Abstract
Acinetobacter baumannii harbours a gene cluster similar to the iac locus of Pseudomonas putida 1290, which can catabolize the plant hormone indole 3-acetic acid (IAA) as an energy source. However, there has been no evidence showing that IAA can be utilized by A. baumannii. This study showed that A. baumannii can grow in M9 minimal medium containing IAA as the sole carbon source. A mutagenesis study indicated that iacA, encoded in the iac locus of A. baumannii, is involved in the catabolism of IAA. As shown by western blotting analysis, the IacA protein was detected in A. baumannii grown in M9 minimal medium with IAA but not with pyruvate, suggesting that the expression of iacA is regulated by the presence of IAA. In vitro studies have shown that IacA can oxidize indole, an IAA-like molecule, converting it to indoxyl, which spontaneously dimerises to form indigo. In this study, we show that the crude extracts from either wild-type A. baumannii or Escherichia coli overexpressing IacA can oxidize IAA. These results imply that the iac gene cluster of A. baumannii is involved in IAA degradation and that the iacA gene is upregulated when cells encounter IAA in their native environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Indigo is an economically important dye in the textile industry. Historically, plants from the genus Indigofera were the original and sole source of the dye but as early as 1928, microorganisms were also reported to produce the pigment (Gray 1928). Biosynthesis of indigo from indole by Escherichia coli carrying an exogenous naphthalene dioxygenase was first described in 1983 (Ensley et al. 1983) and a number of other genes mediating indigo production in E. coli have also been investigated (Ensley et al. 1983; Keil et al. 1987; Li et al. 2000; Drewlo et al. 2001; Nakamura et al. 2001; Choi et al. 2003; Doukyu et al. 2003; Furuya et al. 2004; Lim et al. 2005; van Hellemond et al. 2007; Tischler et al. 2009). The biosynthetic pathways of indigo synthesis in bacteria have been well characterized. In most cases, indole is first regioselectively hydroxylated by an oxygenase to an intermediate, indoxyl, which spontaneously dimerises to form indigo (McClay et al. 2005; Royo et al. 2005; Kwon et al. 2008). Today, most indigo dye is produced synthetically, and its production generates a significant amount of toxic waste. Therefore, biological fermentation by recombinant microorganisms may provide a cleaner alternative for large-scale indigo production (Murdock et al. 1993). In addition to indigo, different kinds of indigoid pigments, including indirubin can also be produced by E. coli expressing indole oxygenases (O’Connor et al. 1997; O’Connor and Hartmans 1998; Nakamura et al. 2001; Meyer et al. 2002; Choi et al. 2003; Furuya et al. 2004; Lim et al. 2005; McClay et al. 2005; Rui et al. 2005). Moreover, human cytochrome P450 and bacterial cytochrome P450 proteins have been engineered to produce a variety of indigoids in E. coli (Gillam et al. 1999; Li et al. 2000; Nakamura et al. 2001; Celik et al. 2005; Wu et al. 2005; Rosic et al. 2007; Rosic 2009).
Indigoids also have the potential to be used for anticancer therapy, but also for Alzheimer’s disease and diabetes (Kunikata et al. 2000; Leclerc et al. 2001; Eisenbrand et al. 2004). Cyclin-dependent kinases (CDKs) are required in the cell cycle control of eukaryotic cells (Morgan 1997). Cyclin-CDK complexes use adenosine triphosphate (ATP) as a phosphate donor for substrate (van den Heuvel and Harlow 1993). Indirubin and its derivates have already been shown to bind the ATP binding site of CDKs and inhibit their activity, resulting in cell cycle arrest and apoptosis in several tumor cell lines Hoessel et al. (1999)Marko et al. 2001; Eisenbrand et al. 2004).
As a consequence of the above, bacterial indigo metabolism is of much interest. While constructing a fosmid library of Acinetobacter baumannii ATCC19606, a blue colony was identified on an LB agar plate. After this pigment was identified as indigo, the iacA gene responsible for its biosynthesis in E. coli was identified by random transposon mutagenesis in this study. In addition, IacA was overexpressed and purified by using an indole synthesis-deficient E. coli as an expression host, which allowed us to analyze the kinetic activity in vitro. To our knowledge, this is the first report to show that A. baumannii ATCC19606 can grow in minimal medium with IAA as the sole carbon source. We also demonstrated that the iac gene cluster is involved in the degradation of IAA and that the IacA protein was strongly expressed in the presence of IAA.
