Abstract
Common buckwheat (Fagopyrum esculentum Moench) is rich in phenolic compounds and may be useful for the treatment of metabolic syndrome in humans. To improve the production of rutin in buckwheat, we overexpressed the flavonol-specific transcription factor, AtMYB12 using Agrobacterium rhizogenes into hairy root culture systems. This induced the expression of flavonoid biosynthetic genes encoding phenylalanine ammonia lyase, cinnamate 4-hydroxylase, 4-coumarate:CoA ligase, chalcone synthase, chalcone isomerase, flavone 3-hydroxylase, flavonoid 3′-hydroxylase, and flavonol synthase. This led to the accumulation of rutin in buckwheat hairy roots up to 0.9 mg/g dry wt. PAP1 expression, however, did not correlate with the production of rutin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many plants contain various antioxidant components, such as phenolic compounds, with flavonoids being the most common and widely distributed group of plant phenolics. Apart from being rich in phenolic compounds, the seeds of common buckwheat (Fagopyrum esculentum Moench), in particular, are a good source of rutin (quercetin-3-rutinoside)—a flavonol glycoside that protects plants from ultraviolet radiation and diseases (Hinneburg and Neubert 2004; Kalinova et al. 2006). Rutin is an antioxidant with many pharmacologically properties in humans, such as anti-inflammatory, anticarcinogenic, antithrombotic, cytoprotective, and vasoprotective effects (Li and Zhang 2001; Kreft et al. 2002). Rutin possesses health-promoting effects, such as attenuating the metabolic syndrome induced by high-carbohydrate/high-fat diets (Panchal et al. 2011). Buckwheat could, thus, be an important industrial source of rutin; a product that should be considered for the treatment of metabolic syndrome in humans.
Various strategies can be employed to increase the production of polyphenolic compounds. For example, overexpression of genes encoding individual biosynthetic enzymes can lead to an increase in the levels of a target compound (Park et al. 2011). This method, however, cannot induce the entire biosynthetic pathway, and, although there is an increase in the amount of end products, this is not enough for the purpose of practical application. An alternative method could use the regulatory genes that directly induce these pathways. For example, members of the MYB transcription factor family are involved in secondary metabolism. AtMYB75/PAP1 and AtMYB90/PAP2 control biosynthesis in vegetative tissues (Borevitz et al. 2000; Gonzalez et al. 2008), while AtMYB123 controls the biosynthesis of proanthocyanidins in the seed coat of Arabidopsis (Lepiniec et al. 2006). Moreover, AtMYB12 is a flavonol-specific activator of flavonoid biosynthesis and a transcriptional regulator of chalcone synthase (CHS) and flavonol synthase (FLS), which are indispensible for the biosynthesis of flavonol (Mehrtens et al. 2005). AtMYB12 could also activate the caffeoylquinic acid (CQA) biosynthetic pathway, when expressed in a tissue-specific manner in tomato (Luo et al. 2008).
Because Luo et al. (2008) indicated that transcription factors might have different specificities for target genes in different plants, we hypothesized that it might be possible to accumulate high levels of rutin in buckwheat, through the overexpression of AtMYB12 and AtMYB75/PAP1. In this study, the hairy root culture system was used to observe the activity of transcription factors. Using transgenic buckwheat hairy roots, we show that flavonoid biosynthetic genes are, indeed, regulated by transcription factors.
Materials and methods
Seed sterilization and germination
De-hulled seeds from common buckwheat were surface-sterilized with 70% (v/v) ethanol for 1 min and 4% (v/v) sodium hypochlorite solution for 10 min. They were then rinsed thrice in sterilized water. Six seeds were placed on 25 ml agar-solidified culture medium in Petri dishes (100 × 15 mm2). The basal medium consisted of MS medium (Murashige and Skoog 1962), solidified with 0.8% (w/v) agar. Prior to addition of agar, the medium was adjusted to pH 5.8, and the complete medium was then sterilized by autoclaving at 121°C for 20 min. The seeds germinated in a growth chamber at 25°C under standard, cool, white fluorescent tubes with a flux rate of 35 μmol/(sm2) and a 16 h photoperiod.
