Abstract
Macrolophus pygmaeus and M. melanotoma (Hemiptera: Miridae) are biological control agents used in greenhouse crops, the former preferring plants of the Solanaceae family and the latter the aster Dittrichia viscosa. The discrimination of these species is of high significance for effective biological pest control, but identification based on morphological characters of the host plant is not always reliable. In this study, sequencing analysis of mitochondrial gene segments 12S rDNA and COI has been combined with crossing experiments and morphological observations to develop new markers for Macrolophus spp. discrimination and to provide new data on their genetic variability. This is the first comprehensive research in Greece on M. pygmaeus and M. melanotoma genetic variability based on sequencing data from 12S rDNA and COI gene segments. The relationship of this variability to host plant preference must be investigated in an agricultural ecosystem.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The main goals of classical biological control are to introduce and establish natural enemies that will effectively control pest populations on plants. Better knowledge of natural enemies and pest groups helps to increase rates of introduction and success of biological control agents (Van Driesche et al. 2008). The correct species classification of a natural enemy is essential for successful application in biological pest control programs (Gordh and Bearsley 1999).
The genus Macrolophus (Hemiptera: Miridae) consists of several omnivorous predatory Palaearctic species, among which Macrolophus pygmaeus (Rambur 1839), M. melanotoma (Costa 1853) (= caliginosus Wagner 1951), and M. costalis Fieber 1858, are used in biological control of crop pests. Macrolophus pygmaeus has been reported on plants of the Solanaceae family, such as Solanum nigrum L. or Solanum lycopersicum L. (Solanales: Solanaceae) (Lykouressis et al. 2000; Castañé et al. 2004), and M. melanotoma is found on Dittrichia viscosa L. (Greuter 1973) (Asterales: Asteraceae) (Josifov 1992; Alomar et al. 1994; Lykouressis et al. 2000; Ingegno et al. 2011; Machtelinckx et al. 2012).
The prey catalog of the genus Macrolophus includes many arthropods that are major pests, such as whiteflies (Trialeurodes vaporariorum Westwood, 1856; Hemiptera: Aleyrodidae), aphids [Myzus persicae (Sulzer 1776), Hemiptera: Aphididae], thrips [Frankliniella occidentalis (Pergande 1895), Thysanoptera: Thripidae], and mites (Tetranychus urticae Koch 1836, Acari: Tetranychidae) (Malausa and Trottin 1996; Perdikis and Lykouressis 2000; Enkegaard et al. 2001; Hillert et al. 2002; Margaritopoulos et al. 2003; Blaeser et al. 2004; Vandekerkhove et al. 2006; Mohd et al. 2009). Macrolophus spp. can also feed either on leaves, sucking their sap (Hommes and Ter Horst 2002; Margaritopoulos et al. 2003; Perdikis and Lykouressis 2004), or on pollen or nectar from plants (Perdikis and Lykouressis 2000; Vandekerkhove and De Clercq 2010; Portillo et al. 2012). Feeding on alternative food, when prey is not available, supports the predators’ survival for a relatively long period (Perdikis and Lykouressis 2000, 2004; Portillo et al. 2012). This enhances their establishment on crops and their potential in biological control (Ehler 1990; Gnanvossou et al. 2005).
Many studies have shown the importance of Macrolophus species as biological control agents on open (outdoors) or protected (greenhouse) cultivation ecosystems (Avilla et al. 2004; Perdikis et al. 2008). These species colonize field and greenhouse crops originating from habitats near the agricultural landscape of the southern region of the European Mediterranean area (Lykouressis et al. 2000; Castañé et al. 2004; Gabarra et al. 2004; Perdikis et al. 2011).
Macrolophus pygmaeus and M. melanotoma are characterized by a simple and similar external morphology. Their classification has been based on the variability of characters such as body size, the height of the black band-shaped macula behind the eyes, and the color (totally black or black with a white zone) of the first antennal segment (FAS). Discrimination of these species based on morphological characters includes a great number of misconceptions and is considered problematic (Josifov 1992; Perdikis et al. 2003; Martinez-Cascales et al. 2006). Indeed, M. pygmaeus on tomatoes has been recently misidentified as M. melanotoma, and it was named M. caliginosus for a long period after this was ascertained by researchers or commercial producers of beneficial insects (Castañé et al. 2011; Pumarino et al. 2011). Therefore, identification of the two species based on the host plants in correlation with their morphological characters does not promote high confidence, as it is also known that both morphological types based on FAS color may coexist on the same host plant.
