Abstract
Objectives
To clone and characterize caffeic acid 3-O-methyltransferase (LcCOMT) from the rhizome of Ligusticum chuanxiong, a traditional medicinal herb having a high content of ferulic acid.
Results
LcCOMT encoded an ORF of 362 amino acids with a calculated MW of 39,935 Da and pI of 5.94. Polygenetic tree indicated that LcCOMT was attributed to a new member of COMTs in plants. The recombinant LcCOMT was expressed in E. coli. HPLC and 1H NMR analyses of purified LcCOMT protein confirmed that it could catalyze caffeic acid to produce ferulic acid in vitro. The further site-mutagenesis proved that His268 was one key catalytic residue. In addition, the substantial changing expression level of LcCOMT under chilling treatment suggested that LcCOMT might play important role in the accumulation of ferulic acid under chilling treatment.
Conclusions
This is the first report of the isolation and characterization of a COMT clone from traditional medicine containing high contents of pharmaceutical ferulic acid.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Caffeic acid 3-O-methyltransferase (COMT), belonging to the S-adenosyl-l-methionine (SAM)-dependent O-methyltransferase family, is an essential enzyme in the biosynthesis of phenylpropanoids, especially in the lignin biosynthetic pathway. COMT catalyzes the methylation of the 3-hydroxyl group of caffeic acid to produce ferulic acid and S-adenosyl-l-homocysteine (Vanholme et al. 2012).
Since the first COMT cDNA was reported in 1991 (Bugos et al. 1991), COMT cDNAs have been cloned and characterized from a number of different plants, including Iris hollandica, Nicotiana tabacum, Arabidopsis thaliana, Zea mays, etc. (Vincent et al. 2005). The crystal structure of COMT from Medicago sativa was reported by Zubieta et al. (2002). COMT catalyzes the rate-limiting step of lignin monomer biosynthesis (Trabucco et al. 2013). Regulation of COMT gene showed impacts on lignin structure and implications for the biosynthesis of G and S lignin (Tschaplinski et al. 2012).
Ligusticum chuanxiong L. (Apiaceae), an important herb mainly cultivated in Sichuan Province, China, has attracted attention as its dry rhizome has been widely used as traditional medicine for the treatment of many diseases, including headache, rheumatic arthralgia, cardiovascular diseases, menstrual disorder and swelling pain in China, Japan and Korea (Li et al. 2012). Among the pharmaceutical compounds, including ferulic acid, essential oil, terpenoids and alkaloids, ferulic acid was the most significant active gradient and regarded as the primary marker to evaluate the pharmaceutical value of L. chuanxiong in the Chinese Pharmacopoeia. In addition to the documented health benefits above, ferulic acid is also used to produce cosmetics, food and health products (Li et al. 2012).
There are several traditional medicines, including L. chuanxiong, Angelica sinensis and Ferula sinkiangensis, containing high contents of ferulic acid (Vincent et al. 2005). To date, however, no reports have been published on the molecular and biochemical characterization of COMTs in the above plants. We have obtained the transcriptome sequence from the rhizome of L. chuanxiong (Song et al. 2015). Based on function annotations, we found a unigene (C41658) with the highest homology with Ammi majus COMT (accession number Q6T1F5). It was named as L. chuanxiong COMT (LcCOMT). It had a fragment of 680 bp length containing 65 nucleotides downstream of the terminal TAG codon but it is an incomplete open reading frame. In this work, our main aim was the full-length molecular cloning and functional expression of LcCOMT cDNA from L. chuanxiong. In addition, the key amino acid residue of LcCOMT was identified by site-mutagenesis. Finally, the expression pattern of LcCOMT under difference stresses was investigated.
