Abstract
DNA shuffling was carried out with two chitosanase genes belonging to glycoside hydrolase family eight from Bacillus cereus KNUC51 and B. cereus KNUC55. The shuffled products, YM18 and YM20, which showed higher activity than the parents at 40°C, were selected for further studies. The 50 kDa chitosanases were purified using affinity chromatography with glutathione-Sepharose 4B. In general, the specific activity of YM18 is enhanced 250% and that of YM20 is 350% compared to the parents. YM20 exhibits a shift of the optimal pH level from 5.5 to 6.5. DNA sequence analysis revealed that YM18 and YM20 contained 2 amino acid substitutions (I13T and A87V for YM18; K66R and N352S for YM20). We presumed that these amino acid substitutions increase the specific activity and change the property of the two variants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chitosan is a linear polysaccharide consisting of d-glucosamine and N-acetyl-d-glucosamine residues. It is widely used in food, agricultural, and medical agents. In particular, chitooligosaccharide has attracted attention as a useful agent in the food and pharmaceutical fields due to its versatile antitumor, antifungal, and antimicrobial activity.
Chitosanase (E.C.3.2.1.132) catalyzes the endohydrolysis of β-1,4-linkages between d-glucosamine and N-acetyl-d-glucosamine in chitosan. It has been found in various organisms including bacteria, fungi, and plants. These chitosanases belong to various glycosyl hydrolase (GH) families including GH-5, GH-8, GH-46, GH-75, and GH-80, according to their amino acid sequences (Gao et al. 2008). Chitosanases produced from Bacillus cereus KNUC51 and KNUC55 (Park et al. 2004), which have been investigated in a previous study, belong to GH-8, a family that displays an (α/α)6 fold and that presently includes xylanases, cellulases, lichenases, and a number of other chitosanases (Collins et al. 2005; De Vos et al. 2006; Lee et al. 2007).
DNA shuffling of low homologous DNA fragments increases the sequence diversity of the library, resulting in an accelerated rate of enzyme functional improvement. It is also a powerful tool for identifying the major structural features of a protein family that confer a desired phenotype. For example, the activity of TEM-1 β-lactamase increased to 32,000 times higher than that of the parental source by the third round of DNA shuffling and the stability of p-nitrophenyl (pNP) esterase against the solvent was increased more than 50-fold compared with that of parental sources (Moore et al. 1997; Stemmer 1994). The molecular evolution of chitosanase to obtain higher activity and stability has rarely been attempted, even though many chitosanases have been screened over a time span of decades.
In this study, DNA shuffling was carried out with two chitosanase genes from isolated B. cereus KNUC51 and B. cereus KNUC55. The shuffled products had several amino acid substitutions and showed improved chitosanase activity. YM20, moreover, had a broader range of pH than the parents.
Materials and methods
Bacterial strains, plasmids, and medium
The chitosanase genes (csn) originating from Bacillus cereus KNUC51 (BO1) and B. cereus KNUC55 (BO5) were selected for directed evolution. Escherichia coli DH5α was used for the expression and screening of the variant chitosanase library. Escherichia coli BL21(DE3)pLysS was used as the host strain for overexpression and purification of recombinant proteins. The plasmids, pGEM T-Easy vector (Promega, Madison, USA) and pGEX-2T (Amersham Biosciences, Uppsala, Sweden), were used as the cloning vector and overexpression vector, respectively. The recombinants of E. coli DH5α were selected using Luria-Bertani (LB) medium supplemented with ampicillin and colloidal chitosan (Bertani 1951). The recombinants of E. coli BL21(DE3)pLysS were cultivated in LB medium supplemented with ampicillin (100 μg/ml) and chloramphenicol (25 μg/ml).
Assay of chitosanase activity
Soluble chitosan (1% w/v) was incubated with the enzyme at pH 5.5 and 40°C for 10 min. The specific activity of chitosanases were determined by modified Schale’s method (Imoto and Yagishita 1971). One unit of chitosanase activity was defined as the amount of enzyme that produced 1 mmol reduced sugar per min. d-Glucosamine was used for the standard curve. The quantity of protein was determined by the Bradford assay and albumin from bovine serum was used as the protein standard. Optimum pH of recombinant chitosanase was measured with 1% soluble chitosan adjusted to various pH conditions.
