Abstract
Rhipicephalus (Boophilus) microplus is an obligate haematophagous arthropod and the major problem for cattle industry due to economic losses it causes. The parasite shows a remarkable adaptability to changing environmental conditions as well as an exceptional ability to survive long-term starvation. This ability has been related to a process of intracellular protein degradation called autophagy. This process in ticks is still poorly understood and only few autophagy-related (ATG) genes have been characterized. The aim of the present study was to examine the ESTs database, BmiGI, of R. microplus searching for ATG homologues. We predicted five putative ATG genes, ATG3, ATG4, ATG6 and two ATG8s. Further characterization led to the identification of RmATG8a and RmATG8b, homologues of GABARAP and MAP1LC3, respectively, and both of them belonging to the ATG8 family. PCR analyses showed that the expression level of RmATG8a and RmATG8b was higher in egg and larval stages when compared to ovary and midgut from adult ticks. This up-regulation coincides with the period in which ticks are in a starvation state, suggesting that autophagy is active in R. microplus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rhipicephalus (Boophilus) microplus is a haematophagous arthropod that represents a major problem for cattle industry due to the economic losses it causes. This tick acts also as pathogen vector, causing diseases such as babesiosis and anaplasmosis (Rosario-Cruz et al. 2005; Ribeiro et al. 2007). Rhipicephalus microplus is a one-host tick and its life cycle occurs in two phases, the first entirely independent of the host. In this phase, the eggs hatch in the environment and the larvae crawl up grass or other plants to find a host. The second phase involves attachment of larvae, nymph and adult (male and female) stages to a bovine host, with all three instars remaining on the host.
Under favorable environmental conditions R. microplus can complete its life cycle in 3–4 weeks. However, in the summer, larvae can survive for as long as 3–4 months without feeding and, in cooler temperatures, may live without food for up to 6 months (Spickler 2007). This remarkable ability to survive long-term starvation has been related to a non-selective process of bulk cytoplasmic degradation called autophagy.
All eukaryotic cells have an autophagy system induced by starvation (Yoshimoto et al. 2004). This system degrades organelles and proteins into small polypeptides to help maintain amino acid pools and energy balance (Lilienbaum 2013; Ryter et al. 2013). However, autophagy in ticks is poorly understood and only few autophagy-related (ATG) genes have been characterized. The HlATG3, HlATG4, HlATG6, HlATG8 and HlATG12 genes were recently reported in the tick Haemaphysalis longicornis (Umemiya et al. 2007, 2008; Umemiya-Shirafuji et al. 2010; Kawano et al. 2011; Umemiya-Shirafuji et al. 2014). Nevertheless, in a study conducted to search for tick homologues of yeast or mammalian ATG genes in eight tick databases, no ATG homologues were found except for H. longicornis and Ixodes scapularis (Malagoli et al. 2010). This prompted us to reexamine the Boophilus microplus gene index, BmiGI, searching for ATG homologues. We found five putative ATG genes (ATG3, ATG4, ATG6 and two ATG8s). In this study, we present the molecular characterization of RmATG8a and RmATG8b, homologues of GABARAP and MAP1LC3, respectively, belonging to the ATG8 family, as well as their expression in different developmental stages and organs in R. microplus.
Materials and methods
Ticks and tissues
Rhipicephalus microplus ticks were obtained from a susceptible strain reared under laboratory conditions at CENID-PAVET-INIFAP, located in the State of Morelos, Mexico. Rearing was done as described by Davey et al. (1980). Fully engorged female ticks were obtained after the spontaneous detachment from calves infested with 10-day old R. microplus larvae. They were transferred to a humidity chamber and kept at 28 °C throughout oviposition. Eggs were collected 3 days after oviposition, whereas larvae were collected 10 days after hatching. Ovarian and intestinal tissues were obtained from engorged female ticks the following day after detachment, as described by Tsuda et al. (2001).
RNA isolation and cDNA synthesis
Eggs, larvae, or adult tissues were pooled (100 mg) and total RNA was extracted using TRIZOL reagent (Invitrogen, Carlsbad, CA, USA). cDNA was synthesized from 3.9 µg of RNA using the SuperScript® III First-Strand Synthesis System (Invitrogen) according to manufacturer’s instructions.
