Abstract
“Masseiras” is an ancient Portuguese agriculture system, where soil was developed from sand dunes enriched with seaweeds over more than a century. Due to the importance for the local economy, this system evolved for greenhouse structures. In this study we compared the bacterial community composition and structure of “Masseiras” soil, aiming at assessing the potential impact of different agricultural practices. The bulk soil of two greenhouses (following or not the recommended agriculture good practices, FGP and NFGP, respectively) was compared based on their physicochemical properties and bacterial community. In both FGP and NFGP, Proteobacteria, Acidobacteria, Actinobacteria, Bacteroidetes, Firmicutes, and Gemmatimonadetes were in a proportion of 5:1:1:1:1:1. However, the bacterial community of soil FGP was richer and more diverse than that of soil NFGP. Members of the classes Bacilli and Gemm-1, with higher relative abundance in NFGP and FGP, respectively, were those contributing most for distinguishing the bacterial communities of both soils. The differences in the structure of the bacterial communities correlated (Mantel test) with some soil physicochemical properties, such as electrical conductivity and nitrate and Zn contents, which were significantly higher in soil NFGP than in soil FGP.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
“Masseiras” fields are ancient and unique agriculture systems developed at the end of the XIX century in the Northwest coast region of Portugal. These systems were manually constructed in dug sand dunes, and protected from the wind by cultures such as vineyards. Soils became enriched in organic matter by using seaweed-based soil amendments. Over the last three decades, this mode of intensive production evolved and the natural wind protection system was substituted by simple plastic greenhouse structures, while seaweed-based amendment was gradually replaced by synthetic fertilizers (Fonseca 2010; Melo et al. 2012). Yet, the intensive horticultural production of tomato, lettuce, pepper, cucumber or melons is still a common practice, involving the use of biological or chemical fertilization and pest control (Melo et al. 2012). This production system offers high yields, being an important supplier of agricultural produce for local and foreign markets and contributing to increase the income of farmers. Although today the roots of the original ancient system production can still be witnessed, the agriculture practices differ among farmers, from highly sustainable procedures to erratic soil amendments. As a consequence, there are risks of soil deterioration, in particular due to intensive fertilization and soil salinization. Soils under intensive agriculture production are known to be subjected to strong human impacts, where land management practices have significant and long-lasting effects on physicochemical soil properties (e.g., Bronick and Lal 2005; Knops and Tilman 2000; van Diepeningen et al. 2006), and also on bacterial communities (Buckley and Schmidt 2003; Lopes et al. 2014, 2015). Two important drivers of properties of agricultural soils include the parent materials, i.e. the initial geological elements that by weathering originate a particular soil, and the history of cultivation and management (Acosta-Martínez et al. 2010; Ulrich and Becker 2006). In addition, soil properties are strongly dependent on soil microbiome composition and structure due to their role in soil biochemistry (Gadd 2010; Huang et al. 2014; Oehl et al. 2001) and soil aggregation (Bronick and Lal 2005; Mager and Thomas 2011), among others. However, soil microbiome can be also influenced by properties such as pH, organic carbon content, total nitrogen, minerals or salinity (Fierer and Jackson 2006; Gadd 2010; Lauber et al. 2009). The particular nature of Masseiras fields suggested that it could hold a unique bacteriome. A hypothesis that was behind the current study was that agriculture practices that do not follow recommended guidelines may contribute to disturb such a unique microbiome. This hypothesis is supported by previous descriptions, made for other microbiomes (Figuerola et al. 2012). Also aware of the importance of an adequate soil management, the European Union Common Agricultural Policy (CAP) guidelines were created with the aim of sustaining the balance between the conservation of the natural environment and farming practices. These guidelines aim to mitigate the adverse impacts that farming may have on natural resources (pollution of soil, water and air; fragmentation of habitats and loss of wildlife) and, thus, on the farming sustainability Monitoring Agriculture ResourceS (MARS) 2015 (http://ec.europa.eu/agriculture/envir/index_en.htm). To test our hypothesis, two Masseiras greenhouses differing in the implementation and compliance with the recommended good practices were selected for this study. The aim was to assess whether the adoption of non-recommended practices would be noticeable in the bacteriome of these Masseiras soils, by comparing soil physicochemical properties and bacterial community structure.
Materials and methods
Site description and sampling
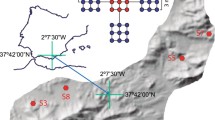
Soil samples were collected from two private Masseiras-type greenhouses (FGP and NFGP) located in Northwest Portugal (41°25′N; 8°45′W). In both greenhouses it is practiced conventional agriculture, although both differ in the frequency, doses, rotation and sequence of phytochemicals used. In greenhouse FGP (ca. 60 × 8 m), the recommended procedures for soil amendments (manure, mushroom growing substrates, N–P–K fertilizers) and pest control (biological insect control and fungicides) are in compliance with the Good Agricultural and Environmental Practices (i.e. supplying of nutritional compounds based on a previous professional assessment of the requirements of the crops versus their availability in soil through periodic analysis of the soil physicochemical parameters; minimize the use of synthetic plant protection products, which, when necessary, are supplied according to recommended guidelines and adequate for the horticultural sector in the region) (https://marswiki.jrc.ec.europa.eu/wikicap/index.php/Main_Page). In contrast, in greenhouse NFGP (ca. 50 × 10 m), these good practices were not observed, being the nutritional and phytosanitary compounds supplied on the basis of trial-and-error methods. In greenhouse FGP different crops (i.e. tomato, lettuce, etc.) are rotated in monoculture. In greenhouse NFGP distinct crops are grown in different windrows (polyculture). These greenhouses were selected because among those giving permission for sampling i) they differed in the management practices and ii) at the sampling period were being cultivated with the same crop. Samples were collected in the area dedicated to tomato (Solanum lycopersicum L.) crops. In each greenhouse three composite samples were collected from three different windrows at 20th July of 2014. Each composite sample consisted of six individual bulk soil cores collected from the upper 0–30 cm of a given windrow at intervals of ca. 2 m, pooled and homogenized.
