Abstract
A Gram-positive, yellow pigmented strain, BKS 3-46T was isolated from a soil sample collected from the rhizosphere of Ficus benghalensis (banyan tree) in Bhitarkanika mangrove forest, in the Indian state of Odisha, and subjected to polyphasic taxonomic study. The 16S rRNA gene sequence of the strain was determined and the sequence analysis showed that strain BKS 3-46T should be assigned to the genus Isoptericola. The chemotaxonomic data supported this taxonomic placement i.e. presence of menaquinone MK-9(H4); major fatty acids anteiso C15:0 and iso-C15:0; and phosphatidylglycerol, diphosphatidylglycerol and phosphatidylinositol (PI) as major polar lipids. Further phylogenetic analysis of the 16S rRNA gene sequence confirmed that the strain BKS 3-46T belongs to the genus Isoptericola and is closely related to Isoptericola halotolerans MTCC 11265T (98.6 %) followed by Isoptericola nanjingensis MTCC 11633T (98.4 %) and Isoptericola chiayiensis MTCC 11634T (98.1 %). However, the DNA–DNA hybridization values obtained between strain BKS 3-46T and other related strains were well below the threshold that is required for the proposal of a novel species. The G+C content of the genomic DNA was determined to be 70.4 mol%. The phenotypic and genotypic data showed that the strain BKS 3-46T merits the recognition as a representative of a novel species of the genus Isoptericola. It is proposed that the isolate should be classified in the genus Isoptericola as a novel species, Isoptericola rhizophila sp. nov. The type strain is BKS 3-46T (=MTCC 11080T=JCM 19252T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Isoptericola was first proposed by Stackebrandt et al. (2004), by re-classification of Cellulosimicrobium variabile Bakalidou et al. 2002 as Isoptericola variabilis gen. nov., comb. nov. At present the genus Isoptericola comprises of seven species with validly published names (http://www.bacterio.cict.fr/a/isoptericola.html) and all these species have been isolated from different environmental sources: Isoptericola chiayiensis (soil, Tseng et al. 2011), Isoptericola dokdonensis (soil, Yoon et al. 2006), Isoptericola halotolerans (saline soil sample, Zhang et al. 2005), Isoptericola hypogeus (Roman catacomb of Domitilla, Groth et al. 2005), Isoptericola jiangsuensis (beach soil, Wu et al. 2010), Isoptericola nanjingensis, and Isoptericola variabilis (hind gut of termites, Stackebrandt et al. 2004). In the present study, an actinobacterial strain BKS 3-46T, which was isolated from a soil sample collected from the rhizosphere of Ficus benghalensis (banyan tree), is reported for the first time and subjected to polyphasic taxonomy. 16S rRNA gene sequence comparison revealed that the isolate is Isoptericola-like organism. The aim of the present study was to determine the exact taxonomic position of the isolate.
Materials and methods
Strains, cultivation and phenotypic characterization
Strain BKS 3-46T was isolated from a soil sample collected from the rhizosphere of F. benghalensis (banyan tree), in Bhitarkanika mangrove forest, in the Indian state of Odisha (86′45″–87′17″E longitude and 20′17″–20′47″N latitude), by dilution plate technique on Zobell Marine Agar (ZMA; HiMedia, India) at 30 °C. To study its phenotypic characters, the isolate was routinely cultivated on TSA medium at 30 °C and maintained as glycerol stocks at −70 °C. The reference type strains I. halotolerans MTCC 11265T, I. nanjingensis MTCC 11633T, I. chiayiensis MTCC 11634T, I. variabilis MTCC 11260T, I. hypogeus MTCC 11257T, I. dokdonensis MTCC 11635T and I. jiangsuensis MTCC 11262T were obtained from Microbial Type Culture Collection & Gene Bank (MTCC), Institute of Microbial Technology, Chandigarh, India.
