Abstract
The death domain (DD), which is a versatle protein interaction module, is the prime mediator of the interactions necessary for apoptosis, innate immunity and the necrosis signaling pathway. Because DD mediated signaling events are associated with critical human diseases, studies in these areas are of great biological importance. Accordingly, many biochemical and structural studies of DD have been conducted in the past decade to investigate apoptotic and innate immune signaling. Evaluation of the molecular structure of DD and their interactions with partners have shown the underlying molecular basis for the assembly of DD mediated complexes and for the regulation of apoptosis and innate immunity. This review summarizes the structure and function of various DDs and DD:DD complexes involved in those signaling pathways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The death domains (DDs) are well-known protein interaction modules that belong to the death domain superfamily, which includes death effector domains (DEDs), caspase recruitment domains (CARDs), and pyrin domains (PYDs) [1–3]. The death domain superfamily is the prime mediator of the interactions necessary for apoptosis, innate immunity and the necrosis signaling pathway [4]. Among these superfamily (DD, DED, CARD and PYD), DDs have been intensively studied biochemically and structurally. This review will focus on structural studies of various DDs and their complexes in the main cellular signaling pathway.
More than 33 DD encoding genes have been identified in mammals, including eight members of the TNF (Tumor Necrosis Factor) receptor family and many intracellular signaling proteins, especially those involved in apoptosis, innate immunity and the necrosis signaling pathway, such as FADD (Fas-Assiciated protein with Death Domain), TRADD (Tumor necrosis factor Receptor type 1-Associated protein with Death Domain), PIDD (p53-induced protein with a DD), RAIDD (RIP-associated ICH-1 homologous protein with a death domain), Myd88 (Myeloid Differentiation primary response gene (88)) IRAKs (IL-1R-associated kinases) and RIP1 (Receptor-Interacting Protein 1) [4–10]. Several DD containing TNF family receptors, including TNFR1(Tumor Necrosis Factor Receptor1), Fas (Apo-1/CD95), DR3 (Death Receptor 3), DR4 (Death Receptor 4), DR5 (Death Receptor 5) and DR6 (Death Receptor 6), are important to the apoptosis signaling pathway [11, 12]. Specifically, their DDs in the cytoplasmic portion of the cells become critical protein–protein interaction modules that oligomerize and recruit death domain containing adaptor proteins, which leads to the activation of initiator caspases such as caspase-8 and -10 [13, 14]. Other initiator caspases such as caspase-9 and -2 are activated in the cytoplasm via the formation of large molecular complexes known as apoptosome and PIDDosome, respectively [15, 16]. The induced proximity of caspases produced by the formation of these large molecular complexes causes the self activation of initiator caspases [17, 18]. DDs play a critical role in mediating the formation of such complexes. For example, PIDDosome was recently identified as a caspase-2 activating complex that contains PIDD, RAIDD, and caspase-2 [16]. Although many uncertainties still exist, PIDDosome formation is important event for caspase-2 dependent apoptosis in a cell-type dependent manner [19–21]. Interestingly, by forming a complex with RIP-1 and Nemo, PIDD can act as a pro-survival factor. Because PIDD is involved in two completely opposite functions, pro- cell survival and pro- cell death, PIDD is considered to be a molecular switch that can control the fate of cells [22, 23]. Recent findings also suggest the involvement of several kinases in the caspase-2 activation and novel functions in non-apoptotic processes, such as cell cycle regulation and DNA repair [24]. DNA-PKcs are PIDD binding partners for DNA repair function in response to γ-irradiation [24]. PIDD is a DD-containing protein [25], while RAIDD is an adaptor molecule that contains a CARD at its amino-terminal region and DD at its carboxy-terminal region [9]. During PIDDosme formation, the RAIDD adaptor acts as a bridge between caspase-2 and PIDD by offering the CARD domain for CARD-CARD interactions and the death domain (DD) for DD-DD interactions, respectively [16]. DD containing proteins also play a role in innate immunity, communicating with Toll-like receptors (TLRs) through adaptor protein MyD88, which contains both a DD and a TIR (Toll/Interleukin-1 Receptor) domain [26–28]. DD of MyD88 serves to recruit a family of kinases known as IRAKs that also contains DD at the N-terminus [28]. DD interaction between MyD88 and IRAKs is important for TLR signaling [26, 29, 30]. Pelle and Tube complexes formed via DD interactions in flies are also involved in the innate immunity response against fungal infection and developmental dorsal–ventral patterning of drosophila [27, 31].
