Abstract
A novel actinomycete, designated strain NEAU-GRX11T, was isolated from muddy soil collected from a stream of Jinlong Mountain in Harbin, north China. The organism was found to have morphological and chemotaxonomic characteristics typical of the genus Micromonospora. The 16S rRNA gene sequence of strain NEAU-GRX11T showed highest similarity to Micromonospora zamorensis CR38T (99.2 %), Micromonospora saelicesensis Lupac 09T (99.0 %), Micromonospora chokoriensis 2-19/6T (98.7 %), Micromonospora coxensis 2-30-b/28T (98.5 %), Micromonospora aurantiaca ATCC 27029T (98.4 %) and Micromonospora lupini lupac 14NT (98.3 %). Phylogenetic analysis based on the 16S rRNA gene and gyrB gene demonstrated that strain NEAU-GRX11T was a member of the genus Micromonospora and supported the closest phylogenetic relationship to M. zamorensis CR38T, M. saelicesensis Lupac 09T, M. chokoriensis 2-19/6T and M. lupini lupac 14NT. A combination of DNA–DNA hybridization and some phenotypic characteristics indicated that the novel strain could be readily distinguished from these closest phylogenetic relatives. Therefore, it is proposed that NEAU-GRX11T represents a novel species of the genus Micromonospora, for which the name Micromonospora jinlongensis sp. nov. is proposed. The type strain is NEAU-GRX11T (=CGMCC 4.7103T=DSM 45876T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Micromonospora was first proposed by Ørskov (1923) for actinomycetes that formed single, non-motile spores directly on the substrate mycelium, but no aerial mycelia were produced. Subsequently, this genus was well classified on the basis of chemotaxonomic characteristics, gyrB and 16S rRNA gene sequence analyses (Kroppenstedt 1985; Koch et al. 1996; Kasai et al. 2000). Micromonospora are widely distributed in nature, inhabiting such disparate environments as soil, water, marine environments and plant tissues (Lǖdemann and Brodsky 1963; Kawamoto 1989; Mincer et al. 2002; Maldonado et al. 2009; Trujillo et al. 2010; Li et al. 2013a). Up to now, the genus Micromonospora comprises 56 species including recently described Micromonospora sediminicola (Supong et al. 2013a), Micromonospora spongicola (Supong et al. 2013b), Micromonospora equina (Everest and Meyers 2013), Micromonospora maritima (Songsumanus et al. 2013), Micromonospora schwarzwaldensis (Gurovic et al. 2013), Micromonospora avicenniae (Li et al. 2013a), Micromonospora sonneratiae (Li et al. 2013b), Micromonospora halotolerans (Carro et al. 2013) and Micromonospora kangleipakensis (Nimaichand et al. 2013). During the investigation of exploring potential sources of novel species and novel natural products, strain NEAU-GRX11T was isolated from muddy soil collected from a stream of Jinlong Mountain in Harbin, north China. In this study, the taxonomic status of this strain is reported based on phylogenetic, chemotaxonomic and physiological evidences. It is proposed that strain NEAU-GRX11T is a new species of the genus Micromonospora.
Materials and methods
Isolation and maintenance of the organism
Strain NEAU-GRX11T was isolated from muddy soil collected from a stream of Jinlong Mountain in Harbin, north China (45°30′N, 127°06′E), using the standard dilution plate method and grown on humic acid-vitamin agar (HV) (Hayakawa and Nonomura 1987) supplemented with nystatin (50 mg l−1) and nalidixic acid (20 mg l−1). After 21 days of aerobic incubation at 28 °C, colonies were transferred and purified on oatmeal agar (International Streptomyces Project (ISP) 3 medium) (Shirling and Gottlieb 1966) and maintained as glycerol suspensions (20 %, v/v) at −80 °C.
