Karlsruhe, Germany). An anti-His-HRP antibody (Miltenyi Biotec, Bergisch Gladbach, Germany) was used in combination with the Immobilon™ Western HRP substrate (Merck, Darmstadt, Germany). Identity of the purified protein was verified by peptide mass fingerprinting (PMF) (data not shown).
Production, purification, substrate specificity and EDTA-tolerance of the non-specific nuclease EcNuc. a Schematic illustration of the fusion protein 2xHIS6-EcNuc construct including twin 6xHIS tag, protease cleavage site and coding region of EcNuc. SDS-PAGE (b) and Western blotting analyses (c) of samples from cell extract to purified EcNuc. E. coli Veggie BL21 (DE3) cells harboring the plasmid pET24d(+)::2xHIS6-EcNuc were grown for 20 h without induction at constant shaking and 37 °C (CE—crude extract). Induction of expression led to lethality. SN supernatant, Pellet pellet fraction, NiNTA eluted fraction after affinity chromatography, AIEX eluted fraction after anion exchange chromatography, Cleaved proteolytically cleaved fusion proteins, NiNTA2 flow through of untagged EcNuc after affinity chromatography. d To determine the substrate specificity, EcNuc was incubated with different types of DNA. The degradation efficiency towards the expression plasmid pET24d(+)::2xHIS6-EcNuc (Plasmid) was compared to unsheared dsDNA from calf thymus, to sheared dsDNA from salmon sperm, to ssDNA and to RNA, respectively. e Activity in EDTA-containing buffer was analyzed at concentrations between 1 and 50 mM EDTA. All assays were performed in 50 mM sodium phosphate buffer, pH 6.0 at 40 °C
Enzyme activity assays
Qualitative assays: about 100–200 ng of purified enzyme was incubated for 30 min in reaction buffer supplemented with DNA at a concentration of 5 µg in a final volume of 20 µl. Protein concentrations were determined with the Pierce™ BCA Protein Assay Kit (Thermo Fisher Scientific, Darmstadt, Germany). Reaction was stopped by the addition of 0.25% (w/v) sodium dodecyl sulfate (SDS) prior to agarose gel analysis and ethidium bromide staining. Assays were performed in 50 mM sodium phosphate buffer at pH 6.0 and 25 °C, while concentrations of detergents, metal ions and chelators were varied. Substrate specificity was tested using circularized plasmid DNA, single-stranded DNA (ssDNA) from calf thymus (Sigma-Aldrich, St. Louis, USA) and double-stranded sheared and unsheared genomic DNA (dsDNA) namely UltraPure™ Salmon Sperm DNA Solution (Thermo Fisher Scientific, Darmstadt, Germany), deoxyribonucleic acid from calf thymus, and MS2 RNA (both Sigma-Aldrich, St. Louis, USA).
Quantitative assays: about 20 ng of purified enzyme was incubated for 10–30 min in reaction buffer supplemented with DNA at a concentration of 0.5–60 µg in a final volume of 100 µl. The formation of acid-soluble nucleotides was determined spectrophotometrically at 260 nm in the VICTOR™ X4 Multilabel Plate Reader. The nucleolytic degradation of plasmid DNA was measured in a continuous approach, while enzymatic activity towards genomic DNA (single- and double-stranded, sheared and unsheared) was determined by end point measurements. One unit of nuclease activity is defined as ΔA260 min−1 per mg protein. A mean molar mass per nucleotide of 330 g mol−1 was defined to calculate µM from concentrations of sheared dsDNA. kcat values were calculated from vmax by the assumption that the degradation of 0.05 mg DNA resulted in a change in absorbance of 0.3 in combination with the general formula kcat = Vmax/[E0] (MacLellan and Forsberg 2001).
To evaluate the application potential of EcNuc for the elimination of nucleic acids from crude protein extracts, E. coli Veggie BL21 (DE3) Singles were transformed using the mock vector pET24d(+) and grown to high cell densities (data not shown). Afterwards, cells were harvested and resuspended in 50 mM sodium acetate buffer, pH 6.0 prior to cell lysis (High-pressure homogenization). One gram of cells were dissolved in 4 ml of buffer. Lysed cells were either supplemented with EcNuc (~ 10 Units) or with a metal-ion dependent nuclease (Benzonase® Nuclease, Merck, Darmstadt, Germany, ~ 10 Units) and incubated with or without 20 mM EDTA for 1 h at 35 °C. Viscosity of protein extracts was tested by gravity flow experiments using 1.2 ml pipette tips (Eppendorf, Wesseling-Berzdorf, Germany).
