Abstract
Background and aims
The endocytoscopic system (ECS) helps in virtual realization of histology and can aid in confirming histological diagnosis in vivo. We propose replacing biopsy-based histology for esophageal squamous cell carcinoma (ESCC) by using the ECS. We applied deep-learning artificial intelligence (AI) to analyse ECS images of the esophagus to determine whether AI can support endoscopists for the replacement of biopsy-based histology.
Methods
A convolutional neural network-based AI was constructed based on GoogLeNet and trained using 4715 ECS images of the esophagus (1141 malignant and 3574 non-malignant images). To evaluate the diagnostic accuracy of the AI, an independent test set of 1520 ECS images, collected from 55 consecutive patients (27 ESCCs and 28 benign esophageal lesions) were examined.
Results
On the basis of the receiver-operating characteristic curve analysis, the areas under the curve of the total images, higher magnification pictures, and lower magnification pictures were 0.85, 0.90, and 0.72, respectively. The AI correctly diagnosed 25 of the 27 ESCC cases, with an overall sensitivity of 92.6%. Twenty-five of the 28 non-cancerous lesions were diagnosed as non-malignant, with a specificity of 89.3% and an overall accuracy of 90.9%. Two cases of malignant lesions, misdiagnosed as non-malignant by the AI, were correctly diagnosed as malignant by the endoscopist. Among the 3 cases of non-cancerous lesions diagnosed as malignant by the AI, 2 were of radiation-related esophagitis and one was of gastroesophageal reflux disease.
Conclusion
AI is expected to support endoscopists in diagnosing ESCC based on ECS images without biopsy-based histological reference.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The endocytoscopic system (ECS) is a novel magnifying endoscopic examination that enables the observation of the surface epithelial cells in real time in vivo via vital staining using compounds such as methylene blue [1,2,3,4,5,6,7,8,9,10]. We were the first to perform the clinical trial of ECS in 2003 and reported the characteristics of surface epithelial cells of the normal squamous epithelium and cancerous tissue of the esophagus [1]. The optical magnification power of fourth-generation ECS (currently available in the Japanese market) is × 500 [8]. In addition, its magnification power can be increased up to × 900 by using digital magnification incorporated in the video processor. From the increase in the number of cases observed using ECS in the screening setting, we also reported the characteristics of benign lesions, including esophagitis, which is a condition considered as a differential diagnosis for esophageal squamous cell carcinoma (ESCC) [4, 6, 7]. We proposed a type classification for the squamous epithelium and a flowchart to attain its efficient application [2, 5]. We observed that the overall diagnostic accuracy of both the endoscopist and pathologist to distinguish benign and malignant lesions by using ECS was approximately 95.0% [6]. On the basis of these results, we reported the possibility of diagnosing ESCC by using ECS without biopsy-based histological reference. However, one limitation needs to be overcome before the clinical application of ECS because the histological diagnosis may be difficult to confirm by the endoscopist in real time without consulting a pathologist. Therefore, creating an adequate software that can support the endoscopist is mandatory.
Recently, artificial intelligence (AI) has made remarkable progress in various medical fields, especially as a system for screening medical images, including those in the field of radiological oncology [11], skin cancer classification [12], diabetic retinopathy [13], and histological classification of gastric biopsy [14]. Misawa et al. applied the AI system for the first time to analyze ECS pictures of the colon and reported high overall sensitivity and specificity [15]. This AI system uses texture analysis. However, deep-learning analysis is the next step to texture-based analysis and can become a more powerful supportive tool to interpret medical images based on a historically accumulated set of unique algorithms. Deep learning allows computational models, which are composed of multiple processing layers, to analyze representations of data with multiple levels of abstraction. We constructed a deep learning AI-based diagnostic system through a convolutional neural network (CNN). We successfully applied the AI system to diagnose Helicobacter pylori infections in the stomach [16] and detect gastric cancer [17]. In this study, we applied this system to the previously collected ECS images of the esophagus and examined the diagnostic ability of this system for esophageal malignancy.