Materials and methods
Materials
NADH, FAD, indole 3-acetic acid (IAA), indole and sodium pyruvate were obtained from Sigma (Sigma–Aldrich, St. Louis, MO, USA). Q-Sepharose High Performance anion-exchange gel resin was purchased from GE Healthcare (Uppsala, Sweden). Restriction endonucleases, BamHI and PstI, and Ex Taq DNA polymerase were purchased from TaKaRa (Otsu, Japan). All chemicals used were of molecular biology grade or higher.
Bacteria and bacterial growth conditions
The E. coli strain CY15000 (Yanofsky and Horn 1981), a tryptophanase-deficient mutant, and A. baumannii ATCC 19606 were obtained from the E. coli Genetic Stock Center and the American Type Culture Collection, respectively. E. coli and A. baumannii were routinely grown using LB broth or agar (Miller 1972) at 37°C. Ampicillin, chloramphenicol and kanamycin were used at 100, 12.5, and 25 μg/mL, respectively. M9 minimal medium (Amresco, Solon, OH) supplied with 5 mM IAA or sodium pyruvate was also used as a culture medium for A. baumannii ATCC 19606. A. baumannii ATCC 19606 was grown in LB overnight. The overnight cultures (0.5 mL) were collected by centrifugation and washed once with M9 medium without a carbon source. The washed bacteria were transferred to M9 medium (50 mL) supplemented with 5 mM of IAA or sodium pyruvate and grown at 37°C with agitation. Growth was monitored by measuring the optical density at 600 nm (OD600). The logarithmic phase bacteria were harvested by centrifugation and the bacterial pellets were then used for western blot analysis.
Fosmid library construction of A. baumannii ATCC19606 and identification of the gene mediating blue pigment production in E. coli
The genomic DNA purification and fosmid library construction of A. baumannii ATCC19606 were performed as previously described (Shu et al. 2009; Wu et al. 2009). Chromosomal DNA of A. baumannii ATCC19606 was prepared using the Wizard Genomic DNA Purification System (Promega) according to the manufacturer’s instructions. The fosmid library of A. baumannii ATCC19606 was constructed with a CopyControl Fosmid Library Production Kit (Epicentre). Briefly, A. baumannii ATCC19606 genomic DNA was sheared into ca. 40 kb fragments by pipetting. The sheared DNA was end-repaired with supplied End-Repair Enzyme Mix to generate blunt and 5′-phosphorylated ends. Next, end-repaired DNA of approximate size 40 kb was isolated and purified from low-melting-point agarose gels. The purified DNA was ligated to a linearized and dephosphorylated pCC1FOS vector. The successfully ligated DNA was subjected to Lambda packaging using MaxPlax Lambda Packaging. Finally, packaged DNA was used to infect E. coli EPI300 and clones were selected on chloramphenicol LB plates. A fosmid clone, pDEH, which turned blue on an LB agar plate at 37°C was identified and the end-sequences of the fosmid DNA were determined with the T7 promoter primer (5′-TAATACGACTCACTATAGGG-3′) and the reverse sequencing primer (5′-CAGGAAACAGCCTAGGAA-3′).
To identify the gene involved in blue pigment production of pDEH, transposon mutagenesis with the EZ-Tn5 <KAN-2> insertion kit (Epicentre, Madison, WI, USA) was performed by following the manufacturer’s protocol. After in vitro transposition, fosmids were transformed into E. coli EPI300 (Epicentre) by electroporation and selected on LB agar plates supplemented with chloramphenicol and kanamycin. Clones that could not produce blue pigment were isolated and the EZ-Tn5 <KAN-2> transposon insertion sites were examined by sequencing with the KAN-2 FP-1 primer (5′-ACCTACAACAAAGCTCTCATCAACC-3′).
Sequence analysis
MacVector (Olson 1994) was used for sequence analysis and homologous proteins were identified using the BLAST program (http://www.ncbi.nlm.nih.gov/blast/). The protein family was predicted by searching the Pfam protein database (Bateman et al. 2004; Finn et al. 2006).
Construction of an expression plasmid
To facilitate amplification by PCR, a pair of primers, 5′-ATCCGGATCC ATGAATAAGTTGTCTAAAATGGAG-3′ (the underlined and the boldface letters indicate the BamHI site and start codon of iacA gene, respectively) and 5′-CCTACTGCAGGCAAAACAACACGCGTAATG-3′ (the underlined letters indicate the PstI site), were designed using the nucleotide sequence of iacA. Oligonucleotides were designed to amplify iacA flanked by BamHI and PstI sites. The coding region of iacA was then amplified by PCR using A. baumannii ATCC19606 genomic DNA as a template. The amplification cycling conditions were as follows: 30 cycles of denaturation at 95°C for 0.5 min, annealing at 45°C for 1 min, and extension at 72°C for 2 min, followed by a 5 min extension at 72°C. The PCR products were gel-purified, digested with BamHI and PstI, and ligated with BamHI/PstI-digested pQE-80L vector (Qiagen, Hilden, Germany). The ligation mixture was used to transform E. coli DH5α. The plasmid that carried the iacA gene was designated pIacA.
