Abstract
Bladder cancer is one of the most predominant tumors of the genitourinary tract. In addition to pathological findings, the molecular modifications that might affect tumorigenesis and tumor outcome should be considered when treating bladder cancer. Accordingly, we aimed to investigate the expression levels of both the ASPM and TEF genes in bladder cancer tissues and their value in disease prognosis. The expression levels of the ASPM and TEF genes were analyzed by quantitative real-time PCR (qRT-PCR) in 90 bladder cancer tissue specimens and 90 specimens of normal urinary bladder tissue taken away from the tumor site. The upregulation of ASPM expression and the downregulation of TEF expression were observed in bladder cancer tissues compared to adjacent normal tissues, and these levels were correlated with high-grade tumors, advanced stage disease and the presence of metastasis. Both genes had the ability to predict metastatic association with sensitivity (84.62%) and specificity (68.42%; *P < 0.001) for the ASPM gene and for the TEF gene with sensitivity (80.77%) and specificity (78.95%; *P < 0.001). Additionally, Kaplan–Meier survival analysis indicated that elevated ASPM expression levels and reduced TEF expression levels significantly correlated with decreased overall survival and progression-free survival. The current analysis concludes that ASPM and TEF expressions might be used as potential biomarkers in bladder cancer patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bladder cancer (BC) is one of the most prevalent malignant tumors and has a phenomenally high rate of recurrence (Kwan et al. 2019). Bladder cancer is the second most common genitourinary tract malignancy worldwide (Torre et al. 2015) and the fourth most common cancer among males (Siegel et al. 2018). Regardless of the advancements in both systemic and local treatment and management protocols, noninvasive bladder cancer still has increased rates of progression and disease recurrence, while invasive and metastatic tumors have poor survival rates (Jiang et al. 2019). Bladder cancer has many risk factors that contribute to tumor development and progression such as genetic risk factors, chemical hazards such as cigarette smoking, and long-standing irritation such as pelvic irradiation (Kaufman et al. 2009). In bladder cancer, ordinary imaging procedures are not of much worth in early diagnosis (Rosenkrantz et al. 2016), and even though cystoscopy can help determine tumor characteristics, such as site and size, and affirm pathological findings, there is still a need for a valuable biomarker for early diagnosis (Dunphy et al. 2017).
The abnormal spindle-like microcephaly associated (ASPM) gene is found on chromosome 1q31. The ASPM gene is primarily found in embryonic neuroblasts but has shown to be extensively present in an assortment of grown-up and embryonic tissues as well; it is likewise known as the abnormal spindle microtubule assembly gene (Kouprina et al. 2005). The encoded ASPM protein is involved in mitotic spindle formation and was at first recognized as a centrosomal protein that controls neurogenesis and brain growth, but it is now known to be generally expressed in a diverse array of normal and cancer tissues (Bikeye et al. 2010). Additionally, expanded cell growth and tumor advancement correlate with increased ASPM expression, indicating that ASPM might have an influence on cellular proliferation (Kouprina et al. 2005; Buchman et al. 2011).
Thyrotroph embryonic factor (TEF) is a protein that belongs to the proline- and acidic amino acid–rich (PAR) bZIP family and controls transcription by acting as transcription factor with DNA binding ability (Inukai et al. 2005). TEF is expressed initially in the embryonic anterior pituitary; however, in adult life, it is thought to be involved in controlling cell cycle and the cellular death of hematopoietic cells, thereby affecting their propagation (Yang et al. 2019). The TEF gene is also concerned with circadian rhythm moderation (Hua et al. 2012). Regarding the ability of TEF to bind to DNA, it can reduce the transcription of interleukin 3 (IL-3) and IL-3 receptor β subunits, affecting cellular apoptosis and preventing cell cycle progression (Inukai et al. 2005). In this study, we aimed to evaluate the expression levels of both the ASPM and TEF genes in BC tissues and their value in the disease prognosis, patient survival, and metastatic prediction.