Morphological, cultural and physiological characteristics
Morphological properties were observed by light microscopy (Nikon ECLIPSE E200) and scanning electron microscopy (Hitachi S-3400 N) using cultures grown on ISP 3 agar for 21 days at 28 °C. Cultural characteristics were determined by growth on Czapek’s agar (Waksman 1967), nutrient agar (Waksman 1961) and ISP media 2–7 (Shirling and Gottlieb 1966) at 28 °C for 14 days. The ISCC–NBS colour charts were used to determine the names and designations of colony colours (Kelly 1964). Growth at different temperatures (4, 10, 15, 18, 22, 28, 32, 37, 40, 45 °C) was determined on ISP 3 agar after incubation for 14 days. Growth tests for pH range (4.0, 5.0, 6.0, 7.0, 8.0, 9.0 and 10.0) were carried out by using the buffer system described by Xie et al. (2012), and NaCl tolerance was determined in GY medium (Jia et al. 2013) supplemented with 0–6 % NaCl (w/v) at 28 °C for 7 days on a rotary shaker. Production of catalase, esterase and urease were tested as described by Smibert and Krieg (1994). The utilization of sole carbon and nitrogen, decomposition of cellulose, hydrolysis of starch and aesculin, reduction of nitrate, peptonization of milk, liquefaction of gelatin and production of H2S were examined as described previously (Gordon et al. 1974; Yokota et al. 1993).
Chemotaxonomic characterization
The freeze-dried cells used for chemotaxonomic analysis were obtained from cultures grown in GY medium on a rotary shaker for 4 days at 28 °C. Cells were harvested by centrifugation, washed with distilled water and freeze-dried. The isomers of diaminopimelic acid (DAP) in peptidoglycan were analysed by HPLC method using Agilent TC-C18 Column (250 × 4.6 mm i.d. 5 μm) with a mobile phase consisting of acetonitrile: 0.05 mol l−1 phosphate buffer pH 7.2 = 15:85 at a flow rate of 0.5 ml min−1. The peak detection used an Agilent G1321A fluorescence detector with a 365 nm excitation and 455 nm longpass emission filters (McKerrow et al. 2000). The N-acyl group of muramic acid in peptidoglycan was determined by the method of Uchida et al. (1999). The whole-organism sugars were analyzed according to the procedures developed by Lechevalier and Lechevalier (1980). Phospholipids in cells were extracted and identified by using the method of Minnikin et al. (1984). Menaquinones were extracted from freeze-dried biomass and purified according to Collins (1985). Extracts were analyzed by HPLC–UV method using Agilent Extend-C18 Column (150 × 4.6 mm, i.d. 5 μm), typically at 270 nm. The mobile phase was acetonitrile–propyl alcohol (60:40, v/v) and the flow rate was set to 1.0 ml min−1 and the run time was 60 min. The injection volume was 20 μl, and the chromatographic column was controlled at 40 °C (Wu et al. 1989). The presence of mycolic acids was checked by the acid methanolysis method as described previously (Minnikin et al. 1980). Cells used for cellular fatty acids analysis were obtained with the same cultural condition above, then cells were harvested by centrifugation (6,000 rpm, 10 min), soaked for 24 h and centrifuged (3,000 rpm, 10 min). The supernatant was concentrated and placed in test tubes. To each tube, 2 ml of 1 M NaOH in methanol was added, and the tubes were sealed with nitrogen gas, vortexed, and placed in a 70 °C water bath for 30 min. The tubes were cooled to room temperature, 3 ml of 20 % boron trifluoride was added, then the tubes were sealed, vortexed, and placed in water bath with the same condition above. The tubes were cooled to room temperature, 5 ml of saturated NaCl and 2 ml n-heptane was added and centrifuged (4,000 rpm, 10 min). The heptane phase was used for cellular fatty acids analysis. The composition was analysed by GC–MS using the method of Xiang et al. (2011).