Given the fact that both DD mediated apoptosis and innate immunity are associated with critical human diseases such as cancer, immune disorders, and neuro-degenerative diseases, studies in these areas are of great biological importance. Biochemical and structural studies of DD have been explored over the past decade to enable a better understanding of apoptotic and innate immunue signaling. As a result, several DD structures have been elucidated, including four DD:DD complex structures [1, 31–40]. The molecular structure of DDs and their interactions with partners have revealed the underlying molecular basis for the assembly of DD mediated complexes and for the regulation of apoptosis and the innate immune signaling pathway (Table 1). This review summarizes the structure and function of DDs.
Structural aspects of death domains (DDs)
Structural studies of DDs have been difficult because they are subject to self-aggregation under physiological conditions. However, solubilization via the introduction of mutations, controlling pH, and refolding techniques have resulted in the elucidation of several structures of important DDs, such as the DDs of Fas [41], FADD [33], TNFR1 [34], p75 [39], IRAK4 [40], RAIDD [35] and three hetero-typic DD-DD complexes including those between Pelle DD and Tube DD, RAIDD DD and PIDD DD, and Fas DD and FADD DD [5, 31, 32, 35, 36, 42]. The first structure of the ternary DD complex, MyD88 DD:IRAK4 DD:IRAK2 DD, was recently elucidated [38].
A structural homology search using DALI [43] showed that most DDs are highly similar to other DDs that exhibit the six-helix bundles fold (Fig. 1a and Table 2). Additionally, structure-based sequence alignment revealed that most of the residues buried in the hydrophobic core of RAIDD DD are also conserved among these highly divergent DD structures, suggesting that there is a conserved hydrophobic core that exists across all DDs and possibly other members of the DD superfamily (Fig. 1b). Interestingly, despite their structural similarity, each DD has its own binding partner. The specificity of each DD is critical to its proper function. To clarify the structural difference between DDs, RAIDD DD was superimposed with various other DDs. Pair-wise structural alignments of RAIDD DD with other DDs showed that the helices in RAIDD DD differ from other DDs in their lengths and orientations (Fig. 2). For example, when compared with FADD DD, helices H2 and H3 of RAIDD DD show dramatic differences in orientation. In addition, H6 of TNFR1 DD and Fas DD are much longer than that of RAIDD DD. These differences indicate that the various lengths and orientations might be critical for determining specificity. Interestingly, several DDs, including RAIDD DD and FADD DD, have an H0. Because the DDs in RAIDD and FADD reside at the C-terminal half of the proteins, H0 may indicate a mode of connection to the previous domain, either a CARD for RAIDD or a DED for FADD. Additional differences between the structures of DDs are at the loops connecting the helices, especially those at H3-H4.
Structural similarity among DDs. a A structural homology search using DALI. The structure of DDs, p84 DD, NGFR DD, Tube DD, Fas DD, IRAK DD, RAIDD DD, exhibiting the six-helix bundles fold. b A structural based sequence alignment. Each structure of the DDs was compared 3 dimensionally using the Dali server and sequences of DDs were aligned based on their structural comparison. Conserved hydrophobic cores are indicated
Comparison of DDs with each other has revealed that the gross features of the electrostatic surfaces are extremely diverse. In contrast to the more charged nature of most other DDs, the RAIDD DD surface is much more hydrophobic (Fig. 3). Moreover, the surface charge of p84 is relatively acidic, whereas that of Pelle is relatively basic (Fig. 3). This is the case for both sides of the molecule. Because DDs are protein interaction modules, their surface features dictate their mode of interactions with partners. This apparent difference in the electrostatic surface may also be critical for their specificity.
Electrostatic surface representation of DDs. The first side of the RAIDD DD surface and the top panels of the remaining DDs are shown in the same orientations as in Fig. 2. The opposite side of RAIDD DD and the bottom plots for the other DDs are rotated by 180° along the vertical axis (Y)
Given the similar structure of DDs, we investigated the potential evolutionary relationship among DDs by constructing phylogenetic trees based on structural similarity (Fig. 4). The phylogenetic tree assigned two large branches consisting of p84 DD and Fas DD in one and different DDs in the other. The structural similarity of RAIDD DD was evolutionally similar to that of TNFR-1 DD. For Fas DD, p84 DD was evolutionally close.