Polymerase chain reaction
RmATG8a and RmATG8b sequences (BmiGI accession numbers TC15563 and TC20934, respectively) were obtained by screening the B. microplus gene index, BmiGI, with HlATG8 sequence (GenBank accession number AB513351) from H. longicornis. The following primers (Integrated DNA Technologies, Coralville, IA, USA) were used: F-RmATG8a 5′-CTC CCT CGT CGT GTT TCG-3′ and R-RmATG8a 5′-CAA TTC AAA AGA AAG AGG AAA TGA-3′; F-RmATG8b 5′-GTC TCG TGC GAG GAG ACA AC-3′ and R-RmATG8b 5′-GCC TTG AAC AGT GAG CAC AAG-3′, using cDNA from eggs, larvae, and adult tissues as template. PCR conditions were 94 °C for 5 min followed by 35 cycles, each consisting of denaturation at 94 °C for 30 s, annealing at 55 °C for 30 s, and extension at 72 °C for 60 s. A final extension step at 72 °C for 7 min was included. The amplified products were run on 1 % agarose gels and were analyzed on a UV transilluminator (Bio-Rad Laboratories, Philadelphia, PA, USA). Control amplification was carried out using the following primers: F-5′-GAC GCA GAT CAT GTT CGA GA-3′ and R-5′-ACA GGT CCT TAC GGA TGT CG-3′, designed from B. microplus actin (GenBank accession number AY255624.1; da Silva Vaz et al. 2005; Nijhof et al. 2009). PCR conditions were 94 °C for 5 min followed by 35 cycles, each consisting of denaturation at 94 °C for 30 s, annealing at 60 °C for 30 s, and extension at 72 °C for 45 s, with a final extension step at 72 °C for 7 min. The results are expressed as a ratio of the density of the RmATG8a and RmATG8b products to the density of the actin products from the same template. Quantitative assessment of band intensity was performed by Image Lab™ software (Bio-Rad, CA, USA).
Cloning and sequencing
To confirm positive PCR products, amplicons were purified and then cloned into the pCR®2.1-TOPO T/A plasmid vector (Invitrogen, Carlsbad, CA, USA) which was used to transform electro-competent Escherichia coli TOP10 cells. The E. coli were then incubated overnight at 37 °C on LB plates containing ampicillin and X-gal. Ten white or light blue colonies were isolated and cultured overnight in LB medium containing 50 µg/ml ampicillin. Plasmid DNA was extracted using QIAprep Spin Miniprep Kit (Qiagen, Hilden, Germany) according to manufacturer’s instructions. The positive transformants were analyzed by PCR using forward and reverse M13 primers. Once the correct clone was identified, the plasmid DNA was submitted to the Laboratorio Nacional de Biotecnología Agrícola, Médica y Ambiental (LANBAMA) of IPICYT (San Luis Potosí, México) for sequence analysis.
Bioinformatic analysis
All primers were designed with the public Primer3 software v. 4.0.0 (http://primer3.wi.mit.edu/; Rozen and Skaletsky 2000). The search for similarity among sequences was made using the BLAST service from the National Center for Biotechnology Information (NCBI, USA) (http://blast.ncbi.nlm.nih.gov/Blast.cgi; Altschul et al. 1990). The nucleotide to protein sequence translation was made using the Translate Tool (http://web.expasy.org/translate/; Gasteiger et al. 2003). Alignments of the amino acid and nucleotide sequences were performed using the multiple sequence alignment program ClustalW (http://www.genome.jp/tools/clustalw/; Larkin et al. 2007). The molecular weight and theoretical isoelectric point were predicted by ProtParam tool (http://web.expasy.org/protparam/; Gasteiger et al. 2005). Phylogenetic analysis was performed using the CLC Main Workbench 6.0 software (CLC bio, Cambridge, MA, USA), with 1,000 bootstrap replicates.
