Abstract
Micro- and nanofabrication has allowed the production of ultra-sensitive, portable, and inexpensive biosensors. These devices generally rely on chemical or biological receptors which recognize a particular compound of interest and relay this recognition event effectively by transduction. Recent advances in RNA and DNA synthesis have enabled the use of aptamers, in vitro generated oligonucleotides, which offer high affinity biomolecular recognition to a theoretically limitless variety of analytes. DNA and RNA aptamers have gained so much attention in the biosensor community, that they have begun competing with more established affinity ligands including enzymes, lectins, and most notably, immunoreceptors such as antibodies. This article reviews the current state-of-the-art of aptasensors, or biosensors that use aptamers as molecular recognition elements, emphasizing the synergy between aptamer-based biosensing and micro- and nanotechnology. Aptasensors developed on micro- and nanoscale platforms based on mass changes, electroanalytical techniques, optical transduction, and purification and separation methods will be covered.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Through the recent scientific efforts in genomics and proteomics, better understanding has been gained of the pathology and mechanisms of disease. In doing so, scientists have also become further experienced with important physiological components such as DNA, RNA, proteins and small biological molecules, all of which have shown to be the conduits of disease progression. Identification as well as thorough investigation of a disease-specific biomarker may prove vital, at an early stage of a disease, in treating it successfully. Unfortunately for medical practitioners, disease related biomarkers are either physiologically present in minute quantity or severely contaminated by non-specific compounds in a patient’s bloodstream or body fluid. Hence, highly sensitive as well as specific recognition elements are required for effective detection of such biomarkers. Micro- and nanotechnology has shown potential in providing innovative paradigm-breaking solutions to a variety of sensing applications (Dittrich et al. 2006). In particular, this potential has been well demonstrated for sensors in biology and medicine. Already, micro- and nanoscale sensors utilizing cantilevers (Shekhawat et al. 2006), electrochemical techniques (Morimoto et al. 2007), quantum dots, and metallic nanoparticles (Wernette et al. 2008) are showing potential for effective detection of biological compounds.
This article will review emerging advances in micro- and nanoscale aptasensors, which are biosensors that exploit aptamers, or affinity binding polynucleotides, for the sensitive and specific detection of biological analytes. A biosensor is a device used to detect the presence of and in particular cases, analyze a given target analyte. Typically, they consist of a molecular recognition component, a transducer changing the binding event into a measurable physical signal (e.g., mass, charge, heat or light), and a signal acquisition system. As an important class of biosensors, aptasensors use aptamers as highly selective recognition elements. As receptor molecules, aptamers allow widespread applicability to a diverse array of target analytes due to their generation process which involves exposing analytes to large DNA or RNA pools, offer enhanced stability at room temperature, and are easily synthesized and chemically modified at terminal sites so as to facilitate attachment to stationary surfaces. Aptasensors realized on micro- and nanoscale platforms afford many potential advantages, such as miniaturized construction, rapid, sensitive and specific detection, high throughput, reduced costs and minimized material consumption. Thus, micro- and nanoaptasensors are highly attractive to a broad range of applications, such as proteomics, metabolomics, environmental monitoring, counter-terrorism, and clinical diagnostics and therapeutics.
2 Aptamers as high affinity and specificity receptors
Aptamers are a class of high-affinity molecules derived from ribonucleic (RNA) or deoxyribonucleic (DNA) acids. They are generated by a synthetic process termed systematic evolution of ligands by exponential enrichment (SELEX), which was first described in 1990 (Ellington and Szostak 1990; Tuerk and Gold 1990). In this process, aptamers are isolated against an analyte of interest from large pools of oligonucleotides via affinity partitioning and amplification in an iterative fashion (Fig. 1). The SELEX method has permitted the synthesis of highly unique aptamers from very large populations of random oligomers that recognize molecules with exceptional affinity and specificity. With such abundance offered by random oligonucleotide sequences, the isolation of aptamers with affinity for a large variety of molecules is readily possible. High-affinity aptamers have been selected against a wide range of targets such as organic dyes (Ellington and Szostak 1992), amino acids (Geiger et al. 1996), antibiotics (Schurer et al. 2001), peptides (Nieuwlandt et al. 1995), proteins (Lupold et al. 2002) and whole cells (Herr et al. 2006; Shamah et al. 2008).
An aptamer can be designed to bind to a target analyte with high specificity by way of precise stacking of moieties, specific hydrogen bonding, and target binding-induced folding to their target molecule. With such complex conformational changes upon binding to analytes, aptamers can incorporate small molecules into their nucleic acid structure or integrate into the structure of larger molecules such as proteins by forming secondary structures that match the corresponding epitope(s) on the target (Hermann and Patel 2000). This creates high affinity interactions between aptamers and their targets exhibited by their relatively low dissociation constants, which generally start at the low micromolar and approach the picomolar level (in some instances). This is comparable to that of antibodies (Jayasena 1999). These innate binding mechanisms allow aptamers to distinguish between their target analytes and non-specific molecules by as subtle a variation as chirality, functional end group (e.g., methyl or hydroxyl), or isoform.
When compared to more conventional high-affinity ligands such as antibodies or enzymes, aptamers afford many advantages when used as affinity receptors. First, DNA and RNA aptamers are more stable at room temperature (leading to longer shelf-lives), easily synthesized and chemically modified at terminal sites so as to facilitate attachment to stationary surfaces (Jayasena 1999; Cho et al. 2004; Langer et al. 1981). Second, aptamer complexes are generally reversible, and the binding between an aptamer and its specific target is influenced by environmental stimuli such as solution pH, electric field, electromagnetic radiation, and temperature (Jayasena 1999). The stimuli-dependent binding reversibility is highly attractive; for example, this can enable controlled release and recovery of analytes in practical applications. This attribute also allows aptamer-functionalized sensor surfaces to be regenerated much more easily than antibody-based surfaces. Third, in contrast to antibodies that have to be produced in vivo, aptamers are synthesized by in vitro methods. This eliminates the need for laboratory animals in receptor development, achieves a minimal variability for batch-to-batch aptamer production, and allows for designed modifications to enhance stability, affinity, and specificity. Finally, aptamers can be made to bind to a specific region of the target and with specific binding properties in different binding conditions. These regions can be particular functional domains of the target, e.g., substrate binding pockets or allosteric sites, thereby manipulating the biological function of the analyte after extraction. This is in contrast to antibodies, where in vivo isolation means that it is the animal immune system that selects the sites on the target to which the antibodies bind. The potential utilization of this property is substantiated by therapeutic applications of aptamers (Nimjee et al. 2005).
3 Aptamer generation and the SELEX process
The selection of aptamers for a specific target is based on the SELEX procedure (Ellington and Szostak 1990; Tuerk and Gold 1990) which is illustrated in Fig. 1. The SELEX process begins with an oligonucleotide library, usually consisting of approximately 1014–1016 oligonucleotide strands with a random sequence of defined size, from which binding sequences for a target analyte are searched. An advantage of the SELEX process is that while this starting set (the library) is relatively sparse compared to the entire sequence space (1030–1060 sequences), aptamer generation against a wide variety of targets is still successful. The amount of searchable sequence space can be expanded by utilizing techniques such as error-prone PCR (Gram et al. 1992) and DNA shuffling (Stemmer 1994). For RNA-derived aptamers, a DNA library is initially transcribed into an RNA pool, which serves the same function. Following incubation of the library with target molecules, a small fraction of nucleic acids with affinity towards the target analytes (including surface-binding non-specific nucleic acids) become bound, and the rest of the library sequences are washed away using one of several separation techniques such as solid phase extraction (SPE) or capillary electrophoresis (CE) (Mendonsa and Bowser 2004; Mosing et al. 2005). Binding sequences can then be eluted by altering the environmental conditions of the solution. In order to remove non-specific binders, the standard SELEX process is often supplemented with techniques such as counter-selection or negative selection (Brown and Gold 1995), in which secondary incubation steps expose the library only to surfaces present during primary incubation. These non-specific binders are then removed from the DNA or RNA pool. Following incubation, nucleic acids are amplified by the polymerase chain reaction (PCR) to yield a mixture of oligomers with variable affinity towards the target analyte. Iteration of the above protocol results in the isolation of a pool of oligomers displaying sequential motifs, which after (typically) 8–15 iterative SELEX runs converge to one or possibly a few binding sequences. The detailed association of aptamers to their targets is based on generating three dimensional structures that involve forming loops and hybridized regions which utilize complementary Watson–Crick base-pairing (Ellington and Szostak 1992). The final identification of the aptamer composition by sequencing and elucidation of the aptamer–substrate tertiary structure by NMR spectroscopy (Feigon et al. 1996) leads then to the selection of a high-affinity binding material from a random composite. The most notable attribute of the SELEX method of synthesizing aptamers is the use of PCR amplification, which grants an exponentially increasing number of replicated aptamers, limited only by the amount of starting material. This is unlike cultivating the more standard affinity receptors such as antibodies and enzymes, which rely on the use of immunological response from living organisms.