Protein expression and purification
To avoid indigo production from indole, pIacA was introduced into E. coli CY15000 for IacA overexpression. E. coli CY15000(pIacA) were grown at 37°C by inoculating a 5-mL overnight culture into 5 L of LB medium containing ampicillin. Incubation was maintained at 30°C until the culture reached an OD600 of 0.6, at which point the temperature was lowered to 15°C. Once the medium was allowed to cool for 15 min, expression was induced by the addition of 0.2% lactose and 50 μg/mL isopropyl-β-d-thiogalactopyranoside (IPTG) (Weng et al. 2006). Following overnight incubation, cells were harvested by centrifugation at 4,000×g for 10 min. Cells were then stored at −20°C.
All purification steps were performed on ice or at 4°C. Frozen cells were resuspended in 30 mL of 50 mM potassium phosphate buffer (pH 7.0) containing 5 mM EDTA and then sonicated (10 min, 30 s on, 30 s off) with a Vibra-Cell (Sonics & Materials, Inc. Newtown, CT). Cell debris was removed by centrifugation at 25,000×g for 15 min. The resulting supernatant solution was designated as crude extract. The crude extract was directly applied to a Q-Sepharose High Performance anion-exchange column (2.6 × 70 cm) equilibrated in 10 mM potassium phosphate buffer of pH 7.0. The eluents used were as follows: eluent A (10 mM potassium phosphate buffer, pH 7.0) and eluent B (10 mM potassium phosphate buffer (pH 7.0) containing 0.5 M KCl). The following profile was used for separation: 80-min isocratic development with 100% A; 520 mL linear gradient from 100% A to 80% B; 40 mL linear gradient from 80% B to 100% B; 120 min isocratic development with 100% B. The flow rate was 2 mL/min, and 8 mL fractions were collected. Fractions containing recombinant oxygenase on examination by SDS-PAGE were pooled and concentrated to 5 mL by ultrafiltration in a stirred cell fitted with a YM-3 membrane. Small aliquots of the protein solution were frozen in liquid nitrogen and stored at −80°C.
Enzymatic assays
In a typical reaction, 5 μM purified IacA was used in a 100 μL solution containing 100 mM Tris buffer (pH 8.5). The K m and k cat of indole were determined in the presence of 20 μM FAD; 2 mM NADH; and 4, 2, 1, 0.5, 0.2, and 0.1 mM substrate. The K m and k cat of FAD were determined in the presence of 5 mM indole; 2 mM NADH; and 80, 40, 20, 10, 5, and 2.5 μM FAD. The K m and k cat of NADH were determined in the presence of 5 mM indole, 20 μM FAD, and 2, 1, 0.5, 0.25 and 0.1 mM NADH. Reactions were initiated by the addition of indole, incubated at room temperature for 60 min, and subsequently stopped by the addition of 500 μL dimethylformamide, which denatured the proteins and increased the solubility of indigo. Oxygenase activity, as measured by the production of indigo, was monitored at OD610 (Seixas de Melo et al. 2004).
Preparation of IacA antibody and western blot analysis
The purified IacA protein was used to generate antibody in rabbit. Four booster injections were given at 2 weeks intervals. The serum was collected and stored at −20°C until use. Bacteria cultured in M9 medium with 5 mM of IAA or sodium pyruvate were harvested by centrifugation and the cells were resuspended in phosphate buffered saline. The bacterial suspension was sonicated at 4°C and protein concentration was determined using a Qubit fluorometer (Invitrogen; Carlsbad, CA) with a Quant-it Protein assay kit (Invitrogen). The sonicated cell lysates of A. baumannii ATCC 19606 were mixed with sample loading buffer and boiled at 100°C for 10 min. Cell extracts of A. baumannii ATCC 19606 (5 μg total protein) were separated by a 10% SDS-PAGE (Laemmli 1970) with Mini-Vertical Units SE 250 (GE Healthcare, Piscataway, NJ) and electroblotted onto nitrocellulose membrane (Immobilon-P; Millipore, Bedford, MA) with a Mini Trans-Blot cell (Bio-Rad, Richmond, CA) according to the methods suggested by the manufacturers. Endogenous IacA protein was detected with IacA polyclonal antibody and with alkaline phosphatase-conjugated anti-rabbit immunoglobulin (Novagen, Madison, WI). The nitrocellulose paper was blocked in TBS buffer (50 mM Tris–HCl [pH 7.6], 0.9% NaCl) containing 5% skimmed milk (BD Diagnostic Systems, Franklin Lakes, NJ) for 30 min. The membrane was first incubated with IacA antibody diluted 1:1,000 in TBS containing 0.5% Tween 20 and 5% skimmed milk for 1 h at room temperature. After six washes with TBS containing 0.5% Tween 20, the membrane was incubated in a 1:10,000 dilution of goat anti-rabbit IgG alkaline phosphatase conjugated (Novagen) for 1 h. After 6 washes with TBS containing 0.5% Tween 20 and an additional 2 washes with TBS, the membrane was developed with BCIT/NBT substrate solution (PerkinElmer, Boston, MA).
