Abstract
We previously found that Wickerhamomyces anomalus (formerly Hansenula anomala, Pichia anomala) was the second most frequently isolated yeast in Belgian artisan bakery sourdoughs and that the yeast dominated laboratory sourdough fermentations. Such findings are of interest in terms of the advantage of W. anomalus over other commonly encountered sourdough yeasts and its potential introduction into the sourdough ecosystem. Here, we provide a brief overview of current knowledge on yeast ecology and diversity in sourdough in the context of the potential natural habitat of W. anomalus. Insight into the population structure of W. anomalus was obtained by comparing internal transcribed spacer rDNA sequences of selected sourdough isolates with publicly available database sequences.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The yeast Wickerhamomyces anomalus (formerly Hansenula anomala, Pichia anomala) (Kurtzman et al. 2008) may be encountered in natural environments and in various fermentations, either in a beneficial or spoilage role. It is a member of the normal or transient microbiota of the human skin and oropharynx (Chakrabarti et al. 2001; Kalenic et al. 2001; Murphy et al. 1986) and it has been reported to cause rare, but mostly clustered, nosocomial infections in patients with predisposing factors (e.g., Sekhon et al. 1992). The yeast is considered of low virulence based on the low number of fatal cases caused by the yeast infection itself and the lack of pronounced antifungal drug resistance (da Matta et al. 2001). W. anomalus lacks some key capacities of primary pathogens, such as the ability to enter a host and the dependency on the host for its own replication and transmission (Falkow 1997). Nevertheless, its ability to grow at 37°C makes it as much an emerging opportunistic pathogen as other yeast species sharing this characteristic, including Saccharomyces cerevisiae (Piarroux et al. 1999).
The presence of W. anomalus in a variety of niches and its wide geographical distribution suggest a generalist character in contrast to endemic microorganisms or microorganisms tightly adapted to a single ecological niche. Its ability to survive adverse conditions and to flourish in stressful environments, even outgrowing well-adapted yeasts and fungi, has been explained largely by its physiological versatility, symbolised by the large number of carbon and nitrogen sources that may be metabolised (Kurtzman 1998). Additionally, the production of killer toxins (e.g., Polonelli et al. 1983), the ability to grow at low water activity, osmotic and pH stresses (Fredlund et al. 2002) as well as in the presence of lactic acid bacteria (LAB) (Vrancken et al. 2010) contribute to its overall competiveness. Consequently, this yeast is attracting increasing interest, although detailed fundamental knowledge concerning its biology is lacking.
In the present paper, we provide a brief overview of the sourdough microbial ecosystem and the factors that determine sourdough microbiota, including the predominance of W. anomalus in Belgian sourdoughs. The isolation of this species from sourdough and other cereal-related sources is discussed, together with an analysis of the population structure of W. anomalus.
The sourdough ecosystem
The use of sourdough is thought to have originated in ancient times, being the first form of leavening cereal dough. Compared to dough raised by pure cultures of the bakers yeast S. cerevisiae, the use of sourdough results in bakery products with a prolonged shelf life and improved sensory qualities. Sourdough develops by spontaneous fermentation in mixtures of cereal flour and water (De Vuyst et al. 2009). A stable microbial community consisting of dominant LAB and yeasts develops during periodic refreshments of the flour–water mixture, whose composition correlates with process factors such as temperature, propagation cycles, and cereal types (Meroth et al. 2003a, b; De Vuyst and Neysens 2005; De Vuyst and Vancanneyt 2007; De Vuyst et al. 2009; Sterr et al. 2009; Zannini et al. 2009; Coda et al. 2010).