Structural comparisons. Phylogenetic tree of the DDs constructed based on structural similarity. The calculation was conducted using the program TraceSuite II (http://wwwcryst.bioc.cam.ac.uk/~jiye/evoltrace/evoltrace.html)
Structure of the RAIDD DD:PIDD DD complex
The crystal structure of the PIDDosome core, which is composed of seven RAIDD DDs and five PIDD DDs, has been elucidated [5]. The unusually constructed structure of this DD complex is divided into three layers from the side, five PIDD DDs at the bottom, five RAIDD DDs in the middle, and two additional RAIDD DDs at the top (Fig. 5a, b). Interestingly, this DD complex does not possess a distinct fivefold symmetry. Instead, the complex consists of two different types of unique screw rotations, one rotating around 84° and translating down the axis and the other rotating around 54° and translating up the axis. The complex was formed by three different types of interactions that were classified based on the regions involved in the interaction (Fig. 5c, Table 3). All three types detected in this complex were similar to the previously identified interaction types between the procaspase-9 CARD:Apaf-1 CARD for type I [44] and the Pelle DD:Tube DD for type II [31], while Type III was unique.
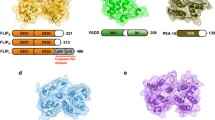
Structures of DD complexes and their interacting interface. The top view (a) and side view (b) of the RAIDD DD: PIDD DD complex structure. The three layers are indicated by grey lines. (c) A schematic diagram of the locations of the three types of interacting interfaces in the RAIDD DD:PIDD DD complex. (d) The structure of the Pelle DD:Tube DD complex. A schematic diagram of the location of the type II interacting interface is also shown. (e) The top view (boxed in black) and side view of the MyDosome (MyD88 DD:IRAK4 DD: IRAK2 DD) complex structure. Four layers are indicated by grey lines. (f) A schematic diagram of the locations of the three types of interacting interfaces in the ternary complex
In the type I interaction, the interacting DDs were related by an approximately 84° rotation. Additionally, about 160 Å2 of the surface area was buried at this interface per partner. Residues at and near H1 and H4 of the first DD interact with residues at H2 and H3 of the second DD. A mixture of hydrophobic, polar and charged interactions occurs at this interface, including the hydrophobic interaction, the salt bridge, and a massive hydrogen bonding network. In the type II interaction, the interacting DDs were found to be related by an approximately 30° rotation. Although the interaction buries a fairly small surface area of approximately 90 Å2 per partner, the shape complementarity score of this interaction was highest among all interactions in the complex according to the SC program in CCP4 [45]. H4 and the loop between H4 and H5 of the first DD and the loop between H5 and H6 and H6 of the second DD mediate the type II interaction. The interface appears to be mostly polar and charged. For the type III interaction, the interacting DDs are related by an approximately 54° rotation. The largest surface area is buried at this interface, approximately 260 Å2 per partner. H3 of the first DD and the H1–H2 and the loop between H3 and H4 of the second DD are primarily involved. Unlike the type I and type II interaction, this interface does not have salt bridges. Instead, many hydrophobic and polar interactions form the interface. However, the nature of the contacts is not conserved within each type of interaction. For instance, the interaction interface (type I interaction) formed by the pro-caspase9 CARD:Apaf-1 CARD complex is complementary in charge. Conversely, in the same type I interaction of the RAIDD DD:PIDD DD complex, a complicated network of hydrophobic contacts and hydrogen bonds mediate the interfaces. Overall, complex charges and hydrophobic interactions are formed in all three types of interaction in the RAIDD DD:PIDD DD complex. The features of each type of interaction are summarized in Table 3.
Structure of the Pelle DD:Tube DD complex
The crystal structure of the heterodimeric Pelle DD:Tube DD complex, which was reported in 1999, was the first complex structure identified [31]. Pelle is a serine/threonine kinase that is recruited to the plasma membrane and activated upon activation of the Toll receptors [46]. The recruitment of Pelle is important for the nuclear localization of Dorsal, which is a key event for dorsal–ventral patterning of Drosophila [47, 48]. Tube is the adaptor protein that facilitates the recruitment of Pelle to the plasma membrane [49].
The most interesting observation associated with these complex structures is the asymmetry of the interaction (Fig. 5d). The complex structure revealed that there are two distinct contact regions between Pelle DD and Tube DD. The first region is formed by the H4 helix and the following loop of Pelle and the groove of Tube that is created by the H1–H2 corner and H6 and the preceding loop. Many charged residues from both sides are involved in this interaction. The second interaction region is formed by the C-terminal tail of Tube-DD and the cavity between the H4–H5 and H2–H3 hairpins of Pelle. Three hydrophobic residues (I169, L171, and L173) on the C-terminal of Tube are clearly inserted into the cavity formed by H5 and the N-terminal of the Pelle DD. Overall, the interaction between the Drosophila proteins Pelle and Tube is mixed, consisting of both hydrophobic and hydrophilic components.