Just as counter- and negative selection can be used to remove non-specific binders from the oligomer pool, other techniques can be adopted as part of a composite SELEX procedure that defines specific aspects of the final sequence pool. By changing the number of SELEX cycles, or by “toggling” the use of different target molecules (Toggle–SELEX) (White et al. 2001), the final pool can be tailored to have a particular degree of selectivity to a number of different targets. In addition, recent work has shown the ability to select “smart aptamers,” or sequences selected with predetermined binding properties such as specific equilibrium dissociation constant (K D). If even more detailed control is required over binding activity, aptamers with specific dynamic association and dissociation constants can be partitioned from a pool of oligonucleotide sequences (Drabovich et al. 2006). Additionally, there is active effort to develop aptamers with specific thermodynamic properties such as change in entropy and enthalpy (Berezovski and Krylov 2005).
4 Aptasensors and their applications
Biological sensing depends greatly on the selectivity and specificity of the receptor molecule to the analyte in question. Aptamers can be used as molecular recognition elements that afford these attributes. As mentioned above, the application of aptamers as sensing receptors offers advantages over methods that are mainly based on standard affinity receptors (e.g., antibodies), such as the possibility of easily regenerating the functionality of immobilized aptamers, their in vitro preparation, and the possibility of using different detection methods due to easy labeling. In the following sections, an overview on the different biosensors realized using aptamers as biorecognition elements is given, taking into consideration also the different transducers used for the development of the biosensors.
4.1 Aptasensors based on detecting mass changes
Sensing methods using mass-based transduction mechanisms have provided an attractive platform for integrating microchip technology with aptamer receptors. Thus, many aptasensors that detect mass changes have recently emerged, including microcantilever, gravimetric, and surface-acoustic wave-based sensors. Large molecular weight molecules and compounds such as proteins, DNA and RNA, or even cells are commonly detected with aptasensors incorporating mass change transduction. Their advances are reviewed in this section.
4.1.1 Micro- and nanocantilever aptasensors
Analytes can be detected by measuring cantilever deflection; a micro- or nanocantilever is functionalized with a receptor molecule, and binding of an analyte by the receptor causes changes in surface stress. This surface stress change causes a deflection of the cantilever, which can then be measured. (de-los-Santos-Alvarez et al. 2008). Due to their miniature dimensions, micro- and nanocantilevers are responsive to extremely small surface stress changes (roughly 10−3 N/m; or, force per unit width), and hence are capable of highly sensitive detection of molecular binding events. Current aptasensors along these lines mostly employ microcantilevers. By employing aptamers as immobilized receptor molecules, the selectivity of microcantilevers toward specific analytes can be improved since non-specific adsorption can be reduced. The work of Savran et al. (2002, 2003, 2004) was the first attempt at such a device (Fig. 2). The sensor cantilever was functionalized with a thiol-modified aptamer specific for Thermus aquaticus (Taq) DNA polymerase. A non-specific oligonucleotide sequence was immobilized on an adjacent reference cantilever. Subsequently, an interrogating light source was focused on interdigitated fingers in between the reference and sensor cantilever creating an observable diffraction pattern. This allowed differential interferometry measurements during binding-induced bending of the sensor cantilever. The system was characterized in terms of sensitivity performing binding experiments at seven different Taq concentrations. Additionally, regeneration of the sensor with a solution of 7 M urea was shown. From these experiments, a K D of 15 pM was calculated for the DNA aptamer and Taq. Moreover, the specificity of the system was confirmed as 75 nM of thrombin was introduced to the cantilever surface to produce no significant signal. Furthermore, the sensor was demonstrated to maintain the ability of detecting the target protein also in a complex mixture, such as E. coli lysate, with comparable signals relative to standard binding conditions.
Schematic of a differential deflection microcantilever utilizing aptamer receptors (adapted from Savran et al. 2004; Copyright permission is being requested). The sensor and reference cantilevers are supported by L-shaped thick structures that connect them to the die. Differential bending is measured using interferometry
More recently, Hwang and Nishikawa (2006) demonstrated sensing of hepatitis C virus (HCV) using an RNA aptamer functionalized onto silicon nitride microcantilevers. The microcantilevers were operated in vibration mode for sensing in which oscillations were induced using a combination of Pt electrodes and piezoelectric (PZT) layers. As HCV was introduced to the vibrating cantilevers, the dynamic response (i.e., frequency shift) was recorded. To this effect, a sensing limit of 100 pg/mL of HCV could be achieved by the device. Additionally, the work calculates the generated surface stress, due to HCV binding to aptamer, on the microcantilever. In the latest work along the lines of a microcantilever aptasensor (Cha et al. 2008), a thin, gold-coated membrane transducer (TMT) fabricated from PDMS on a metal-patterned glass substrate was employed to detect both DNA hybridization and protein recognition. As molecular interactions occurred on the membrane surface, stress-induced deflection was measured by a compact capacitive circuit. Matched and mismatched hybridizations of an immobilized 16-mer DNA oligonucleotide were presented. While the mismatched sample caused little capacitance change, the perfectly matched sample caused a well-defined capacitance decrease versus time due to an upward deformation of the membrane by a compressive surface stress. Additionally, the TMT demonstrated the single nucleotide polymorphism (SNP) capabilities which enabled a detection of mismatching base pairs in the middle of the sequence. Interestingly, this SNP event caused an increase in capacitance (a downward deflection due to tensile stress). Furthermore as a proof-of-concept, thrombin was detected with a 15-mer thrombin specific aptamer.
4.1.2 Acoustic-wave aptasensors
Biochemical binding or adsorption at an interface causes mass changes which can be directly transduced by gravimetric devices using piezoelectric materials. For example, the quartz crystal microbalance (QCM) measures the frequency variation caused by adsorption of a mass on the surface of a crystal oscillator. The quartz is sandwiched between two piezoelectric electrodes on which biomolecules are immobilized. Using this setup, the recognition event can be followed in real time and is compatible with flow-through measurements.
The earliest attempt at combining aptamer and QCM technology was reported by Furtado et al. (1999). A QCM sensor was used to study the interactions of HIV-1 Tar RNA aptamer with Tat protein, which is responsible for the HIV-1 RNA virus infection. Biotinylated Tar RNA was immobilized on the crystal sensor surface using the biotin-neutravidin interaction, and incorporated in an on-line system. Binding of two peptide fragments of Tat on the Tar RNA resulted in a change in both resonance frequency and motional resistance. A second system directly compared the performances obtained using a DNA aptamer and the antibody specific for human immunoglobulin E (IgE) chosen as target analyte (Liss et al. 2002). Both of the receptors were covalently attached on gold-coated QCM and the binding to the IgE target was studied in solution. Comparable detection limits were observed for both the aptamer and antibody (0.5 nM of IgE). However, the linear detection range was tenfold higher when aptamers were used. Additionally, the stability and regeneration characteristics of the aptamer sensor variant proved longer (several weeks) and more complete, respectively.
More recently, work involving aptasensors which utilize Love waves, a particular type of surface acoustic wave, have been used in combination with miniaturization technology (Jung et al. 2007; Schlensog et al. 2004). This is perhaps due to their high sensitivity, which is a consequence of the minimization of the energy loss when working in aqueous solutions. Specifically, these sensors used interdigitated electrodes (chrome/gold/chrome) acting as transmitters which are deposited on top of a piezoelectric crystal (e.g., crystalline silicon dioxide), which acts as a receiver. The frequencies applied to the transmitter cause mechanical stress in the crystal that generates an acoustic wave, which propagates across the surface of the crystal and is detected by the receiver. As the ligand binds to the aptamer (which is immobilized along the wave guide), a change in the resonant frequency or a phase shift occurs.
Taking advantage of the above aspects, Schlensog et al. (2004) and Jung et al. (2007) employed a microfluidic device, fabricated from polymethylmethacrylate on top of a quartz substrate, with aptamer recognition to detect multifunctional serine protease thrombin and Rev peptide (Fig. 3). The sensitivity of the Love-wave biosensor for thrombin and for Rev peptide was calculated by dividing the measured phase shift through the determined mass loading, resulting in 419 and 390 μg/cm2 with a limit of detection of about 72 and 77 pg/cm2 for thrombin and rev peptide, respectively. The authors additionally pointed out the possibility of using this aptamer-based Love-wave sensor in an array format, since the device was demonstrated to achieve highly sensitive analyte detection in parallel fashion. Lastly, a comparison study with the sensor against a commercial surface plasmon resonance (SPR) system (Biacore™ 3000) in terms of reproducibility and signal response proved successful, while the specificity of the system directly correlated with the specificity of the anti-thrombin aptamer for its ligand determined by classical methods, such as filter binding or gel shift assays.