To resolve taxonomy problems, researchers have applied molecular techniques to provide additional reliable information (Berlocher 1984; Simon et al. 1994; Caterino et al. 2000; Yao et al. 2010). By virtue of its simple structure, maternal inheritance, and relatively rapid evolutionary rates, mitochondrial DNA (mtDNA) has become a widely used marker for understanding the structure of animal populations and their genetic variability (Harrison 1989; Hebert and Gregory 2005; Jung et al. 2011; Park et al. 2011).
Barcoding has been developed as a standardized method that is cost- and time-effective to separate or identify species (Hebert et al. 2003). As the number of DNA barcode sequences increases, DNA barcoding is expected to improve studies of systematics, molecular phylogenetics, and population genetics over conventional approaches in DNA, especially when used in combination with additional information such as geographic distribution and genetic diversity (Hajibabaei et al. 2007; Valentini et al. 2009; Jung et al. 2011).
Perdikis et al. (2003) made the first attempt to discriminate M. melanotoma and M. pygmaeus using restriction fragment length polymorphism (RFLP) and randomly amplified polymorphic DNA-polymerase chain reaction (RAPD-PCR) methods. Then, Martinez-Cascales et al. (2006) used sequencing analysis of a cytochrome b mtDNA gene segment as a marker to discriminate these two species. The development of new markers is necessary because there is still some doubt concerning their efficiency.
The aim of this investigation is to contribute to the taxonomic identification of M. melanotoma and M. pygmaeus, using sequencing analysis of cytochrome oxidase subunit I (COI) and 12S ribosomal DNA (12S rDNA) mtDNA gene segments combined with crossing experiments and morphological observations of Macrolophus spp. populations collected from various locations in Greece. The results also provide information regarding the genetic variability of Macrolophus spp. in Greece, as well as some further information of the host-plant preference of these species.
Materials and Methods
Sampling
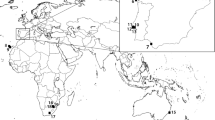
The sampling and preliminary identification of Macrolophus species were based on the hypothesis that they are found on different host plants (Perdikis et al. 2003; Martinez-Cascales et al. 2006). Macrolophus pygmaeus has seldom been reported on D. viscosa, with the exception of a single specimen, according to Machtelinckx et al. (2012). Samples were collected from Solanum host plants in four locations in central and southern Greece and one location in Spain, and from Dittrichia host plants in four Greek locations (Fig. 1; Table 1).
Sampling locations for populations of Macrolophus pygmaeus and M. melanotoma collected in Greece and Spain. Each population consists of insects collected from 5 plants of Solanum nigrum or S. lycopersicum for M. pygmaeus (gray dots) or Dittrichia viscosa for M. melanotoma (black dots). Both species were collected in Kyparissia
At each location, about 20 insects of different ages and both sexes were collected from five different plants (Table 1). Live insects were transferred to the Laboratory of Agricultural Zoology and Entomology and reared on potted plants of the same species as the original host plant (S. nigrum, S. lycopersicum, or D. viscosa). The potted plants were grown in plastic cylindrical cages (11 cm diameter, 30 cm height) under conditions of stabilized temperature (25 ± 1°C), relative humidity (65 ± 5), and photoperiod (16 h light:8 h dark). Eggs of Ephestia kuehniella Zeller (Lepidoptera: Pyralidae) (Entofood–Koppert B.V.) were provided daily as prey. The insects, after completing at least two generations in the cages, were kept for further use in our experiments (morphometric observations, crossing experiments, and sequencing analysis).
Morphometric Observations
Taking into consideration the variation of the ambiguous morphological characters that have been used so far to discriminate between these species (Josifov 1992; Perdikis et al. 2003; Martinez-Cascales et al. 2006), we observed the FAS color (totally black or black with a white zone). Other characters, such as the black macula, were not clearly visible (Martinez-Cascales et al. 2006). Ten insects (five males and five females) were collected from the rearings described (40 M. melanotoma and 50 M. pygmaeus). The FAS color was examined before every testing procedure (crossing experiments and DNA sequencing), causing the minimum inconvenience to the insect. The results were expressed as percentages.