The interaction type of this complex is similar to the type II interaction in the RAIDD DD:PIDD DD complex in that the interaction is primarily mediated between the H4-H5 region of the first DD and the H5–H6 region of the second DD, which is the common interaction mode for type II. Moreover, new interactions between the adjacent H2 region of the Pelle DD and the adjacent H1–H2 region of Tube DD were detected. When compared with other interaction forms in DDs, the type II interaction is mediated by a smaller surface area. In the Pelle DD:Tube DD complex, this interaction formed by the small area is supported by an additional interaction between a long C-terminal tail of Tube DD and the H2–H3 and H4–H5 region of the Pelle DD. The features of the type II interaction detected in the structure of the Pelle DD:Tube DD are summarized in Table 3.
Structure of the MyD88 DD:IRAK4 DD:IRAK2 DD ternary complex
MyD88 is a critical adaptor protein in the TLR/IL-1R signaling pathway and contains DD at the N-terminus and a TIR (Toll/IL-1R homology) domain at the C-terminus. The DD containing kinases, IRAK1, IRAK2 and IRAK4, are positively involved in the signaling by direct interaction with MyD88 via DD-DD interaction. The molecular signaling complex containing the MyD88 and IRAK families is called “MyDDosome”.
Recently, Wu’s group elucidated the crystal structure of MyD88 DD: IRAK4 DD:IRAK2 DD ternary complex [38]. The novel ternary complex structure formed by three different DDs was a tower-shaped structure about 110 Å in height and 70 Å in diameter (Fig. 5e). This structure contained four layers, with MyD88 DD at the bottom two layers, IRAK4 DD in the middle layer and IRAK2 at the top layer. The complex was single stranded left-handed helixes composed of six MyD88 DDs, four IRAK4 DDs, and four IRAK2 DDs (Fig. 5e). The layer interactions were formed between MyD88 DD and IRAK4 DD and between IRAK4 DD and IRAK2 DD. The interfaces formed by those complexes were quite different, providing the specificity in the oligomerization of three different DDs (Fig. 5f). Good charge complementarity between MyD88 DD and IRAK4 DD and poor charge complementarity between the top and bottom surfaces of MyD88 were detected. This ternary DD complex contains all three types of interactions that were previously detected on the complex structure of RAIDD DD:PIDD DD. Similarly a type I interaction is formed by H1 and H4 on one DD and H2 and H3 on the other DD. The type II interaction is formed by the H4-H5 loop on one DD and the H1–H2 loop on the other DD. For the type III interaction H3 of one DD and H1, H2, and loops between H3 and H4 on the other DD are involved. The complex is formed by three type I (MyD88 DD:MyD88 DD, MyD88 DD:IRAK4 DD and IRAK4:IRAK2), three type II (MyD88 DD:MyD88 DD, MyD88 DD:IRAK4 DD and IRAK4:IRAK2, and five type III (MyD88 DD:MyD88 DD, MyD88 DD:IRAK4 DD, IRAK4:IRAK4, IRAK4:IRAK2, and IRAK2:IRAK2) interactions (Fig. 5f). Massive hydrophobic and charged interactions are detected at the surface of each interaction and summarized in Table 3.
Structure of the Fas DD:FADD DD complex
The most recent studies conducted by two different research groups have elucidated Fas DD:FADD DD complex structures and revealed how formation of DISC can be regulated by ligand-receptor clustering [50, 51]. Similar with the previously reported DD complex between RAIDD DD and PIDD DD, Fas DD:FADD DD complex forms an asymmetric oligomeric structure composed of five Fas DDs and five FADD DDs (Fig. 6a, b). This composition of DDs is similar to that of RAIDD DD:PIDD DD. The complex is a two-layered structure with an upper layer of five Fas DD and a lower layer of five FADD DD (Fig. 6a, b). This layer composition is consistent with a similarity between Fas and RAIDD and between FADD and PIDD. The complex also contains all three different types of interactions that were previously detected on the complex structure of RAIDD DD:PIDD DD. Similarly the type I interaction is formed by H1 and H4 on one DD and H2 and H3 on the other DD. The type II interaction is formed by the H4–H5 loop on one DD and the H1–H2 loop on the other DD. For the type III interaction H3 of one DD and H1, H2, and the loops between H3 and H4 on the other DD are involved (Fig. 6c).