4.2 Optical aptasensors
Optical aptasensors make use of the transmission of photons to signal target-binding. Many such sensors, such as molecular beacons, make use of chromophore modifications to the ends of oligonucleotides such as carboxyfluorescein (FAM) and Cyanine (Cy5, Cy3, etc.). Other techniques, such as surface plasmon resonance (SPR) and nanoparticle modifications do not require fluorescent labels for use. Optical signal transduction has been utilized in aptasensors designed for targets such as cocaine (Stojanovic and Landry 2002) and Tat protein (Yamamoto and Kumar 2000), as well as many others. Optical transduction does not place a limit on target selection, however, smaller targets are less likely to impede signal transmission.
4.2.1 Fluorescence-based molecular beacons, light-up probes, and microarrays
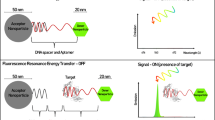
Molecular beacons are oligonucleotide probes designed to undergo structural changes in the presence of a target that will result in a measureable change in output fluorescence (Tyagi et al. 1998). Most beacons are designed with short sequence areas at the 3′ and 5′ termini that are self-complementary; the natural tendency of the oligonucleotide to fold upon itself creates a “hairpin” structure which can be used to bring termini labels such as fluorophores and quenchers into close proximity, which is necessary for their interaction (Fig. 4).
Aptasensing using a fluorescence-based molecular beacon. Initially, self-complementary portions of the oligonucleotide result in a hairpin structure which brings the fluorophore and the quencher into close proximity. Upon introduction of the target, binding-induced conformational change results in an increase in fluorescence
Some molecular beacons are designed as “signal-off,” producing a reduction in fluorescence upon conformational change (Stojanovic and Landry 2002; Ozaki et al. 2006; Stojanovic et al. 2001; Stojanovic and Kolpashchikov 2004). Others can generate increases in fluorescence, sometimes in the presence of a binding complementary strand (Yamamoto and Kumar 2000; Nutiu and Li 2003). Unfortunately, a consequence of dual-termini labeling is a lack of simple immobilization methods due to both ends of the strand being occupied. However, these same aptamer sequences could be immobilized and used as sensors if one end-label were removed and placed on a short complementary strand (Nutiu and Li 2003). In this case, the greater affinity of the aptamer toward its target over a short complementary strand causes the second label to be displaced, resulting in a change in fluorescent intensity. This also creates a free terminus on the primary aptamer strand which could be used for surface immobilization.
To increase throughput and experimental redundancy of conventional bioanalytical assays, microarrays utilizing aptamer receptors and fluorescence detection have been recently reported (Aberem et al. 2006; Bock et al. 2004; Brody et al. 1999; Cho et al. 2006; Collett et al. 2005; Kirby et al. 2004; Mascini et al. 2007; McCauley et al. 2003; Stadtherr et al. 2005; Wang et al. 2006a). Typically, these arrays are fabricated by robotically inking or patterning glass substrates with micrometer-sized spots in a visually addressable matrix. Each spot (or group of spots) consists of a distinct aptamer capable of selective recognition of a particular analyte. This is a similar concept to complementary DNA microarray technology used to increase the efficiency of genomic assays.
One of the first works contributing to aptamer-based microarray technology was reported by McCauley et al. (2003). Using a streptavidin functionalized glass coverslip, they manually spotted biotinylated aptamers specific to cancer-related proteins including inosine monophasphate dehydrogenase (IMPDH), vascular endothelial and basic fibroblast growth factors (VEGF and bFGF). By fluorescence anisotropy detection, all proteins were recognized down to 700 nM with extreme selectivity even in a milieu of complex biological matrices such as human serum or cellular extracts. A significant contribution to microarray research has originated from Ellington et al. (Cho et al. 2006; Collett et al. 2005; Kirby et al. 2004). In one instance where a robotic printer was used to array 400 RNA aptamer candidates onto a streptavidin coated glass slide, lysozyme and IgE proteins (tagged with either Cy3 or Cy5 fluorescence probes) were identified from lysed cells (Collett et al. 2005). Lysozyme could be detected across seven orders of magnitude with a limit of detection of 1 pg/ml. Most interestingly, an attempt was made to incorporate microfluidics and microfabrication technologies with aptamer microarray research (Kirby et al. 2004). A flow cell consisting of a silicon chip with multiple micromachined pyramidal wells was created through which fluid can pass. The flow cell was integrated with a fluid delivery system and coupled to a fluorescent microscope for detection (Fig. 5). Agarose microbeads, on which aptamers for ricin and lyzosyme were coated, were placed into each well. Solutions and analytes were delivered from the widest portion of the wells and transferred out the narrow openings. Ricin was detected at a concentration of 320 ng/ml, which was comparable to conventional antibody assays. Although the array used was relatively small in scale (3 × 3 or 4 × 3), the work demonstrated the feasibility of integrating microdevice technology in an arrayed format.
A schematic illustration showing the bead array chip employed by Kirby et al. (2004), which is packaged into a fluid flow cell. Inset is an SEM blow up of the rectangular-shaped micromachined wells of the aptamer chip array (beads included). The packaged assembly is suitable for both fluorometric (based on epi-fluorescence) and colorimetric (based on transmission) analyses. Adapted from Christodoulides et al. (2002); Copyright permission is being obtained
4.2.2 Colorimetric aptasensors using nanoparticles and quantum dots
Although the flourophore probes described above have been extensively used in biochemical sensing assays, their limitations including narrow excitation spectra, short fluorescence lifetimes, and reduced quantum yield have motivated alternative techniques. Currently, approaches utilizing micro- and nanotechnology, for example, nanoparticles and quantum dots have been explored to mitigate the disadvantages of conventional-scale molecular aptamer beacons and standard fluorescence probes. In principle, metallic nanoparticles and semiconductor quantum dots, when irradiated with electromagnetic radiation, either reflect and scatter light, as in the case with metallic nanoparticles or, emit light (varying wavelengths as a function of particle size) due to quantum confinement such as with quantum dots. Hence, when used in conjunction with aptamers, their high affinity and selectivity and near limitless spectral latitude have allowed benefits over traditional organic dyes on several counts, one of the most immediately obvious being brightness (owing to their high quantum yield) as well as their stability (much less photodestruction).
Colorimetric nanoparticles (Herr et al. 2006; Cho et al. 2006; Farokhzad et al. 2005; He et al. 2007; Li et al. 2007; Liu and Lu 2006; Liu et al. 2006; Pavlov et al. 2004; Polsky et al. 2006; Smith et al. 2007; Turney et al. 2004; Wang et al. 2006b; Wei 2007; Zhao et al. 2007; Zhu et al. 2006) and quantum dots (Bagalkot et al. 2007; Chen 2008; Choi et al. 2006; Dwarakanath et al. 2004, 2005; Hansen et al. 2006; Ikanovic et al. 2007; Levy et al. 2005; Liu et al. 2007; Numnuam et al. 2008; Revesz et al. 2007) have been extensively used with aptamer-based assays offering advantages such as labeling simplicity, independence from expensive analytical instruments, and extremely high extinction coefficients (about 1000 times larger than those of organic dyes). An early attempt at implementing aptamers onto nanoparticles was demonstrated by Pavlov et al. (2004), who enhanced the optical detection of thrombin utilizing aptamer-functionalized gold nanoparticles. In this assay, approximately 80 aptamers were bound to each nanoparticle. Binding of thrombin to the aptamers on adjacent nanoparticles resulted in an aggregated precipitate of nanoparticles, which was subsequently separated from the supernatant by centrifugation, and amplified by catalytic enlargement. This resulted in a detection limit of roughly 2 nM within a tested linear range of 50–120 nM. Additionally, selectivity for thrombin in the presence of BSA and human IgG was demonstrated. Performing a comparative study with ELISA, Li et al. (2007) described the densitometry detection of cytokines which utilized a sandwich assay consisting of an aptamer-functionalized gold nanoparticle and a biotinylated aptamer. Using a microplate analyzer and a scanning wavelength of 630 nm, platelet derived growth factor B-chain homodimer was detected linearly from 1 fM-100 pM with an analytical detection limit of 83 aM. A silver compound was deposited on the gold nanoparticles to enhance absorbance detection. Similarly, an approach reported by Tan et al. (Herr et al. 2006; Smith et al. 2007) used a two nanoparticle assay for the selective recognition of leukemia cells. Aptamer functionalized magnetic nanoparticles were used to specifically capture CCRF–CEM acute leukemia cells, while Rubpy enhanced tetraethyl orthosilicate particles signaled the binding event. Another approach by Chen et al. (2008) combined CdSe/ZnS quantum dots with an anti-tenascin-C DNA aptamer for the highly selective detection of tenascin protein on human glioma cell membranes. The effort demonstrated a proof-of-concept technology for diagnostics and therapy applications with human glioma.