Materials and methods
Molecular cloning of LcCOMT gene
Ligusticum chuanxiong was cultivated in pots containing a soil mixture (nutrient soil:vermiculite:peat, 1.5:1:1, by volume) and grown at 25 °C with 16 h light and 8 h dark. The mature rhizome of L. chuanxiong was collected and frozen in liquid N2 and then stored at −80 °C for RNA isolation. Total RNA was isolated using the RNA plant plus reagent (Omega, USA). The RNA samples were treated with DNase I at 1 unit μg−1 of total RNA to remove DNA contaminate. Purified RNA was then used to synthesize the first-strand cDNA by M-MLV reverse transcriptase (Takara, Japan) using oligo dT-AP primer (5′-GCT GTC AAC GAT ACG CTA CGT AAC GGC ATG ACA GTG TTT TTT TTT TTT TTT TTT-3′). The first PCR for 5′-RACE was performed by Ex Taq (Takara, Japan) based on 1 μl tailed cDNA template in 25 μl reaction buffer using 5′-TGG AGT CTG GAG CCT CTG GAA-3′ and oligo dT-T7 (5′-AGG ACT CAC TAT AGG GCT TTT TTT TTT TVN-3′) as primers. The second PCR for 5′-RACE was carried out by Ex Taq based on 1 μl PCR products from first PCR as template in 50 μl reaction buffer using 5′-ACC GCC AAA TCC GTC ATA-3′ and 5′-GTA ATA CGA CTC ACT ATA GGG C-3′ as primers. All resulting products were subcloned into pMD19-T vectors (Takara) and sequenced (Shanghai Songon, China). The specific primers were designed with Primer Premier 5.0 software. Multiple alignments of deduced amino acid sequences were carried out using DNAMAN software. The phylogenetic relationship of LcCOMT was analyzed with MEGA 6.0 programs (Liu et al. 2015a, b).
Construction of expression plasmids
The full-length ORF of LcCOMT gene was amplified by PCR using the forward primer (5′-CGC GGA TCC ATG AAT ACG GAG CTG ATC CCA CC-3′, the EcoR1 site is underlined and the initiation codon is shown in italics) and reverse primer (5′-CGG AAT TCA CAT TAA GCA GAT GCC AGA CAC CC-3′, the BamH1 site is underlined and the stop codon is shown in italics), to introduce two restriction sites, 5′-EcoR1 and 3′-BamH1, which are immediately followed by the initiation codon and preceded by the stop codon, respectively. The fragment was cloned into pMD19-T vector for sequencing to confirm that no substitutions or deletions occurred. The insert was digested with EcoR1 and BamH1 and gel purified with a gel extraction kit and then integrated into a purified pET28a vector (Amersham) digested with the same enzymes, resulting in a recombinant plasmid pET28a-LcCOMT. The recombinant plasmid pET28a-LcCOMT and empty vector pET28a were transformed into E. coli BL21 (DE3) using standard protocol.
Protein expression and purification
To express LcCOMT fusion protein, the cells transformed with pET28a-LcCOMT were grown at 37 °C in 500 ml LB liquid medium containing 0.1 mg kanamycin ml−1. When the OD600 reached 0.6, IPTG was added to 1 mM and the culture was further incubated for 6 h at 17 °C and 160 rpm. The cells were harvested by centrifugation for 10 min at 4 °C and 5000×g. The sediment was suspended in lysis buffer [20 mM Tris/HCl, pH 7.9, containing 0.1 % (w/v) lysozyme and 150 mM NaCl] and sonicated four times in 20 s bursts at medium intensity. The resulting suspension was centrifuged at 4 °C and 13,000×g for 10 min and the supernatant was collected and loaded onto a Ni-agarose column equilibrated with buffer [20 mM Tris/HCl (pH 8.0), 150 mM NaCl] following the manufacturer’s instructions. The Ni-agarose column loaded recombinant protein was rinsed with buffer [20 mM Tris/HCl (pH 8.0), 150 mM NaCl and 10 mM imidazole]. The LcCOMT proteins bound to the Ni–NTA column were eluted with 0.4 M imidazole, which was subsequently removed by dialysis using distilled water containing 5 % (v/v) glycerol. The purified protein concentration was measured according to Bradford method. Ten μg crude protein extract and 10 μg purified protein were analyzed using 12.5 % SDS-PAGE followed by Coomassie Blue staining.
Enzymatic assay of LcCOMT
The standard reaction mixture consisted of 500 μM caffeic acid, 1 mM SAM and 1 μg purified LcCOMT in 100 μl 100 mM potassium phosphate buffer (pH 7.0) containing 100 mM MgCl2 and 1 mM DTT. After incubation at 37 °C for 120 min, the reaction was terminated by adding 50 μl HCl and the products were then extracted by ethyl acetate four times. The extracted products in ethyl acetate solvent were filtrated with a membrane filter (0.22, 13 mm, Millipore) and analyzed by reverse-phase HPLC with a Hypersil C18 column (4.6 mm × 150 mm). Caffeic acid and ferulic acid in eluted fractions were assayed as described previously by Yoshihara et al. (2008).
