Abstract
Vascular endothelial growth factor (VEGF) is a major angiogenic factor that activates pro-angiogenic molecules to generate new vessels. Recently, we identified a VEGF-A-induced pro-angiogenic gene, BCL-6 associated zinc finger protein (BAZF), in endothelial cells. BAZF interacts with CBF1, a transcriptional regulator of Notch signaling, and downregulates Notch signaling by inducing the degradation of CBF1. A signal inhibition assay with a combination of chemical inhibitors and siRNA revealed that the protein kinase D (PRKD) family, mainly PRKD2, mediated BAZF gene expression by VEGF-A stimulation. A luciferase reporter assay showed that the promoter activity of the BAZF gene was unchanged by VEGF-A stimulation. However, we found that the stability of BAZF mRNA increased in a VEGF-A/PRKD2-dependent manner. In further studies to investigate the underlying mechanism, we successfully identified heat shock protein 90 beta (HSP90β) as a molecule that interacts with and stabilizes BAZF mRNA following VEGF-A/PRKD2 activation. These data suggest that HSP90β may positively regulate angiogenesis, not only as a protein chaperone, but also as an mRNA stabilizer for pro-angiogenic genes, such as BAZF, in a PRKD2 activity-dependent manner.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Angiogenesis is defined as new vascular formation from pre-existing blood vessels. This phenomenon arises in several physiological conditions, such as embryonic development, wound healing, and the female reproductive cycle in adults. Defects in angiogenesis also affect the progression of a wide range of diseases, including rheumatoid arthritis, psoriasis, diabetic retinopathy, and malignant tumors, collectively referred to as “angiogenesis-dependent diseases” [1]. Further understanding of angiogenic mechanisms may lead to novel therapeutics for such diseases. The regulation of angiogenesis depends on a balance between activation and inhibition [2]. When angiogenesis is required, activation can be achieved either by increasing pro-angiogenic activators and/or decreasing angiogenic suppressors. Vascular endothelial growth factor (VEGF), in particular, is a potent pro-angiogenic growth factor that controls most embryonic vessels.
The mammalian VEGF family comprises 5 glycoproteins, VEGF-A, VEGF-B, VEGF-C, VEGF-D (also known as FIGF), and placenta growth factor (PlGF) [3, 4]. VEGF-A is the best characterized member of the VEGF family and is predominantly expressed in many solid human tumors. Members of the VEGF family associate with 3 types of receptor tyrosine kinases, termed VEGFR1 (also known as FLT1), VEGFR2 (also known as Flk1 in mouse or KDR), and VEGFR3 (also known as FLT4). VEGF-A has a high binding affinity with VEGFR1 and VEGFR2. Importantly, VEGFR2 is expressed primarily in the vasculature and is a critical mediator of VEGF-induced angiogenesis.
On the other hand, vessel-stabilizing pathways, such as the Notch signaling pathway, are also essential in angiogenic regulation. Recent studies have demonstrated that angiogenic sprouting is tightly regulated by the Notch signaling circuit of the VEGFR-Notch ligand, Dll4 [5–18]. Interestingly, endothelial cells (ECs) in pre-existing vessels express both Dll4 and Notch, and signaling is thought to be balanced between ECs. Therefore, VEGF-A stimulation could affect the Dll4-Notch balance in pre-existing vessels to rearrange the signaling direction in sprouting areas, upregulating Dll4 in tip cells and inducing Notch signaling in adjacent cells to become stalk cells [19]. However, the precise molecular mechanisms mediating this fine-tuning of the angiogenic “tug-of-war” signaling balance between VEGFR and Notch have remained unclear.
In a recent study, we identified a VEGF-A-induced critical angiogenic factor, BCL6-associated zinc finger protein (BAZF) [20]. VEGF-A-induced BAZF forms a complex with cullin3, an E3 ubiquitin ligase, to degrade CBF-1, a transcriptional factor involved in Notch signaling. The loss of CBF-1 in endothelial cells induces spontaneous angiogenesis in adult mice, revealing that Notch signaling play a key role in the maintenance of angiogenic quiescence [21]. Thus, we propose that BAZF is an important factor that can interrupt the “tug-of-war” balance between VEGFR and Notch signaling to promote angiogenesis.
We have already shown that levels of BAZF mRNA and protein peak in 2–4 h and 4–6 h after a VEGF-A stimulation, respectively [20]. In this study, we investigated the molecular mechanisms mediating VEGF-A-induced BAZF gene expression.
Materials and methods
Reagents and antibodies
Recombinant human VEGF and fibroblast growth factor-2 (FGF-2) were purchased from R&D Systems. RO31-8220, LY294002, PD98059, Gö6976, Gö6983, and protein kinase C beta (PKCβ) inhibitors were purchased from Calbiochem. A23187 and phorbol 12-myristate 13-acetate (PMA) were purchased from Sigma. Actinomycin D was purchased from MP Biomedicals. PU-H71 hydrate was purchased from Santa Cruz Biotechnology. Anti-protein kinase D1 and 2 (PRKD1/2) antibodies were purchased from Santa Cruz Biotechnology. Anti-PRKD3 antibodies were purchased from Bethyl Laboratories Inc. Anti-heat shock protein 90 beta (HSP90β) antibodies were purchased from Millipore, abcam, and EPITOMICS. Anti-HSP27 was purchased from Cell Signaling Technology, and growth factor reduced Matrigel was purchased from BD Biosciences. Anti-green fluorescent protein (GFP) was purchased from abcam.
Cell culture
Human umbilical vein endothelial cells (HUVECs) and human dermal microvascular ECs from Cell System and human coronary artery ECs and human retinal microvascular ECs from Applied Cell Biology Research were cultured in endothelial growth medium, EGM-2 (Lonza). These cells were used for experiments from passages 3–6.
Reverse transcription real-time quantitative PCR (RT-qPCR)
Total RNAs were prepared using TRIzol Reagent (Invitrogen) according to the manufacturer’s protocol. One microgram of RNA was used for first-strand synthesis using the High Capacity RNA-to-cDNA Master Mix (Applied Biosystems). Real-time PCR was performed (FastStart Universal SYBR Green Master ROX; Roche) with the ABI 7300/7500 Real-Time PCR system (Applied Biosystems).
Western blotting
Samples were separated by SDS-PAGE, followed by transfer to a nitrocellulose membrane. The membrane was then blocked with 5 % skim milk in 0.05 % Tween 20/PBS (-) (PBS-T) for 30 min, followed by incubation with a primary antibody. After washing with PBS-T, the membrane was incubated with appropriate horseradish peroxidase conjugated IgG antibodies (Promega). After antibody incubation, proteins were detected by enhanced chemiluminescence (GE Healthcare).
RNA interference
siRNAs targeting human PRKD1, PRKD2, and PRKD3 and a scrambled siRNA control were purchased from QIAGEN. The transfection of siRNA was performed with Lipofectamine RNAiMAX (Invitrogen) at 20 nM according to the manufacturer’s protocol.