One tremendous advantage of coupling nanoparticles with aptamers is the combined effect of signal amplification (imparted by nanoparticles) and target specificity (provided by aptamers) that is not available using conventional fluorescent labels. Moreover, integrating the approach with microfabrication technologies provides the potential for high sensitivity detection and portability. A notable example is the pioneering work of Farokhzad et al. (2005). Aptamers with binding affinity to prostate specific membrane antigen (PSMA), which is a well known protein expressed on the surface of prostate cancer epithelial cells, were conjugated to poly(ethylene glycol) (PEG) functionalized poly(lactic acid) particles. The researchers created a poly(dimethylsiloxane) (PDMS) microchip in which cancerous (LNCaP) and non-cancerous (PC3) cells were cultured and immobilized. Subsequent experiments were performed to characterize the binding properties of free nanoparticle/aptamer conjugates in solution under different shear stress conditions (i.e., flow rates) while introduced to the immobilized cells. Nanoparticle-aptamer bioconjugates selectively adhered to LNCaP but not PC3 cells at static and low shear (<1 dyn/cm2) conditions, but not higher shear (about 4.5 dyn/cm2) conditions. Control nanoparticles and microparticles lacking aptamers and microparticle–aptamer bioconjugates did not adhere to LNCaP cells, even under very low-shear conditions (close to 0.28 dyn/cm2). This work demonstrated that scalability, low cost, reproducibility, and high throughput capability of this technology is potentially beneficial to examining and optimizing a wide array of cell-particle systems prior to in vivo experiments.
4.2.3 Fiber-optic sensors incorporating aptamers
An interesting aptasensing format was presented by Lee and Walt (2000) who functionalized aptamers at the end of an optical fiber for the detection of thrombin. The aptamer was immobilized onto micro silica beads and placed on the distal end of a fiber array. Fluorescein-labeled thrombin was used in a competitive assay format. Although fluorescence signals from the individual aptamer beads showed significant variability, the average signals of 100 beads provided much more precise values. The fiber-optic microarray system had a detection limit of 1 nM for non-labeled thrombin, while testing could be performed in 15 min.
4.2.4 Surface plasmon resonance (SPR) based systems
SPR is a reference optical method where a sensing signal (analyte present on the sensing surface) is compared to a base signal (no analyte present on the sensor surface). In a typical device utilizing SPR sensing, a metal thin-film supporting a surface plasmon is coated with an interrogative receptor (e.g., an aptamer). A sample solution containing the analyte is then injected at a constant flow rate while a detector monitors the changes in the resonance angle, or refractive index which occur at the sensor-chip surface when the analyte in solution binds to the immobilized receptor. This binding event can be monitored in real time in which the signal is proportional to the amount of bound molecules.
A majority of papers have appeared in literature in which conventional scale equipment was used for studying the interaction between selected aptamers and their relative targets (Hwang and Nishikawa 2006; Balamurugan et al. 2006; Gebhardt et al. 2000; Hayashi et al. 2007; Li et al. 2006, 2007; Potyrailo et al. 1998; Tombelli et al. 2005). Notably, Tombelli et al. (2005) described an SPR surface modified with anti-HIV Tat aptamer. The selectivity of the aptamer to HIV Tat as compared to a very similar protein, Rev protein, was on the order of 3–1 with regards to signal response. The researchers observed a detection limit close to 0.12 ppm. Until recently, there had been very little work on integrating SPR sensing of aptamer interactions on a microchip platform. Wang et al. (2007a) demonstrated the potential paradigm shift with a PDMS microfluidic device incorporating thiolated anti-IgE aptamers immobilized on a gold surface. Four distinct oligonucleotide sequences were tested for affinity and selectivity against human IgE until an optimal aptamer candidate emerged (deemed “Aptamer I”) with superior binding properties. Subsequently, a 2 nM detection limit was determined along with a dissociation constant of 0.27 μM. This work demonstrated the utility of integrating a non-labeled analyte detection approach, such as SPR, with the portability and efficiency of microfluidic technology.
4.3 Electroanalytical sensors
In a manner identical to fluorescent tagging, the 3′ and 5′ termini of oligonucleotides can also be modified with electrochemical tags, i.e., redox groups. These markers enable signal detection using established electrochemical instrumentation. A wide variety of aptasensor configurations make use of the movement of electrons to accomplish signal transduction. There are also label-free electroanalytical techniques such as those using impedance measurements, or the interaction of charged ions in solution with the structure of the oligonucleotide itself (Shen et al. 2007; Rodriguez 2005). Electroanalytical transduction-based aptasensors have been used to detect a wide variety of target molecules.
4.3.1 Electrochemical sensors using tags
Electrochemical tags which are commonly used for aptasensor transduction include methylene blue (MB) and ferrocene (Baker et al. 2006; Mir and Katakis 2007; Radi et al. 2006; Radi 2006; Xiao et al. 2005a, b; Zuo et al. 2007). MB and ferrocene have been used as tags in “signal-on” aptasensors for compounds such as cocaine, adenosine triphosphate (ATP), and thrombin. In some cases, the sensors make use of the same binding-induced structural changes used in optical sensors. With an aptamer coupled to an electrode surface, binding-induced conformational changes (due to target interaction) bring the redox tag closer to the electrode surface. Such changes in surface charge density are commonly detected using techniques such as cyclic voltammetry and chronocoulometry. By comparing the results of such tests before and after exposure to a target solution, the amount of immobilized target can be accurately estimated. An alternative to covalently modifying aptamers with electrochemical tags is to introduce charge-bearing particles into solution that will be attracted to the structure of the aptamer. Several sensors have been developed which utilize [Ru(NH3)6]3+ (RuHex), a redox cation which binds to the phosphate backbone of DNA aptamers (Shen et al. 2007; Cheng et al. 2007). In most of these cases, a combination of techniques employing direct current/voltage analysis such as cyclic voltammetry and chronoamperometry are used to determine the surface density of DNA and DNA-target complexes. As a result of their use of surface electrodes for in-solution measurements, these sensors are readily adaptable to microfluidics. For example, it has already been shown in other fields that miniaturized or microfabricated three-electrode cells can be successfully used to monitor DNA hybridization and changes in electrolyte composition (Triroj et al. 2006; Willey and West 2006). Hence, it should be readily feasible to realize microchip aptasensors based on electrochemical tags, and such devices are expected to emerge in the future.
4.3.2 Impedence-based aptasensors
Another method of electroanalytical measurement common in aptasensors is the use of impedance (Degefa and Kwak 2008; Du et al. 2008; Lee et al. 2008; Li et al. 2008; Lohndorf et al. 2005; Xu et al. 2005, 2006). Whereas common electrochemical techniques (e.g., cyclic voltammetry) employ DC current, electrochemical impedance spectroscopy (EIS) uses an identical set of connections with alternating current signals to achieve an accurate measurement of surface charge. Originally found in corrosion studies, this technique has recently become more popular for use in bioanalytical applications. It has been reported that high-frequency impedance measurements are capable of gathering real-time information on aptamer binding events (Lohndorf et al. 2005). Aptasensors using this technique are reported to have detected ATP, thrombin, and platelet-derived growth factor (PDGF) with picomolar and even femtomolar detection limits (Degefa and Kwak 2008; Du et al. 2008; Lee et al. 2008; Li et al. 2008; Xu et al. 2005, 2006). The potential for impedance aptasensors for miniaturization is indicated by the recent miniaturization of an EIS sensor for thrombin (Lee et al. 2008). Here (Fig. 6) gold was used as a counter-electrode, and pyrolized carbon as a working electrode, with DNA aptamers immobilized onto the working electrode via carbodiimide chemistry. A (quasi-) reference electrode was created by inserting a silver wire into the testing solution which contained chloride ions. Testing indicated that the sensor was capable of detecting thrombin concentrations of 500 pM.
Miniaturized electrochemical aptasensor for thrombin. Adapted from Lee et al. (2008); copyright permission is being obtained
4.3.3 Aptasensors based on electrochemiluminescence
Electrogenerated chemiluminescence (ECL) is a sensing principle in which an electrochemical reaction generates an optical signal. This technique benefits from the advantages of strong signal-to-noise ratios common to optical sensors as well as the ease of use of electrochemical sensors. In one example, a cocaine ECL aptasensor was fabricated that employed the same aptamer/target complex as other electroanalytical aptasensors but exhibited an increased sensitivity (Li et al. 2007). It has also been reported that a detection limit of 1.0 fM was reached when using an ECL aptasensor employing displacement of nanoparticle-enhanced strands to detect thrombin (Wang et al. 2007b).