Construction of an iacA mutant
An iacA mutant was generated by random transposon mutagenesis with the EZ-Tn5 <KAN-2> Tnp Transposome Kit (Epicenter) according to previously described methods (Dorsey et al. 2002; Loehfelm et al. 2008). EZ-Tn5 KAN-2 Transposome was electroporated into A. baumannii and independent insertion mutants were selected from LB plates containing kanamycin. Transposon insertion mutants were screened for the loss of the ability to grow on M9 plates with 5 mM IAA as carbon source. A transposon inserted mutant of iacA gene was subsequently identified from the mutants that could not degrade IAA by PCR using iacA specific primers (5′-TTGAAACCAGTGAAGGCTTG-3′, located 125 bp upstream of the iacA gene start condon) and (5′-GCAAAACAACACGCGTAATG-3′, located 64 bp downstream of the iacA gene stop codon). The transposon insertion site of iacA mutant was determined by sequencing with the KAN-2 FP-1 primer.
IAA degradation
Overnight cultures of wild-type and iacA mutant A. baumannii ATCC 19606 were diluted 100-fold into 500 mL flasks containing 150 mL LB and grown at 37°C with agitation. After 6 h of incubation, the cells were collected by centrifugation and resuspended in 5 mL of buffer (100 mM Tris–Cl buffer, pH 8.5). Cells of E. coli CY15000(pIacA) and E. coli CY15000(pQE-80L) were grown using methods described in the section on protein expression and purification. Bacteria from 150 mL of each culture were resuspended in 15 mL of buffer (100 mM Tris–Cl buffer, pH 8.5). Crude extracts were prepared by sonication and cell debris was removed by centrifugation. The supernatant was then filtered through 0.45 μm Millipore filters and the filtered supernatant was used for IAA oxidation.
The reaction mixtures included 70 μL of the supernatant, 20 μM FAD, 2 mM NADH and 5 mM IAA in a total volume of 100 μL. The mixtures were incubated for 18 h at 25°C. Thin-Layer Chromatography (TLC) analysis was used to examine IAA and the IAA derivatives. One microliter of reaction mixture was spotted on a Silica Gel 60 F254 plate (Merck, Darmstadt, Germany) and developed with a solvent of chloroform:methanol:acetic acid (50:5:2, vol/vol). Following chromatography, the TLC plate was dried, and IAA and IAA derivates were visualized with UV light.
Results
Identification of a gene, iacA, involved in indigo production
A fosmid clone, pDEH, which mediated blue pigment production, was selected from an A. baumannii ATCC19606 fosmid library expressed in E. coli. For the identification of the genes involved in the biosynthesis of the blue dye, the EZ-Tn5 <KAN-2> transposon was used to generate mutants. Four transposon insertion clones lacking blue pigment production were isolated on LB plates. The insertion sites of EZ-Tn5 <KAN-2> were determined by sequencing using transposon specific outward-reading primers. Analysis of the transposon insertion sites indicated that one gene of pDEH, here designated iacA, was responsible for producing the blue pigment. Transposons were inserted between nucleotides 97 and 98, 562 and 563, 938 and 939, and 1,074–1,075 downstream of the ATG start site of iacA from these independent mutants.
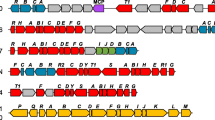
The iacA gene was found to encode a predicted protein of 389 amino acid residues with molecular mass of 42334.5 daltons which was annotated as an acyl-CoA dehydrogenase-like protein in the genomes of A. baumannii ATCC 17978 (accession number ABO12291) and strain ATCC 19606 (accession number EEX04702) in GenBank. Sequence analysis revealed that the IacA protein was 58% identical to its homologue (accession number ABY62757) from Pseudomonas putida 1290. The iacA gene was located in the iac locus of A. baumannii (Leveau and Gerards 2008). A previous study has indicated that the iac locus of P. putida 1290 is required to metabolize IAA and that similar iac gene clusters were found in 22 bacterial species (Leveau and Gerards 2008). Fosmid end sequences of pDEH were sequenced and mapped to the unfinished genome sequence of A. baumannii ATCC19606 available in GenBank. Additionally, pDEH harbored a 42,103-bp genomic DNA fragment that contained partial iacH and intact iacABICDEF genes of iac gene locus of A. baumannii ATCC 19606.