Structural comparison of two controversial Fas DD:FADD DD complex structures. The top view (a) and side view (b) of the Fas DD: FADD DD complex structure solved at neutral pH. (c) A schematic diagram of the locations of the three types of interacting interfaces in the Fas DD:FADD DD complex at neutral pH. The top view (d) and side view (e) of the Fas DD:FADD DD complex structure solved at acidic pH. Four Fas DDs and four PIDD DDs are shown. (f) A schematic diagram of the locations of the three types of interacting interfaces in the Fas DD:FADD DD complex structure solved at low pH condition
Totally different conformation of Fas DD:FADD DD complex structure, solved under acidic conditions, was also recently reported by Scott et al. [36]. Unlike the structure of the Fas DD:FADD DD complex solved in physiological condition at neutral pH and the RAIDD DD:PIDD DD complex, the structure was constructed with four Fas DDs and four FADD DDs (Fig. 6d, e). Interestingly, in the formation of the complex, all of the contacts were mediated by Fas DD (Fig. 6f). The key observation at this complex structure was that Fas DD underwent significant conformational changes when compared to the structure of the uncomplexed Fas DD. The actual interaction between Fas DD and FADD DD for assembly of the complex was via hydrophobic interactions, which were exposed by conformational changes of Fas DD. Hydrophobic patches surrounded by polar residues formed by H1 and H6 of the FADD DD and hydrophobic residues on the stem helix formed by H5 and H6 of the Fas DD are involved in the interaction. Another characteristic of the complex was the relatively weak interaction between Fas DD and FADD DD, despite the large binding surface areas.
The discrepancy between the two Fas DD:FADD DD complex structures (solved at neutral pH and acidic pH) might be due to the structures being solved under different conditions and circumstances. Although recent biochemical, biophysical and intensive mutational studies suggests that Fas/FADD structure solved under acidic condition does not represent the physiological interaction in solution, the relevance of the stoichiometry and function of the Fas DD:FADD DD complex should be more studied.
Future perspectives
The human death domain consists of approximately 32 members, many of which have been studied intensively due to their important role in the central signaling pathways. To date, several DD structures, including four complex structures, Fas DD:FADD DD, RAIDD DD:PIDD DD, Pelle DD:Tube DD, and MyD88 DD:IRAK4 DD:IRAK2 DD, have been elucidated. The details of the assembly of the Fas DD:FADD DD, RAIDD DD:PIDD DD, Pelle DD:Tube DD, and MyD88 DD:IRAK4 DD:IRAK2 DD are quite different, including stiochiometry and the nature of the interacting interface. However, a common helical assembly was detected at Fas DD:FADD DD solved at neutral pH, as well as RAIDD DD:PIDD DD, and MyD88 DD:IRAK4 DD:IRAK2 DD. This might be a representative common assebly mechanism for DD.
The recently reported Fas DD:FADD DD complex structure solved under acidic conditions revealed a totally different regulation mechanism for the assembly of DD [36]. The conditional and conformation change dependent interaction for transient unstable interactions formed by Fas DD:FADD DD within the DISC is obviously an interesting characteristic of the death domain. However, it is worth to keep in mind that the unusual DD complex formation mechanism was based on the structure solved under extremely acidic condition and might be a crystallographic artifact. Many biophysical, biochemical and intensive mutational studies including recently elucidated two more new Fas DD:FADD DD structure support the idea. New Fas DD:FADD DD complex structures, solved at neutral pH, show that there are no structural changes in the C-terminal region of Fas DD, which is observed at Fas DD:FADD DD complex solved at acidic pH and is a critical event for the complex assembly. Moreover, C-terminal tail of Fas DD is dispensable for complex formation between the two DDs in solution under physiological conditions. The new complex solved at neutral pH is constructed by five Fas DDs and five FADD DDs. Although many experiments have been conducted to explain this lack of consistency and to determine which complex is more biologically meaningful, there have been no clear answers to these questions. It is apparent that more studies are required to properly understand the early events involved in DISC formation. Interestingly, the three types of interactions detected in the RAIDD DD:PIDD DD complex structures have also been observed in the Pelle DD: Tube DD complex, the MyD88 DD:IRAK4 DD:IRAK2 DD ternary complex, the Apaf-1 CARD: Caspase-9 CARD complex, and the new Fas DD:FADD DD complex solved at neutral pH, but not in the previously published Fas DD:FADD DD complex solved at acidic pH. Although different assembly mechanisms of DD has been detected under acidic condition, the structures of the Fas DD:FADD DD solved at neutral pH, RAIDD DD:PIDD DD, and MyD88 DD:IRAK4 DD:IRAK2 DD, showing the helical assembly of DDs, provides an excellent model for the DD-mediated assembly of signaling molecules, which could be produced by various types of symmetries and stoichiometries.