Subject and Method
This study was carried out by the Medical Biochemistry & Molecular Biology, Urology, Pathology and Clinical Oncology Departments, Faculty of Medicine, Menoufia University, from January 2017 to June 2019. Specimens were collected from patients who underwent diagnostic cystoscopy and biopsy because of suspicious bladder cancer or after cystectomy. 180 tissue specimens were included in the current analysis. Ninety neoplastic tissue specimens from patients with proven bladder cancer and another 90 specimens of normal urinary bladder tissue taken away from the tumor site were collected.
All patients were examined for the following: full history; general and local urological examination; routine laboratory investigations including urine analysis, serum urea and creatinine; Bilharzial antibodies (Abs); routine radiological investigations, including abdominal pelvic ultrasonography and/or abdominopelvic CT to detect masses in the bladder; and specific investigations including urine cytology.
Patients with performance status ≥ 2, creatinine clearance ≤ 60 ml/min and major comorbidities in the form of organ failure were excluded from the study.
All specimens were received by the Pathology Department, where fresh parts of the tumor mass and adjacent normal bladder tissue were collected and stored in RNA protect Tissue Reagent, RNALater, QIAGEN® ( 10 μl reagent per 1 mg of tissue) at − 80 °C for further molecular analysis of ASPM and TEF mRNA expression levels at Medical Biochemistry & Molecular Biology Department. The rest of the specimens were immersed in formalin and submitted to routine tissue processing to obtain paraffin-embedded samples.
Patients with malignant pathologies were subjected to further metastatic analyses, including CT chest, abdomen, pelvis, and bone scans.
The clinical staging of cancer was based on the TNM American Joint Committee on Cancer-Union International Center Cancer staging system (AJCC-UICC) (Amin et al. 2013). Grading was performed according to World Health Organization (WHO) and the International Society of Urologic Pathology (ISUP) classification (Moch et al. 2016). The malignant tumors were divided into non-muscle-invasive bladder cancer (NMIBC) (stage pTa and pT1) or muscle-invasive bladder cancer (MIBC) (stage pT2, pT3, and pT4) according to the TNM classification for the stage (Amin et al. 2015).
Patients with stage I disease underwent trans-urethral resection of the bladder tumor, followed by intravesical injection. Patients with T2-T4a, cN0MO muscle-invasive bladder cancer were treated with neoadjuvant cisplatin-based chemotherapy followed by radical cystectomy and bilateral pelvic lymph node dissection or chemoradiation. Patients with metastatic disease received platinum-based chemotherapy. Patients were assessed by CTs and cystoscopy, and their response was measured according to RECIST criteria.
Follow-up was completed for all patients until June 2019. The progression-free survival (PFS) rate was defined as the time between the date of diagnosis and the date of progression in the form of local recurrence, newly developed metastases for patients with localized disease, or an increase in size and/or number of metastases in patients with stage IV disease or last visit. The PFS rate was analyzed in relation to different prognostic factors among patients. The overall survival (OS) was considered until date of death or last visit.
Sampling Work-Up
Venipuncture was performed, and seven ml of peripheral blood was taken from each patient. Two ml of blood was assembled into a tube including EDTA as an anticoagulant for CBC (Sysmex XN-1000, Japan (19723), B.M Egypt Company). Five ml was collected into a plain tube, and sera were separated by centrifugation at 5000 R.P.M after blood clotting. The obtained serum was preserved at − 20 °C in order to evaluate serum albumin, urea and creatinine and identification of bilharzial antibodies. The levels of urea in serum and creatinine in both serum and urine were determined by enzymatic colorimetric (DIAMOND Diagnostics, Germany), and creatinine clearance was estimated from the equation: U · V/P; where U is urine creatinine levels, V is the volume of urine by minute, and P is the plasma levels of creatinine (Lavender et al. 1969). The level of albumin was evaluating by exploiting the enhanced specificity of bromocresol green colorimetric analysis (DIAMOND Diagnostic Kit, Germany). Bilharzia or schistosoma antibodies in humans (IgG) were detected using an ELISA (DRG® Schistosoma IgG, USA) (Doenhoff et al. 2004).