DNA preparation, amplification and determination of 16S rRNA and gyrB gene sequences
Extraction of chromosomal DNA and PCR-mediated amplification of the 16S rRNA gene were carried out using a standard procedure (Kim et al. 2000). The PCR product was purified and cloned into the vector pMD19-T (Takara) and sequenced by using an Applied Biosystems DNA sequencer (model 3730XL) and software provided by the manufacturer. Almost full-length 16S rRNA gene sequence (1,507 nt) was obtained and aligned with multiple sequences obtained from the GenBank/EMBL/DDBJ databases using Clustal X 1.83 software. Phylogenetic trees were generated with the neighbour-joining (Saitou and Nei 1987) and maximum-likelihood (Felsenstein 1981) algorithms using Molecular Evolutionary Genetics Analysis (MEGA) software version 5.05 (Tamura et al. 2011). The stability of the clades in the trees was appraised using a bootstrap value with 1,000 repeats (Felsenstein 1985). A distance matrix was generated using Kimura’s two-parameter model (Kimura 1980). All positions containing gaps and missing data were eliminated from the dataset (complete deletion option). 16S rRNA gene sequence similarities between strains were calculated on the basis of pairwise alignment using the EzTaxon-e server (Kim et al. 2012). PCR amplification of the gyrB gene was carried out using primers GYF1 and GYB3 (Garcia et al. 2010) by the PCR program for 16S rRNA gene. Sequencing and phylogenetic analysis was performed as described above. Actinoplanes regularis IFO 12514T was used as an outgroup.
DNA base composition and DNA–DNA hybridization
The G+C content of the genomic DNA was determined by the thermal denaturation (Tm) method as described by Mandel and Marmur (1968), and Escherichia coli JM109 DNA was used as the reference strain. DNA–DNA relatedness tests between isolate NEAU-GRX11T and Micromonospora saelicesensis Lupac 09T, Micromonospora zamorensis CR38T, Micromonospora chokoriensis 2-19/6T and Micromonospora lupini lupac 14NT were carried out as described by De Ley et al. (1970) under consideration of the modifications described by Huss et al. (1983), using a model Cary 100 Bio UV/VIS-spectrophotometer equipped with a Peltier-thermostatted 6 × 6 multicell changer and a temperature controller with in situ temperature probe (Varian).
Results and discussion
The almost-complete 16S rRNA gene sequence (1,507 nt) of strain NEAU-GRX11T was determined and deposited in the GenBank/EMBL/DDBJ databases as KC134254. Based on EzTaxon-e analysis, the species showing most closely related to the novel isolate were M. zamorensis CR38T (99.2 %), M. saelicesensis Lupac 09T (99.0 %), M. chokoriensis 2-19/6T (98.7 %), Micromonospora coxensis 2-30-b/28T (98.5 %), Micromonospora aurantiaca ATCC 27029T (98.4 %) and M. lupini lupac 14NT (98.3 %). The phylogenetic tree (Fig. 1) based on 16S rRNA gene sequences showed that strain NEAU-GRX11T formed a distinct phyletic line with M. zamorensis CR38T, M. saelicesensis Lupac 09T, M. chokoriensis 2-19/6T and M. lupini lupac 14NT, an association that was supported by maximum-likelihood algorithm employed (Supplementary Fig. S1) and by a 79 % bootstrap value in the neighbour-joining analysis. The similarity of gyrB gene (1,162 nt) between strain NEAU-GRX11T and M. zamorensis CR38T, M. saelicesensis Lupac 09T, M. chokoriensis 2-19/6T and M. lupini lupac 14NT were 98.5, 96.7, 95.7 and 97.4 %, respectively. Phylogenetic analysis of gyrB gene sequence supported that strain NEAU-GRX11T was placed in the genus Micromonospora, also near to M. zamorensis CR38T, M. saelicesensis Lupac 09T, M. chokoriensis 2-19/6T and M. lupini lupac 14NT by a 76 % bootstrap value (Supplementary Fig. S2). Further study showed that the DNA relatedness between strain NEAU-GRX11T and M. zamorensis CR38T, M. saelicesensis Lupac 09T, M. chokoriensis 2-19/6T and M. lupini lupac 14NT were 55.5 ± 0.8, 34.6 ± 1.2, 57.3 ± 1.1 and 59.5 ± 2.0 %, respectively. These values were below the threshold value of 70 % recommended by Wayne et al. (1987) for assigning strains to the same species.
