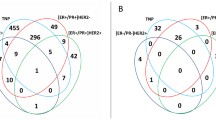
MicroRNA whose expression depends on the receptor status of breast cancer were selected using bioinformatic analysis. The expression of 9 microRNAs (16, 17, 21, 27, 125, 146, 155, 200a, and 221) was analyzed in 76 samples of breast cancer with various receptor phenotypes. The expression of microRNAs 155, 27, and 200a did not differ in various types of breast cancer. The data on positive correlation between the expression of microRNA-21 and microRNA-221 and negative receptor status of the tumor were confirmed. The expression of the tumor suppressing microRNA-125b decreased in samples of breast cancer expressing HER2 and ER and in triple negative breast cancer, which characterizes it as a universal marker of breast cancer. An increase in the expression of microRNA-16 was shown in samples of breast cancer expressing HER2 and ER. The expression of microRNA-17 decreased in triple negative breast cancer and increased in ER+, PR+, and HER+ types of breast cancer. MicroRNAs 16, 17, 21, 125b, 146b, and 221 can be promising markers for differential diagnostics of various phenotypes of breast cancer.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M, Wojcik SE, Aqeilan RI, Zupo S, Dono M, Rassenti L, Alder H, Volinia S, Liu CG, Kipps TJ, Negrini M, Croce CM. miR-15 and miR-16 induce apoptosis by targeting BCL2 Proc. Natl Acad. Sci. USA. 2005;102(39):13,944-13,949.
Dong G, Liang X, Wang D, Gao H, Wang L, Wang L, Liu J, Du Z. High expression of miRNA-21 in triple-negative breast cancers was correlated with a poor prognosis and promoted tumor cell in vitro proliferation. Med. Oncol. 2014;31(7):57. doi: https://doi.org/10.1007/s12032-014-0057-x.
Garcia AI, Buisson M, Bertrand P, Rimokh R, Rouleau E, Lopez BS, Lidereau R, Mikaélian I, Mazoyer S. Down-regulation of BRCA1 expression by miR-146a and miR-146b-5p in triple negative sporadic breast cancers. EMBO Mol. Med. 2011;3(5):279-290.
Gyparaki MT, Basdra EK, Papavassiliou AG. MicroRNAs as regulatory elements in triple negative breast cancer. Cancer Lett. 2014;354(1):1-4.
Heng YJ, Lester SC, Tse GM, Factor RE, Allison KH, Collins LC, Chen YY, Jensen KC, Johnson NB, Jeong JC, Punjabi R, Shin SJ, Singh K, Krings G, Eberhard DA, Tan PH, Korski K, Waldman FM, Gutman DA, Sanders M, Reis-Filho JS, Flanagan SR, Gendoo DM, Chen GM, Haibe-Kains B, Ciriello G, Hoadley KA, Perou CM, Beck AH. The molecular basis of breast cancer pathological phenotypes. J. Pathol. 2017. Vol 241(3):375-391.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S, Magri E, Pedriali M, Fabbri M, Campiglio M, Ménard S, Palazzo JP, Rosenberg A, Musiani P, Volinia S, Nenci I, Calin GA, Querzoli P, Negrini M, Croce CM. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005;65(16):7065-7070.
Lehmann TP, Korski K, Gryczka R, Ibbs M, Thieleman A, Grodecka-Gazdecka S, Jagodziński PP. Relative levels of let-7a, miR-17, miR-27b, miR-125a, miR-125b and miR-206 as potential molecular markers to evaluate grade, receptor status and molecular type in breast cancer. Mol. Med. Rep. 2015;12(3):4692-4702.
Meng L, Xu Y, Xu C, Zhang W. Biomarker discovery to improve prediction of breast cancer survival: using gene expression profiling, meta-analysis, and tissue validation. Onco Targets Ther. 2016;9):6177-6185.
Perou C.M, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lønning PE, Børresen-Dale AL, Brown PO, Botstein D. Molecular portraits of human breast tumours. Nature. 2000;406:747-752.
Piva R, Spandidos DA, Gambari R. From microRNA functions to microRNA therapeutics: novel targets and novel drugs in breast cancer research and treatment (review). Int. J. Oncol. 2013;43(4):985-994.
Saha P, Nanda R. Concepts and targets in triple-negative breast cancer: recent results and clinical implications. Ther. Adv. Med. Oncol. 2016;8(5):351-359.
Sun Y, Liu X, Zhang Q, Mao X, Feng L, Su P, Chen H, Guo Y, Jin F. Oncogenic potential of TSTA3 in breast cancer and its regulation by the tumor suppressors miR-125a-5p and miR-125b. Tumour Biol. 2016;37(4):4963-4972.
Wang J, Ma Z, Carr SA, Mertins P, Zhang H, Zhang Z, Chan DW, Ellis MJ, Townsend RR, Smith RD, McDermott JE, Chen X, Paulovich AG, Boja ES, Mesri M, Kinsinger CR, Rodriguez H, Rodland KD, Liebler DC, Zhang B. Proteome profiling out-performs transcriptome profiling for coexpression based gene function prediction. Mol. Cell Proteomics. 2017;16(1):121-134.
Yang F, Zhang W, Shen Y, Guan X. Identification of dysregulated microRNAs in triple-negative breast cancer (review). Int. J. Oncol. 2015;46(3):927-932.
Zhao JJ, Lin J, Yang H, Kong W, He L, Ma X, Coppola D, Cheng JQ. MicroRNA-221/222 negatively regulates estrogen receptor alpha and is associated with tamoxifen resistance in breast cancer. J. Biol. Chem. 2016;291(43):22859.
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated from Byulleten’ Eksperimental’noi Biologii i Meditsiny, Vol. 163, No. 5, pp. 613-617, May, 2017
Rights and permissions
About this article
Cite this article
Chernyi, V.S., Tarasova, P.V., Kozlov, V.V. et al. Search of MicroRNAs Regulating the Receptor Status of Breast Cancer In Silico and Experimental Confirmation of Their Expression in Tumors. Bull Exp Biol Med 163, 655–659 (2017). https://doi.org/10.1007/s10517-017-3872-1
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10517-017-3872-1