For the cytological analysis of urine, ten ml of morning urine sample was collected, and the sediments were examined for malignant cell content.
ASPM and TEF mRNA Expression Levels
RNA Extraction from Bladder Tissues
Total RNA extractions from all ninety tissue specimens were completed in 48 h to five days maximum from time of tissue collection using the QIAamp RNA Blood MiniKit (Qiagen, USA) as indicated by the producer's instructions (Wang et al. 2000). The quality and quantity of extracted RNA were measured using agarose gel electrophoresis and nanodrop spectrophotometer (NanodropTechnologies).
Two-Stage RT- PCR
First, complementary DNA(cDNA) was synthesized by reverse transcription utilizing a reverse transcriptase kit (SensiFASTcDNA synthesis kit, Bioline Reagents Ltd, United Kingdom). An Applied Biosystems 2720 thermal cycler (Singapore) was used to process the reaction mixture of 20 μl that included 10 μl RNAextract added to a mixture of 1 μl of reverse transcriptase enzyme, 4 μl of 5 × TransAmp buffer and 5 μl of DNase/RNase-free water. The reaction was 10 min at 25 °C for primer annealing, followed by 15 min at 42 °C for reverse transcription, and a final step of 5 min at 85 °C for inhibition of reverse transcriptase enzyme. The produced cDNA was preserved at − 20 °C.
The second stage of cDNA amplification was quantitative real-time PCR (qRT-PCR). It was carried out for both ASPM and TEF mRNA expression relative to the endogenous reference gene β-actin utilizing the 2 × SensiFAST™ SYBR® Lo-ROX Kit (Bioline Reagents Ltd.) and an Applied Biosystems 7500 Real-Time PCR system. The used primers’ accuracy was confirmed utilizing the Primer BLAST program by NCBI and were sequenced as: ASPM: forward primer 5′- AGCATTCCTTTTATCCCAGAAACACCTG-′3 and reverse primer 5′- GCTTGCAGGGGATTTGTGATTTCTTCC-3′; TEF: forward primer 5′- CTGCCTCACAACGACTCCTTTCTCT -′3 and reverse primer 5′- TCGCCTCTGTCTCCTCTTCACCATAG—3′; Β-actin: forward primer 5′-GGCGGCACCACCATGTACCCT-3′ and reverse primer 5′-AGGGGCCGGACTCGTCATACT-3′. A reaction volume of 25 μl for RT-PCR was created from 12.5 μl of 2 × SensiFAST™ SYBR® Lo -ROX Master Mix,1 μl of individual primer, 5.5 μl of DNase/RNase-free water and 5 μl of the previously formed cDNA. The thermal cycler Applied Biosystems 7500 with the software version 2.0.1 used a program of 95 °C for 10 min, then 45 cycling of 15 s at 95 °C and 1 min at 60 °C. The results were interpreted using the comparative Ct method (2−∆∆Ct), and the relative quantification of ASPM and TEF expressions was done by normalizing their expressions to that of the endogenous gene Β-actin.
Statistical Analysis
The IBM SPSS software package version 20.0 (Armonk, NY: IBM Corp) was utilized for the statistical analysis of our data. To compare the two groups, Student’s t test was used for normally distributed quantitative variables, while the Mann–Whitney test, Wilcoxon signed rank test, and Kruskal–Wallis test were used for abnormally distributed quantitative variables. Spearman’s coefficient was used to examine the correlations between quantitative data. In terms of estimating the diagnostic performance of the gene expression levels, receiver operating characteristic curve (ROC) was utilized. Multiple regression analysis was performed. A Kaplan–Meier survival curve was implemented, and Cox regression was performed for significant relation with progression-free survival and the overall survival according to log rank test. Significance of the results was judged at the 5% level.