Although all DDs exhibit a six-helix bundles fold, variations have been detected in the direction and length of the helices. Moreover, the surface features of DDs are completely different, with low sequence homology existing among the DDs. These different features may be responsible for their ability to interact with their own partner. Given the paucity of structural information regarding DD and DD complexes, it is still not known (1) if common modes of interactions may be observed from DDs, (2) whether the three types of interactions introduced by Park et al. (2007) cover all possible interactions generated by DDs (except for conformation change dependent interactions), (3) whether conformation change dependent interactions are common interaction methods or crystallographic artifact. More biochemical and structural studies are required to address these questions and to fully understand the molecular basis of these interactions in the assembly of apoptotic and inflammatory signaling complexes.
References
Park HH, Lo YC, Lin SC, Wang L, Yang JK, Wu H (2007) The death domain superfamily in intracellular signaling of apoptosis and inflammation. Ann Rev Immunol 25:561–586
Weber CH, Vincenz C (2001) The death domain superfamily: a tale of two interfaces? Trends Biochem Sci 26:475–481
Kohl A, Grutter MG (2004) Fire and death: the pyrin domain joins the death-domain superfamily. C R Biol 327:1077–1086
Reed JC, Doctor KS, Godzik A (2004) The domains of apoptosis: a genomics perspective. Sci STKE 239:re9
Jang TH, Zheng C, Wu H, Jeon JH, Park HH (2010) In vitro reconstitution of the interactions in the PIDDosome. Apoptosis 15:1444–1452
Schneider P, Thome M, Burns K et al (1997) TRAIL receptors 1 (DR4) and 2 (DR5) signal FADD-dependent apoptosis and activate NF-kB. Immunity 7:831–836
Hsu H, Xiong J, Goeddel DV (1995) The TNF receptor 1-associated protein TRADD signals cell death and NF-kB activation. Cell 81:495–504
Hsu H, Shu H-B, Pan M-G, Goeddel DV (1996) TRADD-TRAF2 and TRADD-FADD interactions define two distinct TNF receptor 1 signal transduction pathways. Cell 84:299–308
Duan H, Dixit VM (1997) RAIDD is a new ‘death’ adaptor molecule. Nature 385:86–89
Cao Z, Henzel WJ, Gao X (1996) IRAK: a kinase associated with the interleukin-1 receptor. Science 271:1128–1131
Chinnaiyan AM, O’Rourke K, Yu G et al (1996) Signal transduction by DR3, a death domain-containing receptor related to TNF-R1 and CD95. Science 274:990–992
Cleveland JL, Ihle JN (1995) Contenders in FasL/TNF death signaling. Cell 81:479–482
Wajant H (2002) The Fas signaling pathway: more than a paradigm. Science 296:1635–1636
Pop C, Salvesen GS (2009) Human caspases: activation, specificity and regulation. J Biol Chem 284:21777–21781
Zou H, Henzel WJ, Liu X, Lutschg A, Wang X (1997) Apaf-1, a human protein homologous to C. elegans CED-4, participates in cytochrome c-dependent activation of caspase-3. Cell 90:405–413
Tinel A, Tschopp J (2004) The PIDDosome, a protein complex implicated in activation of caspase-2 in response to genotoxic stress. Science 304:843–846
Salvesen GS, Dixit VM (1999) Caspase activation: the induced-proximity model. Proc Natl Acad Sci USA 96:10964–10967
Muzio M, Stockwell BR, Stennicke HR, Salvesen GS, Dixit VM (1998) An induced proximity model for caspase-8 activation. JBC 273:2926–2930
Vakifahmetoglu-Norberg H, Zhivotovsky B (2010) The unpredictable caspase-2: what can it do? Trends Cell Biol 20:150–159
Olsson M, Vakifahmetoglu H, Abruzzo PM, Hogstrand K, Grandien A, Zhivotovsky B (2009) DISC-mediated activation of caspase-2 in DNA damage-induced apoptosis. Oncogene 28:1949–1959
Bergeron L, Perez GI, Macdonald G et al (1998) Defects in regulation of apoptosis in caspase-2-deficient mice. Genes Dev 12:1304–1314
Tinel A, Janssens S, Lippens S et al (2007) Autoproteolysis of PIDD marks the bifurcation between pro-death caspase-2 and pro-survival NF-kappaB pathway. EMBO J 26:197–208
