Abstract
5-Aminolevulinic acid (ALA) synthase (ALAS) HemA from non-sulfur photosynthetic bacteria has been used for the ALA bioproduction, whereas the isoenzyme HemT/HemO is less studied and not used for ALA production. Two ALAS-encoding genes, hemA and hemO from Rhodopseudomonas palustris were cloned, purified and characterized. The ALASs had very high specific activity, 3.6 and 2.7 U/mg, respectively, and strong affinity for one of its substrates, succinyl-CoA, K m with values of 11 and 4.4 μM, respectively. HemO retained up to 60 % maximum activity within a broad range of concentrations of hemin, while HemA kept only 20 % at 10 μM hemin. Escherichia coli overexpressing HemA or HemO produced 5.7 and 6.3 g ALA/l, respectively, in a 5 l bioreactor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
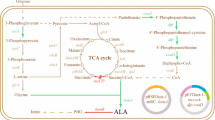
5-Aminolevulinic acid (ALA), the common precursor for the biosynthesis of tetrapyrroles such as heme and chlorophyll, has received extensive attention because of its broad applications as photosensitizer for photodynamic therapy and as a plant growth-promoting substance. Photosynthetic bacteria harboring native 5-aminolevulinic acid synthase (ALAS) HemA, which catalyzes a one-step condensation of succinyl-CoA and glycine (Nishikawa et al. 2002), or recombinant Escherichia coli expressing heterogeneous ALAS (Choi et al. 2008), are the most studied systems for biological production of ALA.
ALAS, encoded by the gene hemA from photosynthetic bacteria such as Rhodobacter sphaeroides and Rhodopseudomonas palustris, shows good characters in terms of specific activity or substrate affinity. The specific activity of the purified HemA from R. sphaeroides was 0.22 U/mg (Bolt et al. 1999), lower than that reported for the recombinant ALAS of R. palustris KUGB306, 0.9 U/mg (Choi et al. 2004). The K m value for succinyl-CoA was 11 μM in R. sphaeroides (van der Werf and Zeikus 1996) and 50 μM in R. palustris KUGB306 (Choi et al. 2004), respectively. The specific activity of GST-affinity purified ALAS from a rhizobia bacterium, Bradyrhizobium japonicum, was 20 U/mg, but no kinetic character was further studied (Lee et al. 2004, 2005). (All the enzyme activities cited here are recalculated based on unified enzyme unit definition, namely 1 U/mg protein is the amount of enzyme required for formation of 1 μmol ALA per min). Searching for ALAS with better enzymatic characters is important as it is obviously the key enzyme for biological production of ALA.
Compared to the well-studied hemA gene product, HemA, the investigation and application of its isozyme HemT isolated from R. sphaeroides is less successful because after overexpression of hemT in E. coli the protein was found neither in the soluble fraction in the cell extract nor had detectable activity in vitro (Bolt et al. 1999). Two ALAS isoforms, hemA and hemO, were recently predicted in the genome sequence of R. palustris CGA009 based on sequence similarity analysis (Larimer et al. 2003). However, further experimental evidence was absent. In the present study, we cloned the hemO gene as well as hemA from R. palustris and overexpressed both genes in E. coli. The study with purified enzymes indicated that their specific activities and K m values for succinyl-CoA are better than those from photosynthetic bacteria according to our knowledge. Fermentation test of E. coli strain MG1655 overexpressing HemO resulted in more than 6 g ALA/l in a 5 l fermentor, indicating the great application potential of the ALAS isomer HemO for ALA bioproduction.
Materials and methods
Strains, plasmids and growth condition
Bacterial strains and plasmids used in this work are listed in Supplementary Table 1. Rhodopseudomonas palustris ATCC 17001 was cultivated as described previously by Choi et al. (2004). For molecular manipulation, E. coli strains harboring plasmids were routinely grown in LB medium supplemented with 0.1 g ampicillin/l at 37 °C. ALA fermentation was conducted with two 5 l jar fermentors at 37 °C, with dissolved O2 at least 30 % (coupled with agitation speed at least 300 rpm), and pH 6.5 (adjusted with 1.84 M H2SO4 and ammonia). Aeration was at 1 vvm.