Colony morphology, cell morphology, motility and Gram’s reaction of the strain were determined by using standard methods (Barrow and Feltham 1993; Murray et al. 1994). Phenotypic characterization was performed using TSA as basal medium and strains were incubated at their optimum growth temperature. Physiological tests such as growth at different temperatures, pH (using biological buffers), NaCl concentrations and acid production from various carbohydrates and other biochemical tests were performed as described (Barrow and Feltham 1993; Smibert and Krieg 1994; Stanier et al. 1966; Takeuchi and Hatano 1998). VITEK® 2-GP cards were used as per the instructions of the manufacturer (bioMérieux).
Chemotaxonomic characterization and DNA base composition
Freeze-dried cells for chemotaxonomic analysis (except for fatty acid analysis) were prepared by harvesting the bacterial cells in the late exponential phase following their growth in Tryptic Soy Broth (TSB; HiMedia, India) at 30 °C for 2 days. Isoprenoid quinones were extracted and purified as described by Saha et al. (2005). For the determination of cellular fatty acids, strain BKS 3-46T and the reference strains were grown on TSB medium at 30 °C for 2 days; cellular fatty acids were extracted, methylated and analysed by using Gas Chromatography according to the instructions of the Sherlock Microbial Identification System (MIDI, USA Version 4.0) as described previously (Sasser 1990; Pandey et al. 2002). Extraction of polar lipids was performed based on the modified protocol of Bligh and Dyer (1959). Two-dimensional TLC was run for identification of polar lipids according to procedures described by Komagata and Suzuki (1987). Lipid spots were detected using the following spray reagents: molybdatophosphoric acid (5 % w/v) in absolute ethanol, molybdenum blue spray reagent (1.3 %, Sigma), ninhydrin (0.2 % w/v) in acetone and anisaldehyde reagent (Sigma) for detection of total lipids, phospholipids, aminolipids and glycolipids respectively. Standard procedures were used to determine the diagnostic cell wall sugars and amino acids (Staneck and Roberts 1974). The absence of mycolic acids was confirmed by TLC (Minnikin and Goodfellow 1976). Total hydrolysis (4.0 N HCl, 100 °C, 16 h) of peptidoglycan for amino acids and peptides were separated by two dimensional ascending TLC as method described by Schumann (2011).
Phylogenetic analysis and genomic relatedness
For 16S rRNA gene sequencing, the genomic DNA extraction and amplification was performed as described previously (Mayilraj et al. 2006). Identification of phylogenetic neighbours and the calculation of pairwise 16S rRNA gene sequence similarities were achieved using the EzTaxon server (Kim et al. 2012) and aligned using Mega version 5.0 (Tamura et al. 2011). Phylogenetic trees were constructed using the neighbour-joining as well as maximum likelihood and maximum parsimony algorithms. Bootstrap analysis was performed to assess the confidence limits of the branching. The G+C content of genomic DNA was determined spectrophotometrically (Lambda 35, Perkin Elmer, Waltham, MA, USA) using the thermal denaturation method (Mandel and Marmur 1968). DNA–DNA hybridization was performed each time with freshly isolated genomic DNA and was repeated three times by the membrane filter method (Tourova and Antonov 1987).
Results and discussion
Phenotypic characterization
The detailed differential phenotypic properties are shown in Table 1 and also given in the species description. Phenotypic data presented in Table 1 indicate that strain BKS 3-46T differs from the closely related species at least by 28 characters, which includes acid production from carbohydrates, sensitivity to antibiotics, casein hydrolysis, nitrate reduction, hydrogen sulphide production, assimilation of different carbon sources and other tests. Growth of the strain on TSA produced yellow pigment after 2 days of incubation.
