Abstract
Cellobiose dehydrogenase (CDH; EC 1.1.99.18) is an extracellular glycosylated protein composed of two distinct domains, a C-terminal catalytic flavin domain and an N-terminal cytochrome-b-type heme domain, which transfers electrons from the flavin domain to external electron acceptors. The soluble flavin domain of the Phanerochaete chrysosporium CDH was successfully expressed in Escherichia coli. The enzyme showed dye-mediated CDH activity higher than that of the complete CDH, composed of flavin domain and heme domain, prepared using Pichia pastoris as the host microorganism. The ability to conveniently express the recombinant CDH flavin domain in E. coli provides great opportunities for the molecular engineering of the catalytic properties of CDH.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cellobiose dehydrogenases (CDHs; EC 1.1.99.18) are extracellular, glycosylated proteins produced by numerous fungi, including the white rot fungi (Cameron and Aust 2001). CDHs are involved in a cellulolytic enzyme pathway of these fungi, although their exact physiological role remains unclear. CDHs catalyze the oxidation of cellobiose (Glc-β-1,4-Glc) and other β-1,4-linked disaccharides or oligosaccharides at the C-1 position to the corresponding lactones (Hallberg et al. 2002). CDHs have recently been paid much attention as anodic catalysts in bio-fuel cells, capable of oxidizing cellulolytic sugars. Enzyme sensors employing CDHs have also been reported (Hilden et al. 2001) for the evaluation of cellulase activities.
CDHs have a unique enzyme structure, composed of two distinct domains: a C-terminal catalytic flavin domain and an N-terminal cytochrome-b-type heme domain, that transfers electrons from the flavin domain to external electron acceptors. The CDH from the white rot fungus, Phanerochaete chrysosporium, is the most extensively studied, including elucidation of the three dimensional crystal structures of each of its two domains, prepared by papain proteolysis (Hallberg et al. 2002). We have recently reported the alteration of substrate specificity by site-directed mutagenesis on the P. chrysosporium CDH recombinantly expressed in Pichia pastoris (Desriani et al. 2010).
Considering that CDHs are glycosylated enzymes with a unique structure, the recombinant production of CDHs have been achieved by using budding yeast, such as P. pastoris (Yoshida et al. 2001; Desriani et al. 2010). However, the recombinant production of CDHs employing Escherichia coli or another prokaryotic host has yet to be reported. The ability to recombinantly produce CDH in a prokaryotic host would accelerate the protein engineering study of CDH, such as the modification of its catalytic properties, which is eagerly awaited by those engaged in CDH applications. In this study, we demonstrate the recombinant production of a functionally active CDH flavin domain from P. chrysosporium, using E. coli as the host. The CDH flavin domain was successfully produced under specific growth conditions as a soluble protein showing dye-mediated CDH activity.
Materials and methods
pET30c/flavin-cdh construction
The structural gene of the Phanerochaete chrysosporium CDH flavin domain was PCR-amplified using as template pPIC9cdh, a vector previously created to express the entire CDH in P. pastoris (Desriani et al. 2010). Because the DNA encoding the flavin domain contained NdeI and NcoI site restriction sites, a ribosome binding site flanked by XbaI and NdeI restriction sites was incorporated into the forward primer (5′-CCCCCTCTAGAAGGAGATATACATATGACACCTTACGATTACATCATC-3′) to facilitate insertion into the pET30c expression vector. An EcoRI restriction site was incorporated into the reverse primer (5′-GGGAATTCAAGGACCTCCCGCAAGCG-3′) downstream of the domain’s stop codon. The flavin domain expression vector pET30c/flavin-cdh was created by digesting the PCR product with XbaI and EcoRI and inserting it into pET30c.