Results
This work was conducted on 90 patients with BC. The sample included 74(82.2%) males and 16(17.8%) females with a mean age of 58 ± 8.7 years. Regarding the histopathological examination of the tumor tissue samples, 54 (60%) of the patients were diagnosed as having conventional urothelial carcinoma (CUC) of the bladder, 24 (26.7%) patients had urothelial carcinoma with squamous differentiation (UCS), and 12 (13.3%) patients were reported to have urothelial carcinoma with other divergent differentiation (UCD) tumors. The majority of our patients were pathologically diagnosed to have muscle-invasive bladder cancer (MIBC), and only 14 (15.6%) patients had non-muscle-invasive bladder cancer (NMIBC). All clinicopathological characters of bladder cancer are listed in Table 1.
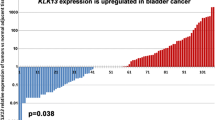
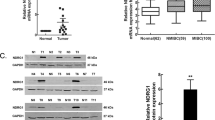
On evaluating the expression level of both ASPM and TEF genes in BC tissues and normal tissues, we found highly significant differences between the two tissues with ASPM gene expression upregulated and TEF gene expression downregulated in BC tissues compared to the normal tissues (P < 0.001*) (Table 2; Fig. 1a, b). Similar results were also obtained up on stratification Patients with BC to NMIBC and MIBC and also evaluating the expression level of both genes in each group separately (Table 3). However, increased ASPM mRNA level and decreased TEF mRNA in malignant tissues appeared to be more predominant in MIBC versus NMIBC (P < 0.001) (Table 6).
Based on the expression level of both the ASPM and TEF genes, a receiver operating characteristics (ROC) curve was utilized. The ASPM gene had a sensitivity of 95.56% and a specificity of 86.67% (P < 0.001*) and the TEF gene had a sensitivity of 91.11% and a specificity of 71.11% (P < 0.001*) to differentiate the BC tissues from normal tissues (Table 4; Fig. 2a). Moreover, both genes had the ability to predict metastasis with a sensitivity of 84.62% and a specificity of 68.42% (P < 0.001*) for the ASPM gene and for the TEF gene a sensitivity of 80.77% and a specificity of 78.95% (P < 0.001*). (Table 5; Fig. 2b). Combination of both the ASPM and TEF genes expression levels has increased the diagnostic accuracy to differentiate the BC tissues from normal tissues. However, this combination reported increased sensitivity to predict metastasis but with declined specificity (Tables 4, 5).
Regarding the ASPM gene, an elevated expression level in malignant tissues appeared to be more predominant in MIBC, high-grade tumors, in advanced stages, higher lymph node involvement and found more prevalently in tumors associated with metastasis. This finding indicates tumor progression. Additionally, smoking patients and patients reported to have positive results for Bilharizal antibodies had significant increased ASPM expression level in BC tissues. On the other hand, decreased expression level of the TEF gene in BC tissues was found to have significant prevalence in smoking patients, in MIBC, in high-grade tumors, in advanced (T3 and T4) tumor size, in advanced tumor stages and in tumors having metastatic lesions (Table 6).
Up on evaluating blood indices and kidney function, the ASPM gene expression level was negatively correlated with Hb% concentration, and TEF gene expression level correlated negatively with the urea concentration in serum (Fig. 3a, b).
In multivariate logistic regression, both ASPM expression level, OR 1.337 (1.106–1.617), and TEF expression level, OR 0.090 (0.007–0.953), were predictors for metastasis. Additionally, ASPM expression level, OR 1.970 (1.246–3.115), and smoking, OR 7.591(1.236–46.624), were independent risk factors of MIBC (Table 7).
When applying the Kaplan–Meier survival curve, log rank analyses revealed that high ASPM gene expression and low TEF gene expression in BC tissue were significantly associated with decreased progression-free overall rates (PFS) (P < 0.001) in BC patients (mean: 13.113; 95% CI (LL-UL) 11.627–14.599; mean: 15.389; 95% CI (LL-UL) 13.689–17.088 for ASPM and TEF genes, respectively). Additionally, high ASPM gene expression and low TEF gene expression in BC tissue showed a significant correlation (P < 0.001) with decreased overall survival (OS) in BC patients (mean: 15.945; 95% CI (LL-UL) 14.990–16.900; mean: 17.046; 95% CI (LL-UL) 15.947–18.45 for the ASPM and TEF genes, respectively) (Fig. 4a–d).
