Abstract
The genus Borrelia contains two groups of organisms: the causative agents of Lyme disease and their relatives and the causative agents of relapsing fever and their relatives. These two groups are morphologically indistinguishable and are difficult to distinguish biochemically. In this work, we have carried out detailed comparative genomic analyses on protein sequences from 38 Borrelia genomes to identify molecular markers in the forms of conserved signature inserts/deletions (CSIs) that are specifically found in the Borrelia homologues, and conserved signature proteins (CSPs) which are uniquely present in Borrelia species. Our analyses have identified 31 CSIs and 82 CSPs that are uniquely shared by all sequenced Borrelia species, providing molecular markers for this group of organisms. In addition, our work has identified 7 CSIs and 21 CSPs which are uniquely found in the Lyme disease Borrelia species and eight CSIs and four CSPs that are specific for members of the relapsing fever Borrelia group. Additionally, 38 other CSIs, in proteins which are uniquely found in Borrelia species, also distinguish these two groups of Borrelia. The identified CSIs and CSPs provide novel and highly specific molecular markers for identification and distinguishing between the Lyme disease Borrelia and the relapsing fever Borrelia species. We also report the results of average nucleotide identity (ANI) analysis on Borrelia genomes and phylogenetic analysis for these species based upon 16S rRNA sequences and concatenated sequences for 25 conserved proteins. These analyses also support the distinctness of the two Borrelia clades. On the basis of the identified molecular markers, the results from ANI and phylogenetic studies, and the distinct pathogenicity profiles and arthropod vectors used by different Borrelia spp. for their transmission, we are proposing a division of the genus Borrelia into two separate genera: an emended genus Borrelia, containing the causative agents of relapsing fever and a novel genus, Borreliella gen. nov., containing the causative agents of Lyme disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Borrelia is an important pathogenic group of helical shaped, motile organisms that form a highly distinct, monophyletic lineage within the phylum Spirochaetes (Paster 2011; Wang and Schwartz 2011). Members of this genus are the causative agents of both Lyme disease, which is currently the most prevalent vector-borne disease in North America and temperate regions of Eurasia, and relapsing fever, which is a disease endemic to many disparate regions of the world (Lindgren and Jaenson 2006; Cutler 2010; Adams et al. 2013). Currently, the genus Borrelia contains 37 species which are carried by arthropod vectors and exhibit varying pathogenicity in mammalian and avian hosts (Margos et al. 2011; Wang and Schwartz 2011; Parte 2014). These species can be separated into two main groups based upon their pathogenicity profiles. The first group, containing the causative agents of Lyme disease, is commonly referred to as the Borrelia burgdorferi sensu lato complex, whereas the other group contains the causative agents of relapsing fever (Postic et al. 1990; Baranton et al. 1992; Wang et al. 1999; Margos et al. 2011; Wang and Schwartz 2011). Although, these two groups are morphologically indistinguishable from each other, their members can be distinguished from each other based on the arthropod vectors which transmit them and by a limited number of biochemical and genetic tests (Wang et al. 1999; Margos et al. 2011; Wang and Schwartz 2011). Our current understanding of the taxonomy and evolutionary relationships among the Borrelia species is based largely on DNA–DNA hybridization studies, 16S rRNA gene sequence analysis and multilocus sequence analysis (MLSA) (Margos et al. 2011; Wang and Schwartz 2011). Although these studies provide evidence suggesting separation of the members of the genus Borrelia into two distinguishable groups, due to lack of other reliable molecular, morphological, or biochemical characteristics that can distinguish these groups, no formal recognition of these two distinct groups of Borrelia has thus far been made (Wang and Schwartz 2011).
Whole genome sequences for members of the genus Borrelia are becoming increasingly available in public databases. There are currently 38 genomes from 18 species of Borrelia available in the NCBI database (NCBI 2014). These genomes provide a valuable resource to gain insight into the evolutionary history of this group of organisms and to identify novel shared molecular characteristics that are specific for this group of organisms. One useful comparative genomic approach, pioneered by our lab, involves the identification of conserved signature indels (CSIs), which are insertions/deletions uniquely present in protein sequences of organisms from the group of interest, and conserved signature proteins (CSPs), which are lineage specific proteins found only in the group of interest (Gupta and Griffiths 2006; Gupta 2010; Naushad et al. 2014). Due to the specificity of these markers (viz. CSIs and CSPs) for particular groups of bacteria, they represent molecular synapomorphies (markers of common evolutionary decent) which can be used to identify and demarcate specific bacterial groups in clear molecular terms. Additionally, whole genome sequences are also enabling the use of other computational algorithms to determine the overall genome similarity among different organisms (Richter and Rosselló-Móra 2009).
Our recent comparative analysis of Spirochaetes genomes has identified 38 CSIs that clearly delimit the major groups within the phylum and were used to revise the taxonomy of the phylum as a whole (Gupta et al. 2013b). In this work, we extend these studies by examining, in detail, the evolutionary relationships among the Borrelia species employing different phylogenetic and comparative genomic approaches. These analyses have identified 31 CSIs and 82 CSPs that are commonly shared by all sequenced Borrelia species. More importantly, these studies have identified of 53 CSIs and 25 CSPs, which serve to clearly distinguish the two main groups of Borrelia species and provide novel molecular markers to demarcate them in definitive terms. The distinctness of these two groups of Borrelia species is also supported by the results of an average nucleotide identity (ANI) analysis of Borrelia genomes and by phylogenetic trees constructed based upon 16S rRNA sequences and concatenated protein sequences. On the basis of the identified molecular markers, phylogenetic studies, and other evidence presented here, it is proposed that the genus Borrelia should be divided into two separate genera: an emended genus Borrelia, containing the causative agents of relapsing fever and a novel genus, Borreliella gen. nov., containing the causative agents of Lyme disease.
Methods
Phylogenetic sequence analysis
Phylogenetic analysis was performed on a concatenated sequence alignment of 25 highly conserved proteins (viz. ArgRS, DnaK, EF-G, EF-Tu, GyrA, GyrB, Hsp60, Hsp70, IleRS, RecA, RpoB, RpoC, SecY, ThrRS, TrpRS, ValRS, and ribosomal proteins L1, L2, L5, L6, S3, S8, S9, S11, and S12) which represent a subset of the core proteins present in all bacteria that are widely used for phylogenetic analysis (Harris et al. 2003; Charlebois and Doolittle 2004; Ciccarelli et al. 2006; Vinuesa 2010; Gao and Gupta 2012b; Gupta et al. 2013b). Sequences for these proteins were obtained from the NCBI database for 38 sequenced Borrelia species (Table 1) and Treponema pallidum Nichols which was used to root the tree. Multiple sequence alignments for these proteins were created using Clustal_X 1.83 (Jeanmougin et al. 1998) and concatenated into a single alignment file. Poorly aligned regions from this alignment file were removed using Gblocks 0.91b (Castresana 2000). The resulting alignment, which contained 12,129 aligned amino acids, was used for phylogenetic analysis. The maximum likelihood tree based on 1,000 bootstrap replicates of this alignment was constructed using MEGA 6.0 (Tamura et al. 2013) employing the Le and Gascuel (Le and Gascuel 2008) substitution model.