Crossing Experiments
In order to examine whether the insects sampled for study belonged to separate species, we performed several crossing experiments according to Perdikis et al. (2003). Samples of immature insects of Macrolophus spp. were collected from the two rearings maintained in our laboratory: (1) M. pygmaeus from S. nigrum collected at Votanikos on June 2009 and (2) M. melanotoma from D. viscosa found at Spata on June 2009. The insects developed individually in a petri dish containing a layer of wet cotton on which a leaf of each respective host plant was placed together with eggs of E. kuehniella. The FAS color of each adult was recorded before its use in the experiments, causing the least disturbance to the insect.
For the insects assigned to M. pygmaeus, only adults with totally black FAS were recorded. For the M. melanotoma insects, adults with totally black FAS were designated B and those with white in the middle of the FAS were designated W. Thus, crossings among the three adult types were performed (Table 2).
Two adult males and two adult females (P generation) of each morphological type were placed in a plastic cylindrical cage containing a plant of either D. viscosa or S. nigrum. Crossings were also performed using the bean Phaseolus vulgaris var. Zargana (Fabaceae). It is known that the genus Macrolophus is able to survive and produce individuals on beans (Tavella and Arzone 1996), but generally Macrolophus are not common on this host plant. For this reason, this plant was selected as a neutral host, so neither of the two species was in a favorable state. Each of the three morphological types (male and female) was crossed with the other types on each of the three host plants (Table 2).
As soon as the first offspring emerged, a second crossing took place. Again, two males and two females (F1 generation) were placed on the host plant. If this second crossing produced an F2 generation (confirming that the F1 generation was fertile), the implicated insects were recorded as belonging to the same species. The possibility of failure of the crossings as a consequence of the presence of any endosymbiotic organism (e.g., Wolbachia spp.; Machtelinckx et al. 2009) is negligible, as the insects were taken from rearings where no abnormality in their reproduction rate was observed. The experiment was performed twice under controlled conditions of photoperiod (16 h light, 8 h dark), temperature (25 ± 1°C), and relative humidity (65 ± 5%).
Molecular Analysis
Female adults of Macrolophus spp. (about 6 per location) were collected alive from the colonies reared in the laboratory and stored at −20°C or in 95% ethanol until used. Total DNA was extracted for each insect using the DNeasy Blood and Tissue Kit (Qiagen) following the manufacturer’s protocol with minor modifications. Fragments of the two mtDNA gene segments were amplified using two sets of primers: 5′-ΑΑΑCΤGGGΑΤΤΑGΑΤΑCCCCΑCΤΑΤ-3′ and 5′-GAGGGTGACGGGCGGTGTGT-3′ for the 12S rDNA gene segment (Palumbi et al. 1991) and the Barcode primers 5′-GGTCAACAAATCATAAAGATATTGG-3′ and 5′-TAAACTTCAGGGTGACCAAAAAATCA-3′ for the COI segment (Folmer et al. 1994).
The PCR for the 12S rDNA segment consisted of 2.4 U Taq polymerase (Invitrogen), 5 μL 10× reaction buffer (Invitrogen), 5 μL dNTP mix (2 mM) (BioLabs), 5 μL MgCl2 (50 mM) (Invitrogen), approximately 100 ng DNA, 0.5 μL of each primer (20 mM) (BioLabs), and sterile water to a total volume of 50 μL. The mixture for amplification of the COI fragment consisted of 1.5 U Taq polymerase (Invitrogen), 5 μL 10× reaction buffer (Invitrogen), 5 μL dNTP mix (2 mM) (BioLabs), 3 μL MgCl2 (50 mM) (Invitrogen), approximately 100 ng DNA, 0.5 μL of each primer (10 μM) (BioLabs), and sterile water to a total volume of 50 μL. Both fragments were amplified using a PEQlab Thermocycler (Primus 25 Advanced). The conditions for 12S rDNA were one preliminary denaturation step at 94°C for 4 min, followed by 35 cycles at 94°C for 1 min, annealing at 44°C for 1 min, and final extension at 72°C for 2 min. For COI, the Barcode protocol was used, with one step of preliminary denaturation at 94°C for 1 min, followed by two groups of cycles. The first group consisted of 6 cycles at 94°C for 1 min denaturation, annealing at 45°C for 1 min 30 s, and extension at 72°C for 1 min 15 s. The second group included 36 cycles at 94°C for 1 min denaturation, annealing at 51°C for 1 min 30 s, and extension at 72°C for 1 min 15 s, and a final extension at 72°C for 5 min (Hebert et al. 2003).