a Kaplan–Meier curve for progression-free survival separated based on low or high ASPM gene expression. b Kaplan–Meier curve for progression-free survival separated based on low or high TEF gene expression. c Kaplan–Meier curve for overall survival separated based on low or high ASPM gene expression. d Kaplan–Meier curve for overall survival separated based on low or high TEF gene expression
Discussion
Even though the incidence rate of BC in Africa is considered one of the lowest worldwide, Egypt has a high incidence of BC with elevated mortality rates, especially in men (Antoni et al. 2017). In the current study, we aimed to investigate the expression levels of both the ASPM and TEF genes in BC tissues as potential markers and estimate their value in prognosis and metastatic prediction.
Regarding ASPM gene expression levels in BC and in agreement with our observations, a recent study conducted on BC using six databases by Xu et al. reported the overexpression of ASPM mRNA in neoplastic tissues and found that its overexpression was related to aggressive malignant features, such as elevated tumor grade and progressive stage, according to the GEO and TCGA databases (Xu et al. 2019). Likewise, Tang et al. examined breast cancer using the GEO dataset and found that there was a significant relationship among ASPM expression, breast cancer development, and poor prognosis (Tang et al. 2019). Increased ASPM transcript level was found within prostate cancer tissues and was linked with progressed tumor stage, incidence of metastasis and poor prognosis (Xie et al. 2017). Additionally, the ASPM mRNA reported to be upregulated in hepatocellular carcinoma and correlated with occurrences of metastases and prompt recurrence rates (Lin et al. 2008).
ASPM is overexpressed in various neoplasms with an observable association with the expression levels of frequent biomarkers of cellular proliferation and related to expanded cell growth and tumor progression (Kouprina et al. 2005). Pai et al. explained that ASPM affects cellular proliferation and the ability of prostate cancer cells to invade were because of its interface with disheveled-3, a moderator of the Wnt pathway, repressing its proteasome degeneration, accordingly enhancing Wnt constancy and permitting the Wnt-stimulated β-catenin transcriptional action (Pai et al. 2019).
The analyses of Wang et al. found that elevated concentrations of both ASPM mRNA and its protein were detected in pancreatic ductal adenocarcinoma (PDAC) cell lines and on further inhibition of ASPM transcription in the malignant cells, the tumor showed reduced cellular growth and a limited ability to produce metastasis (Wang et al. 2013). Similarly, in an in vitro study, higher ASPM expression levels were found to be strongly linked to the aggressive behavior of glioma and tumor recurrence, with its suppression resulting in excessive cellular death (Bikeye et al. 2010).
Consistent with our findings, a high ASPM expression level according to median levels demonstrated a significant correlation with poor OS and PFS in BC tissues (Xu et al. 2019), and in breast cancer, the ASPM expression level was inversely correlated with OS and relapse-free survival (Tang et al. 2019). ASPM upregulation in prostate cancer is associated with reduced OS and might biochemically predict recurrence-free survival (Xie et al. 2017). Moreover, in PDAC, ASPM-silenced mice revealed extended survival (Wang et al. 2013).
With regard to the downregulation of TEF transcription in the neoplastic tissue of BC in contrast to normal tissues, Yang et al. (2019) found similar results in their study on BC based on the TCGA database, as they reported reduced TEF expression levels in tumor tissues, which was further confirmed by their own investigations on BC tissues and cell lines.
The transcription of IL-3 receptor β subunit was effectively suppressed by TEF, resulting in blocking cellular multiplication and prompting cell cycle detention in the G0/G1 stage (Inukai et al. 2005). Furthermore, TEF regulates actin apportionment and cellular configuration in fibroblasts (Gutierrez et al. 2011). P53, a tumor suppressor gene that controls cellular growth, appeared to be mutated in a high percentage of neoplasms (Rivlin et al. 2011), and TEF transcription is thought to be regulated by p53 through the overexpression of microRNA‐125b (Gutierrez et al. 2011), which may explain our results that the TEF expression level is associated with BC development.