Library construction using DNA shuffling
DNA fragments including BO1 and BO5 were cloned into the pGEM T-Easy vector and named TYM51 and TYM55, respectively. The DNA fragments derived from them were amplified by polymerase chain reaction (PCR) using the primers M13-F and M13-R. The purified PCR products were digested with 0.6 U DNase I in the presence of Mn2+ for 10 min at 15°C, as previously described (Lorimer and Pastan 1995). After inactivating DNase I, DNA fragments in the range of 100–150 bp were purified using a Gel Extraction kit (Qiagen, Hilden, Germany). The purified DNA fragments were then subjected to 60 cycles of the first PCR reaction without primers (1 min at 95°C, 1 min at 60°C, and 1 min at 72°C). To generate the chimeric csn libraries, a second PCR reaction was performed for 35 cycles with the primers C1 and C2. The shuffled products were cloned into the pGEM T-Easy vector, and a chimeric library was constructed by transforming into E. coli DH5α. Positive clones showing a larger halo zone than those of the parents were selected.
Construction and expression of the overexpression vector
The complete coding regions of chimeric csn were inserted into glutathione pGEX-2T and transformed into E. coli BL21(DE3)pLysS. Recombinants were cultivated in 5 ml of LB medium with ampicillin for 16 h at 37°C. Cultures were inoculated with 100 ml fresh LB medium with ampicillin, chloramphenicol and incubated for 5–6 h at 25°C. Overexpression of proteins was induced by the addition of 0.1 mM IPTG for 24 h. Cells were harvested by centrifugation at 2,760×g for 10 min.
Purification and characterization of recombinant chitosanase
The cell pellets were washed with PBS and then resuspended in PBS and sonicated. Proteins in the supernatant were obtained by centrifugation at 15,930×g for 10 min at 4°C. The recombinant proteins were purified using affinity chromatography with glutathione-Sepharose 4B (Amersham Biosciences, Uppsala, Sweden). The protein solution was applied to the column, washed 3 times with PBS, and eluted with elution buffer (10 mM glutathione in 50 mM Tris/HCl, pH 8.0). The purified proteins were treated with 5 units of thrombin protease (Amersham Biosciences, Uppsala, Sweden) at 25°C for 16 h to remove the GST region. To confirm the protein purity and determine the molecular weight of the proteins, SDS-PAGE was performed. The Perfect protein marker (Novagen, Madison, USA) was loaded on the gel as the standard size marker.
Prediction of the three-dimensional structure
The three-dimensional structures of WT chitosanase and chimeric chitosanase were predicted using SwissModel (http://swissmodel.expasy.org//SWISS-MODEL.html). The structure of 1V5D (chitosanase of Bacillus sp. K17), which was previously determined by X-ray diffraction, was used for simultaneously modeling chitosanase molecules (Adachi et al. 2004). 1V5D shows 98% and 96% homology in the amino acid sequence with chitosanase from B. cereus KNUC51 and B. cereus KNUC55, respectively.
Results
Construction and screening of the DNA shuffling library
DNA fragments of approximately 1.6 kb were amplified with M13 primers and TYM51 and TYM55 as the template. Supplementary Fig. 1 shows the results of the DNA fragments in all steps of reassembly. DNA fragments of around 100–150 bp were purified and used for reassembly. After the PCR reassembly reaction, weak and dragged bands were observed at around 1.4 kb. The reassembled DNA fragments were used for the PCR amplification of 1.4-kb DNA fragments including the mutated csn gene. The PCR products were purified and ligated with the pGEM T-Easy vector. The plasmids including shuffled products were transformed into E. coli DH5α. Thousands of halo-forming E. coli DH5α appeared on chitosan-containing agar plates. Sixty-seven halo-forming colonies were screened for chitosanase activity of the crude extract. Mutant YM18 and YM20 were found to have improved activity at 40°C. Both mutants were sequenced completely and further characterized.