The IacA protein converts indole to indoxyl with pH optimum of 8.5
The IacA protein was purified by loading the clarified protein solution onto an anion-exchange matrix and eluting with a linear, ascending gradient of potassium chloride, resulting in a protein that was very nearly homogeneous (Fig. 1). The yields and specific activities are summarized in Table 1. The activity of IacA significantly decreased when the pH was below 7.5. A sharp pH optimum was observed in the alkaline range when the pH of the assay solution was varied between 7.5 and 10.0. The maximal rate was achieved at about pH 8.5 (Fig. 2). All further kinetic and substrate specificity experiments were therefore conducted at pH 8.5. The apparent K m values for indole, FAD and NADH were 0.80 ± 0.04 mM, 3.4 ± 0.3 μM, and 0.25 ± 0.06 mM, respectively. The k cat was 0.88 ± 0.02 min−1. Due to the low activity of purified IacA, the kinetic parameters for IAA could not be measured in our assay under the conditions tested.
pH dependence of IacA activity. Assays were performed in 100 mM potassium phosphate buffer at pH 7.0 and 7.5, 100 mM Tris–Cl buffer between pH 8.0 and 10.0, with 5 mM indole, 20 μM FAD, 2 mM NADH, and 5 μM IacA in a total volume of 100 μL. The reaction mixtures were incubated at room temperature for 30 min and quenched by addition of 500 μL DMF. In this comparison, the activity measured at pH 8.5 has been normalized to 100%
IacA expression in A. baumannii
Acinetobacter baumannii ATCC19606 was cultured in minimal M9 media containing 5 mM IAA or sodium pyruvate and subsequent bacterial growth was monitored by measuring OD600. Cells cultured in M9 with IAA and pyruvate reached stationary phase at OD600 values of 0.3 and 0.28, respectively.
The logarithmic growth phase of the bacteria was determined and protein extracts from this phase were analyzed by western blotting. IacA was observed to be expressed only in bacteria cultured in the M9 media with IAA but not with pyruvate (Fig. 3).
Analysis of an iacA mutant
An iacA mutant was identified from an A. baumannii ATCC19606 EZ-Tn5 <KAN-2> transposon library. Approximately 2,500 clones were screened for loss of the ability to grow on an M9 plate with 5 mM IAA and 23 mutants were isolated. One mutant, AB6-2, was then confirmed as an iacA mutant by PCR with iacA specific primers. PCR products amplified from wild type genomic DNA using these primers contain an intact iacA gene (ca. 1,200 bp) whereas the amplicon obtained from the iacA mutant was larger than that obtained from wild-type A. baumannii. The amplicon of AB6-2 was sequenced and the transposon insertion site was identified by aligning the sequences of iacA and the amplicon from AB6-2. EZ-Tn5 <KAN-2> was inserted between nucleotides 970 and 971 of the iacA coding sequence of AB6-2.
Involvement of the iac gene cluster in IAA degradation
Crude cells extracts prepared from E. coli CY15000(pIacA), E. coli CY15000(pQE-80L), wild-type A. baumannii and iacA mutant AB6-2 were used for IAA oxidation reactions. Following incubation for 18 h, the products were separated by TLC and visualized with UV light. IAA was detected on the TLC plate with an R f of 0.67 (Fig. 4). Spots that migrated with R f values of 0.48 and 0.17 were observed from the reaction catalyzed by crude extract of E. coli CY15000(pIacA) and wild-type A. baumannii, respectively. In contrast, the cell crude extracts prepared from E. coli CY15000(pQE-80L) and iacA mutant AB6-2 showed no detectable activity.
Discussion
In this study, we characterized the IacA protein and examined iacA gene expression in A. baumannii ATCC19606. Comparative genomic analysis indicated that the iacA gene was present in the genomes of sequenced strains of A. baumannii. Both IacA of P. putida and A. baumannii contained the N- and C- terminal domains of Acyl-CoA dehydrogenase based on homology searches in the Pfam database. A previous study demonstrated that the two-component styrene monooxygenase (SMO) of P. putida strains S12 and CA-3 can convert indole to indigo (O’Connor et al. 1997). In addition, E. coli DH5α cultured in tryptophan medium expressing a flavoprotein monooxygenase (FMO) of Methylophaga sp. strain SK1 also produced indigo and indirubin (Choi et al. 2003). Unlike these two monooxygenases, IacA and other indigo-producing enzymes belong to the acyl-CoA dehydrogenase family (Table 2). Enzymes encoded by the corresponding genes listed in Table 2 contain Pfam domains Acyl-CoA_dh_N (PF02771) and Acyl-CoA_dh_2 (PF08028), except Bec of Ralstonia eutropha HF39 which contains Acyl-CoA_dh_N (PF02771) and Acyl-CoA_dh_M (PF02770) domains. Like SMO and FMO, acyl-CoA dehydrogenases are flavoproteins (Kim and Miura 2004). However, acyl-CoA dehydrogenases do not contain a conserved FAD-binding sequence (Dym and Eisenberg 2001).