Cloning of hemA and hemO from R. palustris
Fragments containing the ORF and flanking region of hemA or hemO gene were amplified from R. palustris genomic DNA with primers hemA-F1 and hemA-R1, hemO-F1 and hemO-R1 (Supplementary Table 2), respectively, which were designed based on conserved sequences of other R. palustris ALASs. The amplified fragments were purified and subcloned into pEASY-Blunt Cloning Vector (TransGen, Beijing, China). The nucleotide and amino acid sequences were analyzed with the DNA MAN program.
Construction of ALAS recombinant expression vectors
The hemA and hemO genes were amplified from R. palustris genomic DNA with primers hemA-F2 and hemA-R2, hemO-F2 and hemO-R2 (Supplementary Table 2), The amplified products were cleaved with NdeI and HindIII and ligated into pET21a(+) treated with the same enzymes, and then transformed into E. coli Rosetta2 (DE3) for protein expression. The generated plasmid harboring hemA or hemO was named as pET21a–hemA or pET21a–hemO, respectively. For ALA production, the hemA or hemO gene was cleaved from pET21a–hemA or pET21a–hemO with XbaI and HindIII, ligated with pTrc99A treated with the same enzymes and transformed into E. coli MG1655. The obtained plasmid was named as pTrc99A–hemA or pTrc99A–hemO.
Expression and purification of recombinant ALASs
Escherichia coli Rosetta2 (DE3) harboring plasmid pET21a–hemA or pET21a–hemO was cultivated in LB medium containing 0.1 g ampicillin/l at 37 °C until the OD600 reached 0.6–1. Then 0.1 mM IPTG was added and the culture was incubated for 12 h at 28 °C. The cells were harvested by centrifugation at ~8,000*g for 5 min and washed twice with binding buffer (50 mM NaH2PO4, pH 7.4, 30 mM imidazole, 500 mM NaCl). The pellet was resuspended in 10 ml binding buffer and lysed by sonication. After centrifugation at ~10,000*g for 20 min, the supernatant was applied to the purification process with His-Trap columns according to the manufacturer’s instruction. The protein purity was assessed by SDS-PAGE stained with colloidal Brilliant Blue G-250. The protein concentration was measured using Thermo Scientific Pierce BCA Protein Assay Kit.
Enzyme assay and characterization
ALA synthase activity was determined using the method described elsewhere (Burnham 1970). The reaction mixture contained 50 mM Tris/HCl (pH 7.5), 200 mM glycine, 0.2 mM succinyl-CoA, 0.1 mM pyridoxal phosphate (PLP) and 1.78 × 10−3 U (HemA) or 1.32 × 10−3 U (HemO) ALAS. One unit was defined as the amount of enzyme required for formation of 1 μmol ALA per min.
For determination of the kinetic parameters of ALAS, the concentration of the substrates glycine and succinyl-CoA was set in the range of 0.1–10 times of the K m values of other ALASs reported in literature. Chlorohemin was set as appropriate concentrations to examine the inhibitive effect of hemin on the recombinant ALAS while the other substrates were kept excessive (glycine 200 mM and succinyl-CoA 0.2 mM).
Analytical methods
ALA was determined by following the method described by Mauzerall and Granick (1956). All the assays were in triplicate. Glucose concentration was determined with the glucose analyzer SBA-40D. The concentrations of glycine were measured by HPLC using ODS-SP column (Shimadzu) according to the method described by Heinrikson and Meredith (1984). Concentrations of succinic acid were measured by HPLC using Aminex HPX-87H column (BioRad) according to the method of Chinnici et al. (2005).