Janssens S, Tinel A, Lippens S, Tschopp J (2005) PIDD mediates NF-kappaB activation in response to DNA damage. Cell 123:1079–1092
Shi M, Vivian CJ, Lee KJ et al (2009) DNA-PKcs-PIDDosome: a nuclear caspase-2-activating complex with role in G2/M checkpoint maintenance. Cell 136:508–520
Lin Y, Ma W, Benchimol S (2000) Pidd, a new death-domain-containing protein, is induced by p53 and promotes apoptosis. Nat Genet 26:122–127
Fitzgerald KA, Palsson-McDermott EM, Bowie AG et al (2001) Mal (MyD88-adapter-like) is required for Toll-like receptor-4 signal transduction. Nature 413:78–83
Muzio M, Ni J, Feng P, Dixit VM (1997) IRAK (Pelle) family member IRAK-2 and MyD88 as proximal mediators of IL-1 signaling. Science 278:1612–1615
Wesche H, Henzel WJ, Shillinglaw W, Li S, Cao Z (1997) MyD88: an adapter that recruits IRAK to the IL-1 receptor complex. Immunity 7:837–847
Kawai T, Akira S (2006) TLR signaling. Cell Death Differ 13:816–825
Wang Q, Dziarski R, Kirschning CJ, Muzio M, Gupta D (2001) Micrococci and peptidoglycan activate TLR2→MyD88→IRAK→TRAF→NIK→IKK→NF-kappaB signal transduction pathway that induces transcription of interleukin-8. Infect Immun 69:2270–2276
Xiao T, Towb P, Wasserman SA, Sprang SR (1999) Three-dimensional structure of a complex between the death domains of Pelle and Tube. Cell 99:545–555
Salvesen GS, Riedl SJ (2009) Structure of the Fas/FADD complex: a conditional death domain complex mediating signaling by receptor clustering. Cell Cycle 8:2723–2727
Jeong EJ, Bang S, Lee TH, Park YI, Sim WS, Kim KS (1999) The solution structure of FADD death domain. Structural basis of death domain interactions of Fas and FADD. J Biol Chem 274:16337–16342
Sukits SF, Lin LL, Hsu S, Malakian K, Powers R, Xu GY (2001) Solution structure of the tumor necrosis factor receptor-1 death domain. J Mol Biol 310:895–906
Park HH, Wu H (2006) Crystal structure of RAIDD death domain implicates potential mechanism of PIDDosome assembly. J Mol Biol 357:358–364
Scott FL, Stec B, Pop C et al (2009) The Fas-FADD death domain complex structure unravels signalling by receptor clustering. Nature 457:1019–1022
Park HH, Logette E, Rauser S et al (2007) Death domain assembly mechanism revealed by crystal structure of the oligomeric PIDDosome core complex. Cell 128:533–546
Lin SC, Lo YC, Wu H. Helical assembly in the MyD88-IRAK4-IRAK2 complex in TLR/IL-1R signalling. Nature 465: 885-890
Liepinsh E, Ilag LL, Otting G, Ibanez CF (1997) NMR structure of the death domain of the p75 neurotrophin receptor. EMBO J 16:4999–5005
Lasker MV, Gajjar MM, Nair SK (2005) Cutting edge: molecular structure of the IL-1R-associated kinase-4 death domain and its implications for TLR signaling. J Immunol 175:4175–4179
Huang B, Eberstadt M, Olejniczak ET, Meadows RP, Fesik SW (1996) NMR structure and mutagenesis of the Fas (APO-1/CD95) death domain. Nature 384:638–641
Park HH, Wu H (2007) Crystallization and preliminary X-ray crystallographic studies of the oligomeric death-domain complex between PIDD and RAIDD. Acta Crystallogr Sect F Struct Biol Cryst Commun 63:229–232
Holm L, Sander C (1995) Dali: a network tool for protein structure comparison. Trends Biochem Sci 20:478–480
Qin H, Srinivasula SM, Wu G, Fernandes-Alnemri T, Alnemri ES, Shi Y (1999) Structural basis of procaspase-9 recruitment by the apoptotic protease-activating factor 1. Nature 399:549–557
Lawrence MC, Colman PM (1993) Shape complementarity at protein/protein interfaces. J Mol Biol 234:946–950
Belvin MP, Anderson KV (1996) A conserved signaling pathway: the Drosophila toll-dorsal pathway. Annu Rev Cell Dev Biol 12:393–416
Sun H, Towb P, Chiem DN, Foster BA, Wasserman SA (2004) Regulated assembly of the Toll signaling complex drives Drosophila dorsoventral patterning. EMBO J 23:100–110
Grosshans J, Schnorrer F, Nusslein-Volhard C (1999) Oligomerisation of Tube and Pelle leads to nuclear localisation of dorsal. Mech Dev 81:127–138
Sun H, Bristow BN, Qu G, Wasserman SA (2002) A heterotrimeric death domain complex in Toll signaling. Proc Natl Acad Sci USA 99:12871–12876