Statistical analysis
The density ratio data of RmATG8a and RmATG8b products were analyzed by one-way ANOVA using Statgraphics™ 5.1 software. Tukey’s test was used to determine significant differences of each gene at different developmental stages and adult organs in R. microplus. The data represent mean ± SE from three independent biological replicates.
Results
ATG homologues in Rhipicephalus microplus
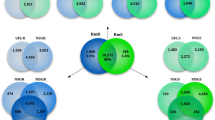
We conducted a search for ATG homologues in R. microplus, screening the BmiGI database (http://compbio.dfci.harvard.edu/tgi/cgi-bin/tgi/gimain.pl?gudb=b_microplus) with the nucleotide sequences of the five ATG genes reported so far from H. longicornis (HlATG3, HlATG4, HlATG6, HlATG8 and HlATG12) plus putative ATG5, ATG7, ATG13 and ATG16 genes from I. scapularis (GenBank accession numbers can be found at Malagoli et al. 2010; Kawano et al. 2011). Based on sequence similarity we predicted five putative ATG genes (ATG3, ATG4, ATG6 and two ATG8s) (Online Resource 1; Table 1). We were unable to find any sequences related to ATG5, ATG7, ATG12, ATG13 and ATG16 genes, most likely due to the incompleteness of the BmiGI database. We were particularly interested in ATG8s because their products perform an important role in the formation of double-membrane autophagosomes, a central step in the intracellular degradation pathway of autophagy. The search of HlATG8 homologues in the BmiGI database revealed three tentative consensus sequences (TCs) (TC16290, TC15563 and TC20934). The ORF prediction and the alignment of the deduced amino acid sequences showed that the first two TCs encode identical residues, which suggests that they come from the same gene. The third TC encodes a similar, though different protein, indicating the presence of another gene. Specifically, the first gene is predicted to be a GABARAP, whereas the second one is more similar to MAP1LC3, although both of them belong to the same ATG8 family. Following the nomenclature adopted by Umemiya-Shirafuji et al. (2010) to name tick ATG genes, we designated these genes as RmATG8a and RmATG8b, respectively. We performed a phylogenetic analysis including all ATG sequences from ticks to date. The phylogenetic tree shows that all ATG8 sequences form a distinctive cluster, suggesting that RmAtg8a and RmAtg8b are homologous to Atg8 from H. longicornis and I. scapularis given their close relatedness (Online Resource 1).
Identification of a cDNA encoding RmATG8a
Given the high sequence similarity between the first two TCs, we chose the TC15563 for further characterization. We designed oligonucleotides and amplified a PCR product of 872 bp length. This amplicon was cloned and sequenced. The flanked region is 830 bp, has an ORF at position 17–371 and codes for a 117-amino acid polypeptide with a predicted molecular mass of 13.91 kDa and a pI of 7.92 (Fig. 1). The deduced polypeptide appears to have no signal peptide. The protein is identical in length to Atg8s from H. longicornis and I. scapularis but shorter than Atg8s from other arthropods. The identified protein was queried against the non-redundant protein database using the BLAST tool, being the best BLAST hits HlAtg8 of H. longicornis (e-value 8e−77), putative γ-aminobutyric acid type A receptor-associated protein (GABARAP) of I. scapularis (e-value 3e−75), Atg8a of Drosophila melanogaster (e-value 3e−74) and GABARAP of Aedes aegypti (e-value 5e−73). BLAST analysis showed 98.3, 96.6, 94.0 and 92.3 % identities between RmAtg8a and HlAtg8 of H. longicornis, putative GABARAP of I. scapularis, Atg8a of D. melanogaster and GABARAP of A. aegypti, respectively. All amino acid sequences compared contain the conserved C-terminal glycine residue (see “Discussion” section) as well as a predicted GABARAP domain (Fig. 2).