In this analysis, we reported that low TEF expression levels are related to shorter progression-free survival and overall survival in BC patients. Similarly, Yang et al. found a significant association with worse survival among those patients (Yang et al. 2019).
The expression levels of both the ASPM and TEF mRNA levels were simultaneously evaluated for the first time in this analysis as both genes thought to affect cell cycle progression. ASPM, which controls mitotic spindle formation, is thought to control mitosis extent and induce cell cycle progression by crossing the G1 point (Capecchi and Pozner 2015). On the other hand, TEF found to have a negative effect on cell cycle progression, the increased TEF mRNA is considerably associated with higher cells proportion in the G0/G1 phase but reduced those in the S phase as hindering G1/S transition. While, TEF silencing has the adverse effect (Yang et al. 2019). Additionally the AKT/FOXOs pathway is controlled by TEF specifically FOXO1 and FOXO4, which are supposed to be tumor suppressors due to their anti‐proliferative properties (Maire et al. 2013). FOXO members exert their tumor suppressing effects by means of transfer to the nucleus and affecting specific genes (Maia et al. 2015). Phosphorylation of these molecules resulting in cytoplasmic transfer abolishing their tumor suppressing effects and enhancing tumor proliferation (Brown and Webb 2018) and TEF‐silenced cells showed increased phosphorylation of AKT and FOXO4 and declined phosphorylation was observed with increased TEF mRNA levels (Yang et al. 2019). However the tumor suppressor FOXOs reported to have an inhibitory effect on ASPM expression and the de-repression of ASPM in Foxo knocked cells results in hyper-proliferative state (Paik et al. 2009). In brief we can suppose that ASPM up regulation is associated with enhanced proliferation and cell cycle progression conversely to TEF expression which in agreement with our finding of the upregulation of the ASPM gene and the downregulation of the TEF gene in cancer tissues compared to their expression level in normal bladder tissues and association with advanced tumor stage and aggressive tumor among BC patients.
In our study, we observed negative correlation between TEF expression and urea levels and between ASPM expression and Hb. The pre-operative renal impairment in BC had16.9% prevalence and reported to be associated with tumor recurrence and bad prognosis (Cao et al. 2016). Additionally, Preoperative anemia was correlated with tumor recurrence and might contribute to BC mortality. Hypoxia may prompt modification in gene expression and enhance tumor growth (Chen et al. 2017). The role of TEF and ASPM in these results is not known and need further studies to reveal this association. The limitation of our study was the MIBC predominance.
Conclusion
This work explored the potential value and prognostic role of the expression of both ASPM and TEF in BC and found that ASPM upregulation and TEF downregulation have a significant role in tumor expansion and the incidence of metastasis. Subsequently, ASPM and TEF seem to be worthy markers in the diagnosis and prognosis of BC and should be considered in determination of therapeutic strategies.