Accession number of nucleotide sequences
The nucleotide sequences of R. palustris ATCC 17001 hemA and hemO genes have been deposited in the GenBank under the accession number JQ048720 and JQ048722, respectively.
Results and discussion
Cloning of ALAS isozymes from R. palustris
The presence of ALAS isoenzymes has been found in mammals and also in the photosynthetic bacterium R. sphaeroides (Bolt et al. 1999; Larimer et al. 2003). So far no experiment supported the existence of ALAS isomers in other photosynthetic bacteria. In the present study, two ALAS genes, hemA and hemO, were cloned, sequenced from R. palustris ATCC 17001, and deposited in the GenBank. Two ORFs were predicted, 1,212 and 1,230 bp, encoding deduced polypeptides of 403 and 409 amino acids with estimated MW of 43.7 and 44.6 kDa and theoretical isoelectric points of 6.41 and 6.45, respectively. The two isozymes share 60 % amino acid sequence identity between each other. The HemA and HemO of R. palustris ATCC 17001 showed amino acid sequence identity of 59.6 and 58.4 % to HemA and HemT of R. sphaeroides, respectively. The HemA of R. palustris ATCC 17001 in this study bore only 59.7 % identity with the HemA cloned from R. palustris strain KUGB306 (Choi et al. 2004) wherein the existence of another isozymer was not studied. However, we found all the seven R. palustris strains with genome release in the NCBI database have two ALAS encoding genes and they showed high amino acid sequence identity with the HemA and HemO cloned in this study, in the range of 84.4–96.8 % and 86.3–95.8 % respectively. The highest similarity was R. palustris BisB5, at 100 % amino acid identity. Moreover, we also successfully cloned two ALAS isozymers from another R. palustris strain ATCC 33872 (data not shown). It seems very common for R. palustris to have two ALAS isozymers.
Overexpression and purification of ALAS isomers in E. coli
To characterize the newly found ALAS isomer, HemO, we overexpressed both the hemO gene as well as hemA individually in E. coli Rosetta2 (DE3). Both hemA and hemO genes expressed their proteins in the soluble fraction in large quantities (Fig. 1a). The recombinant ALAS proteins tagged with 6× His were purified from the cell lysate of E. coli under native conditions according to the manufacturer’s recommendation. The purity was above 95 % (w/w) as measured by SDS-PAGE (Fig. 1b).
a SDS-PAGE analysis of soluble total protein of recombinant E. coli Rosetta2 (DE3) expressing hemA or hemO. The expression was induced by addition of 0.1 mM IPTG at 28 °C. Lane M protein marker, Lane 1 overexpression of HemA, lane 2 overexpression of HemO, Lane 3 negative control of Rosetta2 (DE3) containing pET21a(+). b SDS-PAGE analysis of recombinant ALAS proteins purified by Ni–NTA affinity chromatography. Lane M protein marker, Lane 1 purified HemA, lane 2 purified HemO
Characterization of ALASs
Both purified recombinant enzymes were characterized as described in Methods. The specific activities of HemA and HemO were 3.6 and 2.7 U/mg, respectively, which is one of the highest activities ALAS reported, implying its potential application in ALA bioproduction. The K m values of HemA and HemO were 3.8 and 4.7 mM for glycine, respectively, which are higher than the reported ALAS but still in the same range [(1.89 mM for R. sphaeroides, 2 mM for R. palustris KUGB306 and 9.7 mM for Agrobacterium radiobacter (Choi et al. 2004; Bolt et al. 1999; Lin et al. 2009)]. The K m values of both enzymes were 11 and 4.4 μM for succinyl-CoA which are lower than reported for ALAS from the recombinant R. sphaeroides ALAS HemA at 17 (Bolt et al. 1999), 49.6 μM for the recombinant R. palustris KUGB306 ALAS HemA (Choi et al. 2004) and 257 μM for A. radiobacter (Lin et al. 2009). In particular, HemO exhibits a much stronger affinity to succinyl-CoA.