Construction of promoter-luciferase reporter plasmids
A 5.5-kb fragment containing 5530 bp of the 5′ upstream region and the first basepair of exon I of the human BAZF gene was amplified from the genomic DNA of HUVECs and ligated into the KpnI and NheI sites of the pGL4.1 vector (Promega) to obtain the −5530/+1 reporter construct.
Actinomycin D chase assay
HUVECs were treated with 2 mg/mL actinomycin D for 30 min (added at time 0), followed by treatment with 50 ng/mL VEGF-A and/or chemical inhibitors. HUVECs were collected at the indicated times, and total RNA was isolated as described above. RT-qPCR was performed as described, and mRNA levels were normalized using 18S ribosomal RNA, with the mRNA level at time 0 set at 100 %.
Construction and production of lentivirus vectors
Lentivirus vectors were constructed by inserting mutant cDNAs encoding GFP-fused PRKD2 into the lentiviral expression vector CSII-CMV-MCS-IRES2-Bsd. Lentiviral vectors were produced in 293T cells. Mutant cDNAs for GFP-fused PRKD2 were kindly provided by Dr. Nishimura (Kumamoto University) [22]. Lentiviral expression and packaging vectors were kindly provided by Dr. Miyoshi (RIKEN BioResource Center).
Luciferase assay
The human NF-κB-responsive promoter-luciferase reporter gene was kindly provided by Dr. Yoshimura (Keio University). A reporter plasmid containing the firefly luciferase gene was transfected into HUVECs with pRL-TK (Promega) using Sugarfect (Medgel) according to the manufacturer’s protocol. Luciferase activities were measured using a dual-luciferase reporter assay system (Promega) with a GloMax96 microplate luminometer (Promega) according to the manufacturer’s instructions.
Biotinylated RNA pull-down assay
For biotinylated RNA pull-down assays, PCR fragments containing the T7 RNA polymerase promoter sequence [(T7): CTAATACGACTCACTATAGGGAGA] were used as a template for in vitro transcription of the coding region (CR) and full-length (FL) BAZF using biotinylated CTP. Ten micrograms of biotinylated FL transcript was incubated with 100 μg protein from the cytoplasmic fraction, which was precleared with biotinylated CR transcript, for 1 h at room temperature, followed by isolation of the complexes with streptavidin-coated magnetic Dynabeads (Invitrogen). The pull-down material was separated by SDS-PAGE, and CBB staining was carried out with Bio-Safe Coomassie (BIO-RAD). Protein bands were excised from gels and subjected to in-gel digestion with sequence-grade modified trypsin (Promega), as previously described [20, 23]. Tryptic digests were reconstituted with 0.2 % (v/v) trifluoroacetic acid for mass spectrometric analysis. PMF and MS/MS analysis was performed using a MALDI-TOF/TOF mass spectrometer (Shimadzu AXIMA-TOF2). All MS or MS/MS spectrum data were submitted to the MASCOT program (Matrix Science) in order to obtain a protein candidate for each spot. Database searches were performed against the NCBInr database version 20070629 using the following parameters: (1) unlimited protein molecular weight and pI ranges; and (2) presence of protein modifications, including acrylamide modification of cysteine, methionine oxidation, protein N-terminal acetylation, and pyro-glutamic acid.
Protein-binding RNA detection assay
To detect HSP90β-associated BAZF mRNA, we used a previously reported method, with some modifications [24]. Briefly, VEGF-A-treated HUVECs were lysed using a cell lysis buffer [50 mM Tris (pH7.4), 150 mM NaCl, 1 mM MgCl2, and 0.05 % Nonidet P-40], and cell lysates were then incubated with Dynabeads Protein G (Invitrogen) and anti-HSP90β antibodies (abcam) for 1 h at 4 °C. The beads were then treated with 20 U DNase and 0.5 mg/mL proteinase K for 30 min at 15 °C. Total RNA was isolated from the supernatant using an ethanol precipitation method, and BAZF mRNA was detected by RT-qPCR as described above. The primer sets are listed in Supplemental Table 1.
Two-dimensional gel electrophoresis
After infection of HUVECs with kinase-dead (KD) or constitutively active (CA) PRKD2-expressing lentivirus, cells were collected and lysed with IEF lysis buffer [8.5 M Urea, 4 % CHAPS, and 0.2 % Biolyte 3/10 (Bio-Rad)]. Twenty micrograms of protein from each sample was loaded onto Immobiline Dry Strips (pH 3–10, 13 cm in length; GE Healthcare). Two-dimensional gel electrophoresis was performed as previously described [23]. Western blotting with anti-HSP90β or anti-HSP27 was then carried out as described above.
Endothelial cell network formation assay
Matrigel (BD Biosciences) was prepared in 24-well culture plates (IWAKI). HUVECs were serum starved with EBM-2 containing 0.15 % FBS for 12 h. These cells were seeded onto solidified gels at a concentration of 1 × 104 cells/well in 500 μL of EBM-2 containing 0.15 % FBS and VEGF-A (50 ng/mL) with or without Pu-H71 (1 μM). After 8 h, photographs of the cells were taken using a microscope (Olympus). Total network length was measured and calculated using Image-Pro Plus software (Roper).
Statistical analysis
Data were analyzed by unpaired t tests. Differences were considered significant if p values were less than 0.05.
Results
VEGF-A induces BAZF expression through PRKD2 activation in HUVECs
To clarify which signaling pathway is responsible for the induction of BAZF mRNA in response to VEGF–stimulation in HUVECs, we examined the effects of various kinase inhibitors on VEGF-A-dependent BAZF gene expression. HUVECs were treated with the MEK inhibitor PD98059 (30 μM), pan-PKC inhibitor RO31-8220 (1 μM), phosphatidylinositol 3-kinase inhibitor LY294002 (10 μM), or p38 MAPK inhibitor SB202190 (20 μM) for 30 min before VEGF-A stimulation. Although PD98059 and RO31-8220 significantly inhibited VEGF-A-dependent BAZF mRNA induction, the PKC inhibitor was more effective at suppressing BAZF transcript levels (Fig. 1a). To determine whether PKC activation was involved in VEGF-A-induced BAZF mRNA expression in HUVECs, we examined the effects of PKC activators on BAZF expression. HUVECs were treated with the pan-PKC activator PMA (10 nM) or the cyclic PKC activator A23187 (5 μM) for 2 h. PMA, but not A23187, significantly upregulated BAZF mRNA (Fig. 1b). In addition, we examined which PKC isoform was important for VEGF-A-induced BAZF expression using various PKC inhibitors. HUVECs were treated with RO31-8220, Gö6976 (1 μM), Gö6983 (1 μM), or PKCβ inhibitor (1 μM) for 30 min prior to VEGF-A stimulation. Both RO31-8220 and Gö6976, but neither Gö6983 nor PKCβ inhibitor, significantly inhibited VEGF-A-dependent BAZF mRNA expression (Fig. 1c). Since Gö6976, but not Gö6983, selectively inhibits PKCβ and PRKD enzymes, these data suggested that PRKD activation regulated BAZF gene expression in HUVECs.