The PCR products were purified using the Nucleospin Extract II kit (Macherey–Nagel), according to the supplier’s protocol, and examined using agarose gel electrophoresis. Each sequence was determined via the automated sequencing procedure provided by the Macrogen Company (Seoul, Korea), using the same primers as the amplification procedure. The sequences obtained have been deposited in Genbank for each gene segment separately, with accession numbers HQ707820-HQ707825 (M. melanotoma) and HQ845353-HQ845364 (M. pygmaeus) for 12S rDNA, and HQ707826-HQ707832 (M. melanotoma) and HQ845336-HQ845352 (M. pygmaeus) for COI.
Data and Statistical Analyses
Multiple sequences were aligned using Clustal W2 (Thompson et al. 1994), and they were further checked manually. The authenticity of the mtDNA sequences produced was verified as species of the Miridae family using the Blast program of NCBI (http://blast.ncbi.nlm.nih.gov/). Each gene fragment was analyzed separately. The pairwise genetic distance (p-distance) of the Kimura 2-parameter model (Kimura 1980) was estimated using Mega 5.0 (Tamura et al. 2011). Haplotypes, as well as the number of variable sites and nucleotide diversity, were detected using the DnaSP version 5.10.00 package (Librado and Rozas 2009).
Neighbor-joining, minimum evolution, maximum parsimony, and UPGMA analyses were conducted for each gene separately, using the Mega 5.0 software. The partial mitochondrial sequence that belongs to Lygus lineolaris (Hemiptera: Miridae) was downloaded from NCBI (EU401991) and used as an outgroup for both gene segments, as it contains both of them. The part of the EU401991 sequence that was not similar to either COI or 12S rDNA was excluded and was not used in the analysis. Confidence in the nodes was evaluated by 1,000 bootstrap replicates (Felsenstein 1985).
Results
Based on our morphometric observations (Fig. 2), the FAS color of the M. pygmaeus insects is totally black, but for M. melanotoma this character is not constant, with 35% showing a black FAS and 65% showing a white zone. Thus, the discrimination of M. pygmaeus and M. melanotoma cannot be based on this character.
Morphological types observed in two species of Macrolophus. The first antennal segment (FAS) was observed in five male and five female insects from each sampling location (Fig. 1). All of the M. pygmaeus (top) were found to have a totally black FAS. The M. melanotoma populations (bottom) included both types, totally black or black with a white zone
The results of the crossing experiments (Table 2) do not appear to be affected by the host species. No crosses between M. pygmaeus and M. melanotoma were fertile, confirming that the populations tested belong to two different species.
The 45 mtDNA sequences produced a 328 bp alignment for the 12S rDNA gene segment and revealed 18 haplotypes, 14 of which were unique (Table 3). For COI, the alignment was 621 bp, revealing 24 haplotypes, 21 of which were unique (Table 4). The percentage of variable sites was 11.59% for 12S rDNA (Supplementary Table 1), with an average pairwise genetic distance of 0.044, and 21.58% for COI (Supplementary Table 2), with 0.081 average pairwise genetic distance. That means the COI segment has higher variability.
The greatest divergence (0.112) in 12S rDNA was observed between the populations of M. pygmaeus at Votanikos and M. melanotoma at Penteli, Spata, and Kalamata. In COI the greatest divergence was 0.192, between the two species of Macrolophus at Kyparissia. The haplotype diversity was 0.847 for the 12S rDNA gene segment and 0.914 for COI. The nucleotide diversity was 0.04 for 12S rDNA and 0.07 for COI.