Soil samples were characterised for physicochemical properties (pHH2O, pHKCl, electrical conductivity, organic matter, humic substances and water-extractable (relation of 1:5 w/v) and nutrients content at the Laboratório de Solos e Plantas from the University of Trás-os-Montes. Soil texture was determined according to Guitián Ojea and Carballas (1976). Heavy metals content (Cd, Cr, Cd, Ni, Pb and Zn) were analysed by flame atomic absorption spectrophotometry (AAS) (Perkin Elmer Atomic Absorption Spectrophotometer) after sample digestion with aqua regia (mixture of 65% nitric acid and 37% hydrochloric acid in a ratio of 1:3), followed by filtration. The exchangeable fraction was performed using the first step of the BCR (Community Bureau of Reference) extraction procedure (Rauret et al. 2000), as follows: 1 g of dry soil was introduced into a polypropylene centrifuge tube containing 40 mL of 0.11 M CH3COOH and then shaken for 16 h at room temperature. The aqueous phase, corresponding to the acid soluble/exchangeable fraction, was separated by centrifugation at 3000 rpm for 20 min. Exchangeable metals (Cd, Cr, Cd, Ni, Pb and Zn) were determined by AAS as described above.
Enumeration of bacteria and DNA extraction
In order to determine the load of bacteria in each sampled soil and not only the bacterial diversity and composition, the abundance of bacteria was estimated based on culture-dependent and 16S rRNA gene-based quantitative PCR (qPCR) methods. Fast growing cultivable microorganisms were enumerated based on the membrane filtration method as described by Lopes et al. (2011). Briefly, 5 g of soil was suspended in 45 ml of a sterile solution of sodium hexametaphosphate (1%, w/v) and stirred for 30 min. Serial dilutions of the soil suspensions were filtered through cellulose nitrate membranes (0.22 µm pore size, 47 mm membrane, Sartorius Stedim Biotech, Goettingen, Germany), which were placed onto Plate Count Agar (Merck) and incubated 3 days at 30 °C. Data from triplicates was expressed as colony forming units (CFU) per g of dry soil (oven dried at 105 °C).
Total DNA was extracted from each composite sample using the Power Soil™ DNA Isolation Kit (MO BIO) as described before (Lopes et al. 2011). DNA extracts were stored at −20 °C until further analysis. Quantification of the 16S rRNA gene was performed through qPCR (StepOne™ Real-Time PCR System; Life Technologies, USA) using the primers 1114F (CGGCAACGAGCGCAACCC) and 1275R (CCATTGTAGCACGTGTGTAGCC) as described by Sousa et al. (2016). Briefly, a standard curve was prepared using serial ten-fold dilutions of genomic DNA of Escherichia coli ATCC 25992 extracted with QIAamp DNA Stool Kit (QIAGEN, The Netherlands). The amplification conditions were as follows: 95 °C for 10 min (1 cycle); 95 °C for 15 s, 55 °C for 20 s and 72 °C for 10 s (35 cycles). Melting curves at increments of 0.3 °C from 60 to 95 °C were used to assess the homogeneity of the amplification. Under these conditions, the limit of quantification was 385 gene copies, and 100% efficiency was achieved. Results were analysed using the StepOneTM v.2.3 software (Life Technologies). Data from triplicates was expressed as copy number of 16S rRNA gene per g of dry soil.
Bacterial community analysis
Bacterial community analyses were performed using 454-pyrosequencing, targeting the hypervariable region V3/V4 of the 16S rRNA gene, using fusion primers containing the Roche-454 A and B Titanium sequencing adapters and an eight-base barcode sequence in fusion primer B (V3F—5′-ACTCCTACGGGAGGCAG-3′ and V4R 5′-TACNVRRGTHTCTAATYC-3′; Wang and Qian 2009). PCR reactions were performed as previously described (Lopes et al. 2014). Electrophoresis of the PCR products was undertaken in a 1% (w/v) agarose gel and the ~525 bp amplified fragments were purified using AMPure XP beads (Agencourt, Beckman Coulter, USA) according to manufacturer’s instructions. The amplified DNA was quantified by fluorimetry with PicoGreen dsDNA quantitation kit (Invitrogen, Life Technologies, Carlsbad, California, USA), pooled at equimolar concentrations and sequenced with GS 454 FLX Titanium chemistry, according to manufacturer’s instructions (Roche, 454 Life Sciences, Brandford, CT, USA) at Genoinseq (Cantanhede, Portugal). The raw reads have been deposited into the NCBI short-reads archive database (accession number: SAMN05196279–SAMN05196284).