A 16S rRNA gene sequence based phylogenetic tree was also created based on 53 sequences that included representative strains of all cultured Borrelia species (Supplemental Table 1). 16S rRNA gene sequences larger than 1,200 bp were obtained for all type strains classified under the genus Borrelia in release 115 of the SILVA database (Quast et al. 2013). 16S rRNA gene sequences were also obtained for representative strains from Borrelia species without a cultured type and for T. pallidum Nichols which was used to root the tree. A maximum likelihood tree based on these sequences was created using 1,000 bootstrap replicates of the 16S rRNA sequence alignments in MEGA 6.0 (Tamura et al. 2013) employing the General Time-Reversible (Tavaré 1986) substitution model.
Average nucleotide analysis
Average nucleotide identity values were calculated in order to assess the relatedness of the sequenced Borrelia genomes using the JSpecies v1.2.1 program (Richter and Rosselló-Móra 2009) which utilized an algorithm developed by Goris et al. (2007) to analyze the sequence identity of pairwise genome alignments created using the BLAST v2.2.26 program (Altschul et al. 1997).
Identification of conserved signature indels
To identify CSIs that are commonly shared by the different groups of Borrelia, BLAST searches (Altschul et al. 1997) were performed using each protein in the genome of Borrelia recurrentis A1 as queries. These searches were performed using the default BLAST parameters against all available sequences in the GenBank non-redundant database. For those proteins for whom high scoring homologues (E values < 1e−20) were present in other Borrelia species, multiple sequence alignments were created using the Clustal_X 1.83 program (Jeanmougin et al. 1998). These alignments were visually inspected for the presence of insertions or deletions that were flanked on both sides by at least 5-6 conserved amino acid residues in the neighbouring 30–40 amino acids. Indels that were not flanked by conserved regions were not further considered, as they do not provide useful molecular markers (Gupta 2010; Naushad et al. 2014). The specificity of potentially useful indels for subgroups within of the genus Borrelia was further evaluated by carrying out detailed BLAST searches on short sequence segments containing the indel and the flanking conserved regions (60-100 amino acids long). To ensure that the identified signatures are only present in Borrelia homologues, 250 BLAST hits with the highest similarity to the query sequence were examined for the presence or absence of these CSIs. In this work, we report the results of CSIs that are specific for different groups within the Borrelia and where similar CSIs were not observed in any other bacteria in the top 250 BLAST hits. The sequence alignment files presented here contain sequence information for all sequenced species within the genus Borrelia. However, due to space constraints, different strains of the sequenced species are not shown, but they all displayed similar sequence characteristics.
Identification of conserved signature proteins
To identify proteins that are uniquely present in various groups of Borrelia, BLAST searches (Altschul et al. 1997) were performed using each protein in the genomes of B. burgdorferi B31 and B. recurrentis A1 as queries. These searches were performed using the default BLAST parameters against all available sequences in the GenBank non-redundant database. Proteins were considered CSPs if either all significant hits were from well-defined groups of Borrelia or which involved a large increase in E values from the last hit belonging to a particular group of Borrelia to the first hit from any other bacteria and the E values for the latter hits were >1e−04, indicating weak similarity that could occur by chance (Gao and Gupta 2007; Naushad et al. 2014). In most cases, the lengths of various significant hits were very similar to those of the query proteins.
Results
Genomic characteristics of the sequenced Borrelia
Genome sequences for 38 Borrelia strains comprising 18 different species, which are currently available in the NCBI genome database, were used in these analyses. Some characteristics of these Borrelia genomes are summarized in Table 1. The genomes of most Borrelia species/strains, in addition to containing a linear chromosome, harboured large numbers of linear and circular plasmids, which is very unique among the prokaryotes (Chaconas 2005; Chaconas and Kobryn 2010). The chromosome sizes of the sequenced Borrelia fell within a narrow range between 0.89 and 1.01 Mb, with G+C content ranging between 25.83 and 29.81 %.
Phylogenetic sequence analysis
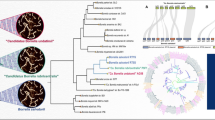
The current understanding of the phylogeny of the genus Borrelia is largely based on phylogenetic trees constructed using 16S rRNA, flagellin or housekeeping gene sequences (Fukunaga et al. 1996; Margos et al. 2009; Wang and Schwartz 2011). In this work, we have constructed a phylogenetic tree of the sequenced Borrelia species using concatenated sequences for 25 conserved housekeeping and ribosomal proteins (Fig. 1). Members of the genus Borrelia have shown some competence for the lateral transfer of tRNA synthetases (Ibba et al. 1997). However, phylogenetic trees based on concatenated sequences for a large number of unlinked and conserved loci minimize the effect of any instances of lateral gene transfer and provide greater resolving power than trees based on any single gene or protein (Rokas et al. 2003; Wu et al. 2009). In the concatenated protein tree, the sequenced Borrelia species clustered into two distinct monophyletic and strongly supported clades, which were separated by long branches. One of these clades consisted of the Lyme disease causing B. burgdorferi species (B. burgdorferi sensu stricto) and its relatives (B. burgdorferi sensu lato), while the other clade was comprised of the relapsing fever Borrelia (B. recurrentis) and its relatives (Fig. 1). These two clades of Borrelia are also clearly distinguished in a phylogenetic tree for 3,737 genome sequenced prokaryotes, which was constructed based upon >400 proteins (Segata et al. 2013).
A maximum likelihood phylogenetic tree of 38 sequenced members of the genus Borrelia based on the concatenated amino acid sequences of 25 conserved proteins. Bootstrap values are shown at branch nodes. The Lyme disease and relapsing fever clades of Borrelia are marked. The letter T refers to the type strain of the species
A phylogenetic tree was also constructed based on the 16S rRNA gene sequences, which included representatives from all cultured Borrelia species (Fig. 2). Except for Borrelia turcica, all Borrelia species were grouped into two distinct clades similar to those seen in the concatenated protein tree. However, an earlier study showed that B. turcica clusters with several unnamed Borrelia isolates in a monophyletic clade related to the relapsing fever Borrelia (Takano et al. 2010). The members of the genus Borrelia have also been observed to branch into two distinct clades in a number of earlier phylogenetic studies based on 16S rRNA and other individual genes/protein sequences (Takano et al. 2010; Margos et al. 2011; Wang and Schwartz 2011).