Overexpression and purification of recombinant chitosanase
The recombinant chitosanases were purified using glutathione affinity chromatography. Most of the GST-tagged chitosanases (Mw 81 kDa) were produced in a soluble form on the SDS-PAGE gel (Supplementary Fig. 2). After digestion using thrombin, the GST-fusion proteins were separated into chitosanases (Mw 50 kDa) and glutathione-S-transferase (Mw 30 kDa).
Characterization of the mutants: biochemical analysis
The profiles of activity at various temperatures are given in Fig. 1. Generally, the specific activity of YM18 is enhanced 250% and the specific activity of YM20 is enhanced 350% compared to the parents. The mutant YM18 chitosanase was most active at 50°C, nearly the same as the parental BO1. The mutant YM20 chitosanase was most active at 70°C, which was also nearly the same as the parental BO5. Chimeric chitosanase shows different aspects of specific activity depending on its origin.
The pH profiles of the activities are shown in Fig. 2. YM20 exhibits a shift of the optimal pH level from 5.5 to 6.5. The relative activities of YM20 at pH 6.5 are approximately 5 times higher than those of the parents, resulting in a broader pH activity profile for YM20. In contrast, the pH activity profile of YM18 is similar to the parents’ profile, with an optimum pH of 5.5.
Characterization of the mutants: sequence analysis and prediction of structure
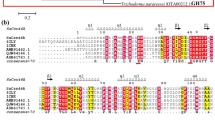
Homology between BO1 and BO5, which consisted of 453 amino acids, is 92%. The dissidence of amino acid between two parents was concentrated at N-terminal and C-terminal of proteins (Fig. 3). YM18 had 96% protein sequence identity with BO1, and YM20 had 95% protein sequence identity with BO5. DNA sequencing revealed that YM18 and YM20 contained two amino acid substitutions (I13T and A87V for YM18; K66R and N352S for YM20).
According to the three-dimensional structures of recombinants predicted by the SwissModel program, two parental chitosanases were similar to 1V5D (chitosanase of Bacillus sp. K17). They have six repetitions of the helix-loop-helix motif that showed a typical α6/α6-double-barrel structure. The I13T of YM18 is not represented on the structure because the query structure does not contain that region. The A87V of YM18 is at the tenth alpha-helix. The two amino acid substitutions, K66R and N352S, exist at the N-terminal region and ninth alpha-helix on the surface of YM20, respectively (Fig. 4).
Ribbon representation of the predicted B. cereus chitosanase structure. The position of the three amino acid replacements that conferred the chitosanase activity and shift of pH performance is shown by sphere representation (YM18, green; YM20, red). The three catalytic residues are shown as a ball-and-stick model
Discussion
To date, many glycoside hydrolases, but not chitosanases, have been used in directed evolution experiments to improve factors such as their activity, thermostability, and pH performance (Dumon et al. 2008; Qin et al. 2008). From various organisms, many kinds of chitosanases, which are classified into three or more subclasses, have been reported, but have not been investigated with regard to improving enzymatic ability.
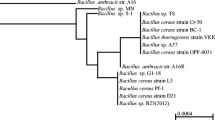
In this study, DNA shuffling was used to improve the specific activity and pH performance of chitosanase belonging to GH-8 from B. cereuse strains. Two variants that show higher activity than the parents at all temperatures were selected for further studies. The sequence relationship is illustrated in a phylogenetic tree showing that the two variants are closely related to each other (data not shown). However, they have distinct characteristics with regard to optimum temperature and pH, which may because they contain three different amino acids (residue 80, 279 and 392) that originated from different parents. Moreover, these two variants contained two individual amino acid substitutions, which are far from the active site residues (Fig. 4). These amino acid substitutions increased the specific activity of the two variants. The substitution I13T of YM18 may enhance the specific activity. The threonine for isoleucine substitution, replacing a hydrophobic residue with a non-charged polar side chain, changes the geometry of its immediate vicinity, presumably creating new hydrogen bonds and tightening the structure of which it is a part (McCarthy et al. 2004). The substitution V387A may not be involved in hydrogen bonding with other residues.