CDH flavin domain preparation
Escherichia coli BL21(DE3) was transformed with pET30c/flavin-cdh and the CDH flavin domain expressed by autoinduction in PA-5052 growth medium, containing 50 mM Na2HPO4, 50 mM KH2PO4, 25 mM (NH4)2SO4, 2 mM MgSO4, trace metals, 0.5% glycerol, 0.05% glucose, 0.2% α-lactose, and 200 mg/l of each amino acid except cysteine and tyrosine (Studier 2005). Cells were collected by centrifugation and resuspended in 50 mM ammonium acetate buffer, pH 5. The cells were disrupted by ultrasonication and centrifuged. The resulting crude extract was fractionated by ammonium sulfate precipitation, and the fraction precipitated from 20 to 60% saturation was suspended in 20 mM Tris/HCl, pH 8, and desalted on a PD-10 column (GE healthcare). The protein sample was then applied on to a 6 ml Resource Q column equilibrated with 20 mM Tris/HCl, pH 8, and eluted with a linear NaCl gradient in the same buffer. The eluted enzyme then underwent size-exclusion chromatography on a Superdex 200 10/300 (GE healthcare) using 20 mM Tris/HCl, pH 8. The prepared CDH flavin domain was then used for kinetic studies.
Characterization of enzymatic properties
The recombinant enzyme activity was determined at room temperature using 0.1 mM 2,6-dichloroindophenol (DCIP) in 50 mM citric acid buffer, pH 5, in the presence of different concentrations of cellobiose as the substrate (Henriksson et al. 1998). The decrease in absorption of DCIP was monitored at 525 nm using a molar absorption coefficient of 8.25 M−1 cm−1.
The effect of pH on the activity and stability of the enzyme was investigated at room temperature in the presence of 5 mM cellobiose using 50 mM sodium citrate buffer in the pH range 3–6 and 50 mM potassium phosphate buffer in the pH range 6–7. The following DCIP molar absorption coefficients were determined experimentally in the appropriate buffers: 8.21 M−1 cm−1 (pH 3), 7.98 M−1 cm−1 (pH 4), 8.25 M−1 cm−1 (pH 5), 8.25 M−1 cm−1 (pH 6), and 9.56 M−1 cm−1 (pH 7).
Results
Recombinant production of CDH flavin domain
The expression of the CDH flavin domain in E. coli was first attempted using IPTG induction at 30, 25, and 20°C. However, no CDH activity was detected from cell free extracts of E. coli harboring pET30c/flavin-cdh under any of the induction conditions investigated. With the aim of producing a functional CDH flavin domain, we searched for milder expression conditions. Studier (2005) reported on several media compositions that facilitate the mild expression under the T7lac promoter (bacteriophage T7 promoter and the lac operator) with continuous cell growth, resulting in high yields of soluble active protein. Dye-mediated CDH activity was detected in cell-free extracts of E. coli grown at different temperatures in PA 5052 medium, an autoinducing minimal medium with amino acids. The observed CDH activity levels in cell-free extracts at each temperature were: 0.14 U/mg (326 U/l) at 20°C, 0.32 U/mg (733 U/l) at 25°C, and 0.08 U/mg (126 U/l) at 30°C. A protein band was observed on SDS-PAGE near the expected molecular weight of 57 kDa at all three incubation temperatures in both the insoluble (Figs. 1, 2, lanes 3 and 5) and soluble fractions (Fig. 2, lanes 4 and 6). The most intense band corresponding to soluble CDH flavin domain was observed in the sample cultivated at 25°C, which is consistent with the observed activity in cell-free extracts.
SDS-PAGE of E. coli BL21(DE3) expressing pET30c/flavin-CDH. Cells grown at 20, 25, or 30°C were disrupted by sonication and the resulting lysate separated by centrifugation into insoluble (lanes 1, 3, 5) and soluble (lanes 2, 4, 6) fractions. Samples were separated by 10% polyacrylamide SDS-PAGE and stained with Coomassie Brilliant Blue R-250. The bands corresponding to the CDH flavin domain are marked with an arrowhead
Characterization of recombinant CDH flavin domain
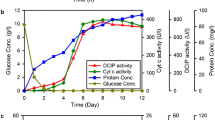
For further enzymatic characterization, the recombinant CDH flavin domain was expressed at 25°C in PA 5052 autoinduction medium and purified. Based on the activity measured in crude extract, the expression level of the CDH flavin domain in E. coli was calculated as 386 U/l culture after 25 h. After the first size-exclusion chromatography, the enzyme was purified to a specific activity of 16.7 U/mg protein at a yield of 7.3% (Table 1).