When the kinetic properties of the enzyme were investigated by assaying the conversion of indole into indigo, the reaction was found to be absolutely dependent upon IacA, NADH, FAD, and indole being present in the assay. No other cofactors were required. The optimum activity of IacA was observed in the alkaline range and the maximal rate was achieved at pH 8.5. This result is consistent with a report of IdoA protein from P. alcaligenes PA-10 (Alemayehu et al. 2004). Both IacA and IdoA are acyl-CoA dehydrogenase-like proteins (Alemayehu et al. 2004). In contrast to wild-type P450 2A6, the k cat of IacA was lower than that of P450 2A6 (Zhang et al. 2009). The K m for indole of IacA was approximately three times higher than the indole K m of P450 2A6 (Zhang et al. 2009).
TLC showed that an IAA derivative (R f 0.48) was produced by the cell crude extract of E. coli CY15000(pIacA). Our results revealed that IacA is responsible for initial attack upon IAA, which is consistent with a previous hypothesis (Leveau and Gerards 2008). The product (R f 0.17) generated by the crude extract of A. baumannii was more hydrophilic than that by the cell crude extract of E. coli CY15000(pIacA) (R f 0.48). Taken together, these data suggest that the initial oxidation of IAA could be catalyzed by IacA, whereas the further degradation of IAA was dependent on enzymes encoded by the iac gene cluster. A previous study also demonstrated that catechol, an intermediate of the IAA degradation pathway, is further metabolized by the enzymes of the β-ketoadipate pathway (Leveau and Gerards 2008). Although several proteins encoded by the iac gene cluster are similar to enzymes for indole degradation, the exact pathway of IAA oxidation is still unknown (Leveau and Gerards 2008). Future research will be needed to unravel the biological functions of iacA and the iac locus.
Western blotting analysis was carried out to examine the expression of IacA protein in A. baumannii ATCC19606 cultured in M9 medium. Our results show that IacA was expressed in the presence of IAA but not pyruvate. These results suggest that the presence of IAA regulates the expression of the iac locus. The amino acid sequences of the proteins of each pair encoded by the iac loci from A. baumannii and P. putida 1290 share over 50% sequence identity, except IacR, with only 32% sequence identity. Previous work has suggested that the iacR gene product is similar to a transcriptional regulator (Leveau and Gerards 2008). Further, the iac loci of A. baumannii ATCC19606 and P. putida 1290 differ in their gene arrangement. Thus, the enzymes involved in IAA catabolism might be similar in both A. baumannii ATCC19606 and P. putida 1290, yet the regulatory pathways may be different.
Our results revealed that: A. baumannii could utilize IAA as a carbon source; the iac locus in A. baumannii was involved in IAA degradation; and the expression of iacA gene was induced by the presence of IAA. These results suggest that IAA was not only an energy source but also regulated gene expression in A. baumannii.