The utilisation of maltose as the most abundant carbohydrate is regarded as a key characteristic of microorganisms during sourdough fermentations. LAB have the ability to produce exceptionally high amounts of energy through heterolactic fermentation of carbohydrates, combined with specialised amino acid catabolism, including the arginine deiminase pathway and branched-chain amino acid conversions (Christensen et al. 1999; Fernández and Zúñiga 2006; Gänzle et al. 2007; De Vuyst et al. 2009). Occasionally, the key enzyme maltose phosphorylase liberates unphosphorylated glucose that can subsequently be used by maltose-negative yeasts in the microbial community (De Vuyst et al. 2009). In turn, yeasts contribute essential amino acids for LAB growth (Gobbetti et al. 1994) and may also produce vitamins that stimulate LAB growth. The metabolic products of LAB carbohydrate metabolism, lactate and acetate, contribute to the acidity of the sourdough and prevent development of spoilage microorganisms. These compounds are also important flavour volatiles and essential texture-forming components. Esters and ornithine from amino acid metabolism are other key contributors to sourdough flavour. Yeasts also contribute in this regard by generating flavour-enhancing fusel alcohols and carbon dioxide. The latter is supplemented with LAB carbon dioxide from heterolactic fermentation causing the dough to rise (reviewed by De Vuyst et al. 2009).
The microbial composition at the start of the fermentation is influenced by the dough ingredients, the production environment, and the “mother sponge” or starter dough. The fate of the microorganisms during sourdough fermentation is influenced by technological parameters such as leavening and storage temperatures, the number and duration of refreshment steps, the ratio of flour to water (dough yield), pH and redox potential, the use of starters and/or bakers yeast, etc. The chemical composition and enzymatic activities of the flour are additional determinants of sourdough microbial ecology. The developing microbial populations are influenced by the metabolic factors mentioned above, which favour stress-tolerant species that utilise available nutrients with high energy gain, while keeping a balanced reduction/oxidation state. Competitive action, dominance, and symbiotic interactions potentially lead to the complementary utilisation of different carbohydrates and other resources (De Vuyst et al. 2009; Gänzle et al. 2009).
An overview of yeast species identified from sourdough fermentations and a selection of commonly found bacteria is given in Fig. 1. The most frequently found co-occurrences of yeast and LAB follow a complementary utilisation of maltose. This disaccharide is not utilised by the yeasts Kazachastania exigua and Candida humilis/milleri, but is essential for co-isolated Lactobacillus sanfranciscensis and L. brevis and is facultatively metabolised by S. cerevisiae and L. plantarum. Based on frequency and complementary use of maltose, these microbial associations are considered particularly stable and favourable for high quality sourdoughs. Numerous other co-occurrences that follow the principle of complementarities, but involve facultative maltose utilisation by the yeasts and/or LAB, have been observed (Infantes and Tourneur 1991; Infantes and Schmidt 1992; Gobbetti et al. 1994; Corsetti et al. 2001; Meroth et al. 2003a, b; Van der Meulen et al. 2007; Scheirlinck et al. 2007, 2008; Iacumin et al. 2009; Osimani et al. 2009; Vrancken et al. 2010).
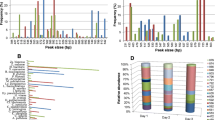
Yeasts and common lactic acid bacteria (LAB) found in sourdoughs. The lines indicate co-isolations of yeasts and LAB that are most frequently found, particularly stable and favorable for a high quality of the end-products. For the yeasts, their ability to ferment maltose (M), acid tolerance (H+), and osmotolerance at 50% glucose (%) are indicated by + positive, − negative, v variable, w weak. Indications followed by an asterisk are based on 1–11 strain records of the Mycothèque de l’Université de Louvain. LAB are grouped according to their metabolic behavior
Wickerhamomyces anomalus in sourdough and cereals
Vrancken et al. (2010) have analysed the yeast species composition of artisan bakery sourdoughs and sourdoughs produced under controlled laboratory conditions by using flours as the only non-sterile component added (Scheirlinck et al. 2007; Van der Meulen et al. 2007). They found W. anomalus as a major component of the yeast population. The dominance of S. cerevisiae (68% of the isolates) in artisan bakery sourdoughs was interpreted by Vrancken et al. (2010) as being caused by the bakery environment, because no S. cerevisiae was intentionally added to these (in some cases 25 years old) traditionally maintained sourdoughs. However, W. anomalus was the second most frequent yeast with 26% of the isolates and was detected in two of 21 artisan sourdoughs as the only yeast species. The dominance of W. anomalus (66% of the isolates) in laboratory sourdough fermentations was associated with its osmotolerance and better acid tolerance, in contrast with the narrower environmental preferences of S. cerevisiae. W. anomalus was introduced into the fermentations by the different cereal flours used, a substrate in which S. cerevisiae is rarely found. The possibility of dominating W. anomalus populations in sourdough should assure an interest in the technological potential of this species including possible contribution to flavour and other product qualities.