Results
A putative endonuclease from Escherichia coli
BLASTP analyses using a characterized endonuclease of the phospholipase D (PLD) family from Salmonella enterica subsp. enterica serovar Typhimurium (formerly known as Salmonella typhimurium, Nuc, PDB: 1BYS_A) as input sequence revealed the presence of a distantly related homologue in the enterobacterium Escherichia coli (KZO82453.1). The unprocessed protein is composed of 169 amino acid residues with a theoretical isoelectric point of pI 9.2 and a predicted molecular mass of 18.4 kDa. A putative signal peptide for secretion into the periplasmic space was predicted to be covered by amino acid residues 1–16. The sequence identity of EcNuc to Nuc is 65.3%. A typical sequence motif of the PLD superfamily HxK(x)4D(x)6GSxN is conserved between His108 and Asn125 (Stuckey and Dixon 1999).
Purification of EcNuc
Expression of the gene in E. coli as fusion protein composed of a 2xHis6 tag, a linker including protease cleavage site and the codon-optimized gene encoding EcNuc without the signal peptide and purification of the recombinant protein was visualized by Coomassie-stained SDS gel and Western blotting analysis using anti-His-HRP antibody. A single band of ~ 30 kDa was detected by Western blot that is conform to the calculated mass of the EcNuc fusion protein (12 + 17 kDa). A cleaved variant that is exclusively composed of the mature enzyme is visible on the SDS gel (Fig. 1b), but not detectable by Western blot analysis (Fig. 1c). A second band of ~ 42 kDa on the SDS gel has been identified by PMF to be E. coli elongation factor Tu1 (data not shown). A final yield of 39% and a purification factor of 171 was obtained for EcNuc (Supplementary Table 1). About 1.5–2.0 mg purified protein was obtained from 1 l high cell density fermentation (data not shown).
Catalytic activity of EcNuc
EcNuc was incubated with different types of nucleic acids, including dsDNA from salmon sperm (sheared) and calf thymus (unsheared), ssDNA from calf thymus and plasmid DNA. All types of nucleic acids were completely degraded indicating that the enzyme acts in an unspecific way (Fig. 1d). Moreover, the enzyme hydrolyzed RNA (Fig. 1d). EcNuc is not affected by low concentrations of EDTA (0–10 mM), but becomes partly inactivated at higher concentrations (Fig. 1e). The nuclease displayed optimal activity at an acidic pH of 5.8 (Fig. 2b) and at a temperature of 41.7 °C (Fig. 2a). The influence of different ions and other supplements on the catalytic activity of EcNuc were evaluated, because salt and detergent tolerance is an important prerequisite for the application of nucleic acid degrading enzymes in industrial downstream processing workflows (Fig. 2c). Michaelis–Menten kinetics revealed an affinity of KM = 202 µM and a maximal reaction velocity of vmax = 4559 U mg−1 towards sheared dsDNA (Table 1).
Effects of temperature, pH and different chemical compounds on the catalytic performance of EcNuc. a An optimal temperature of 41.7 °C was determined when enzymatic activity was tested at different temperatures. b Variations of the pH conditions revealed an optimal pH of 5.8. Circular plasmid DNA was used as substrate. c Various chemical compounds were tested for activating and inhibitory effects. EcNuc was incubated for 60 min on ice in the presence of a specific compound before a quantitative activity assay at 40 °C and pH 6.0 was conducted. The enzyme is inhibited by alcohol solvents (50%(v/v) EtOH and 50% (v/v) MeOH), by chaotropic salts (1 M GuHCl) and by detergents (0.1% (w/v) SDS), respectively. Standard deviations are the result of three independent measurements. EtOH ethanol, MeOH methanol, DTT dithiothreitol, 2-ME 2-mercaptoethanol, GuHCl guanidine hydrochloride, PMSF phenylmethane sulfonyl fluoride, AEBSF 4-(2-aminoethyl)benzenesulfonyl fluoride hydrochloride, and SDS sodium dodecyl sulfate
Application of EcNuc to eliminate nucleic acids from crude protein extracts
Gravity flow experiments were performed to visualize the reduced viscosity of crude protein extracts that were supplemented with EcNuc (Fig. 3a). A metal-ion dependent nuclease (Benzonase) and EcNuc are both capable of degrading nucleic acids derived from lysed cells (Fig. 3b). The metal-ion dependent nuclease is not able to reduce the viscosity in crude protein extracts that were supplemented with 20 mM EDTA, while 76.0% ± 11.3% of a cell sample that was treated with EcNuc passed through the pipette tip in the gravity flow experiment, indicating the catalytic activity of the enzyme under these conditions (Fig. 3c).