The reaction products in ethyl acetate were separated on a Hypersil BDS C18 column with mobile phase of methanol/water (1:4 v/v). The purified peak of product was dissolved in (1:1, v/v) CD3OD and identified using 1H NMR. The NMR spectra were recorded at 25 °C on a Bruker 600-MHz AVANCEIII NMR spectrometer (Bruker, Germany) equipped with a Bruker 5 mm PA BBO probe operated at 600.13 MHz 1H frequency with default parameters.
The optimum pH of LcCOMT was determined by using difference pH buffer ranging from 5.0 to 9.0, and LcCOMT activity was measured at each pH at 37 °C. The optimum temperature of LcCOMT was also assayed in reaction mixtures containing 100 mM potassium phosphate (pH 7.0) at temperatures of 22–42 °C.
Site-directed mutagenesis
The codon CAC (His268) was changed to CTG (Leu) using primer pair (5′-AAG TGG ATA TGT CTG GAT TGG AGC GAT-3′ and 5′-ATC GCT CCA ATC CAG ACA TAT CCA CTT-3′). The mutagenesis was performed using point mutation kit (Tiandz, China). The pET28a-LcCOMT was used as template to produce the mutated recombinant constructs pET28a-LcCOMT H268L. The transformation, expression, purification and enzymatic assays of mutated protein were performed using the same methods as those used for pET28a-LcCOMT.
Expression pattern analysis of LcCOMT under stresses
For chilling stress, L. chuanxiong were cultivated at 4 °C for 0, 1, 6, 12, 24 and 48 h, respectively. For drought and salt stresses, L. chuanxiong were cultivated supplemented with PEG 6000 (100 mg ml−1), or NaCl (200 mM) for 0, 1, 6, 12, 24 and 48 h, respectively. The total RNA of L. chuanxiong leaf was extracted and subjected to cDNA synthesis with the same method used for 5′-RACE.
To analyze LcCOMT expression levels in L. chuanxiong under different stresses, qRT-PCR was performed using SYBR Premix Ex TaqTM II (Takara, Japan) with LightCycler 96 system (Roche Diagnostics, Mannheim, Germany). In each run, 1 μl cDNA template was added to 15 μl reaction buffer using gene-specific primer pairs under the following conditions: pre-denaturation at 95 °C for 30 s, followed by 40 cycles of 5 s at 95 °C, 10 s at 60 °C, and 25 s at 72 °C. Gene-specific primer pairs for real-time PCR of LcCOMT (5′-ACT TTG ATC TCC CTC ATG TTG TTG-3′ and 5′-TCC CCT TTG GGC ACG CTA-3′) and reference gene ribosomal protein L11 (5′-CTC CTT GGT AAC CCT GTG CTG A-3′ and 5′-GTG ATA CTG GAT GTT TTG GCT TTG-3′) were designed using Primer Premier 5.0 based on the full-length cDNA. The relative expression levels of the selected genes were normalized to the reference gene and determined by the ΔΔCt-method. Three biological repeats of each tissue were performed in qRT-PCR assay.
Results and discussion
Cloning and sequence analysis of LcCOMT gene
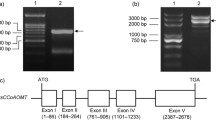
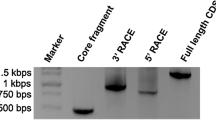
According to BLAST searches, the full length cDNA encoding LcCOMT was obtained by RACE from rhizome of L. chuanxiong. The nucleotide sequence analysis showed that LcCOMT (GenBank accession no. KR106206) contained an ORF of 1171 bp and 362 amino acids with a calculated molecular mass of 39,935 Da and isoelectric point of 5.94. Comparison of the deduced amino acid sequences between LcCOMT and known SAM-dependent OMTs (Supplementary Table 1) showed 60.3–94.3 % identity with COMTs. LcCOMT had the highest identity of 94.3 % with A. majus (Apiaceae) COMT. Further BLAST searches also detected 23.9–40.2 % identity with alkaloid, flavonoid, isoflavonoid and phenylpropanoid OMTs.