Chemotaxonomic characterization and DNA base composition
Most of the chemotaxonomic properties of strain BKS 3-46T (presented in the species description) are typical of members of the genus Isoptericola. The major fatty acids were identified as anteiso-branched C15:0 (56.2 %), iso-branched C15:0 (16.8 %) and C16:0 (14.8 %). The fatty acid compositions of the reference strains analyzed were qualitatively similar to but quantitatively different from those of the novel strain (Table 2). The major polar lipids were identified as phosphatidylglycerol, diphosphatidylglycerol, phosphatidylinositol, three unknown phospholipids, two unknown glycolipids and one phosphoglycolipid (Supplementary Fig. S1). The major menaquinone detected for strain BKS 3-46T was MK-9 (H4). The peptidoglycan type was A4α l-Lys-d-Asp (Supplementary Fig. S2), type A11.31 according to www.peptidoglycan-types.info, which was compared and confirmed with the data presented for I. dokdonensis strain DS-3T (Schumann 2011). The valid proof for the proposed peptidoglycan structure with d-aspartic acid directly linked to l-lysine is the dipeptide d-Asp-l-Lys which is presented at the position labeled as “1”. This peptide d-Asp-l-Lys is resistant towards hydrolysis and is therefore one of the rare exceptions that a peptide can be found in the total hydrolysate, which usually contains only amino acids and no peptides. The DNA G+C content of strain BKS 3-46T was estimated to be 70.4 mol%, a value within the range (70–74.1 mol%) for members of the genus Isoptericola.
Phylogenetic analysis and genomic relatedness
Almost complete sequence (1410 bp) of 16S rRNA gene of strain BKS 3-46T was determined (GenBank accession no. KC 608148) and compared with those of other closely related taxa retrieved from the GenBank database. Based on the 16S rRNA gene sequence identity, the strain could be assigned to the genus Isoptericola. Sequence analysis revealed that strain BKS 3-46T shared 16S rRNA gene sequence identity with I. halotolerans (98.6 %), followed by I. nanjingensis (98.4 %), I. chiayiensis (98.1 %), I. variabilis (98.0 %), I. hypogeus (97.7 %), I. dokdonensis (97.5 %) and I. jiangsuensis (97.3 %). The neighbour-joining phylogenetic tree, demonstrated that strain BKS 3-46T forms a separate lineage along with the closely related species, this was also evident in the phylogenetic tree constructed using the maximum-parsimony and maximum likelihood algorithms, shown as closed circles at the nodes (Fig. 1). The DNA–DNA relatedness values between strain BKS 3-46T and the closely related taxa were found to be I. halotolerans MTCC 11265T (54.6 ± 0.8 %), followed by I. nanjingensis MTCC 11633T (50.4 ± 0.6 %), I. chiayiensis MTCC 11634T (46.0 ± 0.2 %), I. variabilis MTCC 11260T (48.1 ± 0.6 %), I. hypogeus MTCC 11257T (46.8 ± 0.2 %), I. dokdonensis MTCC 11635T (39.4 ± 0.8 %) and I. jiangsuensis MTCC 11262T (42.7 ± 0.8 %). These values are below the standard threshold value for bacterial species delineation (Wayne et al. 1987).
Phylogenetic neighbour-joining tree based on 16S rRNA gene sequences showing the relationship between Isoptericola rhizophila BKS 3-46T and related taxa. Promonomicrospora citrea DSM 43110T (X83808) was used as an out-group. Bootstrap values (expressed as percentages of 100 replications) greater than 70 % are given at nodes. Branches recovered in the maximum-parsimony and maximum likelihood algorithms are indicated by filled circles. Bar 0.005 % sequence variation. GenBank accession numbers are given in parentheses
Conclusion
Based on the differential phenotypic characteristics such as oxidase, hydrolysis of hypoxanthine, utilization of d-xylose, acid production from lactose to trehalose, sensitivity to antibiotics such as ampicillin, penicillin G, novobiocin and rifampicin and genotypic results, strain BKS 3-46T should be regarded as a new species of the genus Isoptericola. Table 1 shows the main features that distinguish strain BKS 3-46T from all the validly named members of the genus. For the first time a species belonging to the genus Isoptericola isolated from rhizosphere of F. benghalensis (banyan tree) is reported. The polyphasic evidence gathered in this study allow the conclusion that strain BKS 3-46T represents a novel species of the genus Isoptericola, for which the name Isoptericola rhizophila sp. nov. is proposed.