A maximum likelihood tree based on the 16S rRNA gene sequences of representative strains of Borrelia. Bootstrap values are shown at branch nodes. The Lyme disease and relapsing fever clades of Borrelia are marked. The letter T refers to the type strain of the species. The accession numbers of the 16S rRNA gene sequences used in this analysis are provided in Supplemental Table 1
Conserved signature indels that distinguish the two clades of Borrelia
CSIs and CSPs that are restricted to a given group of related species provide useful molecular characteristics for evolutionary studies (Gupta 1998; Rokas and Holland 2000; Gao and Gupta 2012a). Recently, CSIs have been used to define novel taxonomic groups and to propose important taxonomic changes for groups of bacteria (viz. Aquificae, Bacillus, Chloroflexi, Neisseriales, Spirochaetes, Synergistetes and Thermotoga) at different taxonomic ranks (Bhandari and Gupta 2012; Adeolu and Gupta 2013; Bhandari et al. 2013; Gupta et al. 2013a, b; Gupta and Lali 2013; Bhandari and Gupta 2014). In this work we have carried out comprehensive comparative analyses of Borrelia genomes in order to identify CSIs that clarify the relationship between the Borrelia. These studies have identified 31 CSIs that are specifically found in protein homologues from members of the genus Borrelia as currently defined and absent in homologues from all other sequenced bacterial groups. Fifteen of these 31 CSIs are identified for the first time in this work, whereas the remaining 16 CSIs were identified in our earlier analysis of the phylum Spirochaetes (Gupta et al. 2013b). One example of a novel CSI that is uniquely found in all of the sequenced species from the genus Borrelia is shown in Fig. 3. In the example shown, a 3 aa insert in a conserved region of the bacterial rod-shaped determining protein MreB is uniquely present in all sequenced Borrelia species, but it is not found in sequences from any other Spirochaetes or other phyla of bacteria (Fig. 3). Sequence information for the 14 other novel CSIs that are also specific for the genus Borrelia is presented in Supp. Fig. 1–14 and a summary of all 31 Borrelia specific CSIs is presented in Table 2.
A partial sequence alignment of the rod shape-determining protein MreB, showing a CSI (boxed) that is uniquely present in all members of the genus Borrelia. Sequence information for a single Borrelia strain from each of the 18 sequenced Borrelia species and a limited number other bacteria is shown here, but unless otherwise indicated similar CSIs were detected in all members of the indicated group and not detected in any other bacterial species in the top 250 BLAST hits. The dashes in the alignments indicate identity with the residue in the top sequence. GenBank identification (GI) numbers for each sequence are indicated in the second column. Sequence information for 30 other CSIs that are specific for all sequenced Borrelia species is provided in Supplemental figures 1–14 and Table 2
Our analyses have also identified 53 CSIs that are specific for or distinguish between the two main clades of Borrelia species, which are observed in the phylogenetic trees. Of these, seven CSIs are specific for the Lyme disease Borrelia clade, whereas another eight novel CSIs are uniquely found in the Borrelia species that are part of the relapsing fever clade. Examples of a CSI specific for the Lyme disease Borrelia clade and a CSI specific for the relapsing fever Borrelia clade are shown in Fig. 4. Figure 4a shows a 1 aa insert in a conserved region of Recombinase A that is uniquely found in all eight sequenced species from the Lyme disease Borrelia clade, whereas Fig. 4b shows a 1 aa deletion in the nicotinamide-nucleotide adenylyltransferase protein that is specific for members of the relapsing fever Borrelia clade. Sequence information for other CSIs that are specific for these two clades of Borrelia species are presented in Supp. Fig. 15–27 and Table 3. In addition to these 15 CSIs found in widely distributed proteins, 38 other CSIs in proteins that are mainly found in Borrelia species also serve to distinguish the Lyme disease Borrelia clade from the relapsing fever Borrelia clade. Because homologues for these proteins, or the conserved regions where these CSIs are present in these proteins, are not found in other bacteria, it is difficult to infer whether these CSIs represent insertions or deletions in the two groups. However, these CSIs still serve to distinguish between the two groups of Borrelia. One example of a 3 aa indel in a Borrelia specific protein of unknown function that distinguishes the Lyme disease Borrelia clade from the relapsing fever Borrelia clade is shown in Fig. 5. Sequence information for 37 other CSIs in different proteins that are of a similar kind is presented in Supp. Fig. 28–64 and Table 4.
Partial sequence alignments of (a) the protein Recombinase A showing a one amino acid insertion (boxed) identified in members of the Lyme disease Borrelia (b) the protein Nicotinamide-nucleotide adenylyltransferase showing a one amino acid deletion identified in members of the relapsing fever Borrelia. These CSIs were not found in the sequence homologues from any other sequenced bacteria. Sequence information for other Lyme disease or relapsing fever Borrelia specific CSIs is presented in Supplemental figures 15–27 and summarized in Table 3
A partial sequence alignment of a Borrelia lineage specific protein with currently unknown function (Hypothetical protein BB0838) showing a three amino acid insertion (boxed) which distinguishes the Lyme disease and relapsing fever Borrelia. Sequence information for other CSIs present in Borrelia lineage specific proteins is presented in Supplemental figures 28–64 and summarized in Table 4
Conserved signature proteins which are specific for Borrelia or distinguish its two clades
Another useful category of molecular markers whose discovery has been enabled by comparative genomic analysis are conserved signature proteins (CSPs) that are uniquely present in different lineages of prokaryotes. Due to the specific presence of these genes/proteins in particular lineages of bacteria, they again provide useful molecular markers of common evolutionary decent for identifying and demarcating different bacterial groups in clear molecular terms. Our analyses of Borrelia genomes in this regard have led to identification of 107 proteins which are uniquely found either in all (or most) sequenced Borrelia species or are specific for only the Lyme disease Borrelia clade or the relapsing fever Borrelia clade. The results of BLAST searches for three CSPs that are specific to either all sequenced Borrelia, members of the Lyme disease Borrelia, or members of the relapsing fever Borrelia are shown in Table 5. As seen from this Table, high scoring homologues for these proteins are only found in different Borrelia species belonging to their specified clades, but not in any other bacterial organism. Thus, similar to the CSIs, these CSPs again are distinctive characteristics of the species from these clades and provide valuable molecular markers for their identification and demarcation. Of the CSPs that we have identified, 82 proteins are uniquely present in all or most of the sequenced Borrelia species and they are likely distinguishing characteristics of all members of the recently described family Borreliaceae (Table 6; Gupta et al. 2013b). In some cases, the homologues of these proteins were not detected in a few isolated strains of Borrelia. However, in every case, the proteins were not present in any other bacterial group, suggesting that the strains lacking these homologues have either undergone gene loss or that they are earlier branching lineages within these clades. In addition to the CSPs that are specific for all Borrelia (or the family Borreliaceae), we have also identified 21 CSPs whose homologues are only found in the Lyme disease Borrelia (Table 7) and four other CSPs, which are restricted to members of the relapsing fever Borrelia (Table 7). Some characteristics of the different CSPs are summarized in Tables 6 and 7. The cellular functions of most of these CSPs are unknown, but they may be related to some of the distinguishing properties exhibited by their specified clades.