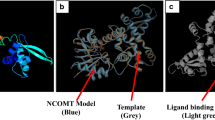
Based on the analysis of crystallized COMT in complex with substrate from M. sativa and homology model of (S)-scoulerine 9-OMT (CjSMT) in complex with substrate from Coptis chinensis, there are three consensus domains proposed for plant OMTs, including the substrate binding domain, the SAM binding domain and the methyl transfer system (Zhu et al. 2012; Zubieta et al. 2002). As shown in Supplementary Fig. 1, multiple alignments of LcCOMT and SAM-dependent OMT family members with known substrate specificity revealed that LcCOMT contained the above three consensus domains. The substrate binding domain was composed of two regions (113–167 and 300–308). Comparative substrate-binding site analysis of COMTs and OMTs with different substrate specificity showed differences in the respective substrate binding residues. For example, residues Met129, Leu135, Ala162, His165, Phe171, Ile315, and Ile318 in LcCOMT were replaced by Phe113, Val119, Phe146, Ala150, Val156, Met164, Pro300, and Leu303 in CjSMT, respectively. These differences in the active site probably affect the conformation and presumably play significant roles in the substrate specificity (Zhou et al. 2010; Zhu et al. 2012). The SAM binding site (residues 200–255) was composed of conserved residues (Asp205, Gly208, Asp230, Leu231, Asp230, Met231 and Lys264). The methyl transfer system was found in the C-terminus of LcCOMT and composed of three conserved catalytic residues (His268, Glu296 and Glu328).
LcCOMT was subjected to phylogenetic analysis with 40 OMTs retrieved from NCBI protein database in order to understand the relationship of LcCOMT with other OMTs (Supplementary Fig. 2). The branching pattern of the OMTs phylogenetic tree showed two distinct clusters. LcCOMT was located to the COMTs cluster near to A. majus COMT (Hehmann et al. 2004). Collectively, LcCOMT was attributed to the COMTs family.
Expression and enzymatic assay of recombinant LcCOMT
The recombinant LcCOMT was expressed in E. coli BL21 (DE3) with an MW of 43 kDa (see Fig. 1). Its molecular weight was higher than that of the calculated one of 39,935 kDa due to the His-tag in the N-terminal fusion peptide. The recombinant LcCOMT was further purified by Ni-agarose column. The purified recombinant LcCOMT with single band in SDS-PAGE was subjected to enzymatic assay.
The soluble fractions of extracts of E. coli expressing LcCOMT were subjected to SAM-dependent OMT assays using caffeic acid and SAM as substrates. As shown in Fig. 2c, when SAM was added to the mixture, ferulic acid was detected in the reaction products. This result confirmed that LcCOMT encodes SAM-dependent COMT, which catalyzes the transfer of the methyl moiety from SAM to caffeic acid to form ferulic acid.
HPLC analysis of reaction products catalyzed by purified recombinant LcCOMT. (a) A control of caffeic acid [HPLC retention time (Rt) 8.597 min], (b) a control of ferulic acid (Rt 15.582 min), (c) the reaction product of LcCOMT (SAM was added to the reaction mixture), and (d) the reaction product of LcCOMT H268L (SAM was added to the reaction mixture)
Furthermore, the purified product peak was analyzed using 1H NMR (Supplementary Fig. 3). The structure information of product was consistent with that of ferulic acid reported in previous literature (Tao et al. 2011), confirming the product was ferulic acid.
Site-directed mutagenesis
Based upon the crystal structure of MsCOMT (Zubieta et al. 2002) and sequence alignments with the large family of plant OMTs, methylation process involved base-assisted deprotonation of the hydroxyl group followed by a nucleophilic attack on the reactive methyl group of SAM. In LcCOMT, the methyl transfer system was composed of His268, Glu296 and Glu328 residues. Glu296 and Glu328 bracket the catalytic His268 residue, ensuring the optimal orientation of the imidazole group of His268 for deprotonation of the 3′-hydroxyl of the caffeic acid. Because the sulfur of SAM is positively charged, the transmethylation process is easily facilitated by the deprotonation step. In order to confirm whether the His268 was the key residue in the methylation, the codon CAC (His268) was mutated to CTG (Leu). Mutation of His268 to Leu completely eliminated methyltransferase activity (Fig. 2d), implicating His268 was indeed an important catalytic residue.
