Abstract
A binary vector containing two reporter gene cassettes has been developed. This vector is ideal for optimising new plant transformation systems. Following optimisation, one of the reporter genes can be replaced with a gene of interest; the second can be used as a marker to confirm transgenic lines, and to estimate locus number and determine zygosity. This allows simple, efficient and economical screening for homozygous single-insert lines and azygous controls.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Development of efficient transformation systems is a rate-limiting step in genetic engineering of plant species for applied biotechnology, genetic complementation and gene expression studies. Agrobacterium is the method of choice for transformation where possible, and has been reviewed in detail (e.g., Gelvin 2003). Agrobacterium-mediated transformation is limited by host range, which originally restricted transformable species to those in the dicot group. However, it is now possible to transform many previously recalcitrant species, in particular important monocot crop species (e.g., rice, corn, wheat, barley, sugarcane, banana). Projects are now underway to develop and optimise Agrobacterium-mediated transformation systems for a large variety of species. In order to do this, universal promoter: reporter gene constructs are used. These consist of a constitutive promoter driving expression of a reporter gene with an easily-assayable phenotype. Once a transformation system has been developed, it is possible to initiate studies on specific genes of interest. However, confirmation of transgenic lines and segregation analysis to determine zygosity can be complex if the phenotype of the gene of interest is unknown or the gene is difficult or costly to assay for. It is therefore of use to have a secondary reporter gene included on the transformation vector for use as a co-segregating marker.
Materials and methods
Generation of expression cassette
To generate a 35S promoter:GFP expression cassette, an EcoRI-cut and blunt-ended fragment containing the synthetic sGFP(S65T) gene (Chiu et al. 1996) from pUBIGFP (Christensen and Quail 1996) was inserted into the plasmid pBI221 (Mitsuhara et al. 1996) replacing the GUS gene to create p35SsGFP. pGFP:GUSPlus (GenBank EF546437) was made by cloning the 35S:GFP:NOS expression cassette from p35SsGFP into pCAMBIA1305.1 (CAMBIA, Canberra, Australia) upstream and in the opposite orientation to the 35S:GUSPlus:NOS cassette, by ligation through the HindIII and EcoRI sites in the multiple cloning sites. In this way, expression of both reporter genes benefits from the enhancer elements (Odell et al. 1988; Omirulleh et al. 1993) found in the two 35S promoters.
Transformation and selection
Tobacco (Nicotiana tabacum cv. Samsun NN) transformation was performed essentially according to Guerineau et al. (1990) with the following changes: MES was omitted from all media; leaf sections were cut while suspended in the Agrobacterium culture; co-cultivation was carried out on solidified medium (0.8% agar); following co-cultivation leaf sections were washed in liquid selection medium; augmentin (200 μg/ml) was used to remove agrobacteria and hygromycin (20 μg/ml) was used as a selective agent in the selective medium; and rooting was performed on a medium identical to selection medium but omitting plant hormones. Primary transgenic (T0) plants were analysed for GUS activity using the histochemical assay as described previously (Jefferson 1987).
Results and discussion
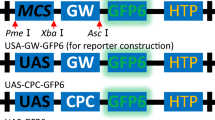
To facilitate development and optimisation of transformation systems, we have constructed a transformation vector, pGFPGUSPlus, which bears expression cassettes for both green fluorescent protein (GFP) (Chalfie et al. 1994) and an improved β-glucuronidase (GUSPlus) (Broothaerts et al. 2005) (Fig. 1). Each reporter gene is controlled by the constitutive cauliflower mosaic virus (CaMV) 35S promoter, which is active in dicot species and many monocot species. Reporter gene cassettes and a hygromycin resistance gene cassette are located between T-DNA borders, allowing Agrobacterium-mediated transformation; this plasmid should also be effective for direct gene transfer. All three expression cassettes can be easily removed and replaced with a gene of interest or a different selectable resistance gene (Fig. 1).