Patients and methods
Preparation of training and test images
Between November 2011 and February 2018, we performed 308 esophageal ECS examinations of 240 patients at Saitama Medical Center, Saitama Medical University. Two prototype ECSs (Olympus Medical Systems, Co., Ltd., Tokyo, Japan), GIF-Y0002 (optical magnification: × 400, with a digital magnification of × 700) and GIF-Y0074 (optical magnification: × 500, with a digital magnification of × 900), were employed in this study. Standard endoscopic video systems (EVIS LUCERA CV-260/CLV-260 and EVIS LUCERA ELITE CV-290/CLV-290SL; Olympus Medical System) were used. For vital staining, most cases were stained with toluidine blue, except for some cases with methylene blue.
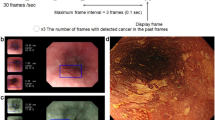
We preserved endoscopic pictures obtained using the maximum optical magnification power of ECS [lower magnification picture (LMP)] and those obtained using the maximum magnification power of ECS in tandem with digital 1.8-fold magnification [higher magnification picture (HMP)] (Fig. 1). Among 308 ECS observations, 4715 pictures from 240 cases obtained between November 2011 and December 2016 retrospectively were recruited for training image to develop an algorithm for analysis of ECS images. The training cases consisted of 126 cases of esophageal cancer (58 superficial and 68 advanced cancer cases, 124 cases were squamous cell carcinoma and 2 cases of advanced Barrett adenocarcinoma), 106 cases of esophagitis [52, radiation-related esophagitis; 45, gastroesophageal reflux disease (GERD); 4, candida esophagitis; 2, eosinophilic esophagitis; and 3, other types of esophagitis], and 8 cases with normal squamous epithelium. All lesions were diagnosed histologically using biopsy histology or resected specimens. Finally, we classified the training images into two different categories, “malignant (number of pictures: 1141 [LMP: 231, HMP: 910])” and “non-malignant (number of pictures: 3574 [LMP: 1150, HMP: 2424]).” We trained the AI system collectively by using the LMP and HMP pictures.
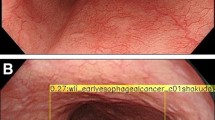
Typical test pictures. a–e Higher magnification pictures (GIF-Y0074, × 900). f–j Lower magnification pictures (GIF-Y0074, × 900). a, f Squamous cell carcinoma (malignant). b, g Normal squamous epithelium (non-malignant). c, h Esophagitis with inflammatory cell infiltration (non-malignant). d, i Regenerative squamous epithelium (non-malignant). e, j White coat with massive aggregation of the white blood cells (non-malignant)
For the evaluation of the diagnostic ability of the constructed AI system, we used 65 consecutive cases with observed surface cell morphology of the esophagus by using ECS obtained between January 2017 and February 2018. We excluded 10 cases in which clear pictures could not be obtained (2 cases of malignant melanoma, 2 cases of advanced esophageal cancer, and 2 cases of esophagitis) and cases with only LMP or HMP (4 cases). Finally, test cases consisted of 27 cases of malignant lesions and 28 cases of non-malignant lesions. We examined 1520 pictures (LMP: 467; HMP: 1053). Median number of ECS pictures of the test set was 24.0 (range 9–59).
All cases of malignant lesions were of esophageal squamous cell carcinoma, 20 were of superficial cancer, and 7 cases were of advanced cancer. Among the non-malignant lesions, 27 cases were of esophagitis (14, radiation-related esophagitis; 9, GERD; and 4, other types of esophagitis) and 1 was of esophageal papilloma.
The ECS images for CNN-training were randomly rotated between 0° and 359°; their black frames were cropped, and augmented approximately 5 times as a result. In this manner, it is known that the robustness of CNN increases.
Construction of a CNN algorithm
To construct an AI-based diagnostic system, we used a deep neural network architecture, called GoogLeNet (https://arxiv.org/abs/1409.4842), which is comprised of 22 layers for image classification. A Caffe deep learning framework, one of the most widely used frameworks originally developed at the Berkeley Vision and Learning Centre, was used to train, validate, and test the CNN.