The phylogenetic trees based on minimum evolution, maximum parsimony, neighbor-joining, and UPGMA analyses exhibited the same topology for 12S rDNA and COI. For this reason, only the minimum evolution tree for 12S rDNA and the maximum parsimony tree for COI are presented here (Fig. 3). The sequences corresponding to M. pygmaeus are grouped together in the same main clade, and those of M. melanotoma are grouped in the second clade, according to the topology of all dendrograms constructed, based on the sequencing data obtained either from COI or 12S rDNA, clearly demonstrating the discrimination between both species. It is noted that the specimens of M. melanotoma separate into two groups. The suggested topology is supported by high bootstrap values.
Phylogenetic relationships of M. pygmaeus and M. melanotoma collected from locations in Greece and Spain. a Minimum evolution tree based on sequencing data obtained from the 12S rDNA gene segment (45 sequences with sequence length 328 bp). b Maximum parsimony tree based on sequencing data obtained from the COI-Barcode gene segment (48 sequences with sequence length 621 bp). Lygus lineolaris (EU401991) was used as outgroup. Location codes as in Table 1. Bootstrap numbers appear to the left of each node
Discussion
The aim of this research is to contribute to the discrimination and identification of two major species of the genus Macrolophus used as biocontrol agents. The discrimination of M. pygmaeus and M. melanotoma has been based on morphological characters of dubious value (Alomar et al. 1994; Lykouressis et al. 2000; Perdikis et al. 2003; Castañé et al. 2004; Martinez-Cascales et al. 2006). Our results further confirm that the morphometric characters cannot be considered diagnostic (Perdikis et al. 2003; Martinez-Cascales et al. 2006); the FAS color, which was supposed to be the best visible character used for distinguishing the two species, does not provide further information, and there is a risk of misidentification.
The crossing experiments confirmed that M. pygmaeus and M. melanotoma are different species. The F2 generation was produced only when M. pygmaeus was mated with M. pygmaeus and M. melanotoma with M. melanotoma, regardless of the host plant on which the insects were crossed. In some cases, it seems that the host plant can be a preliminary basis for discriminating between species, and that M. pygmaeus may have a broader host-plant range than M. melanotoma (Lykouressis et al. 2000; Castañé et al. 2004; Ingegno et al. 2011; Machtelinckx et al. 2012).
The topology of the phylogenetic trees (all of which grouped the two species into two separate clades) shows that they belong to distinct species (Fig. 3). This is also confirmed by the grouping of samples from the same host plant together, regardless of the samples’ countries of origin (Greece and Spain). So, it was shown that populations on different genera of host plants belong to different species: M. pygmaeus on S. nigrum or S. lycopersicum and M. melanotoma on D. viscosa.
Both gene segments analyzed in this study (12S rDNA and COI) can be used for species discrimination because they show sufficient sequence diversity (Ferri et al. 2009; Allegrucci et al. 2011; Muráriková et al. 2011). The highest genetic divergence values between the two mtDNA segments were observed in COI, which may be due to its higher evolutionary rate (Lunt et al. 1996). Nevertheless, there is no clear separation within the populations of each species, even in the case of M. melanotoma collected from the location of Kyparissia; the Kyparissian samples seem to be somehow separated from the other populations, as three of the specimens from the same location (mKYP-1, mKYP-2, mKYP-3) are included in another cluster (Fig. 3). It is not possible, therefore, to detect the location where every individual was found.
Comparing the results of the present research with the others, the sequencing analysis is more accurate than the RAPD-PCR used by Perdikis et al. (2003). Two mtDNA gene segments were analyzed in our study, with a total length of 940 bp. These two mtDNA gene segments can be applied separately for the discrimination of Macrolophus species. This is the first time in the literature that sequencing data for the 12S rDNA mtDNA gene segment are provided for Macrolophus species.
The data obtained from the current research combines different approaches to provide new molecular markers for the discrimination of these species, to contribute to the knowledge of the genetic structure of Macrolophus species, and to detect the genetic variability of each species that occurs in Greece.
Further investigation will be necessary to determine if the intraspecies variability detected is related to the host plant or to other factors. This kind of information is essential in understanding the evolution and the relationships among populations, and it might contribute to the conservation of the insect populations.