Sequences were assigned to samples by the 8-bp barcode using QIIME pipeline (Caporaso et al. 2010b) and sequences shorter than 300 bp, with quality scores lower than 25 and with more than 1 undetermined base were eliminated. Chimeric sequences were identified and removed using USEARCH v6.0 (Edgar et al. 2011). Free-chimeric sequences were further grouped into operational taxonomic units (OTUs) using UCLUST (Edgar 2010) with a phylotype threshold of ≥97% sequence identity and were taxonomically assigned using QIIME defaults. Sequences of each OTU were aligned using PyNAST (Caporaso et al. 2010a) and were classified at the 97% identity level using Ribosomal Database Project (RDP) classifier (Wang et al. 2007) trained on the GreenGenes 16S rRNA database (v13_8 release) (DeSantis et al. 2006). After excluding the sequences not assigned to the domain Bacteria (about 0.5% of the sequences), a total of 47392 high-quality sequences were obtained, distributed in 6064 OTUs. From those, 3400 OTUs were represented by more than one sequence. OTUs represented by only one sequence (singletons) were excluded from further analysis. Since a variable number of sequences per library (sample) was obtained, the number of sequences per sample was normalized according to the smallest library (3040 sequences). Alpha diversity metrics including Shannon index, phylogenetic diversity (PD) whole tree, number of OTUs, and Simpson index were calculated after normalization according to smallest library (3040 sequences) (Faith 1992; Shannon and Weaver 1963; Simpson 1949). Beta diversity patterns were assessed using the weighted UniFrac metric (Lozupone and Knight 2005), which considers both the abundance and phylogenetic distance of each OTU, and the results presented as Principal Coordinate Analysis (PCoA) biplot that defined the position of the bacterial groups contributing most to the variations among the six datasets analysed.
Statistical analysis
Differences between soil physicochemical properties, bacterial abundance and alpha-diversity metrics were achieved by the Student’s t test for independent means using the SPSS software (version 20.0, IBM Software, Chicago, Illinois). Differences of bacterial taxon relative abundance between soils were achieved using a pairwise t-test with the Storey FDR correction for multiple comparisons as implemented in the STAMP software v.2.1.3 (Park et al. 2014). Cohen’s d effect size (Ellis 2010), used to indicate the standardized difference between the two means, was calculated using the Microsoft Excel spreadsheet. Relationships between soil physicochemical properties and bacterial communities were evaluated through the Mantel test (999 iterations), calculated upon the OTU weigthed Unifrac distance matrix and the distance matrix of each physicochemical parameter, obtained using the QIIME pipeline. Significance of the clusters observed in PCoA was assessed through Adonis test (permutational multivariate non-parametric analysis of variance) as implemented in QIIME software (Anderson 2001).
Results
Soil physicochemical properties, soil metal content and bacterial abundance
Most of the physicochemical parameters of soil NFGP were significantly different (p < 0.05) of those of soil FGP (Table 1). Specifically, soil NFGP presented significantly (p < 0.05) lower pH, and sixfold and threefold higher NO3 content and electrical conductivity than soil FGP, respectively. Accordingly, soil NFGP had higher content of soluble ions, such as Ca2+, Mg2+ (fourfold higher) and K+ (1.5-fold higher) than FGP soil. In addition, the total metal content was significantly higher in soil NFGP than in FGP, except in the case of Ni. In the analysis of the fraction of exchangeable metals, only Zn was detected (6.4 ± 0.8 mg kg−1 in FGP and 15.0 ± 1.3 mg kg−1 in NFGP), while the others were below detection limit (Cu <2.0 mg kg−1; Ni <0.3 mg kg−1; Cr <2.3 mg kg−1; Cd <0.3 mg kg−1; Pb <3.7 mg kg−1). In contrast, the abundance of cultivable heterotrophic bacteria and the copy number of 16S rRNA gene per g of dry soil was similar in both greenhouse soils (Table 1).
Bacterial community structure and composition
After normalization, a total of 18,240 sequences were obtained (3040 sequences per library), corresponding to 2908 OTUs (Table 2). Out of these, 1340 OTUs were observed in both soils, and 174 OTUs were present in all the six libraries analysed. The relative abundance (number of the sequences per OTU/total number of normalized sequences) of these 174 core OTUs was similar for both soils (44.2 and 40.3% for soils NFGP and FGP, respectively).