4.3.4 Aptasensors using nanotubes and nanofilaments
A recent innovation in electroanalytical aptasensors has involved the use of nanostructures (Huang et al. 2008; Kase et al. 2004; Maehashi et al. 2007; So et al. 2005, 2008; Yoon et al. 2008) for field-effect transduction of molecular binding. Constructions of one-dimensional semiconductor nanowires and carbon nanotubes have previously been described for the sensitive detection of viruses (Patolsky et al. 2004), small molecules (Wang et al. 2005), and proteins (Star et al. 2003). However, these devices have relied mainly on immunoreceptors as analyte recognition molecules. In such devices, there is a high possibility that the recognition binding occurs outside the electrical double layer (Debye length) in physiological salt concentrations (Schasfoort et al. 1990) since immunoreceptors (antibodies and enzymes) are generally larger than the electrical double layer (>10 nm). Hence, most charged analytes, when binding occurs on the receptor, will be at a distance greater than the Debye length (>3 nm in 10 mM ionic concentrations), making them impossible to detect. Since aptamers are typically only 15–100 nucleotides long, tend to self-complement before and after binding to targets, and form complex binding configurations such as G-quartet formations and single and double hairpin loops, they can reduce their overall size (Maehashi et al. 2007; Yoon et al. 2008; Yang et al. 2008). These phenomena allow particularly small aptamers (15–25 nucleotides) a potential effective length of only 1–2 nm (So et al. 2005, 2008), making it possible that the aptamer-analyte binding event can occur inside the electrical double layer in millimolar salt concentrations.
So et al. (2005) successfully demonstrated the first single-walled carbon nanotube field effect transistor (SWNT-FET) biosensor using aptamers as an alternative to immunoreceptor-based sensing elements. The authors immobilized a 15-mer DNA thrombin-binding aptamer to the side wall of a carbon nanotube transistor pretreated with carbodiimidazole-activated Tween 20 (CDI-Tween) using amine (-NH2) carbodiimidazole chemistry. Conductance measurements were taken across a single functionalized SWNT-FET as varying concentrations of thrombin were introduced (Fig. 7). The device was sensitive to thrombin down to nearly 10 nM. However, the researchers surmised that lower detection limits were possible when using higher quality SWNT-FETs. In addition, selectivity of the thrombin aptamer was tested against elastase (another serine protease with an isoelectric point and molecular weight similar to thrombin) to which the conductance of the SWNT-FET showed no change. Similarly, Maehashi et al. (2007) produced another notable example of an SWNT-FET aptasensor for the detection of IgE protein. Using a 45-mer DNA aptamer, a comparison study was performed against a monoclonal antibody specific to IgE. A detection limit of 250 pM IgE was obtained using the aptamer-based device, whereas with the antibody sensor, a decreased source-drain current was observed under similar conditions resulting in poorer performance.
Binding of thrombin protein on a SWNT-FET-based aptamer sensor. The nanotube is grown between two electrodes for use as a gate. Adapted from So et al. (2005); Copyright permission is being obtained
4.4 Aptasensors using extraction and separation techniques
This section provides a brief overview of the use of aptamers for selectively recognizing and detecting analytes using extraction and separation techniques. Similar to non-aptamer based separation systems, applications of aptamers to this technique allow the detection of a range of analytes from small molecules to larger proteins and even cells (Mascini 2008). Hence, aptamer-based separation techniques allow a variety of applications including drug development, cell extraction or capture, molecular extraction, purification and separation, and electrochromatography. The integration of microtechnology has been limited in this area despite its potential benefits; we will hence review initial efforts in this direction.
4.4.1 Affinity extraction, purification, enrichment and separation
For the most part, aptamers have been immobilized onto surfaces, typically as stationary phases in affinity chromatography, to provide separation of target and non-target compounds (Brumbt et al. 2005; Clark and Remcho 2003; Cole et al. 2007; Connor and McGown 2006; Dangerfield et al. 2006; Deng et al. 2001, 2003; Dick and McGown 2004; German et al. 1998; Michaud et al. 2003, 2004; Ravelet et al. 2006; Romig et al. 1999; Ruta et al. 2007; Zhao et al. 2008). The first reported separation system using aptamers employed a fluorescently labeled DNA aptamer for quantifying human IgE in a competitive assay (German et al. 1998). Samples comprising free aptamers and aptamer–IgE complex were injected into a capillary and quantification of IgE was performed through evaluation of the peak area for the labeled aptamer. Separations of samples containing fluorescently labeled aptamer and IgE were complete in less than 60 s and revealed two zones, one corresponding to free aptamer and the other to aptamer bound to IgE. The free aptamer peak decreased and bound aptamer peak increased in proportion to the amount of IgE in the sample so that IgE could be detected with a linear dynamic range of 105 and a detection limit of 46 pM. The assay was highly selective since IgG was introduced to the IgE-specific aptamer incurring no fluorescence response. Moreover, IgE did not bind to control DNA sequences of the same length as the IgE-specific aptamer. Another example of aptamers applied as capture ligands was the purification of recombinant L-selectin receptor globulin (LS-Rg) from complex cell medium using a human L-selectin-specific DNA aptamer (Romig et al. 1999). A 1,500-fold enrichment of LS-Rg with 83% recovery was demonstrated. A DNA oligonucleotide with scrambled sequence immobilized on the same column material failed to bind detectable levels of LS-Rg, demonstrating the specificity of the system. For elution of LS-Rg, alteration of charge with an EDTA gradient was used. A recent application of this approach involved purification of the host nuclear protein that interacted with an important, highly structured RNA motif encoded within the U3 region of the genome of mouse mammary tumor virus (Dangerfield et al. 2006). The protein was identified by mass spectrometry and elution was carried out by competitive binding with solution of streptomycin and NaCl.
Although microtechnology has been extensively incorporated into standard extraction, purification, and separation phases, little has been accomplished in this field with aptamer affinity media (Crevillen et al. 2007). Kim et al. (Cho et al. 2004; Chung et al. 2005) demonstrated the first significant attempt at scaling an aptamer extraction and purification system to a miniaturized level. A replicase-specific aptamer was conjugated to either magnetic PEG (Cho et al. 2004) or CutiCore® (Chung et al. 2005) microbeads via a photo-cleavable linker in order to concentrate and purify HCV replicase, as a preparatory step before mass spectrometry. Two devices were used for testing. In the first microdevice (Fig. 8a; Cho et al. 2004), magnetic PEG microbeads were immobilized in a PDMS microfluidic device, with integrated pneumatic valving and venting, using a permanent magnet. After specific binding of HCV to the aptamer, washing and removing impurities, UV (360 nm) light irradiation of the microbeads cleaved the linker and subsequently released the aptamer/HCV complex for further analysis by matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS). Hence after each purification step, the device could be regenerated by removing the magnet, flushing the used microbeads and replenishing them with fresh beads. This approach was sufficient to enhance detection sensitivity to an estimated limit of 10 fmol of HCV polymerase. A subsequent device (Fig. 8b; Chung et al. 2005) consisted of glass-silicon construction where CutiCore® microbeads were packed in a fritted microchamber rather than held in place using a permanent magnet. Here, reverse flow was utilized to remove spent microbeads while the chip was washed with an NaOH solution before packing with fresh microbeads for subsequent experimentation.
a A schematic of the microaffinity purification chip used by (Cho et al. 2004). The device consisted of a PDMS on glass construction. A hydrophobic fluorocarbon film was used for enhanced fluid control (Adapted from Cho et al. (2004); copyright permission is being obtained). b Schematic of a second aptamer-based purification chip used by Chung et al. (2005) created from silicon and glass
Recently, we have reported microfluidic systems for the selective extraction, purification, enrichment and isocratic release and detection (i.e., elution using a single mobile phase with constant composition) of biomolecules using aptamer receptors. Our work exploits the property of aptamer systems to reversibly bind to their target analytes in a temperature-dependent manner. This is in contrast to conventional microfluidic extraction and purification systems which employ competitive displacement or potentially harsh solvents to reverse the aptamer-analyte complex. Both methods can be inefficient and harmful to sensitive analytes. Our early-stage work has produced a microfluidic system for thermally controlled analyte purification and enrichment (Nguyen et al. 2009) (Fig. 9a). The device was fabricated from PDMS bonded to glass substrates on which microheaters and temperature sensors were fabricated. Polystyrene microbeads functionalized with aptamer molecules were packed in a microchamber for use as a solid phase. We demonstrated highly specific extraction of a model metabolic analyte, adenosine monophosphate coupled with thiazole orange (TO-AMP) in a 105 dynamic range. Additionally, we demonstrated controlled release and regeneration of the device using a thermal stimulus at 32.5°C (Fig. 9b). Such a low temperature is expected to be safe for thermally sensitive target analytes and aptamer-functionalized surfaces. Most recently, we have extended this approach to demonstrate purification and concentration of trace AMP by roughly 1,000× (T. Nguyen et al., in preparation), and further, the integration of extraction, thermally activated isocratic elution, and label-free MALDI-MS detection on a single microfluidic platform (T. Nguyen et al., in preparation).
a Microfluidic device consisting of an aptamer functionalized packed chamber for selective extraction and purification of TO-AMP. b Thermally induced release of captured TO-AMP. Extraction of TO-AMP from solution occurred at room temperature. Subsequently, the temperature was raised incrementally. Notice the sharp decrease in signal at 32.5°C which continues to vanish as the temperature increases, implying release of the TO-AMP from the ATP-aptamer (adapted from Nguyen et al. 2009; copyright permission is being obtained)
4.5 Conclusion and future outlook
Aptamers have great potential for use in bioanalytical applications, particularly with biosensors. The main impetus for aptamer development is their ability to recognize virtually any target with high affinity and specificity, much like antibodies which are currently the established receptor in many assays because of their wide availability and proven success. However, the SELEX process offers in vitro generation, tighter control of receptor properties, and a milieu of functional modifications not available in standard immunoreceptor cultivation. These advantages afford aptamers significant advantages over their current antibody and enzyme counterparts, while at times even outperforming them. Despite this, aptamers significantly lag behind standard affinity receptors in terms of availability for most pertinent analytes, in particular, small molecules and very complex targets (e.g., viruses, protein complexes, cells and whole organisms). In part, this can be attributed to the relatively short history of aptamers (less than two decades), whereas antibodies and enzymes have dominated affinity receptor research for 70 years (Marrack 1938). Moreover, it is economically straining to generate aptamers with high specificity and selectivity (both important factors to consider when selecting aptamers for a given application) for the previously mentioned complex targets.