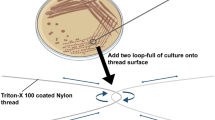
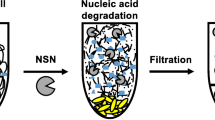
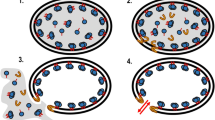
Potential application of EcNuc to degrade nucleic acids in crude protein extracts. a Schematic illustration. b, c Documentation of gravity flow experiments to visualize the viscosity of crude cell extract samples. 1 control without enzyme, 2 metal-ion dependent nuclease (Benzonase), 3 EcNuc. bEcNuc and a metal-ion dependent nuclease degrade the nucleic acids in the sample without EDTA and enable the crude protein extract to run through the pipette tip, while a sample containing no nuclease leads to a plugged pipette tip and the formation of a viscous drop. cEcNuc is also capable to reduce the viscosity of a crude protein extract sample that was supplemented with 20 mM EDTA, while the metal-ion dependent nuclease only displayed reduced activity. Bar charts aside of the pictures indicate the volume that passed through the pipette tips. Values are given as % of volume passed through the tip. The amount of cell sample that was filled into the tip at the beginning of the experiment (about 1 ml of lysed crude protein extract) was set to 100%. Standard deviations are the result of three independent measurements
Discussion
Due to their versatile application potential, non-specific nucleases have attracted considerable interest. There are several enzymes in E. coli that are capable to degrade nucleic acids. The nuclease domain of the bacterial toxin colicin E9 displayed catalytic activity towards dsDNA in the presence of the transition metals Co2+ and Ni2+, while no Ca2+-dependent activation of the enzyme was determined (Pommer et al. 1998). In contrast to the nuclease domain of the bacterial toxin colicin E9, activity of EcNuc is not dependent on the presence of divalent cations. Moreover, EcNuc depolymerized deoxynucleic acid, including closed circular plasmids as well as sheared and unsheared genomic DNA. It is most active at 41.7 °C and at a pH of 5.8, which is comparable to the extracellular nuclease Rv0888 from the pathogenic bacterium Mycobacterium tuberculosis exhibiting optimal activity at 41 °C and at a pH of 6.5 (Dang et al. 2016). Determination of kinetic parameters and comparison with commercial enzymes revealed that EcNuc exhibited higher activity than bovine DNase I, but it is less active compared with NucA from S. marcescens (Table 1).
During purification EcNuc and EF-Tu1 (elongation factor Tu1) coeluted from the column, indicating a putative interaction between both proteins. It has been described earlier that EF-Tu1 undergoes specific polymerization and tightly binds to DNases (Beck et al. 1978). Only a limited number of isozymes that are independent of metal ions in their catalytic region have been experimentally tested in detail and were investigated with regard towards their application potential in biotechnology. Tolerance against high salt concentrations is demanded in downstream processes, because standard buffers for protein chromatography often contain 50–500 mM NaCl. Another positive effect of high salt concentrations is that nucleic acids are enabled to dissociate from proteins at high salt concentrations. Thereby, these nucleic acids become available for degradation by salt tolerating enzymes. EcNuc is slightly inhibited in the presence of high salt concentrations. The nuclease from S. marcescens is not affected by concentrations of NaCl up to 200 mM (Eaves and Jeffries 1963), but becomes quickly inactivated by chelating agents. In contrast, EcNuc might be applicable in EDTA-containing cell lysis buffer, useful to increase yields of recombinant proteins due to inactivation of metal-dependent proteases. However, EDTA does not allow a metal-ion dependent affinity chromatography approach, such as Ni2+-HIS-tag chromatography as a first purification step. Therefore, EcNuc in combination with EDTA-containing buffers is compatible with alternative purification approaches including hydrophobic interaction chromatography, ion exchange chromatography or the recently developed ultrahigh affinity chromatography (Vassylyeva et al. 2017).
Conclusions
A novel metal-ion independent nuclease from E. coli has been produced and characterized. Due to its substrate promiscuity, EcNuc has been shown to be applicable for the removal of different types of nucleic acids in EDTA-containing buffers for the subsequent purification of proteins from crude extracts.