References
Alemayehu D, Gordon LM, O’Mahony MM, O’Leary ND, Dobson AD (2004) Cloning and functional analysis by gene disruption of a novel gene involved in indigo production and fluoranthene metabolism in Pseudomonas alcaligenes PA-10. FEMS Microbiol Lett 239:285–293
Bateman A et al (2004) The Pfam protein families database. Nucleic Acids Res 32:D138–D141
Celik A, Speight RE, Turner NJ (2005) Identification of broad specificity P450CAM variants by primary screening against indole as substrate. Chem Commun (Camb) 29:3652–3654
Choi HS, Kim JK, Cho EH, Kim YC, Kim JI, Kim SW (2003) A novel flavin-containing monooxygenase from Methylophaga sp strain SK1 and its indigo synthesis in Escherichia coli. Biochem Biophys Res Commun 306:930–936
Choi KY, Kim D, Koh SC, So JS, Kim JS, Kim E (2004) Molecular cloning and identification of a novel oxygenase gene specifically induced during the growth of Rhodococcus sp. strain T104 on limonene. J Microbiol 42:160–162
Dorsey CW, Tomaras AP, Actis LA (2002) Genetic and phenotypic analysis of Acinetobacter baumannii insertion derivatives generated with a transposome system. Appl Environ Microbiol 68:6353–6360
Doukyu N, Toyoda K, Aono R (2003) Indigo production by Escherichia coli carrying the phenol hydroxylase gene from Acinetobacter sp. strain ST-550 in a water-organic solvent two-phase system. Appl Microbiol Biotechnol 60:720–725
Drewlo S, Bramer CO, Madkour M, Mayer F, Steinbuchel A (2001) Cloning and expression of a Ralstonia eutropha HF39 gene mediating indigo formation in Escherichia coli. Appl Environ Microbiol 67:1964–1969
Dym O, Eisenberg D (2001) Sequence-structure analysis of FAD-containing proteins. Protein Sci 10:1712–1728
Eisenbrand G, Hippe F, Jakobs S, Muehlbeyer S (2004) Molecular mechanisms of indirubin and its derivatives: novel anticancer molecules with their origin in traditional Chinese phytomedicine. J Cancer Res Clin Oncol 130:627–635
Ensley BD, Ratzkin BJ, Osslund TD, Simon MJ, Wackett LP, Gibson DT (1983) Expression of naphthalene oxidation genes in Escherichia coli results in the biosynthesis of indigo. Science 222:167–169
Finn RD et al (2006) Pfam: clans, web tools and services. Nucleic Acids Res 34:D247–D251
Furuya T, Takahashi S, Ishii Y, Kino K, Kirimura K (2004) Cloning of a gene encoding flavin reductase coupling with dibenzothiophene monooxygenase through coexpression screening using indigo production as selective indication. Biochem Biophys Res Commun 313:570–575
Gillam EM et al (1999) Formation of indigo by recombinant mammalian cytochrome P450. Biochem Biophys Res Commun 265:469–472
Gray PHH (1928) The formation of indigotin from indol by soil bacteria. Proc R Soc Lond Ser B 102:263–280
Hart S, Kirby R, Woods DR (1990) Structure of a Rhodococcus gene encoding pigment production in Escherichia coli. J Gen Microbiol 136:1357–1363
Hoessel R et al (1999) Indirubin, the active constituent of a Chinese antileukaemia medicine, inhibits cyclin-dependent kinases. Nat Cell Biol 1:60–67
Keil H, Saint CM, Williams PA (1987) Gene organization of the first catabolic operon of TOL plasmid pWW53: production of indigo by the xylA gene product. J Bacteriol 169:764–770
Kim JJ, Miura R (2004) Acyl-CoA dehydrogenases and acyl-CoA oxidases. Structural basis for mechanistic similarities and differences. Eur J Biochem 271:483–493
Kunikata T, Tatefuji T, Aga H, Iwaki K, Ikeda M, Kurimoto M (2000) Indirubin inhibits inflammatory reactions in delayed-type hypersensitivity. Eur J Pharmacol 410:93–100
Kwon NR et al (2008) Identification of functionally important amino acids in a novel indigo-producing oxygenase from Rhodococcus sp. strain T104. Appl Microbiol Biotechnol 79:417–422
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Leclerc S et al (2001) Indirubins inhibit glycogen synthase kinase-3 beta and CDK5/p25, two protein kinases involved in abnormal tau phosphorylation in Alzheimer’s disease. A property common to most cyclin-dependent kinase inhibitors? J Biol Chem 276:251–260
Leveau JH, Gerards S (2008) Discovery of a bacterial gene cluster for catabolism of the plant hormone indole 3-acetic acid. FEMS Microbiol Ecol 65:238–250
Li QS, Schwaneberg U, Fischer P, Schmid RD (2000) Directed evolution of the fatty-acid hydroxylase P450 BM-3 into an indole-hydroxylating catalyst. Chemistry 6:1531–1536
Lim HK et al (2005) Characterization of a forest soil metagenome clone that confers indirubin and indigo production on Escherichia coli. Appl Environ Microbiol 71:7768–7777
Loehfelm TW, Luke NR, Campagnari AA (2008) Identification and characterization of an Acinetobacter baumannii biofilm-associated protein. J Bacteriol 190:1036–1044