The layers of CNN were trained by backpropagation algorithm, by which loss gradients for all weights in the network can be efficiently updated. They were fine-tuned using AdaDelta (https://arxiv.org/abs/1212.5701) at a global learning rate of 0.004. Each image was resized to 224 × 224 pixels. We used a pre-trained model that learned natural-image features through ImageNet. These values were set up by trial and error to ensure that all data were compatible.
Outcome measures of AI diagnosis
Per-image analysis
After constructing the CNN by using the training image set, we examined the test images. The specifications for the machine used in this study were CPU: Intel Core i7-7700K and GPU: GeForce GTX1070.
Each test image was calculated from the probability of malignancy. A receiver-operating characteristic (ROC) curve was plotted and the area under the ROC curve (AUC) was calculated. A cutoff index was determined by considering the sensitivity and specificity of the ROC curve.
Per-patient analysis
Each case of the test images was diagnosed as malignant if two or more ECS pictures obtained from the case showed a malignancy. The sensitivity, positive predictive value (PPV), negative predictive value (NPV), and specificity of AI to diagnose esophageal cancer were calculated as follows:
- Sensitivity :
-
Detected number of correct esophageal cancer lesions/histologically confirmed number of esophageal cancer lesions
- PPV :
-
Detected number of correct esophageal cancer lesions/number of lesions diagnosed as esophageal cancer by using the AI
- NPV :
-
Number of lesions correctly diagnosed as non-neoplastic lesions (one or no ECS picture showing a neoplastic lesion)/number of lesions diagnosed as non-neoplastic by using the AI
- Specificity :
-
Number of lesions that were correctly diagnosed as non-neoplastic lesion (one or no ECS picture showing a neoplastic lesion)/number of histologically proved non-neoplastic lesions.
ECS classification
Before the AI-based diagnosis, one experienced endoscopist (YK: examiner in all the ECS examinations) made the diagnosis during the endoscopic procedure. The endoscopist classified all the cases according to the type classification. If the test case included pictures showing lesions diagnosed as different types of ECS, we considered the higher type first. The ECS type classification was as follows:
-
Type 1, Surface epithelial cells have a low nucleus-to-cytoplasm (N/C) ratio and cell density but without evident nuclear abnormality.
-
Type 2, Surface epithelial cells have a high nuclear density but no evident nuclear abnormality and clear borders between cells.
-
Type 3, Surface epithelial cells have evidently increased nuclear density and nuclear abnormality, such as irregular nuclear size and shape, with hyperchromatism (Fig. 2).
We compared the AI-based diagnosis and the diagnosis based on the type classification by the endoscopist.
Results
Pathological diagnosis of the test cases
All esophageal cancer cases were histologically confirmed as lesions of squamous cell carcinoma. Four cases of GERD were histologically diagnosed as regenerative squamous epithelium, 3 were of esophageal ulcers, and 2 were of esophagitis. The histological diagnosis of the radiation-related esophagitis was regenerative squamous epithelium in 2 cases, esophagitis in 9, esophageal ulcer in 1, pseudo-malignant erosion in 1, and necrotic tissue in 1 (Table 1).
Performance of deep learning artificial intelligence
Per-image analysis
The AI structured in this study required 17 s to analyze 1520 images (0.011 s per image). From the plotted ROC curve of the total images, the AUC was 0.85. Furthermore, when HMP and LMP were separately analyzed, the AUC of the HMP and LMP were 0.90 and 0.72, respectively, (Fig. 3).
On the basis of the ROC analysis, we set the cutoff value for the probability of the malignancy at 90%, expecting a high specificity for non-malignant lesions. The sensitivity, specificity, PPV, and NPV for the total images were 39.4, 98.2, 92.9, and 73.2, respectively. When the HMPs and LMPs were analyzed separately, the sensitivity, specificity, PPV, and NPV were, respectively, 46.4, 98.4, 95.2, and 72.6 for the HMPs and 17.4, 98.2, 79.3, and 74.4 for the LMPs.