The bacterial communities in both soils were represented by sequences affiliated to 34 phyla, 10 of which with relative abundances >1%. In both FGP and NFGP, Proteobacteria was the most abundant phylum (average of about 51%) and Alphaproteobacteria was the dominant class (average of 23 and 26% in soils FGP and NFGP, respectively). However, the abundance of other bacterial groups differed in soils FGP and NFGP (Fig. 1a). While in soil FGP Deltaproteobacteria (11.1%), Gammaproteobacteria (9.7%) and Betaproteobacteria (7.6%) were the most abundant classes (Fig. 1a), in soil NFGP Bacilli were dominant (12.0%), accompanied by Gammaproteobacteria (11.5%) and Deltaproteobacteria (8.4%). Indeed, Firmicutes were 2.4 times more abundant in soil NFGP (12.2%) than in soil FGP (p = 0.017; Cohen’s d = 3.8), mainly due to members of the class Bacilli (p = 0.045; Cohen’s d = 3.9) (Fig. 1a). In contrast, Nitrospirae and Gemmatimonadetes, in particular members of the class Gemm-1, were more abundant in soil FGP than in soil NFGP (p = 0.017; Cohen’s d = 5.0 and p = 0.028; Cohen’s d = 6, respectively) (Fig. 1a).
A total of 40.6% of the sequences could be affiliated to a known family, being distributed in a total of 152 families. Of these, 20 families represented more than 1% of the total sequences in soil FGP or NFGP, although a distinct pattern of distribution was observed in both soils (Fig. 1b). The most abundant families in soil FGP were Hyphomicrobiaceae, Rhodospirillaceae (both of class Alphaproteobacteria) and Cytophagaceae (Bacteroidetes), each representing about 4% of the total number of sequences. In soil NFGP, the most prevalent families were Bacillaceae (Firmicutes), Hyphomicrobiaceae, Sphingomonadaceae (both of class Alphaproteobacteria), Xanthomonadaceae (Gamaprotoeobacteria) and Cytophagaceae, with relative abundances varying between 6.5% and 3.8%. Among the most notorious differences were the relative abundance of Bacillaceae and Sphingomonadaceae, 4.3- and 2.5-fold higher in soil NFGP than in soil FGP (p = 0.041; Cohen’s d = 4.2 and p = 0.033; Cohen’s d = 4.9, respectively) (Fig. 1b).
Twenty two per cent of the total sequences could be affiliated to a known genus, being distributed by 180 genera. Only 12 of these genera had a relative abundance higher than 0.5% in at least one of the soils. Figure 2 presents the distribution of genera for the most prevalent families. Within the families Cytophagaceae, Hyphomonadaceae and Rhodospirillaceae more than 80% of the sequences could not be affiliated with any known genus (Fig. 2). On the contrary, most of the reads affiliated to the families Bacillaceae and Sphingomonadaceae had significant identity with known genera (Fig. 2). Most of the members of the family Bacillaceae belonged to the genus Bacillus in both soils (89.7 and 80.8% in FGP and NFGP, respectively). The dominant genus of the family Sphingomonadaceae was Kaistobacter in both soils, although with different relative abundance values, 54.5 and 84.1% in FGP and NFGP soil, respectively (Fig. 2). The abundance of reads affiliated to the genera Sphingobium and Sphingomonas was also different, both more prevalent in soil FGP than in NFGP (18.5 vs. 2.1% and 15.9 vs. 8.6%, respectively). In contrast, soil NFGP had higher prevalence of Luteimonas and Dokdonella, both of the family Xanthomonadaceae, than soil FGP (10.3 vs 0.5% and 29.7 vs 15.1% respectively).
Relative abundance of genera from the more prevalent families (as represented in the PCoA biplot, see Fig. 3). Grey bars represent unclassified sequences at genera level. Percentages presented in y-axis indicate the average abundance of each family relative to the total sequences for each soil (n = 3). Syntrophobacteraceae was not included since none of the reads belonging to this family were classified below family level
Comparison of the bacterial diversity in soils FGP and NFGP and relationship with soil physicochemical properties
In spite of presenting a similar bacterial density and composition, the abundance of several taxonomic groups differed in soils FGP and NFGP. These differences coincided with significant differences in the richness values, calculated as the average number of OTUs as well as the Chao1 estimator of richness. Both richness metrics indicated a significantly higher richness in soil FGP than in NFGP (Table 2). This was in agreement with the observation of significantly higher Shannon and Phylogenetic Diversity indices in soil FGP than in NFGP (Table 2). Beta diversity metrics evidenced also the differences between the bacterial communities of soils FGP and NFGP, as shown in the PCoA biplot based on the weighted Unifrac distances metrics (Fig. 3). The abundance of the taxonomic groups distributed along axis 1 of the PCoA biplot, which explained 61% of the observed variance, supported the distinction of the bacterial communities of the studied soils (Fig. 3). Unclassified bacteria of the order Bacillales, and of the families Bacillaceae, Sphingomonadaceae, Xanthomonadaceae, and Cytophagaceae were more abundant in soil NFGP than in soil FGP. In contrast, unclassified the class Gemm-1 and the families Hyphomonadaceae, Rhodospirillaceae and Syntrophobacteriaceae were more abundant in soil FGP than in soil NFGP. These were the differences that most contributed to distinguish both soils (Fig. 3). The Adonis test of significant grouping (p < 0.01) confirmed that about 58% of the observed variation could be attributed to differences in the bacterial communities of the studied soils. As hypothesised, these differences were significantly correlated (p < 0.05) with several soil properties, as evidenced by the Mantel test based on the weighted UniFrac values (Table 1). Significant correlation factors with values above 0.8 were observed for soil texture, pH, NO3 and Zn content (total and exchangeable values) and between 0.7 and 0.8 for electrical conductivity, soluble Ca and Mg and total Cr content.