Neighbour-joining phylogenetic tree based on nearly complete 16S rRNA gene sequence (1,507 nt) showing the relationship between strain NEAU-GRX11T and all species of the genus Micromonospora. Catellatospora citrea subsp. citrea DSM 44097T was used as an outgroup. Asterisks indicate branches of the tree that were also recovered using the maximum-likelihood method. Bootstrap values >50 % (based on 1,000 replications) were shown at branch points. Bar 0.005 substitutions per nucleotide position
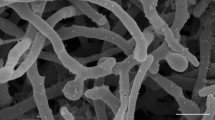
The morphological and cultural properties of strain NEAU-GRX11T are consistent with its classification as a member of the genus Micromonospora (Kawamoto 1989). Strain NEAU-GRX11T produced well-developed and branched substrate hyphae on ISP 3 medium. Non-motile and oval spores (0.6 × 0.8 μm) were borne singly on the substrate mycelium and the spore surface was smooth (Fig. 2). The novel isolate showed good growth on ISP 3 and ISP 6 agar, moderate growth on ISP 4 and ISP 7 agar, poor growth on Czapek’s, nutrient and ISP 5 agar, and no growth on ISP 2 agar. The colour of colonies on different media was very dark purple (ISP 3), dark grayish purple (ISP 6), pale purple (ISP 5), strong orange yellow (ISP 4), pale yellow (ISP 7) and pale greenish yellow (nutrient and Czapek’s agar). No aerial mycelium was seen on these media tested. Deep purple soluble pigment was produced on ISP 3 agar. Melanin was not observed on ISP 6 and ISP 7 agar. Strain NEAU-GRX11T was observed to grow well between pH 6.0–9.0, with an optimum pH of 8.0. The temperature range for growth was determined to be 18–37 °C, with the optimum growth temperature at 28 °C. Strain NEAU-GRX11T was observed to grow in presences of 0–3 % NaCl (w/v). Detailed physiological characteristics are presented in the species description.
Hydrolysates of strain NEAU-GRX11T were found to contain glycine and meso-diaminopimelic acid (DAP). The whole-cell sugars were rhamnose, xylose and glucose (Supplementary Fig. S3). The acyl type of the cell wall polysaccharides was glycolyl. Mycolic acids were not detected. The predominant menaquinones were MK-9(H4) (60.4 %), MK-9(H6) (31.5 %) and MK-9(H8) (8.1 %). The phospholipid profile consisted of diphosphatidylglycerol (DPG), phosphatidylmonomethylethanolamine (PME), phosphatidylethanolamine (PE) and phosphatidylinositol (PI) (Supplementary Fig. S4), corresponding to phospholipid type PII of Lechevalier et al. (1977). The cellular fatty acid profile was determined to be composed of C16:0 (39.3 %), C15:0 (16.3 %), C17:1 ω8c (11.8 %), C18:0 (9.2 %), 10-methyl C17:0 (8.6 %), C17:0 (4.4 %), anteiso-C17:0 (4.3 %), C18:1 ω12c (2.9 %), C16:1 ω9c (1.9 %) and C13:0 (1.2 %) (Supplementary Fig. S5; Table S1). The DNA G+C content of the strain was 70.04 ± 0.21 mol%.
Strain NEAU-GRX11T presented a phenotypic profile that clearly distinguished it from its closest phylogenetic neighbours, in particular soluble pigment on various media, hydrolysis of starch, urea hydrolysis and patterns of carbon and nitrogen utilization, and the phospholipids (presence of PME) and major cellular fatty acid compositions (C16:0 and C15:0) of strain NEAU-GRX11T could differentiate it from its most related type strains of the genus Micromonospora (Table 1). Furthermore, a low level of DNA–DNA relatedness was observed between strain NEAU-GRX11T and its closest phylogenetic relatives. It is evident from the genotypic and phenotypic data presented above that strain NEAU-GRX11T is distinguishable from previously described Micromonospora species. Therefore, strain NEAU-GRX11T represents a novel species of the genus Micromonospora, for which the name Micromonospora jinlongensis sp. nov. is proposed.
Emended description of the genus Micromonospora
All the descriptions are similar to that given by Ørskov (1923), Lechevalier et al. (1977), Kroppenstedt (1985) and Kawamoto (1989) except for the following changes. Micromonospora strains generally have large amounts of iso- and anteiso-fatty acids in their cell membranes, but C16:0 and C15:0 may be major fatty acids. The phospholipid profile may consist of PME.