Average nucleotide analysis
DNA–DNA hybridization is a commonly used method to determine the relatedness of different organisms and for assignment of species to either the same or different genera (Thompson et al. 2013). However, concerns have been raised about the scalability and reproducibility of these studies (Rosselló-Mora 2006). The availability of genome sequences have now made it possible to calculate pairwise ANI values between different genomes, which are analogous to DNA homology values (Richter and Rosselló-Móra 2009). We have compared the ANI values for all available genome sequenced Borrelia species (Fig. 6). The ANI values for different members within the genus Borrelia range between 73.03 and 99.34 % identity. However, based upon the comparisons of the ANI values, the Borrelia species can again be divided into two distinct clusters. One cluster, consisting of the members of the Lyme disease Borrelia, had intercluster ANI values which ranged between 91.33 and 98.06 % identity. The other cluster, which consisted of the members of the relapsing fever Borrelia, had intercluster ANI values which ranged between 82.51 and 99.34 % identity (Fig. 6). In contrast to the high ANI values for species within the two clusters, the ANI values of Borrelia species between the members of the two clusters were significantly lower, ranging between 73.03 and 74.85 % identity, indicating that the members of these clusters are distinct from each other.
Discussion
Genetic differences between the Lyme disease and relapsing fever Borrelia have been observed in a number of earlier studies (Postic et al. 1990; Fukunaga et al. 1996; Ras et al. 1996; Valsangiacomo et al. 1997; Margos et al. 2009). However, due to lack of distinct characteristics that can clearly distinguish the Lyme disease Borrelia from the relapsing fever Borrelia, it has proven difficult to reliably distinguish species from these two groups. This is responsible for the failure to diagnose or misdiagnosis of Lyme disease Borrelia in many individuals and also an underreporting of the overall incidence of this disease in the population (Wright et al. 2012; Ljostad and Mygland 2013). Detailed comparative analyses on genome sequences from Borrelia species that is reported here have identified numerous discrete molecular characteristics that are specifically shared by either members of the Lyme disease Borrelia clade or the relapsing fever Borrelia clade. The molecular markers described in this work provide novel and highly specific means for identification of members of the Lyme disease Borrelia group by either molecular sequence based (e.g. PCR, pyrosequencing, etc.) methods (Ahmod et al. 2011; Dunaj et al. 2013) or immunological methods (Wright et al. 2012; Ljostad and Mygland 2013).
The results reported here from multiple lines of investigations provide compelling evidence that the known Borrelia species are comprised of at least two evolutionary distinct groups of organisms corresponding to the Lyme disease Borrelia clade and the relapsing fever Borrelia clade. The different lines of investigation that support the distinctness of these two clades can be briefly summarized as follows:
-
1.
In phylogenetic trees based on the 16S rRNA gene or concatenated sequences for 25 conserved proteins, the species from these two groups formed distinct and strongly supported clades that are separated from each other by long branches.
-
2.
This work has identified 7 CSIs and 21 CSPs that are uniquely present in all of the genome sequenced species from the Lyme disease Borrelia clade and eight CSIs and four CSPs that are specific for the relapsing fever Borrelia clade. The unique and mutually exclusive presence of these molecular characteristics in these two groups of species provides compelling evidence that they are derived from distinct ancestors. The identified molecular markers also provide reliable means for the demarcation of these two clades in molecular terms.
-
3.
Whole genome ANI analyses of Borrelia genomes show that species from within either the Lyme disease Borrelia group or the relapsing fever Borrelia group had much higher ANI values when compared to other members of their group (range 82.51–99.34 %) than with members of the opposing Borrelia group (range 73.03–74.85 %).
-
4.
The species from these two groups differ in terms of their pathogenicity profiles and the characteristics of the arthropod vectors which are involved in their transmission. The species which are part of the Lyme disease clade are transmitted via arthropod vectors that are hard tick species related to the Ixodes ricinus complex, while a majority of the members of the relapsing fever Borrelia clade are transferred by soft-bodied ticks within the family Argasidae (Table 8).
Table 8 Distinguishing characteristics of the Lyme disease Borrelia (Borreliella) and the relapsing fever Borrelia
Taxonomic implications
The evidence obtained from different lines of investigations summarized above provides compelling evidence that the known Borrelia species are comprised of two main clades corresponding to the “Lyme disease Borrelia and its relatives” and the “relapsing fever Borrelia and its relatives”. Of these two main groups, the Lyme disease Borrelia clade, based upon branching in the 16S rRNA gene tree and concatenated protein tree is comprised of the following 14 validly named species: B. afzelii, B. americana, B. bavariensis, B. burgdorferi, B. carolinensis, B. garinii, B. japonica, B. kurtenbachii, B. lusitaniae, B. sinica, B. spielmanii, B. tanukii, B. turdi, and B. valaisiana. All other currently validly named Borrelia species are part of the relapsing fever Borrelia clade. The observations presented in this work make a strong case for division of the existing genus Borrelia into two different genera corresponding to the Lyme disease Borrelia clade and the relapsing fever Borrelia clade. Ideally, the genus name Borrelia should be retained for the Lyme disease Borrelia clade, which includes the best known species from this genus, B. burgdorferi, the first identified causative agent of Lyme disease (Barbour 1984). However, the type species of the genus Borrelia, Borrelia anserina, is a part of the relapsing fever clade. Hence, the genus name Borrelia must be retained for the relapsing fever clade (Bergey 1925; Lapage et al. 1992; Wang and Schwartz 2011). Therefore, species from the Lyme disease clade must be transferred to a new genus indicating their distinctness from the relapsing fever clade (viz. the emended genus Borrelia). To minimize confusion among scientists and other health care professionals, we are proposing that the species that are part of the Lyme disease clade should be transferred to a new genus, Borreliella gen. nov. The proposed name retains much of the original name of the genus Borrelia, thus it is unlikely that the species with the new names (e.g. B. burgdorferi) could be confused with any other unrelated species. The emended description of the genus Borrelia and a description of the newly proposed genus, Borreliella gen. nov, containing 14 new combinations, are provided below.