References
Allegrucci G, Trucchi E, Sbordoni V (2011) Tempo and mode of species diversification in Dolichopoda cave crickets (Orthoptera, Rhaphidophoridae). Mol Phylogenetics Evol 60:108–121
Alomar O, Goula M, Albajes R (1994) Mirid bugs for biological control: identification, survey in non cultivated winter plants, and colonization of tomato fields. IOBC/WPRS Bull 17:217–223
Avilla J, Albajes R, Alomar O, Castañé C, Gabarra R (2004) Biological control of whiteflies on vegetable crops. In: Heinz KM, Van Driesche RG, Parrella MP (eds) Biocontrol in protected culture. Ball Publishing, Batavia, Ill., pp 171–184
Berlocher SH (1984) Insect molecular systematics. Ann Rev Entomol 29:403–433
Blaeser P, Sengonca C, Zegula T (2004) The potential use of different predatory bug species in the biological control of Frankliniella occidentalis (Pergande) (Thysanoptera: Thripidae). J Pest Sci 77:211–219
Castañé C, Alomar O, Goula M, Gabarra R (2004) Colonization of tomato greenhouses by the predatory mirid bugs Macrolophus caliginosus and Dicyphus tamaninii. Biol Control 30:591–597
Castañé C, Arnó J, Gabarra R, Alomar O (2011) Plant damage to vegetable crops by zoophytophagous mirid predators. Biol Control 59:22–29
Caterino MS, Cho S, Sperling FAH (2000) The current state of insect molecular systematics: a thriving Tower of Babel. Ann Rev Entomol 45:1–54
Costa A (1853) Cimicum Regni Neapolitani. Centuria tertia et quartae fragmentum (Reprint from Atti del Reale Istituto d'Incoraggiamento alle Scienze Naturali di Napoli 8(1855):225–299)
Ehler LE (1990) Introduction strategies in biological control of Insects. In: Mackauer M, Ehler LE, Roland J (eds) Critical issues in biological control. Intercept Limited, UK, pp 111–134
Enkegaard A, Brødsgaard HF, Hansen DL (2001) Macrolophus caliginosus: functional response to whiteflies and preference and switching capacity between whiteflies and spider mites. Entomol Exp Appl 101:81–88
Felsenstein J (1985) Confidence-limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Ferri E, Barbuto M, Bain O, Galimberti A, Uni S, Guerrero R, Ferté H, Bandi C, Martin C, Casiraghi M (2009) Integrated taxonomy: traditional approach and DNA barcoding for the identification of filarioid worms and related parasites (Nematoda). Front Zool 6:1
Fieber FX (1858) Criterien zur generischen Theilung der Phytocoriden (Capsini aut.). Wiener Entomologische Monatschrift 2:289–327, 329–347, 388
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3:294–299
Gabarra R, Alomar O, Castañé C, Goula M, Albajes R (2004) Movement of greenhouse whitefly and its predators between in and outside of Mediterranean greenhouses. Agric Ecosyst Environ 102:341–348
Gnanvossou D, Hanna R, Yaninek JS, Toko M (2005) Comparative life history traits of three neotropical phytoseiid mites maintained on plant-based diets. Biol Control 35:32–39
Gordh G, Bearsley JH (1999) Taxomomy and biological control. In: Bellows TS, Fischer TW (eds) Handbook of biological control. Academic Press, San Diego, pp 45–56
Greuter W (1973) In: Greuter W, von Raab-Straube E (eds) Compositae. Euro+Med Plantbase—the information resource for Euro-Mediterranean plant diversity
Hajibabaei M, Singer GAC, Hebert PDN, Hickey DA (2007) DNA barcoding: how it complements taxonomy, molecular phylogenetics and population genetics. Trends Genet 23:167–172
Harrison RG (1989) Animal mitochondrial DNA as a genetic marker in population and evolutionary biology. Trends Ecol Evol 4:6–11
Hebert PD, Gregory TR (2005) The promise of DNA barcoding for taxonomy. Syst Biol 54:852–859
Hebert P, Cywinska A, Ball S, deWaard J (2003) Biological identifications through DNA barcodes. Proc Royal Soc Lond B Biol Sci 270:313–321