Marko D et al (2001) Inhibition of cyclin-dependent kinase 1 (CDK1) by indirubin derivatives in human tumour cells. Br J Cancer 84:283–289
McClay K, Boss C, Keresztes I, Steffan RJ (2005) Mutations of toluene-4-monooxygenase that alter regiospecificity of indole oxidation and lead to production of novel indigoid pigments. Appl Environ Microbiol 71:5476–5483
Meyer A, Wursten M, Schmid A, Kohler HP, Witholt B (2002) Hydroxylation of indole by laboratory-evolved 2-hydroxybiphenyl 3-monooxygenase. J Biol Chem 277:34161–34167
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor
Morgan DO (1997) Cyclin-dependent kinases: engines, clocks, and microprocessors. Annu Rev Cell Dev Biol 13:261–291
Murdock D, Ensley BD, Serdar C, Thalen M (1993) Construction of metabolic operons catalyzing the de novo biosynthesis of indigo in Escherichia coli. Biotechnology (NY) 11:381–386
Nakamura K, Martin MV, Guengerich FP (2001) Random mutagenesis of human cytochrome p450 2A6 and screening with indole oxidation products. Arch Biochem Biophys 395:25–31
O’Connor KE, Hartmans S (1998) Indigo formation by aromatic hydrocarbon-degrading bacteria. Biotechnol Lett 20:219–223
O’Connor KE, Dobson AD, Hartmans S (1997) Indigo formation by microorganisms expressing styrene monooxygenase activity. Appl Environ Microbiol 63:4287–4291
Olson SA (1994) MacVector: an integrated sequence analysis program for the Macintosh. Methods Mol Biol 25:195–201
Rosic NN (2009) Versatile capacity of shuffled cytochrome P450s for dye production. Appl Microbiol Biotechnol 82:203–210
Rosic NN, Huang W, Johnston WA, DeVoss JJ, Gillam EM (2007) Extending the diversity of cytochrome P450 enzymes by DNA family shuffling. Gene 395:40–48
Royo JL, Moreno-Ruiz E, Cebolla A, Santero E (2005) Stable long-term indigo production by overexpression of dioxygenase genes using a chromosomal integrated cascade expression circuit. J Biotechnol 116:113–124
Rui L, Reardon KF, Wood TK (2005) Protein engineering of toluene ortho-monooxygenase of Burkholderia cepacia G4 for regiospecific hydroxylation of indole to form various indigoid compounds. Appl Microbiol Biotechnol 66:422–429
Seixas de Melo J, Moura AP, Melo MJ (2004) Photophysical and spectroscopic studies of indigo derivatives in their keto and leuco forms. J Phys Chem A 108:6975–6981
Shu HY et al (2009) Genetic diversity of capsular polysaccharide biosynthesis in Klebsiella pneumoniae clinical isolates. Microbiology 155:4170–4183
Tischler D, Eulberg D, Lakner S, Kaschabek SR, van Berkel WJ, Schlomann M (2009) Identification of a novel self-sufficient styrene monooxygenase from Rhodococcus opacus 1CP. J Bacteriol 191:4996–5009
van den Heuvel S, Harlow E (1993) Distinct roles for cyclin-dependent kinases in cell cycle control. Science 262:2050–2054
van Hellemond EW, Janssen DB, Fraaije MW (2007) Discovery of a novel styrene monooxygenase originating from the metagenome. Appl Environ Microbiol 73:5832–5839
Weng YP, Hsu FC, Yang WS, Chen HP (2006) Optimization of the overexpression of glutamate mutase S component under the control of T7 system by using lactose and IPTG as the inducers. Enzyme Microb Technol 38:465–469
Wu ZL, Aryal P, Lozach O, Meijer L, Guengerich FP (2005) Biosynthesis of new indigoid inhibitors of protein kinases using recombinant cytochrome P450 2A6. Chem Biodivers 2:51–65
Wu KM et al (2009) Genome sequencing and comparative analysis of Klebsiella pneumoniae NTUH-K2044, a strain causing liver abscess and meningitis. J Bacteriol 191:4492–4501
Yanofsky C, Horn V (1981) Rifampin resistance mutations that alter the efficiency of transcription termination at the tryptophan operon attenuator. J Bacteriol 145:1334–1341
Zhang ZG, Liu Y, Guengerich FP, Matse JH, Chen J, Wu ZL (2009) Identification of amino acid residues involved in 4-chloroindole 3-hydroxylation by cytochrome P450 2A6 using screening of random libraries. J Biotechnol 139:12–18
Acknowledgments
The authors acknowledge the technical services provided by Sequencing Core Facility of the National Yang-Ming University Genome Research Center (YMGC). The Sequencing Core Facility is supported by National Research Program for Genomic Medicine (NRPGM), National Science Council. This project was supported by National Science Counsel of Taiwan (NSC97-2320-B-309-001 and NSC98-2320-B-309-003-MY3). This study was also supported in part by Tzu Chi University under contract number TCMRC-P-99011.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lin, GH., Chen, HP., Huang, JH. et al. Identification and characterization of an indigo-producing oxygenase involved in indole 3-acetic acid utilization by Acinetobacter baumannii . Antonie van Leeuwenhoek 101, 881–890 (2012). https://doi.org/10.1007/s10482-012-9704-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-012-9704-4