Discussion
One of the objectives of this study was the characterization of the bacterial community of Masseiras soil, based on the analysis of two greenhouse soils. The six most abundant phyla were Proteobacteria, Bacteroidetes, Firmicutes, Acidobacteria, Actinobacteria, and Gemmatimonadetes, which have been reported in different types of soil. However, the observed approximate proportion of the relative abundance of members of Proteobacteria, Bacteroidetes and Acidobacteria (5:1:1) highlighted the difference between Masseiras and other soils. Previous studies that characterized a large number of soils from different origins such as agricultural, forest soils and grasslands reported similar proportions of Proteobacteria and Acidobacteria, and sometimes Bacteroidetes (Fay et al. 2016; Fierer et al. 2012; Lauber et al. 2009; Lopes et al. 2014). Members of the phylum Firmicutes although often present in soils of different origins are reported at low prevalence values, typically less than 5% (Fay et al. 2016; Fierer et al. 2012; Janssen 2006; Lauber et al. 2009; Lopes et al. 2014). These contrasts may be related with the fact that none of the previous studies included greenhouse soils. Indeed, beside the fact that Masseiras is a man-made agricultural system, supposedly with a unique microbial community, intensive agriculture is carried out in these greenhouses. The greenhouse environments are known to create extremes conditions of temperature, among others. This argument is supported by previous studies that show that members of the Firmicutes, namely of family Bacillaceae, can survive desiccation, extreme environmental variation and soil use (Battistuzzi and Hedges 2009). The ability of some Bacillaceae to form endospores that can endure survival under stress conditions, is also part of the explanation for the high abundance of Firmicutes in Masseiras soil (Logan and De Vos 2009). However, also the human intervention may explain the relative high abundance of Firmicutes. Indeed, a significant increase in the abundance of Firmicutes was reported in case studies where forest soils were converted in agricultural soil or pasture (Montecchia et al. 2015; Rodrigues et al. 2013). In general, it seems adequate to conclude that the stress imposed by the intensive agriculture carried out in the studied greenhouses may have contributed to explain the comparatively high relative abundance of Firmicutes in Masseiras soil. In contrast with Firmicutes, the relative abundance of Acidobacteria in the Masseiras soils could be considered low when compared to other soils (Davis et al. 2011; Fierer et al. 2007; Lauber et al. 2009). Again, it may be related with the soil properties, since favourable conditions for Acidobacteria combine low soil pH, clay texture and the advanced soil maturation (Lauber et al. 2009; Russo et al. 2012; Sun et al. 2015). Hence, it can be argued that the physicochemical properties of Masseiras soil may not favour the proliferation of Acidobacteria, while other groups such as Proteobacteria and Firmicutes may gain advantage. Indeed, Russo et al. (2012) suggested that Proteobacteria might become dominant in relation to Acidobacteria in sandy loam soils. Gemmatimonadetes was another phylum with higher relative abundance than that commonly reported in soils. In spite of the wide distribution of members of this phylum, in pasture, crop agriculture and forests (Chan et al. 2008; Lauber et al. 2009; Li et al. 2014; Montecchia et al. 2015), their relative abundance is normally <5%. Although the biology of Gemmatimonadetes is still poorly characterized since only a few isolates were reported to date, their occurrence is apparently associated to low-moisture environments (DeBruyn et al. 2011). Therefore, the low water holding capacity of sandy soils, characteristic of Masseiras may explain the high prevalence of members of this phylum.
A second objective of this study was to preliminarily assess if different management practices, specifically those that do not follow the recommended good practices, could impact the bacterial communities and therefore, disturb the Masseiras soil bacteriome. As hypothesised, the bacterial communities were significantly different in soils FGP and NFGP. Although statistical analyses cannot be used to prove cause-effect relationships but rather to test the veracity of hypotheses, we believe that the differences observed between soil types were related with the management practices. This conclusion is based not only on the alpha diversity indicators (Table 2), which show that soil NFGP is less rich and diverse than soil FGP, but also on the beta diversity comparison (Fig. 3). Greenhouse NFGP has a poorly controlled management, which may explain the significantly higher nitrate content, as well as the high content in soluble ions, including metals, and electrical conductivity of this soil, compared with the soil FGP. These parameters may have influenced the structure and the diversity of the bacterial community of the NFGP soil. Indeed, previous studies have shown that mineral fertilization (N/P/K) disturb the soil bacterial communities, with eventual loss of richness and diversity (Ruppel et al. 2007; Allison and Martiny 2008). The higher abundance of members of the family Bacillaceae as well as of unclassified members of the order Bacillales in soil NFGP, compared with FGP, was one of the most noticeable differences between both greenhouse soils. As pointed out above, members of the family Bacillaceae have been described as resistant and/or tolerant to stressful conditions, including to metals such as Cu, Cr and Zn (Faisal and Hasnain 2004; Sun et al. 2010; Taniguchi et al. 2000), and their abundance has been positively related with metal content in commercial composts (Silva et al. 2016). Hence, the management of the greenhouse NFGP may have imposed an increased stress (nitrates, electrical conductivity, metals) on the microbial community, which could explain, at least in part, the high abundance of the phylum Firmicutes.