Total RNA extraction and quantification of gene expression
Total RNA was isolated from different lines of transgenic buckwheat hairy roots using the RNeasy Plant Mini Kit (Qiagen; Valencia, CA, USA). For qRT-PCR, 1 μg total RNA was reverse-transcribed using the Superscript II First Strand Synthesis Kit (Invitrogen; Carlsbad, CA, USA) and oligo (dT)20 primers.
Transcription levels were analyzed by real-time PCR. The gene-specific primer sets were designed for real-time PCR, as previously described (Li et al. 2010) (Supplementary Table 1). Gene expression was normalized to that of the histone H3 gene, which, in this case, was used as a housekeeping gene (Timotijevic et al. 2010). Real-time PCR was performed in triplicate on a MiniOpticon system (Bio-Rad Laboratories; Hercules, CA) with the Quantitect SYBR Green PCR Kit (Qiagen). The PCR protocol was as follows: denaturation for 5 min at 95°C, followed by 40 cycles of denaturation for 15 s at 95°C, annealing for 15 s at 56°C, and elongation for 20 s at 72°C. The results of PCR were calculated as the mean of 3 replicate treatments. Statistical differences between treatments were evaluated by calculating the standard deviation.
Plasmid construction for the transformation of buckwheat hairy roots
The PAP1 and AtMYB12 overexpression constructs consisted of a cauliflower mosaic virus (CaMV) 35S promoter, the respective gene sequences, and the bar (bialaphos resistance) gene as a selectable marker. PCR was used to amplify the full-length sequences of PAP1 and AtMYB12, which were previously published (Borevitz et al. 2000; Mehrtens et al. 2005). The PCR primers and cloning vectors used for these experiments are shown in Supplementary Table 1. The recombinant genes were PCR amplified with gene specific primer sets, according to the Gateway cloning strategy (Karimi et al. 2002). After a second round of PCR using adapter primer sets, each amplified product was subcloned into the pDONR221 vector (Invitrogen) by recombination. These vectors were then transformed into the E. coli strain TOP10 (Invitrogen), and clones were selected with 50 mg/l kanamycin. Finally, the inserts were shuttled from the entry vectors into the pB7FWG2 binary destination vector (obtained from the Functional Genomics unit of the Department of Plant System Biology; VIB-Ghent University) using Gateway LR Clonase II Plus Enzyme Mix (Invitrogen). The clones were transformed in the E. coli strain TOP10 (Invitrogen) and selected with 50 mg spectinomycin/l.
Transformation of buckwheat hairy roots
The resulting binary vectors, pPAP1 and pAtMYB12, were transformed in the Agrobacterium rhizogenes strain, R1000. A β-glucuronidase overexpression vector (pGUS) was used for the control hairy roots. Young stems of common buckwheat were collected from the plants grown in vitro and cut at the ends into 7 mm sections. The excised stems were dipped into the A. rhizogenes R1000 cultures in liquid inoculation medium for 10 min, blotted dry on sterile filter paper, and incubated in the dark at 25°C on agar-solidified MS medium. After 2 days of co-cultivation, the explanted tissues were transferred to a hormone-free medium containing MS salts and vitamins (0.5 mg nicotinic acid/l, 0.5 mg pyridoxine–HCl/l, 0.1 mg thiamine-HCl/l, and 2 mg glycine/l), 30 g sucrose/l, 500 mg cefotaxime/l, 1 mg glufosinate/l and 8 g agar/l. Numerous hairy roots were observed emerging from the wound sites within 2 weeks. The hairy roots were separated from the explanted tissues and subcultured in the dark at 25°C on agar-solidified MS medium. After repeated transfer to fresh medium, rapidly growing hairy root cultures were obtained. Isolated roots (0.5 g/l) were transferred to 30 ml MS liquid medium, containing 30 g sucrose/l, in 100 ml flasks. The root cultures were maintained at 25°C on a shaker (100 rev/min) in a growth chamber under standard, cool, white fluorescent tubes with a flux rate of 35 μmol/(s/m2) and a 16 h photoperiod. After 30 days, hairy roots were harvested and their dry weight and phenolic compound content was determined. Three flasks were used for every culture condition, and the experiments were performed in duplicate.