References
Amin MB, McKenney JK, Paner GP et al (2013) ICUD-EAU International Consultation on Bladder Cancer 2012: pathology. Eur Urol 63:16–35. https://doi.org/10.1016/j.eururo.2012.09.063
Amin MB, Smith SC, Reuter VE et al (2015) Update for the practicing pathologist: the International Consultation On Urologic Disease-European association of urology consultation on bladder cancer. Mod Pathol 28:612–630. https://doi.org/10.1038/modpathol.2014.158
Antoni S, Ferlay J, Soerjomataram I et al (2017) Bladder cancer incidence and mortality: a global overview and recent trends. Eur Urol 71(1):96–108
Bikeye SNN, Colin C, Marie Y et al (2010) ASPM-associated stem cell proliferation is involved in malignant progression of gliomas and constitutes an attractive therapeutic target. Cancer Cell Int 10:1–9. https://doi.org/10.1186/1475-2867-10-1
Brown AK, Webb AE (2018) Regulation of FOXO factors in mammalian cells. Current topics in developmental biology. Academic Press Inc, Cambridge, pp 165–192
Buchman JJ, Durak O, Tsai LH (2011) ASPM regulates Wnt signaling pathway activity in the developing brain. Genes Dev 25:1909–1914. https://doi.org/10.1101/gad.16830211
Cao J, Zhao X, Zhong Z et al (2016) Prognostic value of pre-operative renal insufficiency in urothelial carcinoma: a systematic review and meta-analysis. Sci Rep. https://doi.org/10.1038/srep35214
Capecchi MR, Pozner A (2015) ASPM regulates symmetric stem cell division by tuning Cyclin E ubiquitination. Nat Commun 6:8763. https://doi.org/10.1038/ncomms9763
Chen C, Hu L, Li X, Hou J (2017) Preoperative anemia as a simple prognostic factor in patients with urinary bladder cancer. Med Sci Monit 23:3528–3535. https://doi.org/10.12659/MSM.902855
Doenhoff MJ, Chiodini PL, Hamilton JV (2004) Specific and sensitive diagnosis of schistosome infection: can it be done with antibodies? Trends Parasitol 20:35–39
Dunphy KM, Garino MC, Shaw NM et al (2017) When the gold standard proves to be fool’s gold-blue-light cystoscopy in a case of high-risk non-muscle-invasive bladder cancer. Urology 110:27–30. https://doi.org/10.1016/j.urology.2017.05.032
Gutierrez O, Berciano MT, Lafarga M, Fernandez-Luna JL (2011) A novel pathway of TEF regulation mediated by microRNA-125b contributes to the control of actin distribution and cell shape in fibroblasts. PLoS ONE 6:e17169. https://doi.org/10.1371/journal.pone.0017169
Hua P, Liu W, Kuo SH et al (2012) Association of Tef polymorphism with depression in Parkinson disease. Mov Disord 27:1694–1697. https://doi.org/10.1002/mds.25195
Inukai T, Inaba T, Dang J et al (2005) TEF, an antiapoptotic bZIP transcription factor related to the oncogenic E2A-HLF chimera, inhibits cell growth by down-regulating expression of the common β chain of cytokine receptors. Blood 105:4437–4444. https://doi.org/10.1182/blood-2004-08-2976
Jiang F, Qi W, Wang Y et al (2019) lncRNA PEG10 promotes cell survival, invasion and migration by sponging miR-134 in human bladder cancer. Biomed Pharmacother 114:108814. https://doi.org/10.1016/j.biopha.2019.108814
Kaufman DS, Shipley WU, Feldman AS (2009) Bladder cancer. Lancet 374:239–249. https://doi.org/10.1016/S0140-6736(09)60491-8
Kouprina N, Pavlicek A, Collins NK et al (2005) The microcephaly ASPM gene is expressed in proliferating tissues and encodes for a mitotic spindle protein. Hum Mol Genet 14:2155–2165. https://doi.org/10.1093/hmg/ddi220
Kwan ML, Garren B, Nielsen ME, Tang L (2019) Lifestyle and nutritional modifiable factors in the prevention and treatment of bladder cancer. Urol Oncol Semin Orig Investig 37:380–386. https://doi.org/10.1016/j.urolonc.2018.03.019
Lavender S, Hilton PJ, Jones NF (1969) The measurement of glomerular filtration-rate in renal disease. Lancet (Lond, Engl) 2:1216–1218. https://doi.org/10.1016/s0140-6736(69)90752-1
Lin SY, Pan HW, Liu SH et al (2008) ASPM\s a novel marker for vascular invasion, early recurrence, and poor prognosis of hepatocellular carcinoma. Clin Cancer Res 14:4814. https://doi.org/10.1158/1078-0432.CCR-07-5262