The question of the potential origin of W. anomalus in sourdough fermentations led us to evaluate selected and well-documented strains of this species, available from culture collections. Considering 89 strains from the CBS, ARS-NRRL, and BCCM/MUCL collections, widely diverse substrates were seen, indicating a generalist nature for W. anomalus (Table 1). However, when categorising the substrates into processed food and food components (55 strains), natural substrates (26 strains), and others (8 strains), a high frequency of food-related substrates was noted. W. anomalus was also associated with natural sources, including flowers, fruits, other plant surfaces and exudates, animals, and mushrooms. Geographically, the collection strains originated mostly from Europe, but also from America, Asia, and Africa. This evaluation is only indicative, as deposits in culture collections are biased by the scope of the collection and the motivation of the depositors. A true analysis of the ecological niche of the species would imply long-term collecting efforts throughout diverse environments and locations.
Focussing on culture collection strains and published reports of W. anomalus isolated from sourdough and associated substrates such as cereals and flours, the species was found in these habitats on ten different occasions (Table 2). Kurtzman et al. (1970) isolated yeasts from wheat and wheat flour from six regions in the USA and showed that basidiomycetous yeasts dominated in grain samples. W. anomalus was isolated from flours, but not from grains, from three regions of the “wheat belt” area in the USA. This species was shown to outgrow other yeasts and established itself as dominant species (68% of isolates) in ensiled high-moisture corn. This was most likely due to its ability to assimilate lactate produced by LAB, another dominating group of microorganisms typically present in silage (Burmeister and Hartman 1966). Similar findings were made in rice stored in a sealed bin (Teunisson 1954) and during storage of high-moisture wheat grain where it inhibited the growth of moulds (Bjönberg and Schnürer 1993). Therefore, the yeast species W. anomalus appears to be associated with the natural microbiota of cereals and this yeast possesses characteristics that confer an advantage over other yeasts in co-culture with LAB. However, the exact nature of the link between W. anomalus and cereals remains unknown.
Strain diversity
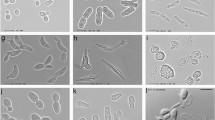
To track the possible origins of W. anomalus populations, molecular tools are needed. The study of Belgian bakery and laboratory sourdoughs provided numerous isolates from different fermentations that were used as a starting material to search for strain-specific markers. One hundred and seventy-seven W. anomalus isolates were identified by PCR-fingerprinting with the mini-satellite specific primer M13, of which a selection was characterised by physiology and morphology, as well as by sequence analysis of the D1/D2 LSU rDNA, the ITS1/5.8S/ITS2 rDNA and partial sequences of the ACT1 gene (Vrancken et al. 2010). PCR-fingerprinting revealed two profiles distinguished by the presence or absence of one bright band of approximately 1,800 bp. To investigate intra-species variation, 27 isolates that represented different bakery and laboratory sourdoughs as well as both PCR-fingerprint profiles were sequenced with respect to their ITS region and the ACT1 gene. The ACT1 gene showed up to seven heterogeneous nucleotide positions, in which the electropherograms showed peaks of two different nucleotides at the same position at similar signal strength. Consequently, these positions could not be identified by direct sequencing of PCR products and, hence, cannot be used for potential strain distinction without prior cloning. Also, ITS sequences did not yield significant variation among most isolates, with the exception of isolate R7-24h1 (MUCL 51241) from a laboratory sourdough fermentation that differed by six substitutions and three insertions or deletions from the type strain of W. anomalus and all other sourdough isolates. A BLAST search showed that four identical sequences had been determined by other researchers, while another 44 ITS sequences, highly similar to the type strain, were present in the databases. Among the four strains with ITS sequences identical to R7-24h1 (MUCL 51241), the assignment of MTCC 462 to the species W. anomalus was confirmed by sequencing of the intergenic spacer (IGS) region (Sutar et al. 2004). The deviating ITS sequences formed a cluster that was well separated from the cluster containing the type strain, including even more variable sequences, all of which were distinct from closely related species such as Wickerhamomyces subpelliculosa, Wickerhamomyces ciferrii, Wickerhamomyces edaphicus, and a yet undescribed Wickerhamomyces species (Fig. 2). This cluster was characterised by a shared motif in the ITS1 region, a fact that raised our interest in a search for potential sequential sequence evolution in this subgroup of W. anomalus.