Alignment of the deduced amino acid sequence of RmAtg8a, HlAtg8 of Haemaphysalis longicornis (accession no. BAI82577), putative GABARAP of Ixodes scapularis (accession no. XP_002408370), Atg8a of Drosophila melanogaster (accession no. NP_727447), and GABARAP of Aedes aegypti (accession no. XP_001652571). Asterisks indicate identical residues. GABARAP domain is shaded in grey. The predicted tubulin binding site is underlined and the predicted ATG7 binding site is shaded in dark grey. The conserved glycine is boxed
Identification of a cDNA encoding RmATG8b
Based on TC20934 sequence we designed oligonucleotides and amplified a PCR product of 500 bp length which was sequenced. The flanked region is 459 bp and it has an ORF at position 49–424 that codes for a 124-amino acid polypeptide with a predicted molecular mass of 14.39 kDa and a pI of 8.58 (Fig. 3). No signal peptide was found in the predicted polypeptide. The protein is similar in length to Atg8s from other species. The deduced sequence was subjected to BLAST analysis. Among sequences that produced significant alignments were Microtubule-associated protein one light chain three (MAP1LC3) of Danio rerio (e-value 2e−48), MAP1LC3 of S. scrofa (e-value 9e−45) and MAP1LC3 of Homo sapiens (e-value 7e−43). BLAST analysis showed 58.06 and 54.54 % identities among RmAtg8b and MAP1LC3 of D. rerio and MAP1LC3 of S. scrofa and H. sapiens, respectively. All compared sequences contain the conserved C-terminal glycine residue as well as a predicted GABARAP domain (Fig. 4).
Alignment of the deduced amino acid sequence of RmAtg8b, MAP1LC3C of Danio rerio (accession no. NP_956592), MAP1LC3A of Sus scrofa (accession no. NP_001164298), and MAP1LC3A of Homo sapiens (accession no. NP_115903). Asterisks indicate identical residues. GABARAP domain is shaded in grey. The predicted tubulin binding site is underlined and the predicted ATG7 binding site is shaded in dark grey. The conserved glycine is boxed
Expression patterns of RmATG8a and RmATG8b in Rhipicephalus microplus
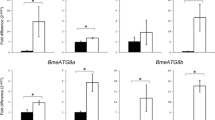
The highest expression level of RmATG8a mRNA was observed at the egg and larval stages. Interestingly, adult tissues (ovary and midgut) coming from fed ticks showed a statistically significant lower expression (Fig. 5a). The expression pattern of RmATG8b mRNA was very similar (Fig. 5b).
RmATG8a (a) and RmATG8b (b) mRNA expression in different stages of Rhipicephalus microplus. RT-PCR reactions were performed of total RNA from egg, larvae and adult tissues (ovary and midgut). Tick actin was used as a control. The data are expressed as a density ratio of RmATG8s and actin products. The data represent mean ± SE from three independent biological replicates
Discussion
The exceptional ability of R. microplus, and in ticks in general, to survive long-term starvation has been related to a process of intracellular protein degradation called autophagy. This process is well conserved in metazoans; however, the study in ticks is still in its infancy. Recently, five ATG genes were characterized in the tick H. longicornis (Umemiya-Shirafuji et al. 2010; Kawano et al. 2011). In this study, we revisited the B. microplus gene index, BmiGI (Guerrero et al. 2005) looking for ATG homologues. We found five putative ATG genes (ATG3, ATG4, ATG6 and two ATG8s). Unfortunately, we could not identify ATG5, ATG7, ATG12, ATG13 and ATG16 homologues in the database. To the best of our knowledge, this is the first time ATG genes are reported on R. microplus.
Atg8 proteins are well conserved in organisms from yeast to higher eukaryotes and are involved in diverse intracellular trafficking and autophagy processes (Le Grand et al. 2013). Animal Atg8 proteins comprise three subfamilies: MAP1LC3, GABARAP and Golgi-associated ATPase enhancer of 16 kDa (GATE-16). Corresponding genes have been both duplicated and lost during evolution (Shpilka et al. 2011). As a consequence, the subfamilies are more or less represented in specific lineages. This is clearly seen in arthropods, where ticks such as H. longicornis have only a GABARAP gene, but the blacklegged tick, I. scapularis, has been predicted to have GATE-16, GABARAP, and MAP1LC3 genes. On the other hand, the honey bee, Apis mellifera, has a GABARAP and a MAP1LC3 gene, and the fruit fly, D. melanogaster, has two GABARAP genes (Shpilka et al. 2011). In this study we identified a GABARAP and a MAP1LC3 gene in the tick R. microplus. These findings are relevant given the central role that their products play in the autophagy process.