Emended description of the genus Borrelia (Swellengrebel 1907) (approved lists 1980)
Organisms are helical, 0.2–3 µm in diameter and 3–180 µm in length. Cells do not have hooked ends. Periplasmic flagella overlap in the central region of the cell. Cells are motile, host-associated and microaerophilic. The diamino acid component of the peptidoglycan is l-ornithine. Organisms are chemo-organotrophic and utilize carbohydrates or amino acids as carbon and energy sources. Members of this genus are the causative agents of relapsing fever. The G+C content of the genomic DNA is 27–32 (mol%). The type species is B. anserina (Bergey 1925) (Approved Lists 1980) (Skerman et al. 1980).
Organisms from this genus are distinguished from all other bacteria examined to date by the CSIs and conserved signature proteins described in this report (Tables 3, 4, 7).
Description of Borreliella gen. nov.
Borreliella (Bor.re’li.el’la. N.L. fem. dim. n. Borreliella, named after Amédée Borrel, a French bacteriologist)
Organisms are helical, 0.2–0.3 µm in diameter and 20–30 µm in length. Cells do not have hooked ends. Periplasmic flagella overlap in the central region of the cell. Cells are motile, host-associated and microaerophilic. The diamino acid component of the peptidoglycan is l-ornithine. Organisms are chemo-organotrophic and utilize carbohydrates or amino acids as carbon and energy sources. Members of this genus are the causative agents of Lyme disease. The G+C content of the genomic DNA is 26–29 (mol%). The type species is B. burgdorferi comb. nov.
Organisms from this genus are distinguished from all other bacteria examined to date by the CSIs and conserved signature proteins described in this report (Tables 3, 4, 7).
Description of Borreliella afzelii comb. nov.
Basonym: Borrelia afzelii (Canica et al. 1994)
The description of the species is the same as the description given for B. afzelii by Canica et al. (1994). The species exhibits the genus properties and contains the CSIs and CSPs indicated in the description of Borreliella.
Type Strain: VS461T (=ATCC 51567T = CIP 103469T = DSM 10508T)
Description of Borreliella americana comb. nov.
Basonym: Borrelia americana (Rudenko et al. 2010)
The description of the species is the same as the description given for B. americana by Rudenko et al. (2010). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: SCW-41T (=ATCC BAA-1877T = DSM 22541T)
Description of Borreliella bavariensis comb. nov.
Basonym: Borrelia bavariensis (Margos et al. 2013b)
The description of the species is the same as the description given for B. bavariensis by Margos et al. (2013a, b). The species exhibits the genus properties and contains the CSIs and CSPs indicated in the description of Borreliella.
Type Strain: PBiT (=DSM 23469T = BAA-2496T)
Description of Borreliella burgdorferi comb. nov.
Basonym: B. burgdorferi (Johnson et al. 1984)
The description of the species is the same as the description given for B. burgdorferi by Johnson et al. (1984). The species exhibits the genus properties and contains the CSIs and CSPs indicated in the description of Borreliella.
Type Strain: B31T (=ATCC 35210T = CIP 102532T = DSM 4680T)
Description of Borreliella carolinensis comb. nov.
Basonym: Borrelia carolinensis (Rudenko et al. 2011)
The description of the species is the same as the description given for B. carolinensis by Rudenko et al. (2011). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: SCW-22T (=ATCC BAA-1773T = DSM 22119T)
Description of Borreliella garinii comb. nov.
Basonym: Borrelia garinii (Baranton et al. 1992)
The description of the species is the same as the description given for B. garinii by Baranton et al. (1992). The species exhibits the genus properties and contains the CSIs and CSPs indicated in the description of Borreliella.
Type Strain: 20047T (=ATCC 51383T = CIP 103362T = DSM 10534T)
Description of Borreliella japonica comb. nov.
Basonym: Borrelia japonica (Kawabata et al. 1994)
The description of the species is the same as the description given for B. japonica by Kawabata et al. (1994). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: HO14T (=ATCC 51557T = JCM 8951T)
Description of Borreliella kurtenbachii comb. nov.
Basonym: Borrelia kurtenbachii (Margos et al. 2013a)
The description of the species is the same as the description given for B. kurtenbachii by Margos et al. (2013a, b). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: 25015T (=ATCC BAA-2495T = DSM 26572T)
Description of Borreliella lusitaniae comb. nov.
Basonym: Borrelia lusitaniae (Le Fleche et al. 1997)
The description of the species is the same as the description given for B. lusitaniae by Le Fleche et al. (1997). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: PotiB2T (=CIP 105366T)
Description of Borreliella sinica comb. nov.
Basonym: Borrelia sinica (Masuzawa et al. 2001)
The description of the species is the same as the description given for B. sinica by Masuzawa et al. (2001). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: CMN3T (=DSM 23262T = JCM 10505T)
Description of Borreliella spielmanii comb. nov.
Basonym: Borrelia spielmanii (Richter et al. 2006)
The description of the species is the same as the description given for B. spielmanii by Richter et al. (2006). The species exhibits the genus properties and contains the CSIs and CSPs indicated in the description of Borreliella.
Type Strain: PC-Eq17N5T (=CIP 108855T = DSM 16813T)
Description of Borreliella tanukii comb. nov.
Basonym: Borrelia tanukii (Fukunaga et al. 1997a)
The description of the species is the same as the description given for B. tanukii by Canica et al. (1994). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: Hk501T (=ATCC BAA-127T = JCM 9662T)
Description of Borreliella turdi comb. nov.
Basonym: Borrelia turdi (Fukunaga et al. 1997b)
The description of the species is the same as the description given for B. turdi by Fukunaga et al. (1997). The species exhibits the genus properties indicated in the description of Borreliella.
Type Strain: Ya501T (=ATCC BAA-126T = JCM 9661T)
Description of Borreliella valaisiana comb. nov.
Basonym: Borrelia valaisiana (Wang et al. 1997)
The description of the species is the same as the description given for B. valaisiana by Wang et al. (1997). The species exhibits the genus properties and contains the CSIs and CSPs indicated in the description of Borreliella.