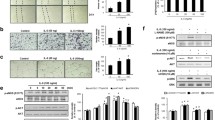
VEGF-A induces BAZF expression through PRKD activation in HUVECs. a Identification of the VEGFR-mediated downstream pathway regulating BAZF gene activation. HUVECs were pretreated with the MEK inhibitor PD98059 (30 μM), PKC inhibitor RO31-8220 (1 μM), PI3 K inhibitor LY294002 (10 μM), or MAPK inhibitor SB202190 (20 μM). Inhibitor-treated HUVECs were stimulated with 50 ng/mL VEGF-A for 2 h, and BAZF mRNA was detected by reverse transcription real-time quantitative PCR (RT-qPCR). b The PKC family pathway stimulates BAZF gene expression. HUVECs were stimulated with 50 ng/mL VEGF-A, 10 nM phorbol 12-myristate 13-acatate (PMA), or 5 μM A23187 Calcium Ionophore for 2 h. c Identification of the PKC member that regulates the BAZF gene. HUVECs were pretreated with different PKC family inhibitors, including RO31-8220 (1 μM), Gö6976 (1 μM), Gö6983 (1 μM), or PKCβ inhibitor (1 μM) for 30 min before VEGF-A stimulation. The pretreated cells were then incubated with 50 ng/mL VEGF-A for 2 h. d Knockdown efficiency of siRNA against the PRKD family. HUVECs were transfected with siRNAs (20 nM) targeting PRKD family members. The knockdown efficiency of each siRNA was checked at the protein level using western blotting analysis. Bands were normalized using a β-actin loading control. e PRKD2 is responsible for BAZF gene activation by VEGF-A. BAZF mRNA was measured in HUVECs after siRNA-mediated knockdown of PRKDs and subsequent stimulation with 50 ng/mL VEGF-A for 2 h. f Induction of BAZF mRNA by PRKD2 activation. HUVECs were infected with various mutant PRKD2-expressing lentiviruses at 1 multiplicity of infection (m.o.i.). Forty-eight hours after infection, the HUVECs were stimulated with 50 ng/mL VEGF-A for 2 h. mAG1; monomeric Azami Green 1, WT wild-type, CA constitutively active mutant, KD; kinase-dead mutant. The experiments were independently performed 3 times. *p < 0.05; ** p < 0.01. g The protein expression levels of exogenous PRKD2 between WT 1PRKD2 and the mutants in the infected HUVECs. The protein level of β-actin was shown as a loading control
The PRKD family includes 3 isoforms, PRKD1, PRKD2, and PRKD3. To assess which PRKD isoform was required for BAZF mRNA induction by VEGF-A in HUVECs, we examined the effects of siRNAs targeting each PRKD mRNA on BAZF mRNA expression. When HUVECs were transfected with human PRKD1, PRKD2, or PRKD3 siRNA at 20 nM, the expression of each PRKD protein significantly decreased (Fig. 1d). The PRKD2 siRNA was also effective at inhibiting PRKD3 mRNA and protein expression. PRKD siRNA-transfected HUVECs were treated with VEGF-A for 2 h. PRKD2 and PRKD3 siRNAs significantly inhibited BAZF mRNA expression both with and without VEGF-A treatment; PRKD2 siRNA suppressed VEGF-A-induced gene expression to a greater extent (Fig. 1e), possibly due to the more broad specificity of this siRNA construct. These data indicated that both PRKD2 and PRKD3 mediated VEGF-A-induced BAZF mRNA expression; in contrast, because PRKD3 knockdown was less effective in a VEGF-A-stimulated state, PRKD3 may specifically regulate the steady state expression of BAZF mRNA. In addition, we examined effects of wild-type (WT), CA, or KD PRKD2 mutants on BAZF mRNA expression. HUVECs were infected with each PRKD2 mutant-expressing lentivirus at 1 multiplicity of infection (m.o.i.). As shown in Fig. 1f and g, the CA PRKD2 mutant induced BAZF mRNA expression in HUVECs. On the other hand, expression of the KD-PRKD2 mutant significantly suppressed the induction of BAZF mRNA. These data indicated that PRKD2 activation mainly regulated VEGF-A-induced BAZF gene expression.
A 5.5-kb BAZF promoter does not respond to VEGF-A stimulation
To analyze transcriptional regulation of the BAZF gene, a 5.5-kb fragment of the 5′ upstream region of human BAZF from the first base of exon 1 was cloned in a luciferase reporter plasmid pGL4.1 [pGL4-BAZF (−5530/+1)]. This reporter plasmid was then transfected into various cell types. Transcriptional activity tended to be strong in endothelial cells (ECs), and promoter activity also appeared to correlate well to BAZF mRNA expression in ECs (Fig. 2a). To determine the critical region for promoter activity in the 5.5-kb fragment, we generated a series of deleted BAZF promoter-containing plasmids. These plasmids were transfected into HUVECs, and the cells were assayed for luciferase activity 24 h post-transfection. pGL4-BAZF (−2548/+1), pGL4-BAZF (−2033/+1), pGL4-BAZF (−1603/+1), and pGL4-BAZF (−1218/+1) exhibited transcriptional activity, while pGL4-BAZF (−510/+1) did not (Fig. 2b). These data suggest that the critical region for promoter activity exists in the −1218/−510 region. Therefore, the region spanning −1218/+1 could be a putative BAZF promoter. Hence, pGL4-BAZF (−1218/+1) was used for further studies. Next, we examined whether the BAZF promoter responded to VEGF-A stimulation. pGL4-BAZF (−1218/+1)-transfected HUVECs were serum-starved for 12 h before VEGF-A stimulation, and luciferase activity was measured at 6 h after VEGF-A stimulation. The BAZF promoter exhibited no response to VEGF-A, while NF-κB-mediating activation of the luciferase reporter responded to VEGF-A stimulation (Fig. 2c). These data suggested that endothelium-specific BAZF expression depended on its promoter activity, which is not enhanced by VEGF-A.
VEGF-A does not affect the transcriptional activity of BAZF. a Cell type-specific activity of the BAZF gene promoter. Several types of human primary endothelial cells (CAEC cardiac coronary artery endothelial cells, DMVEC dermal microvascular endothelial cells, RMVEC retinal microvascular endothelial cells), and non-endothelial cell lines were transfected with a BAZF promoter reporter, and luciferase activity was measured in each cell line. BAZF mRNA levels were measured by RT-qPCR. The level of BAZF mRNA expression was normalized to the 18S ribosomal RNA (rRNA). b HUVECs were transfected with a reporter vector including various deleted BAZF gene promoters. Luciferase activity was assayed at 18 h post-transfection and was normalized to the activity of thymidine kinase promoter-driven renilla luciferase. c The BAZF promoter-containing reporter vector was transfected into HUVECs, and luciferase activity was assayed at 6 h post-VEGF-A stimulation (50 ng/mL). Luciferase activity of the NF-κB reporter was also measured as a positive control of VEGF-A stimulation. The experiments were independently performed at least 3 times. *p < 0.05
VEGF-A stabilizes BAZF mRNA through a PRKD2 signaling pathway
To determine whether BAZF mRNA was regulated by VEGF-A stimulation at post-transcriptional levels, we performed actinomycin D chase experiments. The half-life (t1/2) of BAZF mRNA was analyzed by RT-qPCR following treatment with actinomycin D to inhibit de novo transcription. The t1/2 of BAZF mRNA was found to be significantly longer after VEGF stimulation (t1/2 = 3.47 h) than without treatment (t1/2 = 0.98 h), and RO31-8220 inhibited BAZF mRNA stabilization by VEGF-A (t1/2 = 0.98 h), as shown in Fig. 3a. On the other hand, neither the t1/2 nor the level of GAPDH mRNA was affected by VEGF-A, with or without RO31-8220 treatment (Fig. 3b). These data indicated that BAZF mRNA levels were stabilized by a VEGF-A stimulation through a PRKD-mediated signaling pathway.