Schematic diagram of pGFPGUSPlus (Full sequence and annotation details for pGFPGUSPlus can be found under Genbank accession number EF546437). LB, left border for T-DNA integration; RB, right border for T-DNA integration; HPTII, hygromycin resistance gene; GFP, green fluorescent protein gene; GUSPlus, β-glucuronidase gene (includes catalase intron); 35S, cauliflower mosaic virus 35S promoter; 2 × 35S, 35S promoter containing a double enhancer region (Omirulleh et al. 1993); N, nopaline synthase (nos) terminator region; C, cauliflower mosaic virus 35S terminator region. Restriction enzyme sites are shown (e.g., EcoRI) which are not found in the plasmid backbone outside the border sequences. A kanamycin resistance gene is present on the plasmid backbone for selective amplification in bacteria. The hygromycin gene cassette for selection of transgenic plant tissues can be easily replaced with a selectable resistance gene of choice. Use of GUSPlus as a selectable marker precludes false positives from surviving endogenous agrobacteria because of the presence of an intron in the coding region (Broothaerts et al. 2005; Vancanneyt et al. 1990)
Use of pGFPGUSPlus allows choice of either GUSPlus or GFP, depending on the availability of assay systems in the laboratory. In addition, one of the reporter genes can be replaced by a gene of interest after optimisation has been achieved, allowing use of essentially the same plasmid backbone in subsequent expression analyses. Furthermore, the remaining reporter gene cassette allows secondary selection of transgenic tissues (in addition to antibiotic selection) and facilitates simple locus number and segregation analyses of transgenic lines. This is particularly useful when the function of the gene of interest is unknown, or the assay is costly or complex. Segregation analysis can also be used to identify homozygous and heterozygous parents from T1 generation single-locus lines using the T2 progeny. Generation of single-copy homozygous lines is important in transgenic plant analysis in order to achieve genetic stability in subsequent generations. This method can also be used to identify azygous plants in the T1 generation; azygous plants are ideal controls as they are near-isogenic with transgenic plants.
To test pGFPGUSPlus, we used it for transformation of tobacco. GUSPlus activity was used to identify primary transgenic (T0) plants (Fig. 2A). The locus number of T0 plants was estimated, and azygous T1 plants were identified by segregation analysis in the T1 generation (Fig. 2B). The zygosity of single-locus plants from the T1 generation was determined by segregation of GUSPlus activity in the T2 generation (Fig. 2C–E). In this way, identification of homozygous and azygous plants (for use as near-isogenic controls) is easily facilitated. We are currently using pGFPGUSPlus to optimise a transformation system for soybean (Glycine max cv. Bragg) (Fig. 2F–G), and have also used pGFPGUSPlus for hairy root transformation of G. max (cv. Williams) (data not shown) using Agrobacterium strain K599 (Kereszt et al. 2007). We have successfully used pGFPGUSPlus for functional analysis in transgenic tobacco by replacing the GFP reporter gene with a gene of interest and using GUSPlus as a marker for transgene locus segregation (Vickers et al., manuscript in preparation). In this case, single-locus insertion lines identified by segregation analysis of GUSPlus activity in the T1 generation were confirmed by Southern blot analysis; subsequent to this, the zygosity of T1 plants was determined by segregation analysis of GUSPlus activity in the T2 generation. The assay for the reporter gene (GUSPlus) was much faster and simpler than assaying for the gene of interest. We were also able to generate azygous controls for each homozygous single-locus transgenic line.
Reporter gene activity in transgenic plant tissues transformed with pGFPGUSPlus. (A)–(D) show leaf discs (diam. = 6 mm) from transgenic tobacco plants assayed for GUSPlus activity. Primary transgenic (T0) plants can be identified by GUSPlus activity; four discs from an individual T0 plant are shown in (A). When leaf discs from individual T1 progeny plants exhibit a segregation ratio of ∼1:3 (B), it suggests that the T0 parent plant is a single-locus (SL) insertion line. Azygous (Az) plants can also be identified in the T1 generation. To identify homozygous T1 plants, segregation analysis is performed in the T2 plants. T2 progeny of GUSPlus-positive T1 plants show either no segregation, demonstrating a homozygous (Ho) parent (C), or ∼1:3 segregation, demonstrating a heterozygous (He) parent (E). T2 progeny of Az plants identified in the T1 generation are of course also Az and show no GUSPlus activity (D). Reporter gene activity can also be used to facilitate optimisation of the transformation process; (F) shows GFP in soybean cotyledons 2 d after co-cultivation with Agrobacterium tumefaciens bearing pGFPGUSPlus, and (G) shows GFP in the shoot apex of a transgenic soybean plantlet. GFP was visualised using a Leica MZ6 dissecting microscope fitted with the GFP Plus fluorescence module (Diagnostic Instruments, Sterling Heights, MI, USA)
pGFPGUSPlus is an efficient transformation vector with a number of useful features, which are applicable for facilitating both gene expression studies and the development of plant transformation systems. In combination with transient transformation and gene expression analyses methods (Schenk et al. 1998; Vickers et al. 2003), this vector could also be used for gene promoter analyses, e.g., (Schünmann et al. 2004; Vickers et al. 2006).