The dye-mediated CDH activity of the purified flavin domain showed a cellobiose concentration-dependence and reached saturation around 5 mM cellobiose (Fig. 3). The Vmax value calculated from these measurements is 12.2 U/mg, which is higher than that previously observed with the purified complete CDH, composed of flavin domain and heme domain, expressed in P. pastoris (Desriani et al. in press). The Km for the flavin domain was calculated to be 0.07 mM, almost identical to that of the CDH expressed in P. pastoris. Investigation of the flavin domain’s pH dependence showed that the highest dye-mediated CDH activity was achieved at pH 4, with barely detectable activity at pH 7.
Discussion
CDH is composed of a C-terminal flavin domain and an N-terminal cytochrome-b-type heme domain. The flavin domain of CDH is an FAD-binding catalytic domain, whose structure belongs to the glucose/methanol/choline (GMC) oxidoreductase family (Zamocky et al. 2004). In prokaryotes, FAD-harboring redox enzymes, including GMC oxidoreductases, are produced and folded in the cytosol. In Gram-negative bacteria, some FAD-harboring redox enzymes are secreted into the periplasmic space in the fully folded form via the TAT secretion system, which requires a specific signal sequence (Sargent 2007). E. coli endogenously produces cytochrome b-562 (cyt-b562) with a Sec secretional signal sequence in an unfolded form. After cleavage of the signal sequence and transportation by the Sec secretional system into periplasmic space, cyt-b562 is folded with the incorporation of heme to form mature cyt-b562 (Nikilla et al. 1991). Considering the differences in post translational modification of each domain renders impossible the recombinant production of the complete and functional CDH in E. coli, we attempted to produce the CDH flavin domain alone in E. coli without the heme domain.
Conventional over-expressing induction conditions for the T7lac promoter system with IPTG yielded only an insoluble inactive recombinant product. Because native CDH is a glycosylated protein, the milder conditions provided by the autoinduction method appear to be necessary to avoid the formation of inclusion bodies when expressed in the nonglycosylated form. Native CDH has three glycosylation sites, two in the flavin domain and one in the heme domain, which may partially interfere with the access of the artificial electron acceptor to the flavin cofactor, whose normal function is to transfer electrons to the heme domain. The absence of such sugar moieties in the E. coli-expressed CDH flavin domain may explain the higher dye-mediated dehydrogenase activity compared with the P. pastoris-expressed CDH.
The pH optimum for the native CDH was previously reported to be pH 6 for the dye-mediated activity, and pH 4–4.5 for the direct electron transfer, via the heme domain (Tasca et al. 2009). Because the two domains were assumed to be oppositely charged at pH 4–4.5, the electrostatic attractions between to two domains were thought to favor the internal electron transfer by minimizing the electron transfer distance. However, electrostatic repulsion between the deprotonated surfaces of the two domains at pH 6 favors the dye-mediated electron transfer. Our results clearly demonstrated that the CDH flavin domain showed the highest activity at pH 4 (Fig. 4). It therefore appears that the reported pH 6 optimum for the dye-mediated reaction of the native CDH is mainly due to the dissociation of the flavin domain and heme domain, rather than the optimal catalytic efficiency of the flavin domain.