Atg8 and its orthologs are synthesized as inactive precursors with a C-terminal extension and they undergo a cleavage of the extension to expose a C-terminal glycine residue (Klionsky and Schulman 2014). This cleavage is mediated by the cysteine protease Atg4, which is specific to Atg8 proteins. The cleaved Atg8 is then covalently conjugated to the membrane lipid phosphatidylethanolamine (PE) through its exposed glycine to promote the autophagosome formation and expansion (Shpilka et al. 2011). The proteolysis and posterior conjugation of Atg8 to PE on the autophagic membrane are essential and remain conserved through evolution. Interestingly, RmAtg8a and RmAtg8b identified in the present study show this conserved C-terminal glycine residue, as well as the GABARAP domain, an ubiquitin-like domain shared by all Atg8s (Figs. 2, 4).
With respect to the expression levels of both RmATG8a and RmATG8b, they were higher in both the egg and larvae when the tick is in a starvation state. Whereas in adults, expression of the genes were lower in ovary and midgut. These results are consistent with a previous study in H. longicornis tick where the transcripts of HlATG8 were elevated in egg and unfed larvae and nymphs and decreased after a blood meal in fed larvae and nymphs as well as in the adult midgut (Umemiya-Shirafuji et al. 2010, 2014). Our findings suggest that autophagy occurs in R. microplus as in other species. The process may be initiated in the egg and larval stage, while the tick is unable to feed from host blood, and turned off once the tick reaches a host.
Future studies are needed to demonstrate the participation of RmATG8a and RmATG8b in the autophagy process in R. microplus, which may shed new light on how ticks can survive long-term starvation. As well as whether these proteins have redundant functions or each member participates in the formation of specialized autophagosomes. Finally, autophagic process seen as a survival strategy in ticks could well represent a target from a new class of acaricides.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
da Silva VIJr, Imamura S, Nakajima C, de Cardoso FC, Ferreira CAS, Renard G, Masuda A, Ohashi K, Onuma M (2005) Molecular cloning and sequence analysis of cDNAs encoding for Boophilus microplus, Haemaphysalis longicornis and Rhipicephalus appendiculatus actins. Vet Parasitol 127:147–155
Davey RB, Garza J, Thompson GD, Drummond RO (1980) Ovipositional biology of the southern cattle tick, Boophilus microplus (Acari: Ixodidae), in the laboratory. J Med Entomol 17:117–121
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker John M (ed) The proteomics protocols handbook. Humana Press, New York, pp 571–607
Guerrero FD, Miller RJ, Rousseau ME, Sunkara S, Quackenbush J, Lee Y, Nene V (2005) BmiGI: a database of cDNAs expressed in Boophilus microplus, the tropical/southern cattle tick. Insect Biochem Mol Biol 35:585–595
Kawano S, Umemiya-Shirafuji R, Boldbaatar D, Matsuoka K, Tanaka T, Fujisaki K (2011) Cloning and characterization of the autophagy-related gene 6 from the hard tick, Haemaphysalis longicornis. Parasitol Res 109:1341–1349
Klionsky DJ, Schulman BA (2014) Dynamic regulation of macroautophagy by distinctive ubiquitin-like proteins. Nat Struct Mol Biol 21:336–345
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Le Grand JN, Bon K, Fraichard A, Zhang J, Jouvenot M, Risold PY, Boyer-Guittaut M, Delage-Mourroux R (2013) Specific distribution of the autophagic protein GABARAPL1/GEC1 in the developing and adult mouse brain and identification of neuronal populations expressing GABARAPL1/GEC1. PLoS One 8(5):e63133 15
Lilienbaum A (2013) Relationship between the proteasomal system and autophagy. Int J Biochem Mol Biol 4:1–26