Similarity tree of 103 internal transcribed spacer rDNA sequences of W. anomalus and closely related species. The tree was derived by neighbour joining using p-distances after pairwise deletion of gaps in the software MEGA. These options were chosen to preserve a maximum of information that might be lost by more restrictive sequence evolution models. The alignment consisted of 534 sites. Bootstrap proportions over 70% after 1,000 repetitions are given above branches. Terminals are labelled with the sequence accession number if available, followed by the abbreviated species denomination as shown in the sequence database entry (Pa: Pichia anomala, Pcif: Pichia ciferrii, Psp: Pichia sp., Psub: Pichia subpelliculosa, Wa: Wickerhamomyces anomalus, Wedaph: Wickerhamomyces edaphicus, Wsp: Wickerhamomyces sp., Ssp: Saccharomycete, Saccharomycetales sp., Scsp: Saccharomycodaceae sp.) and the strain number if available. Sourdough isolates from Vrancken et al. (2010) are underlined. Type strain sequences are in bold fonts. The partial sequence alignment shows the ITS1 motif that distinguished group B of W. anomalus isolates from the majority forming group A, represented in the alignment only by the three independently determined type strain sequences and one sourdough isolate. Sequence FJ605112 does not seem to represent a strain of W. anomalus. Dots indicate nucleotides identical to the first line
Sequential sequence evolution should be understood as mutational steps that connect pairs of sequences. It may be hypothetically reconstructed by parsimony network analysis of sequences of closely related strains (Templeton et al. 1992; Crandall and Templeton 1993). In contrast to tree-building methods, it assumes the inclusion of the ancestral haplotype in the sample, it allows for recombining sequences, and the result may include cyclic networks. The detection of sequential sequence evolution is important in this context, as it may confound species assignment by counting the simple number of nucleotide differences to the type strain of the species. Parsimony network analysis has been used to reconstruct the potential connection of strains with higher than expected numbers of nucleotide differences to the type strain via intermediately observed or unseen haplotypes (Lachance et al. 2010). In the present paper, the analysis of ITS sequences, following the methodology of Lachance et al. (2010), indicated that the type strain of W. anomalus, determined independently in three laboratories, was present in the most populated haplotype A, to which also the majority of the sourdough isolates were assigned (Fig. 3). Seven other haplotypes were part of the reconstructed network, meaning that they should be considered as members of the same species based on the analysed data. The sequence of a strain isolated from the lacewing Chrysoperla sillemi collected from cotton in the Nagpur district of the Maharashtra state of India (FJ605112), while included into the W. anomalus cluster by the ITS similarity tree with high bootstrap support but on a long branch, was not accepted into the network. Its species assignment should consequently be reconsidered.