Description of Isoptericola rhizophila sp. nov.
Isoptericola rhizophila (rhi.zo’phi.la.Gr. n. rhiza a root; Gr. adj. philos on loving; N.L. fem. adj. rhizophila root-loving).
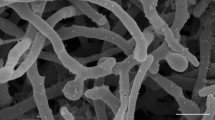
Gram-positive, non-acid-fast, aerobic bacterium. Forms lemon-orange coloured, circular, glistening, opaque colonies with an entire margin on TSA medium. Colony sizes are 0.5–2.0 mm. Cells are irregularly spherical or rods, occurring in pairs and clusters. Cells vary in size (0.6–1.0 μm wide by 1.0–1.5 μm long). Does not form endospores. Non-motile. No mycelial growth phase is observed. Catalase-positive and oxidase-negative. Tolerates up to 7.0 % NaCl and grows at temperatures between 25 and 37 °C, with optimum growth 30 °C. Growth occurs at between pH 6 and 10, with optimum growth at pH 8.0. Positive for decomposition of casein, starch and gelatin. Negative for indole, urease and hydrogen sulfide production, methyl red and Voges–Proskauer reactions, utilization of citrate, decomposition of aesculin and tyrosine, and for nitrate reduction. Acid is produced from l-rhamnose, d-fructose and trehalose. Acid is produced weakly from l-arabinose but not from d-cellobiose, myo-inositol, d-mannitol, d-maltose, d-galactose, glycerol, salicin, sucrose or d-xylose. Utilization of various compounds as sole carbon sources is detailed in Table 1. The dipeptide in cell wall hydrolyzates was A4α l-Lys-d-Asp. The whole-cell sugars are galactose, xylose and mannose. The predominant lipids are diphosphatidylglycerol, phosphatidylglycerol, one unknown glycolipid and other unidentified lipids. No mycolic acids are present. Contains major amounts of anteiso-branched C15:0, iso-branched C15:0 and C16:0 fatty acids. The major menaquinone is MK-9 (H4). The DNA G+C content of the type strain is 70.4 mol%.
The type strain BKS 3-46T (=MTCC 11080T=JCM 19252T) was isolated from a soil sample collected from rhizosphere of F. benghalensis (banyan tree), Bhitarkanika mangrove forest, in the Indian state of Odisha. The GenBank accession number for the 16S rDNA sequence of strain BKS 3-46T is KC 608148.
References
Bakalidou A, Kämpfer P, Berchtold M, Kuhnigk T, Wenzel M, König H (2002) Cellulosimicrobium variabile sp. nov., a cellulolytic bacterium from the hindgut of the termite Mastotermes darwiniensis. Int J Syst Evol Microbiol 52(4):1185–1192
Barrow GI, Feltham RKA (1993) Cowan and Steel’s manual for the identification of medical bacteria, 3rd edn. Cambridge University Press, Cambridge
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37(8):911–917
Groth I, Schumann P, Schütze B, Gonzalez JM, Laiz L, Saiz-Jimenez C, Stackebrandt E (2005) Isoptericola hypogeus sp. nov., isolated from the Roman catacomb of Domitilla. Int J Syst Evol Microbiol 55(4):1715–1719
Huang Z, Sheng XF, Zhao F, He LY, Huang J, Wang Q (2012) Isoptericola nanjingensis sp. nov., a mineral-weathering bacterium. Int J Syst Evol Microbiol 62:971–976
Kim O, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA Gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Komagata K, Suzuki K (1987) Lipid and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–207
Mandel M, Marmur J (1968) Use of ultraviolet absorbance temperature profile for determining the guanine plus cytosine content of DNA. Methods Enzymol 12B:195–206
Mayilraj S, Saha P, Suresh K, Saini HS (2006) Ornithinimicrobium kibberense sp. nov., isolated from the Himalayas,India. Int J Syst Evol Microbiol 56:1657–1661