Heme, the end product of tetrapyrroles biosynthesis pathway, inhibits the activity of the first enzyme of the pathway, 5-ALA synthase (Scholnick et al. 1972). In the present study, hemin was used to test its potential effect on the recombinant HemA and HemO. Both enzymes were extremely sensitive to hemin (Fig. 2). Nearly 40 % of the enzymatic activities lost when 2 μM hemin was added into the reaction system. However, the inhibition pattern of HemA and HemO differed from each other significantly when the concentration of hemin further increased. Only 20 % of HemA activity left at 8 μM hemin and above, however, almost 50 % of HemO still remained active even when the concentration of hemin was raised up to 25 μM. The high residual activity of HemO under hemin excess condition should favor the ALA accumulation in biotechnological applications.
ALA production by the recombinant ALAS in E. coli
To demonstrate the biological function of HemA and HemO in vivo and their application potential for ALA production, we constructed plasmids pTrc99A–hemA and pTrc99A–hemO which are compatible with the fast growing wild type E. coli strains. Escherichia coli MG1655 harboring plasmid pTrc99A–hemA or pTrc99A–hemO was cultivated in a 5 l fermentor (see Section “Materials and Methods”). The precursors of the ALAS, succinic acid and glycine, were added into the medium. After 24 h, 6.3 and 5.7 g ALA/l were produced by the respective recombinant strains expressing HemO or HemA (Fig. 3). The experiments were repeated three times. In all cases, the strain expressing HemO produced more ALA than the strain expressing HemA. Interestingly, the strain expressing HemO produced ALA without coupled consumption of succinate between 12 and 16 h, indicating that the succinyl-CoA from glucose metabolism may supply enough precursors for ALA biosynthesis. For the reports in the literature where ALA yield was relatively high (above 5 g/l), commercial strains with rare codon optimization capacity such as Rosetta (DE3) or BL21 were frequently used and a complicated process optimization were applied (Xie et al. 2003; Choi et al. 2008). In this study, we demonstrated the first time that a high yield of ALA can also be achieved with wild type E. coli strains expressing HemO or HemA from R. palustris with a rather simple process control.
ALA production by E. coli MG1655 carrying pTrc99A–hemA (a) or pTrc99A–hemO (b) in fed batch fermentation. Cultivation was performed in a 5 l fermentor with LB medium supplemented with 9 g succinic acid/l. IPTG and glycine were added at 4 h to 0.025 mM and 4 g/l, respectively. Additional glycine was added at 10 h to a final concentration of 2 g/l. Glucose was continuously fed into the fermentor during the fermentation
Conclusions
In this study, two ALAS isomers hemA and hemO were cloned from R. palustris ATCC 17001 and overexpressed in E. coli. Both recombinant ALASs exhibited higher specific activity and stronger affinity to succinyl-CoA than most of the ALAS reported this far. Interestingly, HemO was less inhibited by high-concentration hemin than HemA. HemO out-performed HemA for accumulation of high titer of ALA and readily overexpressed in wild type E. coli strains, providing new source of ALA synthase for rational metabolic engineering. This is the first time that the co-existence of two functional ALAS isozymes in R. palustris was validated and that the biotechnological application of HemO for high level ALA production was experimentally demonstrated.