Parsimony network analysis following the methodology of Lachance et al. (2010), including the same internal transcribed spacer rDNA sequences as in Fig. 1, but showing only those 87 sequences identified by this analysis as members of the species W. anomalus. Labels include the sequence accession number, strain number, substrate, and country of origin if available. Sourdough isolates from Vrancken et al. (2010) are underlined and type strain sequences are in bold fonts. Lines indicate single nucleotide changes and small circles between them represent missing intermediate haplotypes. Ovals and the square include identical haplotypes (after gap deletion), the square signifying the ancestral haplotype as deduced by the software. The shaded boxes A and B were distinguished by a specific ITS1 sequence motif shown in Fig. 2. Additional single nucleotide changes were distributed throughout the alignment
The ancestral haplotype can be defined as the most common one and the one that led to most diversification, two conditions which are expected to coincide (Crandall and Templeton 1993). In the present study, this coincidence was not observed. While the most populated haplotype A was only connected to two other haplotyes, haplotype B gave rise to five other haplotypes (Fig. 3). The question is why haplotype B, which had apparently more time for diversification, was far less frequently found than haplotype A? It will not be possible to answer this question based on the biased sampling of database sequences and the restricted data originating from a single sequence locus. A potential explanation may be that the underlying assumption for the prediction of ancestry, a geographically structured population with limited gene flow, has not been met. The potential cause of killer activity by mycocins in a particular haplotype seems to be unlikely, as strains in both haplotypes A and B have been specified to possess killer activity. It should be noted that haplotypes A and B have been found in the same environment, for example in a study of clinical and environmental isolates (Reyes et al. 2004) or in laboratory sourdough fermentation R7 (Vrancken et al. 2010). One and the same sourdough sample yielded the only sourdough isolate of haplotype B as well as a second isolate of W. anomalus belonging to haplotype A. The study of 23 sequences of Belgian sourdough isolates can thus be considered as a representative sample of this particular and locally determined environment, which showed the strong overrepresentation of haplotype B in the sample. However, more markers that provide information on the strain level may contribute to a better resolution of populations within the species W. anomalus.
Conclusions
Wickerhamomyces anomalus is well-adapted to the sourdough microbial ecosystem, where it is encountered as the only yeast or in an inter-yeast species community. It was introduced into laboratory sourdough fermentations via cereal flours as the only non-sterile component added. Although W. anomalus is associated with a wide geographic distribution and can be isolated from diverse substrates, it has repeatedly been reported from sourdoughs and other cereal-related sources, suggesting a link between this yeast and cereals. Its broad distribution appears to be due to its competitiveness under stressful environmental conditions, even if initially present in low cell numbers in mixed microbial populations of spontaneous fermentation systems. The W. anomalus ITS sequence-based population structure showed one large and homogenous haplotype and seven far smaller haplotypes, one of which was deduced to take a more central role in the evolution of the species than the quantitatively dominating major haplotype. The major haplotype may form a particularly successful population without regard to geography and with uninterrupted gene flow, coherent with W. anomalus being of generalist character.
Reliable strain typing methods are urgently needed for W. anomalus to further investigate the origins and routes of transfer, competiveness, and other factors. These are especially needed in view of the value of this yeast in food and feed preservation, and its recognition as a potential emerging opportunistic pathogen.