References
Broothaerts W, Mitchell HJ, Weir B et al (2005) Gene transfer to plants by diverse species of bacteria. Nature 433:629–633
Chalfie M, Euskirchen G, Ward WW et al (1994) Green fluorescent protein as a marker for gene expression. Science 236:802–805
Chiu WL, Niwa Y, Zeng W et al (1996) Engineered GFP as a vital reporter in plants. Curr Biol 6:325–330
Christensen AH, Quail PH (1996) Ubiquitin promoter-based vectors for high-level expression of selectable and/or screenable marker genes in monocotyledonous plants. Transgenic Res 5:213–218
Gelvin SB (2003) Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol Mol Biol Rev 67:16–37
Guerineau F, Brooks L, Meadows J et al (1990) Sulfonamide resistance gene for plant transformation. Plant Mol Biol 15:127–136
Jefferson RA (1987) Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol Biol Rep 5:387–405
Kereszt A, Li D, Indrasumunar A et al (2007) Agrobacterium rhizogenes-mediated transformation of soybean to study root biology. Nat Protoc 2(4):948–952
Mitsuhara I, Ugaki M, Hirochika H et al (1996) Efficient promoter cassettes for enhanced expression of foreign genes in dicotyledonous and monocotyledonous plants. Plant Cell Physiol 37:49–59
Odell JT, Knowlton S, Lin W et al (1988) Properties of an isolated transcription stimulating sequence derived from the cauliflower mosaic virus 35S promoter. Plant Mol Biol 10:263–272
Omirulleh S, Ábrahám M, Golovkin M et al (1993) Activity of a chimeric promoter with the doubled CaMV 35S enhancer element in protoplast-derived cells and transgenic plants in maize. Plant Mol Biol 21:415–428
Schenk PM, Elliott AR, Manners JM (1998) Assessment of transient gene expression in plant tissues using the green fluorescent protein as a reference. Plant Mol Biol Rep 16:313–322
Schünmann PHD, Richardson AE, Vickers CE et al (2004) Analysis of the HORvu;Pht1;1 promoter identifies regions controlling root expression and the phosphate response. Plant Physiol 136:4205–4214
Vancanneyt G, Schmidt R, O’Connor-Sanchez A et al (1990) Construction of an intron-containing marker gene: splicing of the intron in transgenic plants and its use in monitoring early events in Agrobacterium-mediated plant transformation. Mol Gen Genet 220:245–250
Vickers CE, Xue G-P, Gresshoff PM (2006) A novel cis-acting element, ESP, contributes to high-level endosperm-specific expression in an oat globulin promoter. Plant Mol Biol 62:195–214
Vickers CE, Xue G-P, Gresshoff PM (2003) A synthetic xylanase as a novel reporter in plants. Plant Cell Rep 22:135–140
Acknowledgements
This research was supported by Biotechnology and Biological Sciences Research Council grant BBS/B/12172 (CEV and PMM), ARC Centre of Excellence grant CEO348212 (PMG and CEV), the University of Queensland Strategic Research Fund and the Queensland Government Smart State Initiative (PMG) and the Cooperative Research Centre for Tropical Plant Protection (PMS).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vickers, C.E., Schenk, P.M., Li, D. et al. pGFPGUSPlus, a new binary vector for gene expression studies and optimising transformation systems in plants. Biotechnol Lett 29, 1793–1796 (2007). https://doi.org/10.1007/s10529-007-9467-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-007-9467-6