VEGF-A stabilizes BAZF mRNA through a PRKD2-mediated pathway. The half-life of BAZF mRNA was measured after actinomycin D treatment to inhibit de novo mRNA synthesis. HUVECs were stimulated by VEGF-A at 30 min after actinomycin D treatment. Total RNA was collected at each indicated time, and BAZF (a) and GAPDH (b) mRNA levels were measured by RT-qPCR. Data were normalized to 18S rRNA levels and represented as a percentage of the mRNA levels before actinomycin D treatment. The experiments were independently performed at least 3 times
The BAZF 3′ UTR mediates regulation of mRNA stability by PRKD2
The 3′ untranslated region (3′UTR) of mRNA is known to be important for mRNA stability and translation. In particular, the AU-rich sequence element (ARE), AUUUA, in the 3′UTR was identified as a correlative region with mRNA decay [25]. BAZF mRNA also includes several ARE sites in its 3′UTR region. To investigate whether the 3′UTR of BAZF mRNA played a role in mRNA stability, we constructed a luciferase reporter plasmid containing the 3′UTR of BAZF mRNA (pCMV-luc-BAZF 3′UTR). Luciferase activity of pCMV-luc-BAZF 3′UTR significantly decreased as compared with the luciferase activity of the plasmid without the 3′UTR (pCMV-luc; Fig. 4a). PMA, a potent activator of PRKD2, significantly increased BAZF mRNA levels (Fig. 1b). Moreover, at 6 h post-PMA stimulation, luciferase activity of pCMV-luc-BAZF 3′UTR, but not that of pCMV-luc, increased, and this activity was attenuated by treatment with the PRKD inhibitor Gö6976 (Fig. 4b). Next, we examined whether the level of luciferase mRNA produced from pCMV-luc-BAZF 3′UTR was stabilized by PMA stimulation. These reporter plasmids were transfected into HUVECs, and the cells were treated with actinomycin D for 30 min before exposure to PMA. At 6 h post-PMA stimulation, the level of luciferase mRNA produced from pCMV-luc-BAZF 3′UTR, but not from pCMV-luc, increased (Fig. 4c). Moreover, downregulation of PRKD2 by siRNA abrogated PMA-dependent luciferase activity of the BAZF 3′UTR-fused luciferase gene (Fig. 4d). On the other hand, overexpression of PRKD2 mutants revealed that expression of the CA PRKD2 construct enhanced the luciferase activity of pCMV-luc-BAZF 3′UTR both without and with actinomycin D treatment (Fig. 4e, f). These data suggest that the 3′UTR of BAZF is responsible for mRNA stability and that PRKD2 activity is involved in BAZF 3′UTR-mediated regulation of mRNA stability.
VEGF-A-induced PRKD2 activation stabilizes BAZF mRNA via the 3′UTR. a HUVECs were transfected with a reporter vector with or without the 3′UTR of BAZF mRNA. Luciferase activity was assayed 18 h post-transfection and was normalized to the activity of thymidine kinase promoter-driven renilla luciferase. b The indicated reporter vector-transfected HUVECs were pretreated with Gö6976 for 30 min before PMA treatment and then stimulated with 10 nM PMA for 6 h. Luciferase activity was normalized to 18S rRNA levels and represented as the fold induction relative to unstimulated cells. c Reporter vector-transfected HUVECs were pretreated with actinomycin D before PMA stimulation. d siRNA targeting PRKD2 and the indicated reporter genes were sequentially transfected into HUVECs. Next, the HUVECs were stimulated with PMA for 6 h. e, f HUVECs were infected with mAG1 or, WT, CA, or KD PRKD2 cDNA-coding lentiviruses. Cells were then transfected with the indicated reporter 30 h later. The HUVECs were incubated without (e) or with (f) actinomycin D. The experiments were independently performed at least 3 times. * p < 0.05; ** p < 0.01; N.S. not significant
HSP90β binds BAZF mRNA and regulates mRNA stability
Previous reports have shown that 3′UTR binding proteins play a critical role in the regulation of mRNA stability [26, 27]. To identify molecules that bind to the 3′UTR of BAZF mRNA and regulate its stability, we precipitated BAZF mRNA binding proteins from HUVEC whole cell lysates with biotinylated BAZF mRNA and streptavidin-coated magnetic beads. The precipitated proteins were separated by SDS-PAGE and analyzed by mass spectrometry. We found that the interaction of HSP90β with BAZF mRNA increased after PMA stimulation (Fig. 5a). Other identified PMA-stimulation-dependent BAZF mRNA binding molecules were cytoskeleton-related proteins (data not shown). A recent report demonstrated that an HSP90 inhibitor destabilizes B cell chronic lymphocytic leukemia/lymphoma 6 (BCL6) mRNA [28], indicating that HSP90β may stabilize BCL6 mRNA; however, the details are still unclear. Therefore, we focused on analysis of HSP90β function in BAZF mRNA stabilization. To confirm binding of HSP90β to BAZF mRNA, we performed an immunoprecipitation (IP) assay using an anti-HSP90β antibody. The association of BAZF mRNA with HSP90β was monitored by RT-qPCR. The BAZF PCR product was enriched in the HSP90β IP sample in a VEGF-A-dependent manner, and an HSP90-specific inhibitor Pu-H71 (1 μM) [29] inhibited the enrichment of BAZF mRNA (Fig. 5b). To determine whether HSP90β was involved in stabilization of BAZF mRNA, we examined the effects of HSP90 inhibitors on BAZF mRNA stability after VEGF-A stimulation. HUVECs were treated with an HSP90-specific inhibitor Pu-H71 (1 μM) or 17-allylaminogeldanamycin (17-AAG; 1 μM) [30] for 30 min before exposure to VEGF-A. Both Pu-H71 and 17-AAG significantly inhibited BAZF mRNA stabilization by VEGF-A (Fig. 5c, d). Next, we performed an actinomycin D chase assay to investigate the effects of an HSP90β inhibitor on the t1/2 of BAZF mRNA. The t1/2 of BAZF mRNA significantly decreased from 1.72 to 0.74 h with Pu-H71 treatment (Fig. 5e). Next, we analyzed the influence of Pu-H71 on pCMV-luc-BAZF 3′UTR luciferase activity. Pu-H71 significantly inhibited PMA- and CA PRKD2-dependent luciferase activity of pCMV-luc-BAZF 3′UTR (Fig. 5f, g). These data suggested that HSP90β regulated the stability of BAZF mRNA through binding to the 3′UTR of BAZF mRNA in a VEGF-A/PRKD2 signaling pathway-dependent manner.