In this regard, miniaturization technology exhibits great promise. The advantages of conventional aptamer-based biosensors have already successfully driven their initial integration with micro- and nanotechnologies, as we have elaborated above. This allows higher throughput processing, multiplexed designs, faster analysis times, portability and economy in addition to higher selectivity. More importantly, however, it should become possible to incorporate the SELEX process on-chip so as to reduce experimental costs (from limiting reagent as well as sample consumption and waste), reaction time (and thus overall time), and therefore greatly increase the efficiency of performing SELEX. In addition, miniaturization of the SELEX process allows for potentially simultaneous selections, exploring numerous variables and modifications, to increase the chances of obtaining an optimal aptamer for any application. Hence, improving SELEX promises to realize its theoretically limitless aptamer selection capability and offers expansive applicability of miniaturized aptamer devices to many practical situations, including biosensing and beyond. This is an important attribute considering the growing demands of medical and biological fields like proteomics, metabolomics, and medical diagnostics and therapeutics, and bioterrorism in which thousands, if not millions, of individual samples of varying target analytes can be required to be processed simultaneously.
For example, researchers can utilize micro- and nanotechnology in conjunction with aptamers to provide in vivo sensors, i.e., nano-sized devices envisioned to be ingested or injected where they could act as reporters of in vivo concentrations of hundreds or thousands of key analytes (LaVan et al. 2003). Afterwards, transmission of the identification and recognition of analytes with clinical relevance to an external data capture system will occur. Additionally, aptamer nanoanalytical devices can potentially enhance compartmental analyses of metabolite levels and metabolic activity. This should drive “diagnostic methodologies,” which has already been demonstrated conventionally in Yeast and mammalian cell cultures for determination of carbohydrate homeostasis in living cells with subcellular resolution (Fehr et al. 2005). Furthermore, analytical micro- and nanodevices employing aptamers are increasingly sought after to support the fight against bioterrorism (Fischer et al. 2007). Owing once again to their synthetic selection, in addition to the ability to chemically synthesize and modify them on demand, and finally their structural stability, aptasensors hold great promise for providing the much needed tools to enhance our ability to detect and characterize biothreats.
The bar has been set high, perhaps unrealistically so, for miniaturized aptamer devices. Yet, similar to all budding technologies, such great expectations are a natural result of successfully demonstrated proof-of-concept. Realizing the full potential, as is often the case, will depend on translating the promising results found in the presented literature to actual practical devices. However, with advances in both aptamer generation against a growing number of targets and their corresponding sensing systems, combined with advances in micro- and nanotechnology, robust, efficient and most importantly, practical aptasensors are on the horizon.
References
Aberem MB et al (2006) Protein detecting arrays based on cationic polythiophene-DNA-aptamer complexes. Adv Mater 18(20):2703–2705
Bagalkot V et al (2007) Quantum dot–aptamer conjugates for synchronous cancer imaging, therapy, and sensing of drug delivery based on bi-fluorescence resonance energy transfer. Nano Lett 7(10):3065–3070
Baker BR et al (2006) An electronic, aptamer-based small-molecule sensor for the rapid, label-free detection of cocaine in adulterated samples and biological fluids. J Am Chem Soc 128(10):3138–3139
Balamurugan S et al (2006) Designing highly specific biosensing surfaces using aptamer monolayers on gold. Langmuir 22(14):6446–6453
Berezovski M, Krylov SN (2005) Thermochemistry of protein-DNA interaction studied with temperature-controlled nonequilibrium capillary electrophoresis of equilibrium mixtures. Anal Chem 77(5):1526–1529
Bock C et al (2004) Photoaptamer arrays applied to multiplexed proteomic analysis. Proteomics 4(3):609–618
Brody EN et al (1999) The use of aptamers in large arrays for molecular diagnostics. Mol Diagn 4(4):381–388
Brown D, Gold L (1995) Template recognition by an RNA-dependent RNA-polymerase: identification and characterization of two RNA binding sites on Q beta replicase. Biochemistry 34(45):14765–14774
Brumbt A et al (2005) Chiral stationary phase based on a biostable L-RNA aptamer. Anal Chem 77(7):1993–1998
Cha M et al (2008) Biomolecular detection with a thin membrane transducer. Lab Chip 8(6):932–937
Chen XC et al (2008) Quantum dot-labeled aptamer nanoprobes specifically targeting glioma cells. Nanotechnology 19(23)
Cheng AKH, Ge B, Yu HZ (2007) Aptamer-based biosensors for label-free voltammetric detection of lysozyme. Anal Chem 79(14):5158–5164
Cho S et al (2004) Microbead-based affinity chromatography chip using RNA aptamer modified with photocleavable linker. Electrophoresis 25(21–22):3730–3739
Cho EJ et al (2006) Optimization of aptamer microarray technology for multiple protein targets. Anal Chim Acta 564(1):82–90
Choi JH, Chen KH, Strano MS (2006) Aptamer-capped nanocrystal quantum dots: a new method for label-free protein detection. J Am Chem Soc 128(49):15584–15585
Christodoulides N et al (2002) A microchip-based multianalyte assay system for the assessment of cardiac risk. Anal Chem 74(13):3030–3036
Chung WJ et al (2005) Microaffinity purification of proteins based on photolytic elution: toward an efficient microbead affinity chromatography on a chip. Electrophoresis 26(3):694–702
Clark SL, Remcho VT (2003) Electrochromatographic retention studies on a flavin-binding RNA aptamer sorbent. Anal Chem 75(21):5692–5696
Cole JR et al (2007) affinity capture and detection of immunoglobulin E in human serum using an aptamer-modified surface in matrix-assisted laser desorption/ionization mass spectrometry. Anal Chem 79(1):273–279
Collett JR, Cho EJ, Ellington AD (2005) Production and processing of aptamer microarrays. Methods 37(1):4–15
Connor AC, McGown LB (2006) Aptamer stationary phase for protein capture in affinity capillary chromatography. J Chromatogr A 1111(2):115–119
Crevillen AG et al (2007) Real sample analysis on microfluidic devices. Talanta 74(3):342–357
Dangerfield JA et al (2006) Enhancement of the StreptoTag method for isolation of endogenously expressed proteins with complex RNA binding targets. Electrophoresis 27(10):1874–1877
Degefa TH, Kwak J (2008) Label-free aptasensor for platelet-derived growth factor (PDGF) protein. Anal Chim Acta 613(2):163–168
de-los-Santos-Alvarez N et al (2008) Aptamers as recognition elements for label-free analytical devices. Trac-Trends Anal Chem 27(5):437–446
Deng Q et al (2001) Retention and separation of adenosine and analogues by affinity chromatography with an aptamer stationary phase. Anal Chem 73(22):5415–5421
Deng Q, Watson CJ, Kennedy RT (2003) Aptamer affinity chromatography for rapid assay of adenosine in microdialysis samples collected in vivo. J Chromatogr A 1005(1–2):123–130
Dick LW, McGown LB (2004) Aptamer-enhanced laser desorption/ionization for affinity mass spectrometry. Anal Chem 76(11):3037–3041
Dittrich PS, Tachikawa K, Manz A (2006) Micro total analysis systems latest advancements and trends. Anal Chem 78(12):3887–3907
Drabovich AP et al (2006) Selection of smart aptamers by methods of kinetic capillary electrophoresis. Anal Chem 78(9):3171–3178
Du Y et al (2008) Multifunctional label-free electrochemical biosensor based on an integrated aptamer. Anal Chem 80(13):5110–5117
Dwarakanath S et al (2004) Quantum dot-antibody and aptamer conjugates shift fluorescence upon binding bacteria. Biochem Biophys Res Commun 325(3):739–743
Dwarakanath S et al (2005) Quantum dot–antibody and aptamer conjugates shift fluorescence upon binding bacteria (vol 325, p 739, 2004). Biochem Biophys Res Commun 329(1):416–416