Minnikin DE, Goodfellow M (1976) Lipid composition in the classification and identification of Nocardia and related taxa. In: Goodfellow M, Brownell GH, Serrano JA (eds) In the biology of the Nocardiaceae. Academic Press, London, pp 160–219
Murray RGE, Doetsch RN, Robinow CF (1994) Determinative and cytological light microscopy. In: Gerhard P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 21–41
Pandey KK, Mayilraj S, Chakraborti T (2002) Pseudomonas indica sp. nov., a novel butane utilizing species. Int J Syst Evol Microbiol 52:1559–1567
Saha P, Mondal AK, Mayilraj S, Krishnamurthi S, Bhattacharya A, Chakrabarti T (2005) Paenibacillus assamensis sp. nov., a novel bacterium isolated from a warm spring in Assam, India. Int J Syst Evol Microbiol 55:2577–2581
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids, MIDI technical note 101. MIDI Inc, Newark
Schumann P (2011) Peptidoglycan structure. In: Rainey FA, Oren A (eds) Taxonomy of prokaryotes. Academic Press, Chennai, pp 101–129
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhard P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Stackebrandt E, Schumann P, X-L Cui (2004) Reclassification of Cellulosimicrobium variabile Bakalidou, 2002 as Isoptericola variabilis gen. nov., comb. nov. Int J Syst Evol Microbiol 54:685–688
Staneck JL, Roberts GD (1974) Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl Microbiol 28:226–231
Stanier RY, Palleroni NJ, Doudoroff M (1966) The aerobic pseudomonads: a taxonomic study. J Gen Microbiol 43:159–271
Takeuchi M, Hatano K (1998) Union of the genera Microbacterium Orla-Jensen and Aureobacterium Collins et al. in a redifined genus Microbacterium. Int J Syst Bacteriol 48:739–747
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: evolutionary genetics analysis using maximum likelihood, evolutionary distance and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tourova TP, Antonov AS (1987) Identification of microorganisms by rapid DNA–DNA hybridization. Methods Microbiol 19:333–355
Tseng M, Liao HC, Chiang WP, Yuan GF (2011) Isoptericola chiayiensis sp. nov., isolated from mangrove soil. Int J Syst Evol Microbiol 61:1667–1670
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Truper HG (1987) Report of the adhoc committee on reconciliation of approaches to bacterial systematic. Int J Syst Bacteriol 37:463–464
Wu Y, Li WJ, Tian W, Zhang LP, Xu L, Shen QR, Shen B (2010) Isoptericola jiangsuensis sp. nov., a chitin-degrading bacterium. Int J Syst Evol Microbiol 60:904–908
Yoon JH, Schumann P, Kang SJ, Jung SY, Oh TK (2006) Isoptericola dokdonensis sp. nov., isolated from soil. Int J Syst Evol Microbiol 56:2893–2897
Zhang YQ, Schumann P, Li WJ, Chen GZ, Tian XP, Stackebrandt E, Xu LH, Jiang CL (2005) Isoptericola halotolerans sp. nov., a novel actinobacterium isolated from saline soil from Qinghai Province, north-west China. Int J Syst Evol Microbiol 55:1867–1870
Acknowledgments
We thank Mr. Malkit Singh for his excellent technical assistance. We also thank Mr. Nitin Kumar Singh, Ms. Ishwinder Kaur and Ms. Monu Bala for their assistance in collection and preliminary work. This work was supported by Council of Scientific and Industrial Research and Department of Biotechnology, Government of India. This is IMTECH communication number 58/2013.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kaur, N., Rajendran, M.K., Kaur, G. et al. Isoptericola rhizophila sp. nov., a novel actinobacterium isolated from rhizosphere soil. Antonie van Leeuwenhoek 106, 301–307 (2014). https://doi.org/10.1007/s10482-014-0197-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-014-0197-1