Maia ARR, De Man J, Boon U et al (2015) Inhibition of the spindle assembly checkpoint kinase TTK enhances the efficacy of docetaxel in a triple-negative breast cancer model. Ann Oncol 26:2180–2192. https://doi.org/10.1093/annonc/mdv293
Maire V, Baldeyron C, Richardson M et al (2013) TTK/hMPS1 Is an attractive therapeutic target for triple-negative breast cancer. PLoS ONE 8:e63712. https://doi.org/10.1371/journal.pone.0063712
Moch H, Humphrey PA, Ulbright TM et al (2016) Tumor of urinary tract. In: Moch H, Humphrey PA, Ulbright TM, Reuter V (eds) WHO Classification of Tumours of the Urinary System and Male Genital Organs, 4th edn. International Agency for Research on Cancer, Lyon, pp 78–133
Pai VC, Hsu CC, Chan TS et al (2019) ASPM promotes prostate cancer stemness and progression by augmenting Wnt−Dvl-3−β-catenin signaling. Oncogene 38:1340–1353. https://doi.org/10.1038/s41388-018-0497-4
Paik J, Ding Z, Narurkar R et al (2009) FoxOs cooperatively regulate diverse pathways governing neural stem cell homeostasis. Cell Stem Cell 5:540–553. https://doi.org/10.1016/j.stem.2009.09.013
Rivlin N, Brosh R, Oren M, Rotter V (2011) Mutations in the p53 tumor suppressor gene: important milestones at the various steps of tumorigenesis. Genes Cancer 2:466–474
Rosenkrantz AB, Ego-Osuala IO, Khalef V et al (2016) Investigation of multisequence magnetic resonance imaging for detection of recurrent tumor after transurethral resection for bladder cancer. J Comput Assist Tomogr 40:201–205. https://doi.org/10.1097/RCT.0000000000000363
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin 68:7–30. https://doi.org/10.3322/caac.21442
Tang J, Lu M, Cui Q et al (2019) Overexpression of ASPM, CDC20, and TTK confer a poorer prognosis in breast cancer identified by gene co-expression network analysis. Front Oncol 9:1–14. https://doi.org/10.3389/fonc.2019.00310
Torre LA, Bray F, Siegel RL et al (2015) Global cancer statistics, 2012. CA Cancer J Clin 65:87–108. https://doi.org/10.3322/caac.21262
Wang E, Miller LD, Ohnmacht GA et al (2000) High-fidelity mRNA amplification for gene profiling. Nat Biotechnol 18:457–459. https://doi.org/10.1038/74546
Wang WY, Hsu CC, Wang TY et al (2013) A gene expression signature of epithelial tubulogenesis and a role for ASPM in pancreatic tumor progression. Gastroenterology 145:1110–1120. https://doi.org/10.1053/j.gastro.2013.07.040
Xie J-J, Zhuo Y-J, Zheng Y et al (2017) High expression of ASPM correlates with tumor progression and predicts poor outcome in patients with prostate cancer. Int Urol Nephrol 49:817–823. https://doi.org/10.1007/s11255-017-1545-7
Xu Z, Zhang QI, Luh F et al (2019) Overexpression of the ASPM gene is associated with aggressiveness and poor outcome in bladder cancer. Oncol Lett 17:1865–1876. https://doi.org/10.3892/ol.2018.9762
Yang J, Wang B, Chen H et al (2019) Thyrotroph embryonic factor is downregulated in bladder cancer and suppresses proliferation and tumorigenesis via the AKT/FOXOs signalling pathway. Cell Prolif 52:1–13. https://doi.org/10.1111/cpr.12560
Funding
There was no funding for this analysis.
Author information
Authors and Affiliations
Contributions
All the authors have contributed to the study to an adequate degree to be named as authors. AAS and SES performed the lab investigation and the molecular analysis and selected the study design. SG and ME were responsible for samples and data collections and evaluation of the involved patients. AS Performed the pathological examinations and all authors participate in writing and revision of the paper and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflict of interest.
Ethical Approval
This work was performed based on the Declaration of Helsinki and the principles of the “Ethical Committee of Medical Research”, Faculty of Medicine, Menoufia University. A written consent was provided by all participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saleh, A.A., Gohar, S.F., Hemida, A.S. et al. Evaluation of ASPM and TEF Gene Expressions as Potential Biomarkers for Bladder Cancer. Biochem Genet 58, 490–507 (2020). https://doi.org/10.1007/s10528-020-09962-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-020-09962-1