Ellington AD, Szostak JW (1990) Invitro selection of RNA molecules that bind specific ligands. Nature 346(6287):818–822
Ellington AD, Szostak JW (1992) Selection invitro of single-stranded-dna molecules that fold into specific ligand-binding structures. Nature 355(6363):850–852
Farokhzad OC et al (2005) Microfluidic system for studying the interaction of nanoparticles and microparticles with cells. Anal Chem 77(17):5453–5459
Fehr M et al (2005) Development and use of fluorescent nanosensors for metabolite imaging in living cells. Biochem Soc Trans 33:287–290
Feigon J, Dieckmann T, Smith FW (1996) Aptamer structures from A to zeta. Chem Biol 3(8):611–617
Fischer NO, Tarasow TM, Tok JBH (2007) Aptasensors for biosecurity applications. Curr Opin Chem Biol 11(3):316–328
Furtado LM et al (1999) Interactions of HIV-1 TAR RNA with Tat-derived peptides discriminated by on-line acoustic wave detector. Anal Chem 71(6):1167–1175
Gebhardt K et al (2000) RNA aptamers to S-adenosylhomocysteine: kinetic properties, divalent cation dependency, and comparison with anti-S-adenosylhomocysteine antibody. Biochemistry 39(24):7255–7265
Geiger A et al (1996) RNA aptamers that bind l-arginine with sub-micromolar dissociation constants and high enantioselectivity. Nucleic Acids Res 24(6):1029–1036
German I, Buchanan DD, Kennedy RT (1998) Aptamers as ligands in affinity probe capillary electrophoresis. Anal Chem 70(21):4540–4545
Gram H et al (1992) Invitro selection and affinity maturation of antibodies from a naive combinatorial immunoglobulin library. Proc Natl Acad Sci USA 89(8):3576–3580
Hansen JA et al (2006) Quantum-dot/aptamer-based ultrasensitive multi-analyte electrochemical biosensor. J Am Chem Soc 128(7):2228–2229
Hayashi G et al (2007) Reversible regulation of binding between a photoresponsive peptide and its RNA aptamer. Nucleic Acids Symp Ser (Oxf) (51):93–94
He P et al (2007) Ultrasensitive electrochemical detection of proteins by amplification of aptamer-nanoparticle bio bar codes. Anal Chem 79(21):8024–8029
Hermann T, Patel DJ (2000) Adaptive recognition by nucleic acid aptamers. Science 287:820–825
Herr JK et al (2006) Aptamer-conjugated nanoparticles for selective collection and detection of cancer cells. Anal Chem 78(9):2918–2924
Huang YF, Chang HT, Tan WH (2008) Cancer cell targeting using multiple aptamers conjugated on nanorods. Anal Chem 80:567–572
Hwang J, Nishikawa S (2006) Novel approach to analyzing RNA aptamer-protein interactions: toward further applications of aptamers. J Biomol Screen 11(6):599–605
Ikanovic M et al (2007) Fluorescence assay based on aptamer-quantum dot binding to Bacillus thuringiensis spores. J Fluoresc 17(2):193–199
Jayasena SD (1999) Aptamers: an emerging class of molecules that rival antibodies in diagnostics. Clin Chem 45(9):1628–1650
Jung A et al (2007) Biofunctional structural design of SAW sensor chip surfaces in a microfluidic sensor system. Sens Actuators B Chem 124(1):46–52
Kase D et al (2004) Affinity selection of peptide phage libraries against single-wall carbon nanohorns identifies a peptide aptamer with conformational variability. Langmuir 20(20):8939–8941
Kirby R et al (2004) Aptamer-based sensor arrays for the detection and quantitation of proteins. Anal Chem 76(14):4066–4075
Langer P, Waldrop A, Ward D (1981) Enzymatic synthesis of biotin-labeled polynucleotides: novel nucleic acid affinity probes. Proc Natl Acad Sci 78:6633–6637
LaVan DA, McGuire T, Langer R (2003) Small-scale systems for in vivo drug delivery. Nat Biotechnol 21(10):1184–1191
Lee M, Walt DR (2000) A fiber-optic microarray biosensor using aptamers as receptors. Anal Biochem 282(1):142–146
Lee JA et al (2008) An electrochemical impedance biosensor with aptamer-modified pyrolyzed carbon electrode for label-free protein detection. Sens Actuators B Chem 129:372–379
Levy M, Cater SF, Ellington AD (2005) Quantum-dot aptamer beacons for the detection of proteins. ChemBioChem 6(12):2163–2166
Li Y, Lee HJ, Corn RM (2006) Fabrication and characterization of RNA aptamer microarrays for the study of protein-aptamer interactions with SPR imaging. Nucleic Acids Res 34(22):6416–6424
Li Y-Y et al (2007a) Ultrasensitive densitometry detection of cytokines with nanoparticle-modified aptamers. Clin Chem 53(6):1061–1066
Li Y, Lee HJ, Corn RM (2007b) Detection of protein biomarkers using RNA aptamer microarrays and enzymatically amplified surface plasmon resonance imaging. Anal Chem 79(3):1082–1088
Li Y et al (2007c) Electrogenerated chemiluminescence aptamer-based biosensor for the determination of cocaine. Electrochem Commun 9(10):2571–2575
Li X et al (2008) Electrochemical impedance spectroscopy for study of aptamer-thrombin interfacial interactions. Biosens Bioelectron 23(11):1624–1630
Liss M et al (2002) An aptamer-based quartz crystal protein biosensor. Anal Chem 74(17):4488–4495
Liu J, Lu Y (2006) Fast colorimetric sensing of adenosine and cocaine based on a general sensor design involving aptamers and nanoparticles. Angew Chem Int Edit 45(1):90–94
Liu J, Mazumdar D, Lu Y (2006) A Simple and sensitive “dipstick” test in serum based on lateral flow separation of aptamer-linked nanostructures. Angew Chem Int Edit 45(47):7955–7959
Liu JW, Lee JH, Lu Y (2007) Quantum dot encoding of aptamer-linked nanostructures for one-pot simultaneous detection of multiple analytes. Anal Chem 79:4120–4125
Lohndorf M et al (2005) Microfabricated high-performance microwave impedance biosensors for detection of aptamer-protein interactions. Appl Phys Lett 87(24)
Lupold SE et al (2002) Identification and characterization of nuclease-stabilized RNA molecules that bind human prostate cancer cells via the prostate-specific membrane antigen. Cancer Res 62(14):4029–4033
Maehashi K et al (2007) Label-free protein biosensor based on aptamer-modified carbon nanotube field-effect transistors. Anal Chem 79(2):782–787
Marrack JR (1938) The chemistry of antigens and antibodies. H.M. Stationery Office, London
Mascini M (2008) Aptamers and their applications. Anal Bioanal Chem 390(4):987–988
Mascini M et al (2007) Colorimetric microarray detection system for ghrelin using aptamer-technology. Anal Lett 40(7):1386–1399
McCauley TG, Hamaguchi N, Stanton M (2003) Aptamer-based biosensor arrays for detection and quantification of biological macromolecules. Anal Biochem 319(2):244–250
Mendonsa SD, Bowser MT (2004) In vitro selection of high-affinity DNA ligands for human IgE using capillary electrophoresis. Anal Chem 76(18):5387–5392
Michaud M et al (2003) A DNA aptamer as a new target-specific chiral selector for HPLC. J Am Chem Soc 125(28):8672–8679
Michaud M et al (2004) Immobilized DNA aptamers as target-specific chiral stationary phases for resolution of nucleoside and amino acid derivative enantiomers. Anal Chem 76(4):1015–1020
Mir M, Katakis I (2007) Aptamers as elements of bioelectronic devices. Mol Biosyst 3(9):620–622
Morimoto K et al (2007) Electrochemical microsystem with porous matrix packed-beds for enzyme analysis. Sens Actuators B Chem 124(2):477–485
Mosing RK, Mendonsa SD, Bowser MT (2005) Capillary electrophoresis-SELEX selection of aptamers with affinity for HIV-1 reverse transcriptase. Anal Chem 77(19):6107–6112
Nguyen T et al (2009) An aptamer-based microfluidic device for thermally controlled affinity extraction. Microfluid Nanofluidics (in press)
Nguyen T et al, An integrated microfluidic system for affinity extraction and concentration of biomolecules coupled to MALDI-MS (in preparation)
Nguyen T et al, Highly selective concentration and thermally activated release of analytes on a microchip (in preparation)
Nieuwlandt D, Wecker M, Gold L (1995) In vitro selection of RNA ligands to substance-P. Biochemistry 34(16):5651–5659
Nimjee SM et al (2005) The potential of aptamers as anticoagulants. Trends cardiovasc med 15(1):41–45
Numnuam A et al (2008) Aptamer-based potentiometric measurements of proteins using ion-selective microelectrodes. Anal Chem 80:707–712
Nutiu R, Li Y (2003) Structure-switching signaling aptamers. J Am Chem Soc 125(16):4771–4778