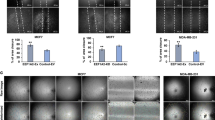
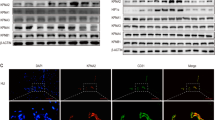
PRKD2 activation induces interaction of HSP 90β and BAZF mRNA. a HUVECs were treated with PMA for 2 h, and the lysate was precipitated with biotinylated BAZF mRNA. The precipitated proteins were separated by SDS-PAGE and identified by mass spectrometry. The arrow indicates HSP90β. b The cell lysate from HUVECs with the indicated treatments was incubated with anti-HSP90β. RNA was isolated from the immunoprecipitates, and β2-microgloblin (β2M) or BAZF mRNA was detected by RT-qPCR. The mRNA abundance was normalized by the HSP90β protein level which was analyzed by ImageJ software. The bottom panel shows the protein level of immunoprecipitated HSP90β in each cell lysate. c, d HUVECs were pretreated with Pu-H71 (1 μM) (c) or 17AAD (1 μM) (d). HSP90β-inhibitor-treated HUVECs were stimulated with VEGF-A for 2 h. e The half-life of BAZF mRNA was measured by incubating HUVECs with actinomycin D. Total RNA was collected at each of the indicated times after actinomycin D treatment, and BAZF mRNA levels were measured by RT-qPCR. Thirty minutes after actinomycin D pretreatment, HUVECs were treated with Pu-H71. Data were normalized to the 18S rRNA and represented as a percentage of the mRNA levels before actinomycin D treatment. (F) HUVECs transfected with reporter vectors were pretreated with Pu-H71 for 30 min and then exposed to 10 nM PMA for 6 h. Luciferase activity was assayed at 18 h post-transfection and was normalized as described above. Data were presented as the fold induction relative to the luciferase activity of the control. g mAG1 or CA PRKD2-expressing HUVECs were transfected with each reporter vector. HUVECs were treated with Pu-H71 at 6 h after transfection, and luciferase activity was measured at 18 h post-transfection. h Cell lysates from KD and CA PRKD2-expressing HUVECs were subjected to 2-dimensional gel electrophoresis. HSP90β and HSP27 were detected by appropriate antibodies. i HUVECs were infected with green fluorescence protein (GFP)- or BAZF-expressing adenoviruses at 250 m.o.i. Transfected cells were seeded on Matrigel with or without Pu-H71 treatment. The total length of each network was measured and calculated in Image-Pro Plus software (Roper). The experiments were independently performed at least 3 times. * p < 0.05; ** p < 0.01
To clarify whether HSP90β is a substrate of PRKD2, we performed 2-dimensional electrophoresis and western blotting of KD or CA PRKD2-expressing HUVEC lysates. The data showed that there was no major protein shift for HSP90β between KD and CA PRKD2-expressing HUVECs, while acidic spots representing HSP27 increased in CA PRKD2-expressing HUVEC (Fig. 5h). In an in vitro kinase assay, we also found that the phosphorylation level of HSP90β did not increase in a CA PRKD2-dependent manner (data not shown). To further elucidate the relationship between HSP90β and BAZF in angiogenesis, we performed a BAZF-expressing endothelial cell network formation assay using the HSP90 inhibitor Pu-H71. HUVECs were infected with adenovirus containing the cDNA of BAZF without the 3′UTR (AdBAZF) or of GFP (AdGFP). Adenovirus-infected HUVECs (250 m.o.i.) were seeded on Matrigel with or without Pu-H71. Network formation was completely inhibited by Pu-H71 in AdGFP-infected HUVECs (Fig. 5i). Moreover, the inhibitory effect of Pu-H71 on network formation was partially rescued by BAZF-overexpression. These data suggest that HSP90β may mediate BAZF expression via a downstream angiogenic pathway.
Discussion
VEGF-A induces PRKD2 activation to regulate BAZF mRNA stability
The PRKD family is composed of 3 members: PRKD1 (also known as PKCμ [31, 32]), PRKD2 [33], and PRKD3 (also known as PKCν [34]). PRKD1, 2, and 3 share a similar molecular structure, including a tandem repeat of zinc finger-like cysteine-rich motifs at the N-terminus that display high affinity for diacylglycerol (DAG) or phorbol esters, a pleckstrin homology domain, and a catalytic domain in the C-terminal region that shares homology with the calmodulin-dependent kinases [35–37]. PRKD1 is the best characterized isoform of the family and is activated by a variety of growth factors and other stimuli [31, 32]. However, the biological functions of PRKD2 and PRKD3 remain unclear. In a recent study, researchers demonstrated that activation of PRKD1 and PRKD2 was required for VEGF-induced phosphorylation of HSP27, which mediated the migration of ECs [38, 39]. Hao et al. [40] proposed that PRKD2, but not PRKD1, plays a pivotal role in EC proliferation and migration and is necessary for angiogenesis, demonstrating that VEGF-activated PRKD2 modulates VEGFR2 and FGFR1 expression. In our current study, we found that PRKD2 signaling, but not PRKD1 signaling, modulated the expression of pro-angiogenic factor BAZF. This finding describes a novel function for PRKD2 and also suggests that PRKD2 activation represents a critical angiogenic signaling molecule, in accordance with the hypothesis of Hao et al.
Numerous mRNAs containing AREs are involved in angiogenesis
AREs are critical cis-acting short sequences in the 3′UTR that assemble a set of RNA-binding proteins and regulate mRNA stability [41]. Close to 15 % of the human transcriptome is composed of ARE-containing mRNAs [42], which include functionally diverse groups, such as those involved in the inflammatory and immune response, transcription, cell proliferation, RNA metabolism, development, and signal transduction [43]. AREs in the 3′UTR of mRNAs coding angiogenic genes, such as VEGF [44–48], basic FGF (bFGF) [49], interleukin-8 (IL-8) [47], hypoxia induced factor-α (HIF1-α) [50], endothelial nitric oxide synthetase (eNOS) [51], and cyclooxygenase (COX-2) [47, 52–54], regulate mRNA stability. BAZF mRNA also has 3 AREs in the 3′UTR, and we showed that VEGF-induced BAZF gene expression is regulated by stabilization of mRNA (Fig. 3a). Indeed, the regulation of pro-angiogenic gene expression through mRNA stabilization seems to be essential for angiogenesis.
The stability of mRNA is determined by not only cis-acting elements, such as AREs, but also trans-acting RNA-binding proteins that regulate the decay of targeted mRNA. Human antigen R (HuR) has been the most extensively studied mRNA stability factor. HuR is a member of the mammalian homologs of embryonic lethal abnormal vision (ELAV) family proteins. HuR binds to the AREs of several mRNAs coding for angiogenic genes, such as VEGF, TNF-alpha, Cox-2, and IL-8, to inhibit mRNA decay [45, 47, 48, 55]. In this report, we also identified HuR as a BAZF mRNA-binding protein (data not shown). However, PMA stimulation did not affect the interaction of HuR with the 3′UTR of BAZF mRNA. Taken together, these data suggest that HuR may not be a central player in the regulation of angiogenic gene expression by PRKD2 activation.