Hillert O, Jäckel B, Plate HP (2002) Macrolophus pygmaeus (Rambur 1839) (Heteroptera: Miridae): ein interressanter Nützling im biologischen Pflanzenschutz. Gesunde Pflanzen 54:66–73
Hommes M, Ter Horst S (2002) Development and life span of Macrolophus pygmaeus Rambur at different temperatures and influence of host plants and prey. IOBC/WPRS Bull 25:103–106
Ingegno BL, Pansa MG, Tavella L (2011) Plant preference in the zoophytophagous generalist predator Macrolophus pygmaeus (Heteroptera: Miridae). Biol Control 58:174–181
Josifov M (1992) Zur taxonomie der palaearktischen Macrolophus-Arten (Insecta, Heteroptera, Miridae). Reichenbachia 29:1–4
Jung S, Duwal RK, Lee S (2011) COI barcoding of true bugs (Insecta, Heteroptera). Mol Ecol Res 11:266–270
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Koch CL (1836) Deutschlands Crustaceen, Myriapoden und Arachniden. Ein Beitrag zur deutschen Fauna. Friedrich Pustet, Regensburg
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Lunt DH, Zhang DX, Szymura JM, Hewitt GM (1996) The insect cytochrome oxidase I gene: evolutionary patterns and conserved primers for phylogenetic studies. Insect Mol Biol 5:153–165
Lykouressis D, Perdikis D, Tsagarakis A (2000) Polyphagous mirids in Greece: host plants and abundance in traps placed in some crops. Boll Lab Entomol Agrar F S 56:57–68
Machtelinckx T, Van Leeuwen T, Vanholme B, Gehesquiere B, Dermauw W, Vandekerkhove B, Gheysen G, De Clercq P (2009) Wolbachia induces strong cytoplasmic incompatibility in the predatory bug Macrolophus pygmaeus. Insect Mol Biol 18:373–381
Machtelinckx T, Van Leeuwen T, Van De Wiele T, Boon N, De Vos WH, Sanchez JA, Nannini M, Gheysen G, De Clercq P (2012) Microbial community of predatory bugs of the genus Macrolophus (Hemiptera: Miridae). BMC Microbiol 12:S9
Malausa JC, Trottin CY (1996) Advances in the strategy of use of the predaceous bug Macrolophus caliginosus (Heteroptera: Miridae) in glasshouse crops. In: Alomar O, Wiedenmann RN (eds) Zoophytophagous Heteroptera: Implications for life history and integrated pest management. Thomas Say Publications in Entomology, Md, USA, pp 178–189
Margaritopoulos JT, Tsitsipis JA, Perdikis DCh (2003) Biological characteristics of the mirids Macrolophus costalis and Macrolophus pygmaeus preying on the tobacco form of Myzus persicae (Hemiptera: Aphididae). Bull Entomol Res 93:39–45
Martinez-Cascales JI, Cenis JL, Cassis G, Sanchez JA (2006) Species identity of Macrolophus melanotoma (Costa 1853) and Macrolophus pygmaeus (Rambur 1839) (Insecta: Heteroptera: Miridae) based on morphological and molecular data and bionomic implications. Insect Syst Evol 37:385–404
Mohd RZ, Fauziah I, Wan Mohamad WAK, Salmah MR, Kamaruzaman J (2009) Biology of Macrolophus caliginosus (Heteroptera: Miridae) predator of Trialeurodes vaporariorum (Homoptera: Aleyrodidae). Int J Biol 1:63–70
Muráriková N, Vaňhara J, Tóthová A, Havel J (2011) Polyphasic approach applying artificial neural networks, molecular analysis and postabdomen morphology to West Palaearctic Tachina spp. (Diptera, Tachinidae). Bull Entomol Res 101:165–175
Palumbi S, Martin A, Romano S, McMillan WO, Stice L, Grabowski G (1991) The simple fool’s guide to PCR, v 2.0. University of Hawaii, Hawaii, p 47
Park DS, Foottit R, Maw E, Hebert PD (2011) Barcoding bugs: DNA-based identification of the true bugs (Insecta: Hemiptera: Heteroptera). PLoS ONE 6:e18749
Perdikis D, Lykouressis D (2000) Effects of various items, host plants and temperatures on the development and survival of Macrolophus pygmaeus Rambur (Hemiptera: Miridae). Biol Control 17:55–60
Perdikis D, Lykouressis D (2004) Myzus persicae (Homoptera: Aphididae) as a suitable prey for Macrolophus pygmaeus (Hemiptera: Miridae) population increase on pepper plants. Environ Entomol 33:499–505