The activities of LcCOMT were analyzed at various pHs and temperatures using caffeic acid and SAM as substrates. The highest activity was observed at pH 7 in 100 mM potassium phosphate buffer (Fig. 3a). This matched with the optimum pH for the COMT activities of sweet basil and I. hollandica, which ranged from pH 6 to 8 (Gang et al. 2002; Yoshihara et al. 2008). The highest activity was reached at 37 °C in 100 mM potassium phosphate buffer (pH 7.0; Fig. 3b). LcCOMT lost its activity over 42 °C and only ~5 % activity remained. This agreed with the optimum temperature for COMT activity for caffeic acid in Thalictrum tuberosum (Frick and Kutchan 1999). In addition, considering the highest homology between LcCOMT and A. majus COMT, the optimal pH and temperature of both COMTs were compared. As a result, A. majus COMT showed the same optimum pH (pH 7.0), but lower optimum temperature (32 °C) (Hehmann et al. 2004).
Expression pattern of LcCOMT under stresses
Hehmann et al. (2004) reported that the expression of COMT was not affected by the fungal elicitor. In this study, the potted L. chuanxiong was subjected to different environmental stresses and then the expression pattern of LcCOMT was assayed by RT-PCR. As shown in Fig. 4, LcCOMT had different expression levels under different stresses. The expression of LcCOMT was relatively stable under salt stress, suggesting that salt stress did not affect the transcription of LcCOMT. The expression level of LcCOMT kept increasing in the first 12 h of drought treatment but increased no more than 1.5-fold at 12 h compared with that of control plant, suggesting that drought treatment have minor influence on the transcription of LcCOMT. This agreed with the result from Vincent et al. (2005) who reported that drought was scarcely related to lignin biosynthesis. The expression level of LcCOMT under chilling stress showed similar tendency with that under drought treat, however, the highest expression level of LcCOMT appeared at 12 h with more than 6-fold compared with that of control plant. It was interesting that L. chuanxiong was an annual herb, and its growing period could be divided into six stages. From ancient times until today, in order to improve its pharmaceutical quality, L. chuanxiong would be transplanted to areas with high altitude and suffer chilling treatment during the “senescence” stage (Li et al. 2012). In our study, the increased expression level (6-fold) of LcCOMT under chilling treatment suggested that LcCOMT might be utilized as a target to improve the content of ferulic acid of L. chuanxiong.
Conclusions
This is the first report of the cloning of the COMT gene. A new member of COMTs from L. chuanxiong, with the highest similarity to A. majus COMT, possessed conserved residues in the three consensus domains, which matched with other COMTs in plants. The sequence diversities in the substrate binding domain might be used to predict the substrate specificity of a novel OMT. The conserved substrate binding residues and catalytic residues found in LcCOMT suggest that LcCOMT possesses the potential to catalyze the methylation of caffeic acid, this speculation was confirmed by further enzymatic experiment of recombinant LcCOMT. His268 was identified as the key residue involved in the methyl transfer system. In addition, the substantial increased expression level of LcCOMT under cold stress suggested that LcCOMT might also play important roles in the accumulation of ferulic acid during chilling stress. Hopefully, LcCOMT could be used to construct transgenic plants in vivo or immobilized enzyme to produce ferulic acid in vitro in the future.