Quantitative analysis of rutin using HPLC
Rutin was determined by HPLC. The extract was filtered through a 0.45 μm poly-filter, diluted twofold with methanol. Analysis used a C18 column (250 × 4.6 mm2, 5 μm) at 30°C with a gradient of 0.15% acetic acid and methanol as elutant at 1 ml/min. Detection was at 280 nm. The results were quantified using rutin from Sigma as standard.
Results
Establishment of transgenic hairy root lines
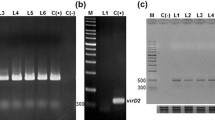
Explants of buckwheat stems were infected with A. rhizogenes strain R1000 harboring the pPAP1, pAtMYB12, or pGUS (β-glucuronidase, for control hairy roots) binary vector. The transgenic hairy root cultures were examined by PCR, with primers for the bar (bialaphos resistance) gene and the rol genes (data not shown). The rol genes of the Ri-plasmid in A. rhizogenes are responsible for the induction of hairy roots from plant species. By using qRT-PCR to measure the expression levels of PAP1 or AtMYB12 in the transgenic hairy root lines and GUS-control hairy root lines, 6 PAP1-overexpressing (P) and 6 AtMYB12-overexpressing (M) hairy root lines were identified. PAP1-overexpressing hairy root lines were designated as P-1, 3, 4, 6, 7, and 8, while AtMYB12-overexpressing (M) hairy root lines were designated as M-1, 4, 7, 8, 10, and 14 (Figs. 1a, 2a).
Analysis of gene expression in buckwheat hairy root lines overexpressing AtMYB12. a The expression level of AtMYB12 relative to H3 in buckwheat hairy roots overexpressing AtMYB12 was confirmed by quantitative real-time PCR. b The expression level of flavonoid biosynthetic genes (FePAL, FeC4H, Fe4CL, FeCHS, FeCHI, FeF3H, FeF3′H, FeFLS, FeDFR, and FeANS) relative to H3 in buckwheat hairy roots overexpressing AtMYB12 was confirmed by quantitative real-time PCR. The top ends of the bars and the error bars indicate the mean and standard deviation (n = 3), respectively. G a control hairy root line transformed with the β-glucuronidase (GUS) overexpression vector pGUS; M-n transgenic hairy root lines (“n” indicates the line number) transformed with the AtMYB12 overexpression vector, pAtMYB12
Analysis of gene expression in buckwheat hairy root lines overexpressing PAP1. a The expression level of PAP1 relative to H3 in buckwheat hairy roots overexpressing PAP1 was confirmed by quantitative real-time PCR. b The expression level of flavonoid biosynthetic genes (FePAL, FeC4H, Fe4CL, FeCHS, FeCHI, FeF3H, FeF3′H, FeFLS, FeDFR, and FeANS) relative to H3 in buckwheat hairy roots overexpressing PAP1 was confirmed by quantitative real-time PCR. The top ends of the bars and the error bars indicate the mean and standard deviation (n = 3), respectively. G a control hairy root line transformed with the β-glucuronidase (GUS) overexpression vector pGUS; P-n transgenic hairy root lines (“n” indicates the line number) transformed with the PAP1 overexpression vector, pPAP1
Effect of AtMYB12 on the expression levels of flavonoid biosynthetic genes
To analyze the effect of AtMYB12 on the expression of flavonoid biosynthesis genes in buckwheat hairy roots, real-time qRT-PCR analyses were carried out. Figures 1b and 2b depict the results of the qRT-PCR analyses. AtMYB12 was strongly overexpressed in AtMYB12-overexpressing hairy root lines with the relative expression levels ranging from 35 to 60-fold more than that of GUS-overexpressing hairy root lines. The flavonoid biosynthetic genes encoding phenylalanine ammonia lyase (PAL), cinnamate 4-hydroxylase (C4H), 4-coumarate:CoA ligase (4CL) 1 and 2, CHS, chalcone isomerase (CHI), flavone 3-hydroxylase (F3H), flavonoid 3′-hydroxylase (F3′H) and flavonol synthase (FLS) 1 and 2 showed a correlation to the AtMYB12 expression level. In general, the expression level of flavonoid biosynthetic genes (FePAL, FeC4H, Fe4CL1, Fe4CL2, FeCHS, FeCHI, FeF3H, FeF3′H, FeFLS1, and FeFLS2) increased in response to elevated AtMYB12 expression. However, the expression level of FeDFR and FeANS was more or less unaffected by changes in AtMYB12 expression; FeDFR expression increased slightly in AtMYB12-overexpressing hairy root lines.