References
Bao Y, Higgins L, Zhang P, Chan SH, Laget S, Sweeney S, Lunnen K, Xu SY (2008) Expression and purification of BmrI restriction endonuclease and its N-terminal cleavage domain variants. Protein Expr Purif 58:42–52
Beck BD, Arscott PG, Jacobson A (1978) Novel properties of bacterial elongation factor Tu. Proc Natl Acad Sci USA 75:1250–1254
Belkebir A, Azeddoug H (2012) Characterization of LlaKI, a new metal ion-independent restriction endonuclease from Lactococcus lactis KLDS4. ISRN Biochem 2012:287230
Benedik MJ, Strych U (1998) Serratia marcescens and its extracellular nuclease. FEMS Microbiol Lett 165:1–13
Brown A, Horobin A, Blount DG, Hill PJ, English J, Rich A, Williams PM, Pritchard DI (2012) Blow fly Lucilia sericata nuclease digests DNA associated with wound slough/eschar and with Pseudomonas aeruginosa biofilm. Med Vet Entomol 26:432–439
Dang G, Cao J, Cui Y, Song N, Chen L, Pang H, Liu S (2016) Characterization of Rv0888, a novel extracellular nuclease from Mycobacterium tuberculosis. Sci Rep 6:19033
Eaves GN, Jeffries CD (1963) Isolation and properties of an exocellular nuclease of Serratia marcescens. J Bacteriol 85:273–278
Liao C, Liu M, Bai X, Liu P, Wang X, Li T, Tang B, Gao H, Sun Q, Liu X, Zhao Y, Wang F, Wu X, Boireau P, Liu X (2014) Characterisation of a plancitoxin-1-like DNase II gene in Trichinella spiralis. PLoS Negl Trop Dis 8:e3097
MacLellan SR, Forsberg CW (2001) Properties of the major non-specific endonuclease from the strict anaerobe Fibrobacter succinogenes and evidence for disulfide bond formation in vivo. Microbiology 147:315–323
Motulsky H (2016) GraphPad Curve Fitting Guide. http://www.graphpad.com/guides/prism/7/curve-fitting/index.htm. Accessed 25 September 2018
Pommer AJ, Wallis R, Moore GR, James R, Kleanthous C (1998) Enzymological characterization of the nuclease domain from the bacterial toxin colicin E9 from Escherichia coli. Biochem J 334(Pt 2):387–392
Rangarajan ES, Shankar V (2001) Sugar non-specific endonucleases. FEMS Microbiol Rev 25:583–613
Rudolph AE, Stuckey JA, Zhao Y, Matthews HR, Patton WA, Moss J, Dixon JE (1999) Expression, characterization, and mutagenesis of the Yersinia pestis murine toxin, a phospholipase D superfamily member. J Biol Chem 274:11824–11831
Song Q, Zhang X (2008) Characterization of a novel non-specific nuclease from thermophilic bacteriophage GBSV1. BMC Biotechnol 8:43
Stuckey JA, Dixon JE (1999) Crystal structure of a phospholipase D family member. Nat Struct Biol 6:278–284
Vassylyeva MN, Klyuyev S, Vassylyev AD, Wesson H, Zhang Z, Renfrow MB, Wang H, Higgins NP, Chow LT, Vassylyev DG (2017) Efficient, ultra-high-affinity chromatography in a one-step purification of complex proteins. Proc Natl Acad Sci USA 114:E5138–E5147
Wang D, Miyazono KI, Tanokura M (2016) Tetrameric structure of the restriction DNA glycosylase RPabI in complex with nonspecific double-stranded DNA. Sci Rep 6:35197
Yang W (2011) Nucleases: diversity of structure, function and mechanism. Q Rev Biophys 44:1–93
Acknowledgements
We thank Stefan Edelburg for the help with the VICTOR™ X4 Multilabel Plate Reader.
Supporting information
Supplementary Figure 1—Codon usage optimized nucleotide sequence of EcNuc without predicted signal peptide. The deduced amino acid sequence is indicated below in 1-letter amino acid code.
Supplementary Table 1—Purification of recombinant EcNuc after expression in E. coli Veggie BL21 (DE3).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Schmitz, S., Nölle, V. & Elleuche, S. A non-specific nucleolytic enzyme and its application potential in EDTA-containing buffer solutions. Biotechnol Lett 41, 129–136 (2019). https://doi.org/10.1007/s10529-018-2618-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-018-2618-0