Wang L, Yang JK, Kabaleeswaran V et al (2010) The Fas-FADD death domain complex structure reveals the basis of DISC assembly and disease mutations. Nat Struct Mol Biol 17:1324–1329
Esposito D, Sankar A, Morgner N, Robinson CV, Rittinger K, Driscoll PC (2010) Solution NMR investigation of the CD95/FADD homotypic death domain complex suggests lack of engagement of the CD95 C terminus. Structure 18:1378–1390
Park A, Baichwal VR (1996) Systematic mutational analysis of the death domain of the tumor necrosis factor receptor 1-associated protein TRADD. J Biol Chem 271:9858–9862
Micheau O, Tschopp J (2003) Induction of TNF receptor I-mediated apoptosis via two sequential signaling complexes. Cell 114:181–190
Stanger BZ, Leder P, Lee TH, Kim E, Seed B (1995) RIP: a novel protein containing a death domain that interacts with Fas/APO-1 (CD95) in yeast and causes cell death. Cell 81:513–523
Hsu H, Huang J, Shu HB, Baichwal V, Goeddel DV (1996) TNF-dependent recruitment of the protein kinase RIP to the TNF receptor-1 signaling complex. Immunity 4:387–396
Gajate C, Mollinedo F (2005) Cytoskeleton-mediated death receptor and ligand concentration in lipid rafts forms apoptosis-promoting clusters in cancer chemotherapy. J Biol Chem 280:11641–11647
Darnay BG, Aggarwal BB (1997) Early events in TNF signaling: a story of associations and dissociations. J Leukoc Biol 61:559–566
Pan G, O’Rourke K, Chinnaiyan AM et al (1997) The receptor for the cytotoxic ligand TRAIL. Science 276:111–113
Chinnaiyan AM, O’Rourke K, Tewari M, Dixit VM (1995) FADD, a novel death domain-containing protein, interacts with the death domain of Fas and initiates apoptosis. Cell 81:505–512
Juo P, Kuo CJ, Yuan J, Blenis J (1998) Essential requirement for caspase-8/FLICE in the initiation of the Fas-induced apoptotic cascade. Curr Biol 8:1001–1008
Kitson J, Raven T, Jiang YP et al (1996) A death-domain-containing receptor that mediates apoptosis. Nature 384:372–375
Chaudhary PM, Eby M, Jasmin A, Bookwalter A, Murray J, Hood L (1997) Death receptor 5, a new member of the TNFR family, and DR4 induce FADD-dependent apoptosis and activate the NF-kappaB pathway. Immunity 7:821–830
Jin Z, Li Y, Pitti R et al (2009) Cullin3-based polyubiquitination and p62-dependent aggregation of caspase-8 mediate extrinsic apoptosis signaling. Cell 137:721–735
Sprick MR, Weigand MA, Rieser E et al (2000) FADD/MORT1 and caspase-8 are recruited to TRAIL receptors 1 and 2 and are essential for apoptosis mediated by TRAIL receptor 2. Immunity 12:599–609
Pan G, Bauer JH, Haridas V et al (1998) Identification and functional characterization of DR6, a novel death domain-containing TNF receptor. FEBS Lett 431:351–356
Boldin MP, Goncharov TM, Goltsev YV, Wallach D (1996) Involvement of MACH, a novel MORT1/FADD-interacting protease, in Fas/APO-1- and TNF receptor-induced cell death. Cell 85:803–815
Ahmad M, Srinivasula SM, Wang L et al (1997) CRADD, a novel human apoptotic adaptor molecule for caspase-2, and FasL/tumor necrosis factor receptor-interacting protein RIP. Cancer Res 57:615–619
Burns K, Clatworthy J, Martin L et al (2000) Tollip, a new component of the IL-1RI pathway, links IRAK to the IL-1 receptor. Nat Cell Biol 2:346–351
Li S, Strelow A, Fontana EJ, Wesche H (2002) IRAK-4: a novel member of the IRAK family with the properties of an IRAK-kinase. Proc Natl Acad Sci USA 99:5567–5572
Berglund H, Olerenshaw D, Sankar A, Federwisch M, McDonald NQ, Driscoll PC (2000) The three-dimensional solution structure and dynamic properties of the human FADD death domain. J Mol Biol 302:171–188
Acknowledgments
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the ministry of Education, Science and Technology (2010-0020962).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, H.H. Structural analyses of death domains and their interactions. Apoptosis 16, 209–220 (2011). https://doi.org/10.1007/s10495-010-0571-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10495-010-0571-z