We also analyzed the effect of PAP1 on the expression of flavonoid biosynthesis genes in buckwheat hairy roots. qRT-PCR analysis showed that PAP1 was overexpressed in PAP1-overexpressing hairy root lines, with the relative expression levels ranging from 50 to 100-fold more than that of GUS-overexpressing hairy root lines. However, the flavonoid biosynthetic genes did not seem to be correlated to PAP1 expression level. The expression level of all flavonoid biosynthetic genes analyzed in this study was unchanged in PAP1-overexpressing hairy root lines.
Effect of AtMYB12 on rutin levels
To investigate the effect of AtMYB12 and PAP1 on the levels of rutin, the AtMYB12- and PAP1-overexpressing hairy roots were analyzed by HPLC (Fig. 3). High levels of rutin were detected in the hairy roots lines expressing AtMYB12, compared to the control hairy roots expressing GUS. AtMYB12-overexpressing hairy root lines produced 2.2–3.7 times more rutin (0.53–0.88 μg/mg) than the GUS-control hairy root line. In contrast, no significant difference between the control and PAP1-overexpressing hairy root lines could be detected by HPLC analysis.
Rutin production in buckwheat hairy root lines overexpressing AtMYB12 and PAP1. Production of rutin in a AtMYB12- and b PAP1-overexpressing hairy root lines was analyzed by HPLC. The top ends of the bars and the error bars indicate the mean and standard deviation (n = 3), respectively. G a control hairy root line transformed with the β-glucuronidase (GUS) overexpression vector pGUS; M-n and P-n transgenic hairy root lines (“n” indicates the line number) transformed with the AtMYB12- and PAP1-overexpression vectors pAtMYB12 and pPAP1, respectively
Discussion
The MYB transcription factor family is widely distributed in higher plants. MYB family members have been found to be involved in diverse physiological and biochemical processes, including the control of cell morphogenesis and response to environmental stress and phytochromes, as well as the regulation of secondary metabolism (Du et al. 2009).
This study demonstrates that the expression of the transcription factor AtMYB12, together with AtMYB75/PAP1 expression, enhances the levels of rutin in buckwheat hairy roots. AtMYB12 was originally identified in Arabidopsis, where its overexpression resulted in increased accumulation of flavonol (Mehrtens et al. 2005). AtMYB12 induced gene expression in both flavonol and CQA biosynthetic pathways in tomato as well as in the flavonoid biosynthetic pathway in tobacco. In the tomato fruit, in particular, high levels of flavonol and CQA accumulated as a result of AtMYB12 overexpression (Luo et al. 2008). In our study, AtMYB12 induced the expression of flavonoid biosynthetic genes such as the PAL gene, which was not a target of AtMYB12 in Arabidopsis (Mehrtens et al. 2005), but was an AtMYB12 target in both tobacco and tomato (Luo et al. 2008). In PAP1-overexpressing buckwheat hairy roots, rutin levels and gene expression in the flavonoid biosynthetic pathway did not increase by PAP1 expression. PAP1 was reported to be an anthocyanin activator in Arabidopsis, and its overexpression upregulated many transcripts involved in the formation of anthocyanins (Borevitz et al. 2000; Tohge et al. 2005).