References
Bjönberg A, Schnürer J (1993) Inhibition of the growth of grain-storage moulds in vitro by the yeast Pichia anomala (Hansen) Kurtzman. Can J Microbiol 39:623–628
Burmeister HR, Hartman PA (1966) Yeasts in ensiled high-moisture corn. Appl Microbiol 14:35–38
Castelli T (1933) Su alcune Hansenule della fermentazione panaria. Arch Microbiol 4:514–529
Chakrabarti A, Singh K, Narang A, Singhi S, Batra R, Rao KLM, Ray P, Gopalan S, Das S, Gupta V, Gupta AK, Bose SM, McNeil MM (2001) Outbreak of Pichia anomala infection in the pediatric service of a tertiary-care center in Northern India. J Clin Microbiol 39:1702–1706
Christensen JE, Dudley EG, Pederson JR, Steele JL (1999) Peptidases and amino acid catabolism in lactic acid bacteria. Antonie van Leeuwenhoek J Microbiol 76:217–246
Coda R, Nionelli L, Rizzello CG, De Angelis M, Tossut P, Gobbetti M (2010) Spelt and emmer flours: characterization of the lactic acid bacteria microbiota and selection of mixed starters for bread making. J Appl Microbiol 108:925–935
Corsetti A, Lavermicocca P, Morea M, Baruzzi F, Tosti N, Gobbetti M (2001) Phenotypic and molecular identification and clustering of lactic acid bacteria and yeasts from wheat (species Triticum durum and Triticum aestivum) sourdoughs of southern Italy. Int J Food Microbiol 64:95–104
Crandall KA, Templeton AR (1993) Empirical tests of some predictions from coalescence theory with applications to intraspecific phylogeny reconstruction. Genetics 134:959–969
da Matta VLR, de Souza Carvalho Melhem M, Colombo AL, Moretti ML, Rodero L, de Almeida GMD, dos Anjos Martins M, Costa SF, Souza Dias MBG, Nucci M, Levin AS (2001) Antifungal drug susceptibility profile of Pichia anomala isolates from patients presenting with nosocomial fungemia. Antimicrob Agents Chemother 51:1573–1576
De Vuyst L, Neysens P (2005) The sourdough microflora: biodiversity and metabolic interactions. Trends Food Sci Technol 16:43–56
De Vuyst L, Vancanneyt M (2007) Biodiversity and identification of sourdough lactic acid bacteria. Food Microbiol 24:120–127
De Vuyst L, Vrancken G, Ravyts F, Rimaux T, Weckx S (2009) Biodiversity, ecological determinants, and metabolic exploitation of sourdough microbiota. Food Microbiol 26:666–675
Falkow S (1997) What is a pathogen? ASM News 63:359–365
Fernández M, Zúñiga M (2006) Amino acid catabolic pathways of lactic acid bacteria. Crit Rev Microbiol 32:155–183
Foschino R, Galli A (1997) Italian style of life: pane, amore e… lievito naturale!. Tecnol Aliment 1:42–59
Foschino R, Gallina S, Andrighetto C, Rossetti L, Galli A (2004) Comparison of cultural methods for the identification and molecular investigation of yeasts from sourdoughs for Italian sweet baked products. FEMS Yeast Res 4:609–618
Fredlund E, Druvefors U, Boysen ME, Lingsten KH, Schnürer J (2002) Physiological characteristics of the biocontrol yeast Pichia anomala J121. FEMS Yeast Res 2:395–402
Gänzle MG, Vermeulen N, Vogel RF (2007) Carbohydrate, peptide and lipid metabolism of lactic acid bacteria in sourdough. Food Microbiol 24:128–138
Gänzle MG, Zhang C, Monang B-S, Lee V, Schwab C (2009) Novel metabolites from cereal-associated lactobacilli—novel functionalities for cereal products? Food Microbiol 26:712–719
Gobbetti M (1998) The sourdough microflora: interactions of lactic acid bacteria and yeasts. Trends Food Sci Technol 9:267–274
Gobbetti M, Corsetti A, Rossi J (1994) The sourdough microflora—interactions between lactic acid bacteria and yeasts—metabolism of carbohydrates. Appl Microbiol Biotechnol 41:456–460
Gullo M, Romano AD, Pulvirenti A, Giudici P (2003) Candida humilis—dominant species in sourdoughs for the production of durum wheat bran flour bread. Int J Food Microbiol 80:55–59