Perdikis DCh, Margaritopoulos JT, Stamatis C, Mamuris Z, Lykouressis DP, Tsitsipis JA, Pekas A (2003) Discrimination of the closely related biocontrol agents Macrolophus melanotoma (Hemiptera: Miridae) and M. pygmaeus using mitochondrial DNA analysis. Bull Entomol Res 93:507–514
Perdikis D, Kapaxidi E, Papadoulis G (2008) Biological control of insect and mite pests in greenhouse solanaceous crops. Eur J Plant Sci Biotech 2:125–144
Perdikis D, Fantinou A, Lykouressis D (2011) Enhancing pest control in annual crops by conservation of predatory Heteroptera. Biol Control 59:13–21
Pergande T (1895) Observation on Certain Thripidae. Insect Life 7:390–395
Portillo N, Alomar O, Wäckers F (2012) Nectarivory by the plant-tissue feeding predator Macrolophus pygmaeus Rambur (Heteroptera: Miridae): nutritional redundancy or nutritional benefit? J Insect Physiol 58:397–401
Pumarino L, Alomar O, Agustí N (2011) Development of specific ITS markers for plant DNA identification within herbivorous insects. Bull Entomol Res 101:271–276
Rambur MP (1839) Faune entomologique de l'Andalousie, vol 2. Arthus Bertrand, Paris, pp 97–176
Simon C, Frati F, Bechenbach A, Crespi B, Liu H, Flook P (1994) Evolution, weighting, and phylogenetic utility of mitochondrial gene sequence and compilation of conserved polymerase chain reaction primers. Ann Entomol Soc Am 87:651–701
Sulzer JH (1776) Abgekürzte Geschichte der Insecten nach dem Linnaeischen System, vol 1. H. Steiner u. Comp., Winterthur, p 105
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) Mega 5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tavella L, Arzone A (1996) Development of Macrolophus caliginosus and Dicyphus errans on different diets (Rynchota: Miridae). Proc 20th International Congress on Entomology, Firenze, Italy, pp 25–31
Thompson J, Higgins D, Gibson T (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680
Valentini A, Pompanon F, Taberlet P (2009) DNA barcoding for ecologists. Trends in Ecol Evol 24:110–117
Van Driesche R, Hoddle M, Center T (2008) Control of pests and weeds by natural enemies: an introduction to biological control. Wiley-Blackwell, Malden, p 484
Vandekerkhove B, De Clercq P (2010) Pollen as an alternative or supplemental food for the mirid predator Macrolophus pygmaeus. Biol Control 53:238–242
Vandekerkhove B, Van Baal E, Bolckmans K, De Clercq P (2006) Effect of diet and mating status on ovarian development and oviposition in the polyphagous predator Macrolophus caliginosus (Heteroptera: Miridae). Biol Control 39:532–538
Wagner E (1951) Contributo alla conoscenza della fauna emitterologica italiana. II. Zwei neue Miriden- Arten und eine bisher übersehene Art aus Italien (Hem. Het.). B. Macrolophus caliginosus n. sp., eine neue Miridenart aus Italien (Hem. Het.). Bollettino dell’Associazione Romana di Entomologia 5:29–31
Westwood JO (1856) The new Aleyrodes of the greenhouse. Gardeners’ Chron. 1856:852
Yao H, Song J, Liu C, Luo K, Han J, Li Y, Pang X, Xu H, Zhu Y, Xiao P, Chen S (2010) Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS ONE 5:e13102
Acknowledgments
We would like to thank Dr. Antonio Sanchez and Mr. Antonio Thomadakis for kindly providing the Macrolophus samples from Spain and Crete (Greece), respectively.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Evangelou, V.I., Bouga, M., Emmanouel, N.G. et al. Discrimination of Two Natural Biocontrol Agents in the Mediterranean Region Based on Mitochondrial DNA Sequencing Data. Biochem Genet 51, 825–840 (2013). https://doi.org/10.1007/s10528-013-9610-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-013-9610-0