According to Turunen et al. (2002), the introduction of arginines into the Ser/Thr surface resulted in a shift of the pH profile to an alkaline pH by 0.5–1.0 pH units. The authors suggest that the K66R amino acid substitution causes a shift of the optimal pH level from 5.5 to 6.5 of YM20. However, an increase in the number of arginines is not in itself sufficient to cause an acidic enzyme to function in highly alkaline conditions. Basic residue Lys66 might play an important role in the pH-dependent catalytic reaction. Moreover, it might drive YM20 to increase specific activity. Further study based on site-directed mutagenesis will elucidate the role of each mutation in controlling the catalytic activity and pH activity.
References
Adachi W, Sakihama Y, Shimizu S et al (2004) Crystal structure of family GH-8 chitosanase with subclass II specificity from Bacillus sp. K17. J Mol Biol 343:785–795
Bertani G (1951) Studies on lysogenesis. I. The mode of phage liberation by lysogenic Escherichia coli. J Bacteriol 62:293–300
Collins T, De Vos D, Hoyoux A et al (2005) Study of the active site residues of a glycoside hydrolase family 8 xylanase. J Mol Biol 354:425–435
De Vos D, Collins T, Nerinckx W et al (2006) Oligosaccharide binding in family 8 glycosidases: crystal structures of active-site mutants of the β-1,4-xylanase pXyl from Pseudoaltermonas haloplanktis TAH3a in complex with substrate and product. Biochemistry 45:4797–4807
Dumon C, Varvak A, Wall MA et al (2008) Engineering hyperthermostability into a GH11 xylanase is mediated by subtle changes to protein structure. J Biol Chem 283:22557–22564
Gao X-A, Ju W-T, Jung W-J et al (2008) Purification and characterization of chitosanase from Bacillus cereus D-11. Carbohydr Polym 72:513–520
Imoto T, Yagishita K (1971) A simple activity measurement of lysozyme. Agric Biol Chem 35:1154–1156
Lee H-S, Jang JS, Choi S-K et al (2007) Identification and expression of GH-8 family chitosanases from several Bacillus thuringiensis subspecies. FEMS Microbiol Lett 277:133–141
Lorimer IA, Pastan I (1995) Random recombination of antibody single chain Fv sequences after fragmentation with DNaseI in the presence of Mn2+. Nucleic Acids Res 23:3067–3068
McCarthy JK, Uzelac A, Davis DF et al (2004) Improved catalytic efficiency and active site modification of 1,4-(beta)-d-glucan glucohydrolase A from Thermotoga neapolitana by directed evolution. J Biol Chem 279:11495–11502
Moore JC, Jin HM, Kuchner O et al (1997) Strategies for the in vitro evolution of protein function: enzyme evolution by random recombination of improved sequences. J Mol Biol 272:336–347
Park YM, Chang HL, Huh TL et al (2004) Molecular cloning of chitosanase gene and quantitative production of chitosan oligomer. Korean J Microbiol Biotechnol 32:16–21
Qin Y, Wei X, Song X et al (2008) Engineering endoglucanase II from Trichoderma reesei to improve the catalytic efficiency at a higher pH optimum. J Biotechnol 135:190–195
Stemmer WP (1994) Rapid evolution of a protein in vitro by DNA shuffling. Nature 370:389–391
Turunen O, Vuorio M, Fenel F et al (2002) Engineering of multiple arginines into the Ser/Thr surface of Trichoderma reesei endo-1,4-(beta)-xylanase II increases the thermotolerance and shifts the pH optimum towards alkaline pH. Protein Eng 15:141–145
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Park, YM., Ghim, SY. Enhancement of the activity and pH-performance of chitosanase from Bacillus cereus strains by DNA shuffling. Biotechnol Lett 31, 1463–1467 (2009). https://doi.org/10.1007/s10529-009-0017-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-009-0017-2