Iacumin L, Cecchini F, Manzano M, Osualdini M, Boscolo D, Orlic S, Comi G (2009) Description of the microflora of sourdoughs by culture-dependent and culture-independent methods. Food Microbiol 26:128–135
Infantes M, Schmidt JL (1992) Characterisation of the yeast flora of natural sourdough located in various French areas. Sci Aliment 12:271–287
Infantes M, Tourneur C (1991) Survey on the lactic flora of natural sourdoughs located in various French areas. Sci Aliment 11:527–545
Kalenic S, Jandrlic M, Vegar V, Zuech N, Sekulic A, Mlinaric-Missoni E (2001) Hansenula anomala outbreak at a surgical intensive care unit: a search for risk factors. Eur J Epidemiol 17:491–496
Kurtzman CP (1998) Pichia E.C. Hansen emend. Kurtzman. In: Kurtzman CP, Fell JW (eds) The yeasts: a taxonomic study, 4th edn. Elsevier, Amsterdam, The Netherlands, pp 273–352
Kurtzman CP, Wickerham LJ, Hesseltine CW (1970) Yeasts from wheat and flour. Mycologia 62:542–547
Kurtzman CP, Robnett CJ, Basehoar-Powers E (2008) Phylogenetic relationships among species of Pichia, Issatchenkia and Williopsis determined from multigene sequence analysis, and the proposal of Barnettozyma gen. nov., Lindnera gen. nov. and Wickerhamomyces gen. nov. FEMS Yeast Res 8:939–954
Lachance MA, Dobson J, Wijayanayaka DN, Smith AME (2010) The use of parsimony network analysis for the formal delineation of phylogenetic species in yeasts: Candida apicola, Candida azyma, and Candida parazyma sp. nov., cosmopolitan yeasts associated with floricolous insects. Antonie van Leeuwenhoek J Microbiol 97:155–170
Meroth CB, Hammes WP, Hertel C (2003a) Identification and population dynamics of yeasts in sourdough fermentation processes by PCR-denaturing gradient gel electrophoresis. Appl Environ Microbiol 69:7453–7461
Meroth CB, Walter J, Hertel C, Brandt MJ, Hammes WP (2003b) Monitoring the bacterial population dynamics in sourdough fermentation processes by using PCR-denaturing gradient gel electrophoresis. Appl Environ Microbiol 69:475–482
Murphy N, Damjanovic V, Hart CA, Buchanan CR, Whitaker R, Cooke RWI (1986) Infection and colonization of neonates by Hansenula anomala. Lancet 327:291–293
Osimani A, Zannini E, Aquilanti L, Mannazzu I, Comitini F, Clementi F (2009) Lactic acid bacteria and yeasts from wheat sourdoughs of the Marche region. Ital J Food Sci 21:269–286
Piarroux R, Millon L, Bardonnet K, Vagner O, Koenig H (1999) Are live Saccharomyces yeasts harmful to patients? Lancet 353:1851–1852
Polonelli L, Archibusacci C, Sestito M, Morace G (1983) Killer system: a simple method for differentiating Candida albicans strains. J Clin Microbiol 17:774–780
Pulvirenti A, Caggia C, Restuccia C, Gullo M, Giudici P (2001) DNA fingerprinting methods used for identification of yeasts isolated from Sicilian sourdoughs. Ann Microbiol 51:107–120
Pulvirenti A, Solieri L, Gullo M, De Vero L, Giudici P (2004) Occurrence and dominance of yeast species in sourdough. Lett Appl Microbiol 38:113–117
Reyes E, Barahona S, Fischman O, Niklitschek M, Baeza M, Cifuentes V (2004) Genetic polymorphism of clinical and environmental strains of Pichia anomala. Biol Res 37(Suppl A):747–757
Rocha JM, Malcata FX (1999) On the microbiological profile of traditional Portuguese sourdough. J Food Prot 62:1416–1429
Rossi J (1996) The yeasts in sourdough. Adv Food Sci 18:201–211
Salovaara H, Savolainen J (1984) Yeast type isolated from Finnish sour rye dough starters. Acta Aliment Pol 10:241–246
Scheirlinck I, Van der Meulen R, Van Schoor A, Vancanneyt M, De Vuyst L, Vandamme P, Huys G (2007) Influence of geographical origin and flour type on diversity of lactic acid bacteria in traditional Belgian sourdoughs. Appl Environ Microbiol 73:6262–6269