Type Strain: VS116T (=CIP 105367T)
References
Adams DA, Gallagher KM, Jajosky RA, Kriseman J, Sharp P, Anderson WJ, Aranas AE, Mayes M, Wodajo MS, Onweh DH et al (2013) Summary of notifiable diseases—United States, 2011. MMWR Morb Mortal Wkly Rep 60(53):1–117
Adeolu M, Gupta RS (2013) Phylogenomics and molecular signatures for the order Neisseriales: proposal for division of the order Neisseriales into the emended family Neisseriaceae and Chromobacteriaceae fam nov. Antonie Van Leeuwenhoek Int J G 104(1):1–24
Ahmod NZ, Gupta RS, Shah HN (2011) Identification of a Bacillus anthracis specific indel in the yeaC gene and development of a rapid pyrosequencing assay for distinguishing B. anthracis from the B. cereus group. J Microbiol Methods 87(3):278–285
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Baranton G, Postic D, Saint Girons I, Boerlin P, Piffaretti J-C, Assous M, Grimont PAD (1992) Delineation of Borrelia burgdorferi Sensu Stricto, Borrelia garinii sp. nov., and Group VS461 associated with Lyme borreliosis. Int J Syst Bacteriol 42(3):378–383
Barbour AG (1984) Isolation and cultivation of Lyme disease spirochetes. Yale J Biol Med 57(4):521
Barbour AG (2005) Relapsing fever. In: Goodman JL, Dennis DT, Sonenshine DE (eds) Tick-borne diseases of humans. ASM Press, Washington, pp 268–291
Barbour AG, Miller SC (2014) Genome sequence of Borrelia parkeri, an agent of enzootic relapsing fever in Western North America. Genome Announc 2(1):e00018
Bergey DH (1925) Bergey’s manual of determinative bacteriology, 2nd edn. The Williams and Wilkins Co, Baltimore
Bhandari V, Gupta RS (2012) Molecular signatures for the phylum Synergistetes and some of its subclades. Antonie Van Leeuwenhoek 102(4):517–540
Bhandari V, Gupta RS (2014) Molecular signatures for the phylum (class) Thermotogae and a proposal for its division into three orders (Thermotogales, Kosmotogales ord. nov. and Petrotogales ord. nov.) containing four families (Thermotogaceae, Fervidobacteriaceae fam. nov., Kosmotogaceae fam. nov. and Petrotogaceae fam. nov.) and a new genus Pseudothermotoga gen. nov. with five new combinations. Antonie Van Leeuwenhoek 105(1):143–168
Bhandari V, Ahmod NZ, Shah HN, Gupta RS (2013) Molecular signatures for Bacillus species: demarcation of the Bacillus subtilis and Bacillus cereus clades in molecular terms and proposal to limit the placement of new species into the genus Bacillus. Int J Syst Evol Microbiol 63:2712–2726
Brenner EV, Kurilshikov AM, Stronin OV, Fomenko NV (2012) Whole-genome sequencing of Borrelia garinii BgVir, isolated from Taiga ticks (Ixodes persulcatus). J Bacteriol 194(20):5713
Canica MM, Du Merle L, Mazie JC, Baranton G, Postic D (1994) Borrelia afzelii sp. nov. Validation of the publication of new names and new combinations previously effectively published outside the IJSB, list no 48. Int J Syst Bacteriol 44:182–183
Casjens SR, Fraser-Liggett CM, Mongodin EF, Qiu WG, Dunn JJ, Luft BJ, Schutzer SE (2011a) Whole genome sequence of an unusual Borrelia burgdorferi sensu lato isolate. J Bacteriol 193(6):1489–1490
Casjens SR, Mongodin EF, Qiu WG, Dunn JJ, Luft BJ, Fraser-Liggett CM, Schutzer SE (2011b) Whole-genome sequences of two Borrelia afzelii and two Borrelia garinii Lyme disease agent isolates. J Bacteriol 193(24):6995–6996
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17(4):540–552
Chaconas G (2005) Hairpin telomeres and genome plasticity in Borrelia: all mixed up in the end. Mol Microbiol 58(3):625–635
Chaconas G, Kobryn K (2010) Structure, function, and evolution of linear replicons in Borrelia. Annu Rev Microbiol 64:185–202
Charlebois RL, Doolittle WF (2004) Computing prokaryotic gene ubiquity: rescuing the core from extinction. Genome Res 14(12):2469–2477
Ciccarelli FD, Doerks T, Von Mering C, Creevey CJ, Snel B, Bork P (2006) Toward automatic reconstruction of a highly resolved tree of life. Science 311(5765):1283–1287
Cutler SJ (2010) Relapsing fever: a forgotten disease revealed. J Appl Microbiol 108(4):1115–1122
Dai Q, Restrepo BI, Porcella SF, Raffel SJ, Schwan TG, Barbour AG (2006) Antigenic variation by Borrelia hermsii occurs through recombination between extragenic repetitive elements on linear plasmids. Mol Microbiol 60(6):1329–1343
Dunaj J, Moniuszko A, Zajkowska J, Pancewicz S (2013) The role of PCR in diagnostics of Lyme borreliosis. Przegl Epidemiol 67(1): 35–39, 119–123
Elbir H, Gimenez G, Robert C, Bergström S, Cutler S, Raoult D, Drancourt M (2012) Complete genome sequence of Borrelia crocidurae. J Bacteriol 194(14):3723–3724
Elbir H, Larsson P, Normark J, Upreti M, Korenberg E, Larsson C, Bergstrom S (2014a) Genome sequence of the Asiatic species Borrelia persica. Genome Announc 2(1):e01127
Elbir H, Larsson P, Upreti M, Normark J, Bergstrom S (2014b) Genome sequence of the relapsing fever borreliosis species Borrelia hispanica. Genome Announc 2(1):e01171
Fraser CM, Casjens S, Huang WM, Sutton GG, Clayton R, Lathigra R, White O, Ketchum KA, Dodson R, Hickey EK et al (1997) Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi. Nature 390(6660):580–586
Fukunaga M, Okada K, Nakao M, Konishi T, Sato Y (1996) Phylogenetic analysis of Borrelia species based on flagellin gene sequences and its application for molecular typing of Lyme disease borreliae. Int J Syst Bacteriol 46(4):898–905
Fukunaga M, Hamase A, Okada K, Nakao M (1997a) Borrelia tanukii sp. nov. Validation of the publication of new names and new combinations previously effectively published outside the IJSB, list no 63. Int J Syst Bacteriol 47:1274
Fukunaga M, Hamase A, Okada K, Nakao M (1997b) Borrelia turdi sp. nov. Validation of the publication of new names and new combinations previously effectively published outside the IJSB, list no 63. Int J Syst Bacteriol 47:1274