References
Bugos RC, Chiang VL, Campbell WH (1991) cDNA cloning, sequence analysis and seasonal expression of lignin-bispecific caffeic acid/5-hydroxyferulic acid O-methyltransferase of aspen. Plant Mol Biol 17:1203–1215
Gang DR, Lavid N, Zubieta C, Chen F, Beuerle T, Lewinsohn E, Noel JP, Pichersky E (2002) Characterization of phenylpropene O-methyltransferases from sweet basil: facile change of substrate specificity and convergent evolution within a plant O-methyltransferase family. Plant Cell 14:505–519
Hehmann M, Lukacin R, Ekiert H, Matern U (2004) Furanocoumarin biosynthesis in Ammi majus L. Cloning of bergaptol O-methyltransferase. Eur J Biochem 271:932–940
Li W, Tang Y, Chen Y, Duan JA (2012) Advances in the chemical analysis and biological activities of Chuanxiong. Molecules 17:10614–10651
Liu Z, Zhu Q, Li J, Zhang G, Jiamahate A, Zhou J, Liao H (2015a) Isolation, structure modeling and function characterization of a trypsin inhibitor from Cassia obtusifolia. Biotechnol Lett 37:863–869
Liu Z, Zhu Q, Li Y, Yu J, Wang W, Tan R, Zhou J, Liao H (2015b) Isolation and in silico characterization of a shikimate kinase from Cassia obtusifolia. Acta Physiol Plant 37:85
Morita Y, Saitoh M, Hoshino A, Nitasaka E, Iida S (2006) Isolation of cDNAs for R2R3-MYB, bHLH and WDR transcriptional regulators and identification of c and ca mutations conferring white flowers in the Japanese morning glory. Plant Cell Physiol 47:457–470
Song T, Liu ZB, Li JJ, Zhu QK, Tan R, Chen JS, Zhou JY, Liao H (2015) Comparative transcriptome of rhizome and leaf in Ligusticum chuanxiong. Plant Syst Evol. doi:10.1007/s00606-015-1211-4
Tao HM, Wang LS, Zhao DQ, Zhu QH, Liu YH (2011) Phenolic compounds from roots of Asparagus filicinus. Chin Tradit Herb Drugs 42:2181–2185 (in Chinese)
Trabucco GM, Matos DA, Lee SJ, Saathoff AJ, Priest HD, Mockler TC, Sarath G, Hazen SP (2013) Functional characterization of cinnamyl alcohol dehydrogenase and caffeic acid O-methyltransferase in Brachypodium distachyon. BMC Biotechnol 13:61
Tschaplinski TJ, Standaert RF, Engle NL, Martin MZ, Sangha AK, Parks JM, Smith JC, Samuel R, Jiang N, Pu Y, Ragauskas AJ, Hamilton CY, Fu C, Wang ZY, Davison BH, Dixon RA, Mielenz JR (2012) Down-regulation of the caffeic acid O-methyltransferase gene in switchgrass reveals a novel monolignol analog. Biotechnol Biofuels 5:71
Vanholme R, Morreel K, Darrah C, Oyarce P, Grabber JH, Ralph J, Boerjan W (2012) Metabolic engineering of novel lignin in biomass crops. N Phytol 196:978–1000
Vincent D, Lapierre C, Pollet B, Cornic G, Negroni L, Zivy M (2005) Water deficits affect caffeate O-methyltransferase, lignification, and related enzymes in maize leaves. Plant Physiol 137:949–960
Yoshihara N, Fukuchi-Mizutani M, Okuhara H, Tanaka Y, Yabuya T (2008) Molecular cloning and characterization of O-methyltransferases from the flower buds of Iris hollandica. J Plant Physiol 165:415–422
Zhou JM, Lee E, Kanapathy-Sinnaiaha F, Park Y, Kornblatt JA, Lim Y, Ibrahim RK (2010) Structure–function relationships of wheat flavone O-methyltransferase: homology modeling and site-directed mutagenesis. BMC Plant Biol 10:156
Zhu QK, Zhou JY, Zhang G, Liao H (2012) Homology modeling and molecular docking studies of Coptis chinensis (S)-scoulerine 9-O-methyltransferase. Chin J Chem 30:2533–2538
Zubieta C, Kota P, Ferrer JL, Dixon RA, Noel JP (2002) Structural basis for the modulation of lignin monomer methylation by caffeic acid/5-hydroxyferulic acid 3/5-O methyltransferase. Plant Cell 14:1265–1277
Acknowledgments
This work was supported by the Grant (No. 31371232) from National Natural Science Foundation of China, Grant (2014ZX09304307001-019) from National Science and Technology Major Project and Grant (No. 2682014RC14) from Fundamental Research Funds for the Central Universities of China.
Supporting information
Supplementary Fig. 1—Sequence alignment of LcCOMT with other SAM-dependent OMT family members.
Supplementary Fig. 2—Phylogenetic tree of LcCOMT and other OMTs constructed by neighbor-joining algorithm.
Supplementary Fig. 3—1H NMR chemical shifts (ppm) for ferulic acid.
Supplementary Table 1—Comparison of the deduced amino acid sequences between LcCOMT and known SAM-dependent O-methyltransferases.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We declare that we have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, JJ., Zhang, G., Yu, Jh. et al. Molecular cloning and characterization of caffeic acid 3-O-methyltransferase from the rhizome of Ligusticum chuanxiong . Biotechnol Lett 37, 2295–2302 (2015). https://doi.org/10.1007/s10529-015-1917-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1917-y