Despite the worldwide increasing greenhouse production of vegetables, studies on the effect of this type of intensive agricultural management on soil microbiota is still scarce. Further studies are necessary to assess if the particular conditions of greenhouse farming may contribute to the deterioration of the soil microbial diversity, essential for soil fertility and protection against pathogenic or otherwise adverse microorganisms.
Conclusions
A unique pattern of bacterial phyla was observed for the greenhouse soil of Masseiras, which comprised Proteobacteria, Acidobacteria, Actinobacteria, Bacteroidetes, Firmicutes, and Gemmatimonadetes in a proportion of 5:1:1:1:1:1. This composition is probably a result of the man-made nature of this sandy soil combined with the fact that it represents a greenhouse system.
The influence of fertilization procedures on the soil microbiota was inferred from the comparative analyses of two greenhouses soils with distinct management practices while producing the same agricultural product. As could be inferred from the Mantel test, specific physicochemical parameters, such as nitrate and Zn content and electrical conductivity were correlated with shifts of some bacterial community members, e.g. Bacillaceae.
Although no general conclusions can be retrieved from a single study, these are interesting results, evidencing the relevance of studies to assess microbiome disturbances in greenhouse systems, mainly when good agriculture practices are disregarded. An interesting implication of this kind of studies is that by demonstrating the disturbance of the microbial communities it is possible to provide science-based evidences of soil deterioration due to inadequate soil fertilization and management, putting in cause soil fertility and, eventually, the safety of the food products. Both aspects with highly relevant societal and economic implications.
References
Acosta-Martínez V, Dowd SE, Sun Y, Wester D, Allen V (2010) Pyrosequencing analysis for characterization of soil bacterial populations as affected by an integrated livestock-cotton production system. Appl Soil Ecol 45:13–25. doi:10.1016/j.apsoil.2010.01.005
Allison SD, Martiny JBH (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA 105:11512–11519. doi:10.1073/pnas.0801925105
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Aust J Ecol 26:32–46. doi:10.1111/j.1442-9993.2001.01070.pp.x
Battistuzzi FU, Hedges SB (2009) A major clade of prokaryotes with ancient adaptations to life on land. Mol Biol Evol 26:335–343. doi:10.1093/molbev/msn247
Bronick CJ, Lal R (2005) Soil structure and management: a review. Geoderma 124:3–22. doi:10.1016/j.geoderma.2004.03.005
Buckley D, Schmidt T (2003) Diversity and dynamics of microbial communities in soils from agro-ecosystems. Environ Microbiol 5:441–452. doi:10.1046/j.1462-2920.2003.00404.x
Caporaso JG, Bittinger K, Bushman FD, DeSantis T, Andersen GL, Knight R (2010a) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267. doi:10.1093/bioinformatics/btp636
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Dushman FD, Costello EK, Fierer N, González Peña A, Goodrich JK, Gordon JK, Huttley GA, Kelley ST, Knights D, Koening JE, Ley RE, Lozupone C, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010b) QIIME allow analysis of high-thoughput community sequencing data. Nat Methods 7:335–336. doi:10.1038/Nmeth.F.303
Chan OC, Casper P, Sha LQ, Feng ZL, Fu Y, Yang XD, Ulrich A, Zou XM (2008) Vegetation cover of forest shrub and pasture strongly influences soil bacterial community structure as revealed by 16S rRNA gene T-RFLP analysis. FEMS Microbiol Ecol 64:449–458. doi:10.1111/j.1574-6941.2008.00488.x
Davis KE, Sangwan P, Janssen PH (2011) Acidobacteria Rubrobacteridae and Chloroflexi are abundant among very slow-growing and mini-colony-forming soil bacteria. Environ Microbiol 13:798–805. doi:10.1111/j.1462-2920.2010.02384.x
DeBruyn JM, Nixon LT, Fawaz MN, Johnson AM, Radosevich M (2011) Global biogeography and quantitative seasonal dynamics of Gemmatimonadetes in soil. Appl Environ Microb 77:6295–6300. doi:10.1128/aem.05005-11
DeSantis T, Hugenholtz P, Larsen N, Rojas M, Brodie E, Keller K, Huber T, Dalevi D, Hu P, Andersen G (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072. doi:10.1128/Aem.03006-05
Edgar R (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. doi:10.1093/bioinformatics/btq461
Edgar R, Haas B, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. doi:10.1093/bioinformatics/btr381
Ellis PD (2010) The essential guide to effect sizes. Statistical power meta-analysis and the interpretation of research results. Cambridge University Press, New York