Gao B, Gupta R (2007) Phylogenomic analysis of proteins that are distinctive of Archaea and its main subgroups and the origin of methanogenesis. BMC Genom 8(1):86
Gao B, Gupta RS (2012a) Microbial systematics in the post-genomics era. Antonie Van Leeuwenhoek 101(1):45–54
Gao B, Gupta RS (2012b) Phylogenetic framework and molecular signatures for the main clades of the phylum Actinobacteria. Microbiol Mol Biol Rev 76(1):66–112
Glöckner G, Lehmann R, Romualdi A, Pradella S, Schulte-Spechtel U, Schilhabel M, Wilske B, Sühnel J, Platzer M (2004) Comparative analysis of the Borrelia garinii genome. Nucleic Acids Res 32(20):6038–6046
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57(1):81–91
Gupta RS (1998) Protein phylogenies and signature sequences: a reappraisal of evolutionary relationships among archaebacteria, eubacteria, and eukaryotes. Microbiol Mol Biol Rev 62(4):1435
Gupta RS (2010) Applications of conserved indels for understanding microbial phylogeny. In: Oren A, Papke RT (eds) Molecular phylogeny of microorganisms. Caister Academic Press, Norfolk, pp 135–150
Gupta RS, Griffiths E (2006) Chlamydiae-specific proteins and indels: novel tools for studies. Trends Microbiol 14(12):527–535
Gupta RS, Lali R (2013) Molecular signatures for the phylum Aquificae and its different clades: proposal for division of the phylum Aquificae into the emended order Aquificales, containing the families Aquificaceae and Hydrogenothermaceae, and a new order Desulfurobacteriales ord. nov., containing the family Desulfurobacteriaceae. Antonie Van Leeuwenhoek 104(3):349–368
Gupta RS, Chander P, George S (2013a) Phylogenetic framework and molecular signatures for the class Chloroflexi and its different clades; proposal for division of the class Chloroflexia class. nov. [corrected] into the suborder Chloroflexineae subord. nov., consisting of the emended family Oscillochloridaceae and the family Chloroflexaceae fam. nov., and the suborder Roseiflexineae subord. nov., containing the family Roseiflexaceae fam. nov. Antonie Van Leeuwenhoek 103(1):99–119
Gupta RS, Mahmood S, Adeolu M (2013b) A phylogenomic and molecular signature based approach for characterization of the phylum Spirochaetes and its major clades: proposal for a taxonomic revision of the phylum. Front Microbiol 4:217
Harris JK, Kelley ST, Spiegelman GB, Pace NR (2003) The genetic core of the universal ancestor. Genome Res 13(3):407–412
Hue F, Ghalyanchi Langeroudi A, Barbour AG (2013) Chromosome sequence of Borrelia miyamotoi, an uncultivable tick-borne agent of human infection. Genome Announc 1(5):e00713
Ibba M, Bono JL, Rosa PA, Soll D (1997) Archaeal-type lysyl-tRNA synthetase in the Lyme disease spirochete Borrelia burgdorferi. Proc Natl Acad Sci USA 94(26):14383–14388
Jeanmougin F, Thompson JD, Gouy M, Higgins DG, Gibson TJ (1998) Multiple sequence alignment with Clustal X. Trends Biochem Sci 23(10):403
Jiang B, Yao H, Tong Y, Yang X, Huang Y, Jiang J, Cao W (2012a) Genome sequence of Borrelia garinii strain NMJW1, isolated from China. J Bacteriol 194(23):6660–6661
Jiang BG, Zheng YC, Tong YG, Jia N, Huo QB, Fan H, Ni XB, Ma L, Yang XF, Jiang JF et al (2012b) Genome sequence of Borrelia afzelii Strain HLJ01, isolated from a patient in China. J Bacteriol 194(24):7014–7015
Johnson RC, Schmid GP, Hyde FW, Steigerwalt AG, Brenner DJ (1984) Borrelia burgdorferi sp. nov.: etiologic agent of Lyme disease. Int J Syst Bacteriol 34(4):496–497
Kawabata H, Masuzawa T, Yanagihara Y (1994) Borrelia japonica sp. nov. Validation of the publication of new names and new combinations previously effectively published outside the IJSB, list no 50. Int J Syst Bacteriol 44:595
Lapage SP, Sneath PHA, Lessel EF, Skerman VBD, Seeliger HPR, Clark WA (1992) International code of nomenclature of bacteria: bacteriological code, 1990 revision. ASM Press International Union of Microbiological Societies, Washington
Le Fleche A, Postic D, Girardet K, Peter O, Baranton G (1997) Characterization of Borrelia lusitaniae sp. nov. by 16S ribosomal DNA sequence analysis. Int J Syst Bacteriol 47(4):921–925
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25(7):1307–1320
Lescot M, Audic S, Robert C, Nguyen TT, Blanc G, Cutler SJ, Wincker P, Couloux A, Claverie JM, Raoult D (2008) The genome of Borrelia recurrentis, the agent of deadly louse-borne relapsing fever, is a degraded subset of tick-borne Borrelia duttonii. PLoS Genet 4(9):e1000185
Lindgren E, Jaenson TG (2006) Lyme borreliosis in Europe: influences of climate and climate change, epidemiology, ecology and adaptation measures. WHO Regional Office for Europe, Copenhagen
Ljostad U, Mygland A (2013) Chronic Lyme; diagnostic and therapeutic challenges. Acta Neurol Scand 127 Suppl(196):38–47
Margos G, Vollmer SA, Cornet M, Garnier M, Fingerle V, Wilske B, Bormane A, Vitorino L, Collares-Pereira M, Drancourt M et al (2009) A new Borrelia species defined by multilocus sequence analysis of housekeeping genes. Appl Environ Microbiol 75(16):5410–5416
Margos G, Vollmer SA, Ogden NH, Fish D (2011) Population genetics, taxonomy, phylogeny and evolution of Borrelia burgdorferi sensu lato. Infect Genet Evol 11(7):1545–1563
Margos G, Piesman J, Lane RS, Ogden NH, Sing A, Straubinger RK, Fingerle V (2013a). Borrelia kurtenbachii sp. nov.: a widely distributed member of the Borrelia burgdorferi sensu lato species complex in North America. Int J Syst Evol Microbiol 64(Pt 1):128–30. doi:10.1099/ijs.0.054593-0
Margos G, Wilske B, Sing A, Hizo-Teufel C, Cao WC, Chu C, Scholz H, Straubinger RK, Fingerle V (2013b) Borrelia bavariensis sp. nov. is widely distributed in Europe and Asia. Int J Syst Evol Microbiol 63(Pt 11):4284–4288
Masuzawa T, Takada N, Kudeken M, Fukui T, Yano Y, Ishiguro F, Kawamura Y, Imai Y, Ezaki T (2001) Borrelia sinica sp. nov., a lyme disease-related Borrelia species isolated in China. Int J Syst Evol Microbiol 51(Pt 5):1817–1824