Although AtMYB12 resulted in significant enhancement of rutin levels in transgenic buckwheat hairy roots, more effective engineered metabolism is needed to identify transcription factors from buckwheat, and to ensure their appropriate specificity in this plant. We are now working to determine and characterize such transcription factors from buckwheat, with the ultimate aim to be able to increase rutin production in buckwheat.
References
Borevitz JO, Xia Y, Blount J, Dixon RA, Lamb C (2000) Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12:2383–2394
Du H, Zhang L, Liu L, Tang X-F, Yang W-J, Wu Y-M, Huang Y-B, Tang Y-X (2009) Biochemical and molecular characterization of plant MYB transcription factor family. Biochemistry 74:1–11
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53:814–827
Hinneburg I, Neubert RHH (2004) Influence of extraction parameters on the phytochemical characteristics of extracts from buckwheat (Fagopyrum esculentum) herb. J Agric Food Chem 53:3–7
Kalinova J, Triska J, Vrchotova N (2006) Distribution of vitamin E, squalene, epicatechin, and rutin in common buckwheat plants (Fagopyrum esculentum Moench). J Agric Food Chem 54:5330–5335
Karimi M, Inzé D, Depicker A (2002) GATEWAY™ vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7:193–195
Kreft S, Strukelj B, Gaberscik A, Kreft I (2002) Rutin in buckwheat herbs grown at different UV-B radiation levels: comparison of two UV spectrophotometric and an HPLC method. J Exp Bot 53:1801–1804
Lepiniec L, Debeaujon I, Routaboul J-M, Baudry A, Pourcel L, Nesi N, Caboche M (2006) Genetics and biochemistry of seed flavonoids. Annu Rev Plant Biol 57:405–430
Li S-q, Zhang QH (2001) Advances in the development of functional foods from buckwheat. Crit Rev Food Sci Nutr 41:451–464
Li X, Park NI, Xu H, Woo S-H, Park CH, Park SU (2010) Differential expression of flavonoid biosynthesis genes and accumulation of phenolic compounds in common buckwheat (Fagopyrum esculentum). J Agric Food Chem 58:12176–12181
Luo J, Butelli E, Hill L, Parr A, Niggeweg R, Bailey P, Weisshaar B, Martin C (2008) AtMYB12 regulates caffeoyl quinic acid and flavonol synthesis in tomato: expression in fruit results in very high levels of both types of polyphenol. Plant J 56:316–326
Mehrtens F, Kranz H, Bednarek P, Weisshaar B (2005) The arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol 138:1083–1096
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 15:473–497
Panchal SK, Poudyal H, Arumugam TV, Brown L (2011) Rutin attenuates metabolic changes, nonalcoholic steatohepatitis, and cardiovascular remodeling in high-carbohydrate, high-fat diet-fed rats. J Nutr 141:1062–1069
Park N, Xu H, Li X, Kim S-J, Park S (2011) Enhancement of flavone levels through overexpression of chalcone isomerase in hairy root cultures of Scutellaria baicalensis. Funct Integr Genomics 11:491–496
Timotijevic GS, Milisavljevic MD, Radovic SR, Konstantinovic MM, Maksimovic VR (2010) Ubiquitous aspartic proteinase as an actor in the stress response in buckwheat. J Plant Physiol 167:61–68
Tohge T, Nishiyama Y, Hirai MY, Yano M, Nakajima J-i, Awazuhara M, Inoue E, Takahashi H, Goodenowe DB, Kitayama M, Noji M, Yamazaki M, Saito K (2005) Functional genomics by integrated analysis of metabolome and transcriptome of Arabidopsis plants over-expressing an MYB transcription factor. Plant J 42:218–235
Acknowledgments
This study was supported by a grant of the Korea Healthcare technology R&D Project, Ministry for Health, Welfare & Family Affairs, Republic of Korea (A091220).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Park, N.I., Li, X., Thwe, A.A. et al. Enhancement of rutin in Fagopyrum esculentum hairy root cultures by the Arabidopsis transcription factor AtMYB12. Biotechnol Lett 34, 577–583 (2012). https://doi.org/10.1007/s10529-011-0807-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-011-0807-1