Malagoli D, Abdalla FC, Cao Y, Feng Q, Fujisaki K, Gregorc A, Matsuo T, Nezis IP, Papassideri IS, Sass M, Silva-Zacarin ECM, Tettamanti G, Umemiya-Shirafuji R (2010) Autophagy and its physiological relevance in arthropods: current knowledge and perspectives. Autophagy 6:575–588
Nijhof AM, Balk JA, Postigo M, Jongejan F (2009) Selection of reference genes for quantitative RT-PCR studies in Rhipicephalus (Boophilus) microplus and Rhipicephalus appendiculatus ticks and determination of the expression profile of Bm86. BMC Mol Biol 10:112
Ribeiro VLS, Toigo E, Bordignon AL, Gonçalves K, von Poser G (2007) Acaricidal properties of extracts from the aerial parts of Hypericum polyanthemum on the cattle tick Boophilus microplus. Vet Parasitol 147:199–203
Rosario-Cruz R, Guerrero FD, Miller RJ, Rodriguez-Vivas RI, Dominguez-García DI, Cornel AJ, Hernandez-Ortiz R, George JE (2005) Roles played by esterase activity and by a sodium channel mutation involved in pyrethroid resistance in populations of Boophilus microplus (Acari: Ixodidae) collected from Yucatan, Mexico. J Med Entomol 42:1020–1025
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana, Totowa, pp 365–386
Ryter SW, Cloonan SM, Choi AMK (2013) Autophagy: a critical regulator of cellular metabolism and homeostasis. Mol Cells 36:7–16
Shpilka T, Weidberg H, Pietrokovski S, Elazar Z (2011) Atg8: an autophagy-related ubiquitin-like protein family. Genome Biol 12(7):226
Spickler AR (2007) Rhipicephalus (Boophilus) microplus. http://www.cfsph.iastate.edu/DiseaseInfo/factsheets.php. Accessed 3 Oct 2013
Tsuda A, Mulenga A, Sugimoto C, Nakajima M, Ohashi K, Onuma M (2001) cDNA cloning, characterization and vaccine effect analysis of Haemaphysalis longicornis tick saliva proteins. Vaccine 19:4287–4296
Umemiya R, Matsuo T, Hatta T, Sakakibara S, Boldbaatar D, Fujisaki K (2007) Cloning and characterization of an autophagy-related gene, ATG12, from the three-host tick Haemaphysalis longicornis. Insect Biochem Mol Biol 37:975–984
Umemiya R, Matsuo T, Hatta T, Sakakibara S, Boldbaatar D, Fujisaki K (2008) Autophagy-related genes from a tick, Haemaphysalis longicornis. Autophagy 4:79–81
Umemiya-Shirafuji R, Matsuo T, Liao M, Boldbaatar D, Battur B, Suzuki H, Fujisaki K (2010) Increased expression of ATG genes during nonfeeding periods in the tick Haemaphysalis longicornis. Autophagy 6:473–481
Umemiya-Shirafuji R, Linggatong RG, Maeda H, Kawano S, Tanaka T, Fukumoto S, Suzuki H, Tsuji N, Fujisaki K (2014) Expression analysis of autophagy-related genes in the hard tick Haemaphysalis longicornis. Vet Parasitol 201:169–175
Yoshimoto K, Hanaoka H, Sato S, Kato T, Tabata S, Noda T, Ohsumi Y (2004) Processing of ATG8s, ubiquitin-like proteins, and their deconjugation by ATG4 s are essential for plant autophagy. Plant Cell 16:2967–2983
Acknowledgments
This work was supported by a Grant SEP-CONACYT-CB-2008-01-104006 from Consejo Nacional de Ciencia y Tecnología México to M.M.V. J.M.F.F. received a scholarship from CONACYT. We thank Tina Coop of Peace Corps México for the critical review of the language of the manuscript. The authors declare that they have no conflict of interest. All experiments comply with the current laws of Mexico.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Flores Fernández, J.M., Gutiérrez Ortega, A., Rosario Cruz, R. et al. Molecular cloning and characterization of two novel autophagy-related genes belonging to the ATG8 family from the cattle tick Rhipicephalus (Boophilus) microplus (Acari: Ixodidae). Exp Appl Acarol 64, 533–542 (2014). https://doi.org/10.1007/s10493-014-9838-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10493-014-9838-3