HSP90β regulates mRNA stability of the pro-angiogenic factor BAZF via a VEGF/PRKD2 pathway in ECs
In the current study, we found that the BAZF 3′UTR interacted with HSP90β, which is regulated by VEGF-A. HSPs are molecular chaperones responsible for the maintenance of cell homeostasis and regulation of cell survival [56]. HSP90, a 90-kDa molecular chaperone, is one of many HSPs in eukaryotes and comprises 1–2 % of the total cellular protein under normal conditions [57–59]. HSP90 is the main functional component of a critical chaperone complex, and many tumors exhibit high expression of HSP90, allowing the activation of tumor-specific signaling pathways and buffering stress conditions in the tumor microenvironment [60]. Therefore, several groups have been working to develop HSP90 inhibitors to target a wide range of malignant tumors [61–63]. Additionally, HSP90 binds ATP in the N-terminal region. However, this ATP-binding function was not required in several chaperone assays, and ATP-nonbinding mutants, although compromised, retain some residual luciferase refolding activity [64]. Moreover, even though HSP90 contains an ATPase domain, the ATP-binding domain shows low intrinsic ATPase activity [65, 66]. These data from previous studies indicate that ATP interactions with HSP90 may play roles other than ATPase and protein refolding. In the current study, we demonstrated that the HSP90 inhibitors 17-AAG and Pu-H71 inhibited PRKD2-mediated stabilization of BAZF mRNA. 17-AAD binds to the ATP-binding pocket of the N-terminal region in HSP90 to associate with adenosine di- or triphosphate [67, 68]. On the other hand, purine scaffolds, such as Pu-H71, also interact with the ATP-recognition site in the N-terminal region of HSP90 [68]. Nucleotides such as mRNA could also interact with the N-terminal region of HSP90β, and these HSP90 inhibitors may compete with mRNA to bind to HSP90β.
A recent study revealed that HSP70 regulates the stability of Bim mRNA in a cytokine-dependent manner [69]. HSP70 binds to AREs in the 3′UTR of specific mRNAs and enhances the stability. Another report also indicated that HSP90 can stabilize BCL-6 mRNA and protein in lymphoma cells [28]. The 3′UTR of BCL-6 also has several AREs. These studies suggest that HSP family proteins work not only as protein chaperones, but also as mRNA stabilizers.
Phosphorylation of serine and/or threonine residues on HSP90 has been shown to regulate HSP90 activity to induce the conformational maturation of oncogenic proteins such as ErbB2, c-Src, Akt, and Raf-1 [70, 71]. HSP90β has several potential PRKD phosphorylation sites. However, 2-dimensional gel electrophoresis revealed that the phosphorylation state of HSP90β did not change with either CA or KD PRKD2 overexpression (Fig. 5h). HSP90β may be indirectly regulated by PRKD2 activation. As a next step to understanding PRKD2-HSP90β-regulated mRNA stabilization, we need to identify all of the components complexed with HSP90β after PRKD2 activation.
Scheme of the molecular mechanism. VEGF-A stimulation induces activation of PRKD2 through VEGFR activation. Activation of PRKD2 leads to recruitment of several molecules, including HSP90β, to the 3′UTR of BAZF mRNA, inhibiting the decay of mRNA. As a result, increased BAZF translation promotes angiogenesis progression
In this report, we found that pro-angiogenic BAZF mRNA stability was regulated by a PRKD2-HSP90β-dependent pathway (Fig. 6). The current study suggested that activated ECs could quickly enter into an angiogenic state from angiogenic quiescence through stabilization of mRNA coding for pro-angiogenic genes, such as BAZF. Further studies on the molecular mechanisms of this mRNA stabilization system may help to identify new targets for the treatment of angiogenesis-associated diseases.
References
Folkman J (2007) Angiogenesis: an organizing principle for drug discovery? Nat Rev Drug Discov 6:273–286
Hanahan D, Folkman J (1996) Patterns and emerging mechanisms of the angiogenic switch during tumorigenesis. Cell 86:353–364
Dvorak HF (2002) Vascular permeability factor/vascular endothelial growth factor: a critical cytokine in tumor angiogenesis and a potential target for diagnosis and therapy. J Clin Oncol 20:4368–4380
Hicklin DJ, Ellis LM (2005) Role of the vascular endothelial growth factor pathway in tumor growth and angiogenesis. J Clin Oncol 23:1011–1027
Hellstrom M, Phng LK, Hofmann JJ, Wallgard E, Coultas L, Lindblom P, Alva J, Nilsson AK, Karlsson L, Gaiano N, Yoon K, Rossant J, Iruela-Arispe ML, Kalen M, Gerhardt H, Betsholtz C (2007) Dll4 signalling through Notch1 regulates formation of tip cells during angiogenesis. Nature 445:776–780
Jakobsson L, Franco CA, Bentley K, Collins RT, Ponsioen B, Aspalter IM, Rosewell I, Busse M, Thurston G, Medvinsky A, Schulte-Merker S, Gerhardt H (2010) Endothelial cells dynamically compete for the tip cell position during angiogenic sprouting. Nat Cell Biol 12:943–953
Benedito R, Roca C, Sorensen I, Adams S, Gossler A, Fruttiger M, Adams RH (2009) The notch ligands Dll4 and Jagged1 have opposing effects on angiogenesis. Cell 137:1124–1135
Benedito R, Trindade A, Hirashima M, Henrique D, da Costa LL, Rossant J, Gill PS, Duarte A (2008) Loss of Notch signalling induced by Dll4 causes arterial calibre reduction by increasing endothelial cell response to angiogenic stimuli. BMC Dev Biol 8:117
Leslie JD, Ariza-McNaughton L, Bermange AL, McAdow R, Johnson SL, Lewis J (2007) Endothelial signalling by the Notch ligand Delta-like 4 restricts angiogenesis. Development 134:839–844
Lobov IB, Renard RA, Papadopoulos N, Gale NW, Thurston G, Yancopoulos GD, Wiegand SJ (2007) Delta-like ligand 4 (Dll4) is induced by VEGF as a negative regulator of angiogenic sprouting. Proc Natl Acad Sci USA 104:3219–3224
Noguera-Troise I, Daly C, Papadopoulos NJ, Coetzee S, Boland P, Gale NW, Lin HC, Yancopoulos GD, Thurston G (2006) Blockade of Dll4 inhibits tumor growth by promoting non-productive angiogenesis. Nature 444:1032–1037
Ridgway J, Zhang G, Wu Y, Stawicki S, Liang WC, Chanthery Y, Kowalski J, Watts RJ, Callahan C, Kasman I, Singh M, Chien M, Tan C, Hongo JA, de Sauvage F, Plowman G, Yan M (2006) Inhibition of Dll4 signalling inhibits tumor growth by deregulating angiogenesis. Nature 444:1083–1087