Faisal M, Hasnain S (2004) Comparative study of Cr(VI) uptake and reduction in industrial effluent by Ochrobactrum intermedium and Brevibacterium sp. Biotechnol Lett 26:1623–1628. doi:10.1007/s10529-004-3184-1
Faith D (1992) Conservation evualuation and phylogenetic diversity. Biol Conserv 61:1–10. doi:10.1016/0006-3207(92)91201-3
Fay BJ, Corrigan A, Murphy RA (2016) Short-term effects of mechanical drainage on fungal and bacterial community structure in a managed grassland soil. Appl Soil Ecol 101:93–100. doi:10.1016/j.apsoil.2016.01.014
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103:626–631. doi:10.1073/pnas.0507535103
Fierer N, Bradford MA, Jackson RB (2007) Toward and ecological classification of soil bacteria. Ecology 88:1354–1364. doi:10.1890/05-1839
Fierer N, Leff JW, Adams BJ, Nielsen UN, Bates ST, Lauber CL, Owens S, Gilbert JA, Wall DH, Caporaso JG (2012) Cross-biome mtagenomic analyses of soil microbial communities and their functional attributes. PNAS 109:21390–21395. doi:10.1073/pnas.1215210110
Figuerola EL, Guerrero LD, Rosa SM, Simonetti L, Duval ME, Galantini JA, Bedano JC, Wall LG, Erijman L (2012) Bacterial indicator of agricultural management for soil under no-till crop production. PLoS ONE 7:e51075. doi:10.1371/journal.pone.0051075
Fonseca P (2010) Qualidade das águas subterráneas do aquífero livre de Esposende—Vila do Conde (NW de Portugal). Universidade do Minho p, Ordenamento e Valorização de Recursos Geológicos, p 86
Gadd GM (2010) Metals, minerals and microbes: geomicrobiology and bioremediation. Microbiology 156:609–643. doi:10.1099/mic.0.037143-0
Guitián Ojea F, Carballas T (1976) Técnicas de análisis de suelos. Pico Sacro Editorial Santiago de Compostela
Huang J, Sheng XF, Xi J, He LY, Huang Z, Wang Q, Zhang ZD (2014) Depth-related changes in community structure of culturable mineral weathering bacteria and in weathering patterns caused by them along two contrasting soil profiles. Appl Environ Microbiol 80:29–42. doi:10.1128/AEM.02335-13
Janssen PH (2006) Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl Environ Microbiol 72:1719–1728. doi:10.1128/aem.72.3.1719-1728.2006
Knops JMH, Tilman D (2000) Dynamics of soil nitrogen and carbon accumulation for 61 years after agricultural abandonment. Ecology 81:88–98. doi:10.1890/0012-9658(2000)081
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120. doi:10.1128/aem.00335-09
Li H, Ye D, Wang X, Settles ML, Wang J, Hao Z, Zhou L, Dong P, Jiang Y, Ma Z (2014) Soil bacterial communities of different natural forest types in Northeast China. Plant Soil 383:203–216. doi:10.1007/s11104-014-2165-y
Logan NA, De Vos P (2009) Family I. Bacillaceae. In: Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman W (eds) The Firmicutes. Bergey´s manual of systematic bacteriology, vol 3, 2nd edn., Bergey’s manual trustSpringer, New York
Lopes AR, Faria C, Prieto-Fernández Á, Trasar-Cepeda C, Manaia CM, Nunes OC (2011) Comparative study of the microbial diversity of bulk paddy soil of two rice fields subjected to organic and conventional farming. Soil Biol Biochem 43:115–125. doi:10.1016/j.soilbio.2010.09.021
Lopes AR, Manaia CM, Nunes OC (2014) Bacterial community variations in an alfalfa-rice rotation system revealed by 16S rRNA gene 454-pyrosequencing. FEMS Microbiol Ecol 87:650–663. doi:10.1111/1574-6941.12253
Lopes AR, Bello D, Prieto-Fernández Á, Trasar-Cepeda C, Manaia CM, Nunes OC (2015) Relationships among bulk soil physicochemical biochemical and microbiological parameters in an organic alfalfa-rice rotation system. Environ Sci Pollut Res 22:11690–11699. doi:10.1007/s11356-015-4410-1
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235. doi:10.1128/Aem.71.12.8228-8235.2005
Mager DM, Thomas AD (2011) Extracellular polysaccharides from cyanobacterial soil crusts: a review of their role in dryland soil processes. J Arid Environ 75:91–97. doi:10.1016/j.jaridenv.2010.10.001
Melo A, Pinto E, Aguiar A, Mansilha C, Pinho O, Ferreira IM (2012) Impact of intensive horticulture practices on groundwater content of nitrates, sodium, potassium, and pesticides. Environ Monit Assess 184:4539–4551. doi:10.1007/s10661-011-2283-4
Monitoring Agriculture ResourceS (MARS) 2015. Joint research centre Institute for Environment and Sustainability (IES). https://marswiki.jrc.ec.europa.eu/wikicap/index.php/Main_Page. Accessed 26 May, 2016
Montecchia MS, Tosi M, Soria MA, Vogrig JA, Sydorenko O, Correa OS (2015) Pyrosequencing reveals changes in soil bacterial communities after conversion of Yungas forests to agriculture. PLoS ONE 10:e0119426. doi:10.1371/journal.pone.0119426
Oehl F, Oberson A, Probst M, Fliessbach A, Roth HR, Frossard E (2001) Kinetics of microbial phosphorus uptake in cultivated soils. Biol Fert Soils 34:31–41. doi:10.1007/s003740100362