Naushad HS, Lee B, Gupta RS (2014) Conserved signature indels and signature proteins as novel tools for understanding microbial phylogeny and systematics: identification of molecular signatures that are specific for the phytopathogenic genera Dickeya, Pectobacterium and Brenneria. Int J Syst Evol Microbiol 64(2):366–383
NCBI (2014) NCBI genome database. http://www.ncbi.nlm.nih.gov/genome/
Parte AC (2014) LPSN—list of prokaryotic names with standing in nomenclature. Nucleic Acids Res 42(D1):D613–D616
Paster BJ (2011) Phylum XV. Spirochaetes Garrity and Holt 2001. In Brenner DJ, Krieg NR, Garrity GM, Staley JT (eds) Bergey’s Manual of Systematic Bacteriology, 2nd edn, vol 3. Springer, New York, pp 471–471 (reprinted from: not in File)
Postic D, Edlinger C, Richaud C, Grimont F, Dufresne Y, Perolat P, Baranton G, Grimont PAD (1990) Two genomic species in Borrelia burgdorferi. Res Microbiol 141(4):465–475
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):D590–D596
Ras NM, Lascola B, Postic D, Cutler SJ, Rodhain F, Baranton G, Raoult D (1996) Phylogenesis of relapsing fever Borrelia spp. Int J Syst Bacteriol 46(4):859–865
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci 106(45):19126–19131
Richter D, Postic D, Sertour N, Livey I, Matuschka FR, Baranton G (2006) Delineation of Borrelia burgdorferi sensu lato species by multilocus sequence analysis and confirmation of the delineation of Borrelia spielmanii sp. nov. Int J Syst Evol Microbiol 56(Pt 4):873–881
Rokas A, Holland PWH (2000) Rare genomic changes as a tool for phylogenetics. Trends Ecol Evol 15(11):454–459
Rokas A, Williams BL, King N, Carroll SB (2003) Genome-scale approaches to resolving incongruence in molecular phylogenies. Nature 425(6960):798–804
Rosselló-Mora R (2006) DNA–DNA reassociation methods applied to microbial taxonomy and their critical evaluation. In: Stackebrandt E (ed) Molecular identification, systematics, and population structure of prokaryotes. Springer, Berlin, pp 23–50
Rudenko N, Golovchenko M, Lin T, Gao L, Grubhoffer L, Oliver JH Jr (2010) Borrelia americana sp. nov. List of new names and new combinations previously effectively, but not validly, published, list no 135. Int J Syst Evol Microbiol 60:1985–1986
Rudenko N, Golovchenko M, Grubhoffer L, Oliver JH Jr (2011) Borrelia carolinensis sp. nov., a novel species of the Borrelia burgdorferi sensu lato complex isolated from rodents and a tick from the south-eastern USA. Int J Syst Evol Microbiol 61(Pt 2):381–383
Schutzer SE, Fraser-Liggett CM, Casjens SR, Qiu WG, Dunn JJ, Mongodin EF, Luft BJ (2011) Whole-genome sequences of thirteen isolates of Borrelia burgdorferi. J Bacteriol 193(4):1018–1020
Schutzer SE, Fraser-Liggett CM, Qiu WG, Kraiczy P, Mongodin EF, Dunn JJ, Luft BJ, Casjens SR (2012) Whole-genome sequences of Borrelia bissettii, Borrelia valaisiana, and Borrelia spielmanii. J Bacteriol 194(2):545–546
Segata N, Bornigen D, Morgan XC, Huttenhower C (2013) PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat Commun 4:2304
Skerman VBD, McGowan V, Sneath PHA (1980) Approved lists of bacterial names. Int J Syst Bacteriol 30(1):225–420
Swellengrebel NH (1907) Sur la cytologie comparée des spirochètes et des spirilles. Ann Inst Pasteur (Paris) 21:562–586
Takano A, Goka K, Une Y, Shimada Y, Fujita H, Shiino T, Watabane H, Kawabata H (2010) Isolation and characterization of a novel Borrelia group of tick-borne borreliae from imported reptiles and their associated ticks. Environ Microbiol 12(1):134–146
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729
Tavaré S (1986) Some probabilistic and statistical problems in the analysis of DNA sequences. In Miura RM (ed) Lectures on mathematics in the life sciences, 17th edn. American Mathematical Society, Providence, pp 57–86 (reprinted from: not in file)
Thompson CC, Chimetto L, Edwards RA, Swings J, Stackebrandt E, Thompson FL (2013) Microbial genomic taxonomy. BMC Genom 14(1):913
Valsangiacomo C, Balmelli T, Piffaretti JC (1997) A phylogenetic analysis of Borrelia burgdorferi sensu lato based on sequence information from the hbb gene, coding for a histone-like protein. Int J Syst Bacteriol 47(1):1–10
Vinuesa P (2010) Multilocus sequence analysis and bacterial species phylogeny estimation. In: Oren A, Papke RT (eds) Molecular phylogeny of microorganisms. Caister Academic Press, Norfolk, pp 41–64
Wang G, Schwartz I (2011) Genus II. Borrelia Swellengrebel 1907, 582AL. In: Brenner DJ, Krieg NR, Garrity GM, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 3, 2nd edn. Springer, New York, pp 484–498
Wang G, van Dam AP, Le Fleche A, Postic D, Peter O, Baranton G, de Boer R, Spanjaard L, Dankert J (1997) Genetic and phenotypic analysis of Borrelia valaisiana sp. nov. (Borrelia genomic groups VS116 and M19). Int J Syst Bacteriol 47(4):926–932
Wang G, van Dam AP, Schwartz I, Dankert J (1999) Molecular typing of Borrelia burgdorferi sensu lato: taxonomic, epidemiological, and clinical implications. Clin Microbiol Rev 12(4):633–653
Wright WF, Riedel DJ, Talwani R, Gilliam BL (2012) Diagnosis and management of Lyme disease. Am Fam Physician 85(11):1086–1093
Wu D, Hugenholtz P, Mavromatis K, Pukall R, Dalin E, Ivanova NN, Kunin V, Goodwin L, Wu M, Tindall BJ (2009) A phylogeny-driven genomic encyclopaedia of bacteria and archaea. Nature 462(7276):1056–1060
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Adeolu, M., Gupta, R.S. A phylogenomic and molecular marker based proposal for the division of the genus Borrelia into two genera: the emended genus Borrelia containing only the members of the relapsing fever Borrelia, and the genus Borreliella gen. nov. containing the members of the Lyme disease Borrelia (Borrelia burgdorferi sensu lato complex). Antonie van Leeuwenhoek 105, 1049–1072 (2014). https://doi.org/10.1007/s10482-014-0164-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-014-0164-x