Sainson RC, Aoto J, Nakatsu MN, Holderfield M, Conn E, Koller E, Hughes CC (2005) Cell-autonomous notch signaling regulated endothelial cell branching and proliferation during vascular tubulogenesis. FASEB J 19:1027–1029
Siekmann AF, Lawson ND (2007) Notch signalling limits angiogenic cell behaviour in developing zebrafish arteries. Nature 445:781–784
Suchting S, Freitas C, le Noble F, Benedito R, Breant C, Duarte A, Eichmann A (2007) The Notch ligand Delta-like 4 negatively regulates endothelial tip cell formation and vessel branching. Proc Natl Acad USA 104:3225–3230
Tammela T, Zarkada G, Wallgard E, Murtomaki A, Suchting S, Wirzenius M, Waltari M, Hellstrom M, Schomber T, Peltonen R, Freitas C, Duarte A, Isoniemi H, Laakkonen P, Christofori G, Yla-Herttuala S, Shibuya M, Pytowski B, Eichmann A, Betsholtz C, Alitalo K (2008) Blocking VEGFR-3 suppresses angiogenic sprouting and vascular network formation. Nature 454:656–660
Caolo V, van den Akker NM, Verbruggen S, Donners MM, Swennen G, Schulten H, Waltenberger J, Post MJ, Molin DG (2010) Feed-forward signaling by membrane-bound ligand receptor circuit: the case of NOTCH DELTA-like 4 ligand in endothelial cells. J Biol Chem 285:40681–40689
Roukens MG, Alloul-Ramdhani M, Baan B, Kobayashi K, Peterson-Maduro J, van Dam H, Schulte-Merker S, Baker DA (2010) Control of endothelial sprouting by a Tel-CtBP complex. Nat Cell Biol 12:933–942
Eilken HM, Adams RH (2010) Dynamics of endothelial cell behavior in sprouting angiogenesis. Curr Opin Cell Biol 22:617–625
Ohnuki H, Inoue H, Takemori N, Nakayama H, Sakaue T, Fukuda S, Miwa D, Nishiwaki E, Hatano M, Tokuhisa T, Endo Y, Nose M, Higashiyama S (2012) BAZF, a novel component of cullin3-based E3 ligase complex, mediates VEGFR and Notch cross-signaling in angiogenesis. Blood 119:2688–2698
Dou GR, Wang YC, Hu XB, Hou LH, Wang CM, Xu JF, Wang YS, Liang YM, Yao LB, Yang AG, Han H (2008) RBP-J, the transcription factor downstream of Notch receptors, is essential for the maintenance of vascular homeostasis in adult mice. FASEB J. 22:1606–1617
Irie A, Harada K, Tsukamoto H, Kim JR, Araki N, Nishimura Y (2006) Protein kinase D2 contributes to either IL-2 promoter regulation or induction of cell death upon TCR stimulation depending on its activity in Jurkat cells. Int Immunol 18:1737–1747
Miura N, Takemori N, Kikugawa T, Tanji N, Higashiyama S, Yokoyama M (2012) Adseverin: a novel cisplatin-resistant marker in the human bladder cancer cell line HT1376 identified by quantitative proteomic analysis. Mol Oncol 6:311–322
Tenenbaum SA, Lager PJ, Carson CC, Keene JD (2002) Ribonomics: identifying mRNA subsets in mRNP complexes using antibodies to RNA-binding proteins and genomic arrays. Methods 26:191–19825
Yang E, van Nimwegen E, Zavolan M, Rajewsky N, Schroeder M, Magnasco M, Darnell JE Jr (2003) Decay of human mRNAs: correlation with functional characteristics and sequence attributes. Genome Res 13:1863–1872
Barreau C, Paillard L, Osborne HB (2005) AU-rich elements and associated factors: are there unifying principles? Nucleic Acids Res 33:7138–7150
Khabar KS (2010) Post-transcriptional control during chronic inflammation and cancer: a focus on AU-rich elements. Cell Mol Life Sci 67:2937–2955
Cerchietti LC, Lopes EC, Yang SN, Hatzi K, Bunting KL, Tsikitas LA, Mallik A, Robles AI, Walling J, Varticovski L, Shaknovich R, Bhalla KN, Chiosis G, Melnick A (2009) A purine scaffold Hsp90 inhibitor destabilizes BCL-6 and has specific antitumor activity in BCL-6-dependent B cell lymphomas. Nat Med 15:1369–1376
He H, Zatorska D, Kim J, Aguirre J, Liauger L, She Y, Wu N, Immormino RM, Gewirth DT, Chiosis G (2006) Identification of potent water soluble purine-scaffold inhibitors of the heat shock protein 90. J Med Chem 49:381–390
Whitesell L, Mimnaugh EG, De Costa B, Myers CE, Neckers LM (1994) Inhibition of heat shock protein HSP90-pp 60v-src heteroprotein complex formation by benzoquinone ansamycins: essential role for stress proteins in oncogenic transformation. Proc Natl Acad Sci USA 91:8324–8328
Johannes FJ, Prestle J, Eis S, Oberhagemann P, Pfizenmaier K (1994) PKCμ is a novel, atypical member of the protein kinase C family. J Biol Chem 269:6140–6148
Valverde AM, Sinnett-Smith J, Van Lint J, Rozengurt E (1994) Molecular cloning and characterization of protein kinase D: a target for diacylglycerol and phorbol esters with a distinctive catalytic domain. Proc Natl Acad Sci USA 91:8572–8576
Sturany S, Van Lint J, Muller F, Wilda M, Hameister H, Hocker M, Brey A, Gern U, Vandenheede J, Gress T, Adler G, Seufferlein T (2001) Molecular cloning and characterization of the human protein kinase D2. A novel member of the protein kinase D family of serine threonine kinases. J Biol Chem 276:3310–3318
Hayashi A, Seki N, Hattori A, Kozuma S, Saito T (1999) PKCnu, a new member of the protein kinase C family, composes a fourth subfamily with PKCmu. Biochim Biophys Acta 1450:99–106
Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S (2002) The protein kinase complement of the human genome. Science 298:1912–1934
Rykx A, De Kimpe L, Mikhalap S, Vantus T, Seufferlein T, Vandenheede JR, Van Lint J (2003) Protein kinase D: a family affair. FEBS Lett 546:81–86
Rozengurt E, Rey O, Waldron RT (2005) Protein kinase D signaling. J Biol Chem 280:13205–13208
Rousseau S, Houle F, Landry J, Huot J (1997) p38 MAP kinase activation by vascular endothelial growth factor mediates actin reorganization and cell migration in human endothelial cells. Oncogene 15:2169–2177
Evans IM, Britton G, Zachary IC (2008) Vascular endothelial growth factor induces heat shock protein (HSP) 27 serine 82 phosphorylation and endothelial tubulogenesis via protein kinase D and independent of p38 kinase. Cell Signal 20:1375–1384
Hao Q, Wang L, Zhao ZJ, Tang H (2009) Identification of protein kinase D2 as a pivotal regulator of endothelial cell proliferation, migration, and angiogenesis. J Biol Chem 284:799–806
Chen CY, Shyu AB (1995) AU-rich elements: characterization and importance in mRNA degradation. Trends Biochem Sci 20:465–470