Description of Micromonospora jinlongensis sp. nov
Micromonospora jinlongensis (jin.long.en′sis. N.L. masc. adj. jinlongensis of or pertaining to Jinlong, the name of a mountain in Harbin, Heilongjiang Province, north China, from where the type strain was isolated).
Aerobic, Gram-stain positive actinomycete that forms well-developed and branched substrate hyphae. Colonies are in the yellow and purple color-series. Grows well on ISP 3 and ISP 6 agar; moderately on ISP 4 and ISP 7 agar; poorly on Czapek’s, nutrient and ISP 5 agar; no growth is observed on ISP 2 agar. Deep purple soluble pigment is produced on ISP 3 agar. Single, non-motile and oval spores (0.6 × 0.8 μm) with a smooth surface are produced. Positive for production of catalase and esterase, hydrolysis of aesculin and starch and liquefaction of gelatin and negative for decomposition of cellulose and urea, reduction of nitrate, peptonization of milk and production of H2S. d-Glucose, d-sorbitol, d-maltose, d-sucrose, d-mannitol, inositol, l-rhamnose, l-arabinose, d-raffinose and lactose are utilized as sole carbon sources but d-mannose, d-ribose, d-fructose, d-xylose and d-galactose are not utilized. l-Arginine, l-asparagine, l-glutamine, l-glutamic acid, l-aspartic acid, l-serine and l-threonine are utilized as sole nitrogen sources but creatine, l-alanine, glycine and l-tyrosine are not utilized. Tolerates up to 3 % NaCl and grows at temperatures between 18 and 37 °C, with an optimum temperature of 28 °C. Growth occurs at initial pH values between 6.0 and 9.0, the optimum being pH 8.0. Cell walls contain meso-diaminopimelic acid as diagnostic diamino acid and the whole cell sugars are rhamnose, xylose and glucose. The acyl type of the cell wall polysaccharides is glycolyl. Mycolic acids are not detected. The predominant menaquinones are MK-9(H4) and MK-9(H6). The phospholipid profile consists of DPG, PME, PE and PI. The predominant fatty acids are C16:0, C15:0, C17:1 ω8c, C18:0 and 10-methyl C17:0 (>5.0 %). The DNA G+C content of the type strain is 70.04 ± 0.21 mol%.
The type strain is NEAU-GRX11T (=CGMCC 4.7103T=DSM 45876T), which was isolated from muddy soil collected on Jinlong Mountain in Harbin, China. The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene and gyrB gene of strain NEAU-GRX11T are KC134254 and KF318646, respectively.
References
Ara I, Kudo T (2007) Two new species of the genus Micromonospora: micromonospora chokoriensis sp. nov. and Micromonospora coxensis sp. nov., isolated from sandy soil. J Gen Appl Microbiol 53:29–37
Carro L, Pukall R, Spröer C, Kroppenstedt RM, Trujillo ME (2012) Micromonospora cremea sp. nov. and Micromonospora zamorensis sp. nov., isolated from the rhizosphere of Pisum sativum. Int J Syst Evol Microbiol 62:2971–2977
Carro L, Pukall R, Spröer C, Kroppenstedt RM, Trujillo ME (2013) Micromonospora halotolerans sp. nov., isolated from the rhizosphere of a Pisum sativum plant. Antonie Leeuwenhoek Int J Gen 103:1245–1254
Collins MD (1985) Isoprenoid quinone analyses in bacterial classification and identification. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic Press, London, pp 267–284
De Ley J, Cattoi H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Everest GJ, Meyers PR (2013) Micromonospora equina sp. nov., isolated from soil from a racecourse. Int J Syst Evol Microbiol 63:879–885
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Garcia LC, Martínez-Molina E, Trujillo ME (2010) Micromonospora pisi sp. nov., isolated from root nodules of Pisum sativum. Int J Syst Evol Microbiol 60:331–337
Gordon RE, Barnett DA, Handerhan JE, Pang C (1974) Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Bacteriol 24:54–63