Scheirlinck I, Van der Meulen R, Van Schoor A, Vancanneyt M, De Vuyst L, Vandamme P, Huys G (2008) Taxonomic structure and stability of the bacterial community in Belgian sourdough ecosystems as assessed by culture and population fingerprinting. Appl Environ Microbiol 74:2414–2423
Sekhon AS, Kowalewska-Grochowska K, Garg AK, Vaudry W (1992) Hansenula anomala fungemia in an infant with gastric and cardiac complications with a review of the literature. Eur J Epidemiol 8:305–308
Sterr Y, Weiss A, Schmidt H (2009) Evaluation of lactic acid bacteria for sourdough fermentation of amaranth. Int J Food Microbiol 136:75–82
Succi M, Reale A, Andrighetto C, Lombardi A, Sorrentino E, Coppola R (2003) Presence of yeasts in southern Italian sourdoughs from Triticum aestivum flour. FEMS Microbiol Lett 225:143–148
Sugihara TF, Kline L, Miller MW (1971) Microorganisms of the San Francisco sour dough bread process. Appl Microbiol 21:456–458
Sutar R, David JK, Ganesan K, Ghosh AK, Singhi S, Chakrabarti A, Bachhawat AK (2004) Comparison of ITS and IGS1 regions for strain typing of clinical and non-clinical isolates of Pichia anomala. J Med Microbiol 53:1–5
Templeton AR, Crandall KA, Sing CF (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III Cladogram estimation. Genetics 132:619–633
Teunisson DJ (1954) Yeasts from freshly combined rough rice stored in a sealed bin. Appl Microbiol 2:215–220
Valmorri S, Tofalo R, Settanni L, Corsetti A, Suzzi G (2010) Yeast microbiota associated with spontaneous sourdough fermentations in the production of traditional wheat sourdough breads of the Abruzzo region (Italy). Antonie van Leeuwenhoek J Microbiol 97:119–129
Van der Meulen R, Scheirlinck I, Van Schoor A, Huys G, Vancanneyt M, Vandamme P, De Vuyst L (2007) Population dynamics and metabolite target analysis of lactic acid bacteria during laboratory fermentations of wheat and spelt sourdoughs. Appl Environ Microbiol 73:4741–4750
Vernocchi P, Valmorri S, Gatto V, Torriani S, Gianotti A, Suzzi G, Guerzoni ME, Gardini F (2004) A survey on yeast microbiota associated with an Italian traditional wheat-leavened baked good fermentation. Food Res Int 37:469–476
Vrancken G, De Vuyst L, Van der Meulen R, Huys G, Vandamme P, Daniel HM (2010) Yeast species composition differs between artisan bakery and spontaneous laboratory sourdoughs. FEMS Yeast Res 10:471–481
Zannini E, Garofalo C, Aquilanti L, Santarelli S, Silvestri G, Clementi F (2009) Microbiological and technological characterization of sourdoughs destined for bread-making with barley flour. Food Microbiol 26:744–753
Acknowledgments
The authors would like to acknowledge their financial support from the Research Council of the Vrije Universiteit Brussel (OZR, GOA, and IOF projects), the Fund for Scientific Research—Flanders, the Institute for the Promotion of Innovation through Science and Technology in Flanders, in particular the SBO project “New Strategy for the Development of Functional and Performance Starter Cultures for Foods in Function of ‘Food Qualitomics’ ”, and the Federal Research Policy (contract BCCM C3/10/003), in particular the project of the Action for the Promotion of and Cooperation with the Belgian Coordinated Collections of Microorganisms.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Daniel, HM., Moons, MC., Huret, S. et al. Wickerhamomyces anomalus in the sourdough microbial ecosystem. Antonie van Leeuwenhoek 99, 63–73 (2011). https://doi.org/10.1007/s10482-010-9517-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-010-9517-2