Ozaki H et al (2006) Biomolecular sensor based on fluorescence-labeled aptamer. Bioorg Med Chem Lett 16(16):4381–4384
Patolsky F et al (2004) Electrical detection of single viruses. Proc Natl Acad Sci USA 101(39):14017–14022
Pavlov V et al (2004) Aptamer-functionalized au nanoparticles for the amplified optical detection of thrombin. J Am Chem Soc 126(38):11768–11769
Polsky R et al (2006) Nucleic acid-functionalized Pt nanoparticles: catalytic labels for the amplified electrochemical detection of biomolecules. Anal Chem 78(7):2268–2271
Potyrailo RA et al (1998) Adapting selected nucleic acid ligands (aptamers) to biosensors. Anal Chem 70(16):3419–3425
Radi AE, O’Sullivan CK (2006) Aptamer conformational switch as sensitive electrochemical biosensor for potassium ion recognition. Chem Commun (32):3432–3434
Radi AE et al (2006) Reagentless, reusable, ultrasensitive electrochemical molecular beacon aptasensor. J Am Chem Soc 128(1):117–124
Ravelet C, Grosset C, Peyrin E (2006) Liquid chromatography, electrochromatography and capillary electrophoresis applications of DNA and RNA aptamers. J Chromatogr A 1117(1):1–10
Revesz E, Lai EPC, DeRosa MC (2007) Towards a FRET-based ATP biosensor using quantum dot/aptamer-based complexes. J Biomol Struct Dyn 24(6):645–645
Rodriguez MC, Kawde AN, Wang J (2005) Aptamer biosensor for label-free impedance spectroscopy detection of proteins based on recognition-induced switching of the surface charge. Chem Commun (34):4267–4269
Romig TS, Bell C, Drolet DW (1999) Aptamer affinity chromatography: combinatorial chemistry applied to protein purification. J Chromatogr B Anal Technol Biomed Life Sci 731(2):275–284
Ruta J et al (2007) Chiral resolution of histidine using an anti-d-histidine L-RNA aptamer microbore column. J Chromatogr B Anal Technol Biomed Life Sci 845(2):186–190
Savran CA et al (2002) Fabrication and characterization of a micromechanical sensor for differential detection of nanoscale motions. J Microelectromech Syst 11(6):703–708
Savran CA et al (2003) Microfabricated mechanical biosensor with inherently differential readout. Appl Phys Lett 83(8):1659–1661
Savran CA et al (2004) Micromechanical detection of proteins using aptamer-based receptor molecules. Anal Chem 76(11):3194–3198
Schasfoort RBM et al (1990) Possibilities and limitations of direct detection of protein charges by means of an immunological field-effect transistor. Anal Chim Acta 238(2):323–329
Schlensog MD et al (2004) A Love-wave biosensor using nucleic acids as ligands. Sens Actuators B Chem 101(3):308–315
Schurer H et al (2001) Aptamers that bind to the antibiotic moenomycin A. Bioorg Med Chem 9(10):2557–2563
Shamah SM, Healy JM, Cload ST (2008) Complex target SELEX. Acc Chem Res 41:130–138
Shen L et al (2007) A chronocoulometric aptamer sensor for adenosine monophosphate. Chem Commun (21):2169–2171
Shekhawat G, Tark SH, Dravid VP (2006) MOSFET-embedded microcantilevers for measuring deflection in biomolecular sensors. Science 311(5767):1592–1595
Smith JE et al (2007) Aptamer-conjugated nanoparticles for the collection and detection of multiple cancer cells. Anal Chem 79(8):3075–3082
So HM et al (2005) Single-walled carbon nanotube biosensors using aptamers as molecular recognition elements. J Am Chem Soc 127(34):11906–11907
So HM et al (2008) Detection and titer estimation of Escherichia coli using aptamer-functionalized single-walled carbon-nanotube field-effect transistors. Small 4(2):197–201
Stadtherr K, Wolf H, Lindner P (2005) An aptamer-based protein biochip. Anal Chem 77(11):3437–3443
Star A et al (2003) Electronic detection of specific protein binding using nanotube FET devices. Nano Lett 3(4):459–463
Stemmer WPC (1994) Rapid evolution of a protein in-vitro by DNA shuffling. Nature 370(6488):389–391
Stojanovic MN, Kolpashchikov DM (2004) Modular aptameric sensors. J Am Chem Soc 126(30):9266–9270
Stojanovic MN, Landry DW (2002) Aptamer-based colorimetric probe for cocaine. J Am Chem Soc 124(33):9678–9679
Stojanovic MN, de Prada P, Landry DW (2001) Aptamer-based folding fluorescent sensor for cocaine. J Am Chem Soc 123(21):4928–4931
Tombelli S et al (2005) Aptamer-based biosensors for the detection of HIV-1 Tat protein. Bioelectrochemistry 67(2):135–141
Triroj N et al (2006) Microfluidic three-electrode cell array for low-current electrochemical detection. Sens J IEEE 6(6):1395–1402
Tuerk C, Gold L (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249:505–510
Turney K et al (2004) Functionalized nanoparticles for liquid atmospheric pressure matrix-assisted laser desorption/ionization peptide analysis. Rapid Commun Mass Spectrom 18(20):2367–2374
Tyagi S, Bratu DP, Kramer FR (1998) Multicolor molecular beacons for allele discrimination. Nat Biotech 16(1):49–53
Wang WU et al (2005) Label-free detection of small-molecule-protein interactions by using nanowire nanosensors. Proc Natl Acad Sci USA 102(9):3208–3212
Wang H et al (2006a) Identification of proteins bound to a thioaptamer probe on a proteomics array. Biochem Biophys Res Commun 347(3):586–593
Wang LH et al (2006b) Unmodified gold nanoparticles as a colorimetric probe for potassium DNA aptamers. Chem Commun (36):3780–3782
Wang ZZ et al (2007a) Surface plasmon resonance imaging for affinity analysis of aptamer–protein interactions with PDMS microfluidic chips. Anal Bioanal Chem 389:819–825
Wang X et al (2007b) Detection of thrombin using electrogenerated chemiluminescence based on Ru(bpy)32+-doped silica nanoparticle aptasensor via target protein-induced strand displacement. Anal Chim Acta 598(2):242–248
Wei H et al (2007) Simple and sensitive aptamer-based colorimetric sensing of protein using unmodified gold nanoparticle probes. Chem Commun (36):3735–3737
Wernette DP et al (2008) Functional-DNA-based nanoscale materials and devices for sensing trace contaminants in water. MRS Bull 33(1):34–41
White R et al (2001) Generation of species cross-reactive aptamers using “Toggle” SELEX. Mol Ther 4(6):567–573
Willey MJ, West AC (2006) A microfluidic device to measure electrode response to changes in electrolyte composition. Electrochem Solid State Lett 9(7):E17–E21
Xiao Y et al (2005a) Label-free electronic detection of thrombin in blood serum by using an aptamer-based sensor. Angew Chem Int Edit 44(34):5456–5459
Xiao Y et al (2005b) A reagentless signal-on architecture for electronic, aptamer-based sensors via target-induced strand displacement. J Am Chem Soc 127(51):17990–17991
Xu D et al (2005) Label-free electrochemical detection for aptamer-based array electrodes. Anal Chem 77(16):5107–5113
Xu Y et al (2006) An aptamer-based protein biosensor by detecting the amplified impedance signal. Electroanalysis 18(15):1449–1456
Yamamoto R, Kumar PKR (2000) Molecular beacon aptamer fluoresces in the presence of Tat protein of HIV–1. Genes Cells 5(5):389–396
Yang RH et al (2008) Noncovalent assembly of carbon nanotubes and single-stranded DNA: an effective sensing platform for probing biomolecular interactions. Anal Chem 80(19):7408–7413
Yoon H et al (2008) A novel sensor platform based on aptamer-conjugated polypyrrole nanotubes for label-free electrochemical protein detection. Chembiochem 9(4):634–641
Zhao W et al (2007) Simple and rapid colorimetric biosensors based on DNA aptamer and noncrosslinking gold nanoparticle aggregation. ChemBioChem 8(7):727–731
Zhao Q, Li XF, Le XC (2008) Aptamer-modified monolithic capillary chromatography for protein separation and detection. Anal Chem 80(10):3915–3920
Zhu HY et al (2006) Aptamer based microsphere biosensor for thrombin detection. Sensors 6(8):785–795
Zuo X et al (2007) A target-responsive electrochemical aptamer switch (TREAS) for reagentless detection of nanomolar ATP. J Am Chem Soc 129(5):1042–1043
Acknowledgments
This work was funded by the National Science Foundation (Grant Numbers CBET-0693274, DBI-0650020, and ECCS-0707748).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nguyen, T., Hilton, J.P. & Lin, Q. Emerging applications of aptamers to micro- and nanoscale biosensing. Microfluid Nanofluid 6, 347–362 (2009). https://doi.org/10.1007/s10404-008-0400-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10404-008-0400-7