Gurovic MS, Müller S, Domin N, Seccareccia I, Nietzsche S, Martin K, Nett M (2013) Micromonospora schwarzwaldensis sp. nov., a producer of telomycin, isolated from soil in the Black Forest. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.051623-0
Hayakawa M, Nonomura H (1987) Humic acid-vitamin agar, a new medium for the selective isolation of soil actinomycetes. J Ferment Technol 65:501–509
Huss VAR, Festl H, Schleifer KH (1983) Studies on the spectrometric determination of DNA hybridisation from renaturation rates. Syst Appl Microbiol 4:184–192
Jia FY, Liu CX, Wang XJ, Zhao JW, Liu QF, Zhang J, Gao RX, Xiang WS (2013) Wangella harbinensis gen. nov., sp. nov., a new member of the family Micromonosporaceae. Antonie Leeuwenhoek Int J Gen 103:399–408
Kasai H, Tamura T, Harayama S (2000) Intrageneric relationships among Micromonospora species deduced from gyrB-based phylogeny and DNA relatedness. Int J Syst Evol Microbiol 50:127–134
Kawamoto I (1989) Genus micromonospora Orskov 1923 147AL. In: Williams ST, Sharpe ME, Holt JG (eds) Ber-gey’s manual of systematic bacteriology, vol 4. Williams and Wilkins, Baltimore, pp 2442–2450
Kelly KL (1964) Inter-Society Color Council–National Bureau of Standards color name Charts illustrated with centroid colors. US Government Printing Office, Washington, DC
Kim SB, Brown R, Oldfield C, Gilbert SC, Iliarionov S, Goodfellow M (2000) Gordonia amicalis sp. nov., a novel dibenzothiophene-desulphurizing actinomycete. Int J Syst Evol Microbiol 50:2031–2036
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Koch C, Kroppenstedt RM, Stackebrandt E (1996) Intrageneric relationships of the actinomycete genus Micromonospora. Int J Syst Bacteriol 46:383–387
Kroppenstedt RM (1985) Fatty acid and menaquinone analysis of actinomycetes and related organisms. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic press, London, pp 173–199
Lechevalier MP, Lechevalier HA (1980) The chemotaxonomy of actinomycetes. In: Dietz A, Thayer DW (eds) Actinomycete taxonomy special publication, vol 6. Society of Industrial Microbiology, Arlington, pp 227–291
Lechevalier MP, De Bièvre C, Lechevalier HA (1977) Chemotaxonomy of aerobic actinomycetes: phospholipid composition. Biochem Syst Ecol 5:249–260
Li L, Mao YJ, Xie QY, Deng Z, Hong K (2013a) Micromonospora avicenniae sp. nov., isolated from a root of Avicennia marina. Antonie Leeuwenhoek Int J Gen 103:1089–1096
Li L, Tang YL, Wei B, Xie QY, Deng Z, Hong K (2013b) Micromonospora sonneratiae sp. nov., isolated from a root of Sonneratia apetala. Int J Syst Evol Microbiol 63:2383–2388
Lǖdemann GM, Brodsky BC (1963) Taxonomy of gentamicin-producing Micromonospora. Antimicrob Agents Chemother 161:116–124
Maldonado LA, Fragoso-Yáñez D, Pérez-García A, Rosellón-Druker J, Quintana ET (2009) Actinobacterial diversity from marine sediments collected in Mexico. Antonie Leeuwenhoek Int J Gen 95:111–120
Mandel M, Marmur J (1968) Use of ultraviolet absorbance temperature profile for determining the guanine plus cytosine content of DNA. Methods Enzymol 12B:195–206
McKerrow J, Vagg S, McKinney T, Seviour EM, Maszenan AM, Brooks P, Seviour RJ (2000) A simple HPLC method for analysing diaminopimelic acid diastereomers in cell walls of Gram-positive bacteria. Lett Appl Microbiol 30:178–182
Mincer TJ, Jensen PR, Kauffman CA, Fenical W (2002) Widespread and persistent populations of a major actinomycete taxon in ocean sediments. Appl Environ Microbiol 68:5005–5011
Minnikin DE, Hutchinson IG, Caldicott AB, Goodfellow M (1980) Thin-layer chromatography of methanolysates of mycolic acid-containing bacteria. J Chromatogr 188:221–233