Park CS, Tyson G, Hugenholtz P, Beiko RG (2014) STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124. doi:10.1093/bioinformatics/btu494
Rauret G, Lopez-Sanchez JF, Sahuquillo A, Barahona E, Lachica M, Ure AM, Davidson CM, Gomez A, Luck D, Bacon J, Yli-Halla M, Muntau H, Quevauviller P (2000) Application of a modified BCR sequential extraction (three-step) procedure for the determination of extractable trace metal contents in a sewage sludge amended soil reference material (CRM 483), complemented by a three-year stability study of acetic acid and EDTA extractable metal content. J Environ Monit 2:228–233. doi:10.1039/B001496F
Rodrigues JL, Pellizari VH, Mueller R, Baek K, Jesus EDC, Paula FS, Mirza B, Hamaoui GS, Tsai SM, Feigl B (2013) Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. P Natl Acad Sci USA 110:988–993. doi:10.1073/pnas.1220608110
Ruppel S, Torsvik V, Daae FL, Øvreås L, Rühlmann J (2007) Nitrogen availability decreases prokaryotic diversity in sandy soils. Biol Fertil Soils 43:449–459. doi:10.1007/s00374-006-0122-5
Russo SE, Legge R, Weber KA, Brodie EL, Goldfarb KC, Benson AK, Tan S (2012) Bacterial community structure of contrasting soils underlying Bornean rain forests: inferences from microarray and next-generation sequencing methods. Soil Biol Biochem 55:48–59. doi:10.1016/j.soilbio.2012.05.021
Shannon C, Weaver W (1963) The mathematical theory of communication. University of Illinois Press, Urbana
Silva ME, Lopes AR, Cunha-Queda AC, Nunes OC (2016) Comparison of the bacterial composition of two commercial composts with different physicochemical stability and maturity properties. Waste Manag 50:20–30. doi:10.1016/j.wasman.2016.02.023
Simpson EH (1949) Measurment of diversity. Nature 168:688
Sousa JM, Macedo G, Pedrosa M, Becerra-Castro C, Castro-Silva S, Fernando M, Pereira R, Silva AMT, Nunes OC, Manaia CM (2016) Ozonation and UV254 nm radiation for the removal of microorganisms and antibiotic resistance genes from urban wastewater. J Hazard Mater. doi:10.1016/j.jhazmat.2016.03.096
Sun LN, Zhang YF, He LY, Chen ZJ, Wang QY, Qian M, Sheng XF (2010) Genetic diversity and characterization of heavy metal-resistant-endophytic bacteria from two copper-tolerant plant species on copper mine wasteland. Bioresour Technol 101:501–509. doi:10.1016/j.biortech.2009.08.011
Sun L, Gao J, Huang T, Kendall JR, Shen Q, Zhang R (2015) Parental material and cultivation determine soil bacterial community structure and fertility. FEMS Microbiol Ecol 91:1–10. doi:10.1093/femsec/fiu010
Taniguchi J, Hemmi H, Tanahashi K, Amano N, Nakayama T, Nishino T (2000) Zinc biosorption by a zinc-resistant bacterium, Brevibacterium sp. strain HZM-1. Appl Microbiol Biot 54:581–588. doi:10.1007/s002530000415
Ulrich A, Becker R (2006) Soil parent material is a key determinant of the bacterial community structure in arable soils. FEMS Microbiol Ecol 56:430–443. doi:10.1111/j.1574-6941.2006.00085.x
van Diepeningen AD, de Vos OJ, Korthals GW, van Bruggen AHC (2006) Effects of organic versus conventional management on chemical and biological parameters in agricultural soils. Appl Soil Ecol 31:120–135. doi:10.1016/j.apsoil.2005.03.003
Wang Y, Qian PY (2009) Conservative fragments in bacterial 16S rRNA gene and primer design for 16S ribosomal DNA amplicons in metagenomic studies. PLoS ONE 4:e7401. doi:10.1371/journal.pone.0007401.t001
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. doi:10.1128/Aem.00062-07
Acknowledgements
Authors gratefully acknowledge Mr. Manuel Flores and Ms. Maria Lopes, owners of the studied greenhouses, for valuable help in soil sampling and for giving all the information concerning the farming management.
Funding
This work was financially supported by Project POCI-01-0145-FEDER-006939 (Laboratory for Process Engineering, Environment, Biotechnology and Energy—LEPABE funded by FEDER funds through COMPETE2020—Programa Operacional Competitividade e Internacionalização (POCI)—and by national funds through FCT—Fundação para a Ciência e a Tecnologia. This work was also supported by National Funds from FCT through the project UID/Multi/50016/2013-CBQF and the Grants SFRH/BPD/87152/2012 (CBC) and SFRH/BPD/92894/2013 (ARL).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
This is a posthumous publication of our dear colleague Dr. Cristina Becerra-Castro.
Rights and permissions
About this article
Cite this article
Becerra-Castro, C., Lopes, A.R., Teixeira, S. et al. Characterization of bacterial communities from Masseiras, a unique Portuguese greenhouse agricultural system. Antonie van Leeuwenhoek 110, 665–676 (2017). https://doi.org/10.1007/s10482-017-0833-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-017-0833-7