Halees AS, El-Badrawi R, Khabar KS (2008) ARED Organism: expansion of ARED reveals AU-rich element cluster variations between human and mouse. Nucleic Acids Res 36:D137–D140
Bakheet T, Williams BR, Khabar KS (2006) ARED 3.0: the large and diverse AU-rich transcriptome. Nucleic Acids Res 34:D111–D114
Claffey KP, Shih SC, Mullen A, Dziennis S, Cusick JL, Abrams KR, Lee SW, Detmar M (1998) Identification of a human VPF/VEGF 3′ untranslated region mediating hypoxia-induced mRNA stability. Mol Biol Cell 9:469–481
Levy NS, Chung S, Furneaux H, Levy AP (1998) Hypoxic stabilization of vascular endothelial growth factor mRNA by the RNA-binding protein HuR. J Biol Chem 273:6417–6423
King PH (2000) RNA-binding analyses of HuC and HuD with the VEGF and c-myc 3′-untranslated regions using a novel ELISA-based assay. Nucleic Acids Res 28:E20
Dixon DA, Tolley ND, King PH, Nabors LB, McIntyre TM, Zimmerman GA, Prescott SM (2001) Altered expression of the mRNA stability factor HuR promotes cyclooxygenase-2 expression in colon cancer cells. J. Clin. Invest. 108:1657–1665
Onesto C, Berra E, Grepin R, Pages G (2004) Poly(A)-binding protein-interacting protein 2, a strong regulator of vascular endothelial growth factor mRNA. J Biol Chem 279:34217–34226
Touriol C, Morillon A, Gensac MC, Prats H, Prats AC (1999) Expression of human fibroblast growth factor 2 mRNA is post-transcriptionally controlled by a unique destabilizing element present in the 3′-untranslated region between alternative polyadenylation sites. J Biol Chem 274:21402–21408
Kim TW, Yim S, Choi BJ, Jang Y, Lee JJ, Sohn BH, Yoo HS, Yeom YI, Park KC (2010) Tristetraprolin regulates the stability of HIF-1alpha mRNA during prolonged hypoxia. Biochem Biophys Res Commun 391:963–968
Lai PF, Mohamed F, Monge JC, Stewart DJ (2003) Downregulation of eNOS mRNA expression by TNFalpha: identification and functional characterization of RNA-protein interactions in the 3′UTR. Cardiovasc Res 59:160–168
Ristimaki A, Narko K, Hla T (1996) Down-regulation of cytokine-induced cyclo-oxygenase-2 transcript isoforms by dexamethasone: evidence for post-transcriptional regulation. Biochem. J. 318(Pt 1):325–331
Gou Q, Liu CH, Ben-Av P, Hla T (1998) Dissociation of basal turnover and cytokine-induced transcript stabilization of the human cyclooxygenase-2 mRNA by mutagenesis of the 3′-untranslated region. Biochem Biophys Res Commun 242:508–512
Xu K, Robida AM, Murphy TJ (2000) Immediate-early MEK-1-dependent stabilization of rat smooth muscle cell cyclooxygenase-2 mRNA by Galpha(q)-coupled receptor signaling. J Biol Chem 275:23012–23019
Sengupta S, Jang BC, Wu MT, Paik JH, Furneaux H, Hla T (2003) The RNA-binding protein HuR regulates the expression of cyclooxygenase-2. J Biol Chem 278:25227–25233
Takayama S, Reed JC, Homma S (2003) Heat-shock proteins as regulators of apoptosis. Oncogene 22:9041–9047
Pearl LH, Prodromou C (2001) Structure, function, and mechanism of the Hsp90 molecular chaperone. Adv Protein Chem 59:157–186
Picard D (2002) Heat-shock protein 90, a chaperone for folding and regulation. Cell Mol Life Sci 59:1640–1648
Pratt WB, Toft DO (2003) Regulation of signaling protein function and trafficking by the hsp90/hsp70-based chaperone machinery. Exp Biol Med (Maywood) 228:111–133
Whitesell L, Bagatell R, Falsey R (2003) The stress response: implications for the clinical development of hsp90 inhibitors. Curr Cancer Drug Targets 3:349–358
Isaacs JS (2005) Heat-shock protein 90 inhibitors in antineoplastic therapy: is it all wrapped up? Expert Opin Investig Drugs 14:569–589
Powers MV, Workman P (2006) Targeting of multiple signaling pathways by heat shock protein 90 molecular chaperone inhibitors. Endocr Relat Cancer 13(Suppl 1):S125–S135
Neckers L (2002) Hsp90 inhibitors as novel cancer chemotherapeutic agents. Trends Mol Med 8:S55–S61
Grenert JP, Johnson BD, Toft DO (1999) The importance of ATP binding and hydrolysis by hsp90 in formation and function of protein heterocomplexes. J Biol Chem 274:17525–17533
Obermann WM, Sondermann H, Russo AA, Pavletich NP, Hartl FU (1998) In vivo function of Hsp90 is dependent on ATP binding and ATP hydrolysis. J Cell Biol 143:901–910
Weikl T, Muschler P, Richter K, Veit T, Reinstein J, Buchner J (2000) C-terminal regions of Hsp90 are important for trapping the nucleotide during the ATPase cycle. J Mol Biol 303:583–592
Isaacs JS, Xu W, Neckers L (2003) Heat shock protein 90 as a molecular target for cancer therapeutics. Cancer Cell 3:213–217
Whitesell L, Lindquist SL (2005) HSP90 and the chaperoning of cancer. Nat Rev Cancer 5:761–772
Matsui H, Asou H, Inaba T (2007) Cytokines direct the regulation of Bim mRNA stability by heat-shock cognate protein 70. Mol Cell 25:99–112
Mimnaugh EG, Worland PJ, Whitesell L, Neckers LM (1995) Possible role for serine/threonine phosphorylation in the regulation of the heteroprotein complex between the hsp90 stress protein and the pp 60v tyrosine kinase. J Biol Chem 270:28654–28659
Zhao YG, Gilmore R, Leone G, Coffey MC, Weber B, Lee PW (2001) Hsp90 phosphorylation is linked to its chaperoning function. Assembly of the retrovirus cell attachment protein. J Biol Chem 276:32822–32827
Acknowledgments
We would like to thank Drs. Yoshimura (Keio University), Nishimura (Kumamoto University), and Miyoshi (RIKEN BioResource Center) for kindly providing the plasmids used in this study. We gratefully appreciate discussions with Drs. Matsushita, Ohnuki, and Nakayama of Ehime University. This study was supported by a grant from Ehime University (D.M.: 054402060) and a Grant-in-Aid for Scientific Research (23112513 to S. Higashiyama, 22590501 to H. Inoue) from the Ministry of Education, Culture, Sports, Science, and Technology, Japan.
Conflict of interest
The authors declare that there are no conflicts of interest that would prejudice the impartiality of this scientific work.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Miwa, D., Sakaue, T., Inoue, H. et al. Protein kinase D2 and heat shock protein 90 beta are required for BCL6-associated zinc finger protein mRNA stabilization induced by vascular endothelial growth factor-A. Angiogenesis 16, 675–688 (2013). https://doi.org/10.1007/s10456-013-9345-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10456-013-9345-x