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JK (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Nimaichand S, Zhang YG, Cheng J, Li L, Zhang DF, Zhou EM, Dong L, Ningthoujam DS, Li WJ (2013) Micromonospora kangleipakensis sp. nov., isolated from a sample of Limestone quarry. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.052746-0
Ørskov J (1923) Investigations into the morphology of the ray fungi. Levin and Munksgaard, Enhagen
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington DC, pp 607–654
Songsumanus A, Tanasupawat S, Igarashi Y, Kudo T (2013) Micromonospora maritima sp. nov., isolated from mangrove soil. Int J Syst Evol Microbiol 63:554–559
Supong K, Suriyachadkun C, Pittayakhajonwut P, Suwanborirux K, Thawai C (2013a) Micromonospora spongicola sp. nov., an actinomycete isolated from a marine sponge in the Gulf of Thailand. J Antibiot (Tokyo). doi:10.1038/ja.2013.35
Supong K, Suriyachadkun C, Tanasupawat S, Suwanborirux K, Pittayakhajonwut P, Kudo T, Thawai C (2013b) Micromonospora sediminicola sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 63:570–575
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Trujillo ME, Kroppenstedt RM, Fernández-Molinero C, Schumann P, Martínez-Molina E (2007) Micromonospora lupini sp. nov. and Micromonospora saelicesensis sp. nov., isolated from root nodules of Lupinus angustifolius. Int J Syst Evol Microbiol 57:2799–2804
Trujillo ME, Alonso-Vega P, Rodríguez R, Carro L, Cerda E, Alonso P, Martínez-Molina E (2010) The genus Micromonospora is widespread in legume root nodules: the example of Lupinus angustifolius. ISME J 4:1265–1281
Uchida K, Kudo T, Suzuki K, Nakase T (1999) A new rapid method of glycolate test by diethyl ether extraction, which is applicable to a small amount of bacterial cells of less than one milligram. J Gen Appl Microbiol 45:49–56
Waksman SA (1961) The Actinomycetes, vol. 2, Classification, identification and descriptions of genera and species. Williams and Wilkins, Baltimore
Waksman SA (1967) The Actinomycetes. A summary of current knowledge, Ronald
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE et al (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Wu C, Lu X, Qin M, Wang Y, Ruan J (1989) Analysis of menaquinone compound in microbial cells by HPLC. Microbiology [English translation of Microbiology (Beijing)] 16:176–178
Xiang WS, Liu CX, Wang XJ, Du J, Xi LJ, Huang Y (2011) Actinoalloteichus nanshanensis sp. nov., isolated from the rhizosphere of a fig tree (Ficus religiosa). Int J Syst Evol Microbiol 61:1165–1169
Xie QY, Lin HP, Li L, Brown R, Goodfellow M, Deng ZX, Hong K (2012) Verrucosispora wenchangensis sp. nov., isolated from mangrove soil. Antonie Leeuwenhoek Int J Gen 102:1–7
Yokota A, Tamura T, Hasegawa T, Huang LH (1993) Catenuloplanes japonicas gen. nov., sp. nov., nom. rev., a new genus of the order Actinomycetales. Int J Syst Bacteriol 43:805–812
Acknowledgments
This work was supported in part by grants from the Opening Fund of Key Laboratory of Soybean Biology in Chinese Ministry of Education (No. SB12B04), the National Key Project for Basic Research (No. 2010CB126102), the National Outstanding Youth Foundation (No. 31225024), the Special Foundation for Scientific and Technological Innovation Research of Harbin (No. 2011RFXXN038) and the Natural Science Foundation of Heilongjiang Province (No. C201029).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Ruixia Gao and Chongxi Liu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gao, R., Liu, C., Zhao, J. et al. Micromonospora jinlongensis sp. nov., isolated from muddy soil in China and emended description of the genus Micromonospora . Antonie van Leeuwenhoek 105, 307–315 (2014). https://doi.org/10.1007/s10482-013-0074-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-013-0074-3