pH-dependence of the dye-mediated activity of the CDH flavin domain expressed in E. coli. The effect of pH on the enzyme activity was investigated at room temperature in the presence of 5 mM cellobiose using 50 mM sodium citrate buffer in the pH range 3–6 (black squares) and 50 mM potassium phosphate buffer in the pH range 6–7 (white circles)
Conclusions
As current mutagenesis studies of CDH are only achievable employing eukaryotic host microorganisms, such as P. pastoris (Desriani et al. in press), the routine screening of mutated enzymes such as directed evolutional approaches are difficult. This is the first study demonstrating the recombinant production of soluble and active CDH flavin domain in a prokaryote. Our achievement provides a simple method to prepare engineered or randomly mutated CDH flavin domain for routine screening, which is likely to contribute to further engineering studies of CDH.
References
Cameron MD, Aust SD (2001) Cellobiose dehydrogenase an extracellular fungal flavocytochrome. Enzym Microb Technol 28:129–138
Desriani, Ferri S, Sode K (2010) Amino acid substitution at the substrate binding subsite alters the specificity of the Phanerochaete chrysosporium cellobiose dehydrogenase. Biochem Biophy Res Commun (in press)
Hallberg BM, Henriksson G, Pettersson G, Divne C (2002) Crystal structure of the flavoprotein domain of the extracellular flavocytochrome cellobiose dehydrogenase. J Mol Biol 315:421–434
Henriksson G, Pettersson G, Johansson G, Ruiz A, Uzcategui E (1991) Cellobiose oxidase from Phanerochaete chrysosporium can be cleaved by papain into two domains. Eur J Biochem 196:101–106
Henriksson G, Sild V, Szabo IJ, Pettersson G, Johansson G (1998) Substrate specificity of cellobiose dehydrogenase from Phanerochaete chrysosporium. Biochim Biophys Acta 1383:48–54
Hilden L, Eng L, Johansson G, Lindqvist SE, Pettersson G (2001) An amperometric cellobiose dehydrogenase based biosensor can be used for measurement of cellulose activity. Anal Biochem 290:245–250
Larsson T, Lindgren A, Ruzgas T, Linquist SE, Gorton L (2000) Bioelectrochemical characterization of cellobiose dehydrogenase modified graphite electrodes: ionic strength and pH dependences. J Electroanal Chem 482:1–10
Nikilla H, Gennis RB, Sligar SG (1991) Cloning and expression of the gene encoding the soluble cytochrome b562 of Escherichia coli. Eur J Biochem 202:309–313
Rostaert FAJ, Renganathan V, Gold MH (2003) Role of the flavin domain residues, his689 and asn732, in the catalytic mechanism of cellobiose dehydrogenase from Phanerochaete chrysosporium. Biochemistry 42:4049–4056
Sargent F (2007) The twin-arginine transport system: moving folded proteins across membranes. Biochem Soc Trans 35:835–847
Studier FW (2005) Protein production by auto-induction in high density shaking cultures. Protein Expr Purif 41:207–234
Tasca F, Gorton L, Harreither W, Haltrich D, Ludwig R, Noll G (2009) Comparison of direct and mediated electron transfer for cellobiose dehydrogenase from Phanerochaete chrysosporium. Anal Chem 81:2791–2798
Yoshida M, Ohira T, Igarashi K, Nagasawa H, Aida K, Hallberg BM, Divne C, Nishino T, Samejima M (2001) Production and characterization of recombinant Phanerochaete chrysosporium cellobiose dehydrogenase in the methylotrophic yeast Pichia pastoris. Biosci Biotechnol Biochem 65:2050–2057
Zamocky M, Hallber M, Ludwig R, Divne C, Haltrich D (2004) Ancestral gene fusion in cellobiose dehydrogenase a specific evolution of GMC oxidoreductase in fungi. Gene 338:1–14
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Desriani, Ferri, S. & Sode, K. Functional expression of Phanerochaete chrysosporium cellobiose dehydrogenase flavin domain in Escherichia coli . Biotechnol Lett 32, 855–859 (2010). https://doi.org/10.1007/s10529-010-0215-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-010-0215-y