References
Bolt EL, Kryszak L, Zeilstra-Ryalls J, Shoolingin-Jordan PM, Warren MJ (1999) Characterization of the Rhodobacter sphaeroides 5-aminolaevulinic acid synthase isoenzymes, HemA and HemT, isolated from recombinant Escherichia coli. Eur J Biochem 265(1):290–299
Burnham BF (1970) δ-Aminolevulinic acid synthase (Rhodopseudomonas sphaeroides). Methods Enzymol 17A:195–204
Chinnici F, Spinabelli U, Riponi C, Amati A (2005) Optimization of the determination of organic acids and sugars in fruit juices by ion-exclusion liquid chromatography. J Fd Comp Anal 18:121–130
Choi HP, Hong JW, Rhee KH, Sung HC (2004) Cloning, expression, and characterization of 5-aminolevulinic acid synthase from Rhodopseudomonas palustris KUGB306. FEMS Microbiol Lett 236(2):175–181
Choi HP, Lee YM, Yun CW, Sung HC (2008) Extracellular 5-aminolevulinic acid production by Escherichia coli containing the Rhodopseudomonas palustris KUGB306 hemA gene. J Microbiol Biotechnol 18(6):1136–1140
Heinrikson RL, Meredith SC (1984) Amino acid analysis by reverse-phase high-performance liquid chromatography: precolumn derivatization with phenylisothiocyanate. Anal Biochem 136(1):65–74
Larimer FW, Chain P, Hauser L, Lamerdin J, Malfatti S, Do L, Land ML, Pelletier DA, Beatty JT, Lang AS (2003) Complete genome sequence of the metabolically versatile photosynthetic bacterium Rhodopseudomonas palustris. Nat Biotechnol 22(1):55–61
Lee DH, Jun WJ, Yoon JW, Cho HY, Hong BS (2004) Process strategies to enhance the production of 5-aminolevulinic acid with recombinant E. coli. J Microbiol Biotechnol 14(6):1310–1317
Lee DH, Jun WJ, Shin DH, Cho HY, Hong BS (2005) Effect of culture conditions on production of 5-aminolevulinic acid by recombinant Escherichia coli. Biosci Biotechnol Biochem 69(3):470–476
Lin J, Fu W, Cen P (2009) Characterization of 5-aminolevulinate synthase from Agrobacterium radiobacter, screening new inhibitors for 5-aminolevulinate dehydratase from Escherichia coli and their potential use for high 5-aminolevulinate production. Bioresour Technol 100(7):2293–2297
Mauzerall D, Granick S (1956) The occurrence and determination of δ-aminolevulinic acid and porphobilinogen in urine. J Biol Chem 219:435–446
Nishikawa S, Tanaka T, Kaminaga T, Watanabe DKWCW, Kikuo LH, Miyachi N, Watanabe K, Hotta Y (2002) Microorganisms producing 5-aminolevulinic acid and processes for producing 5-aminolevulinic acid by using the same. US Patent 6342377
Scholnick PL, Hammaker LE, Marver HS (1972) Soluble-aminolevulinic acid synthetase of rat liver. II. Studies related to the mechanism of enzyme action and hemin inhibition. J Biol Chem 247(13):4132–4137
van der Werf MJ, Zeikus JG (1996) 5-Aminolevulinate production by Escherichia coli containing the Rhodobacter sphaeroides hemA gene. Appl Environ Microbiol 62(10):3560–3566
Xie L, Hall D, Eiteman MA, Altman E (2003) Optimization of recombinant aminolevulinate synthase production in Escherichia coli using factorial design. Appl Microbiol Biotechnol 63(3):267–273
Acknowledgments
This work was supported by the Knowledge Innovation Program of the Chinese Academy of Sciences (Grant No. KSCX2-E-W-Q-13), Tianjin Research Program of Application Foundation and Advanced Technology (No. 10JCZDJC16200), National Natural Science Foundation of China (No. 31070037), and Key Technologies Research and Development Program of Tianjin (No. 11ZCZDSY08500).
Author information
Authors and Affiliations
Corresponding author
Additional information
Lilu Zhang, Jiuzhou Chen dedicated equally to this manuscript.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, L., Chen, J., Chen, N. et al. Cloning of two 5-aminolevulinic acid synthase isozymes HemA and HemO from Rhodopseudomonas palustris with favorable characteristics for 5-aminolevulinic acid production. Biotechnol Lett 35, 763–768 (2013). https://doi.org/10.1007/s10529-013-1143-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-013-1143-4