Among 622 non-cancerous HMPs, 10 pictures (1.6%) exceeded 90% of the probability of malignancy. Seven pictures were obtained for the white coat, 2 pictures showed atypical cells in the pseudo-malignant erosive lesion, and 1 picture showed a normal epithelium. Among 332 LMPs of non-cancerous lesions, 7 (2.1%) obtained for esophagitis exceeded 90% of the probability of malignancy.
Per-patient analysis
Among 27 cases of histologically proven ESCC, 25 were accurately diagnosed as malignant by using the AI (including > 2 pictures with a probability rate for malignancy of > 90%; sensitivity: 92.6%). The median percentage (range) of the number of pictures showing malignancy that were correctly diagnosed as malignant by using the AI in 27 malignant lesions was 40.9% (0–88.9). AI accurately diagnosed 25 of the 28 cases of non-cancerous lesions as non-malignant (specificity: 89.3%). The PPV, NPV, and overall accuracy were 89.3, 92.6, and 90.9%, respectively (Table 2).
The endoscopist diagnosed all malignant cases as type 3. 13 cases of non-cancerous lesions as type 1, 12 cases as type 2, and 3 lesions as type 3. If type 3 is considered to correspond to malignancy, the sensitivity, specificity, PPV, NPV, and overall accuracy of the endoscopist’s diagnosis were 100%, 89.3%, 90.0%, 100%, and 94.5%, respectively.
Two cases of malignant lesions misdiagnosed as non-malignant by the AI were correctly diagnosed as type 3 by the endoscopist. These two cases were of superficial cancer, and increases in nuclear density and nuclear abnormality were apparent. However, nuclear enlargement was not clear. Among 3 cases diagnosed as malignant by the AI for non-malignant lesions, 2 were of radiation-related esophagitis and one was of grade C GERD. The endoscopist did not consider these lesions as malignant from the non-magnified view of these lesions. However, the endoscopist classified 1 case of radiation-related esophagitis as type 3 because of the apparent increase in nuclear density and abnormality. The histological examination of these lesions revealed pseudo-malignant erosions.
Discussion
This is the first report to evaluate ECS images of the esophagus by using deep-learning AI. ECS is a novel endoscopic technology that aids in the realization of the concept of “optical biopsy,” a term for the assessment of tissue by imaging technology to acquire a histologically equivalent image without removing tissue at the time of endoscopy [18]. The most promising aim of the ECS for the esophagus is diagnosis of ESCC without histological biopsy [2, 5]. In our investigation, a high AUC was noted from the ROC curve. This observation was apparent especially in the result from HMP. Concerning this issue, we tested whether a pathologist who is shown only ECS cell images and blinded to all other information can distinguish the ECS pictures that show malignancy. The pathologist diagnosed approximately two-thirds of the esophagitis cases as malignant by using LMP [4]. In these cases, the nuclear density was observed to be increased, but the nuclear abnormality could not be recognized because of the low magnification power. Therefore, we applied digital 1.8-fold magnification (HMP), expecting better recognition of nuclear morphology. The sensitivity and specificity provided by the pathologist dramatically improved to > 90% [6, 7]. In our present study, all LMPs of non-cancerous lesions misdiagnosed as malignant by the AI were obtained from the esophagitis cases. These pictures revealed high nuclear density, but nuclear abnormality could not be evaluated by the endoscopist because of the low magnification power. On the basis of these results, our present study also recommends referring to HMP rather than LMP for better recognition of nuclear abnormality. On the other hand, most HMPs of non-cancerous lesions misdiagnosed as malignant by the AI were obtained from the observation of white coat. The ECS features of the white coat showed aggregation of small inflammatory cells. We speculated that these pictures were misunderstood by the AI as LMPs of ESCCs (Fig. 1). We should establish two separate AI systems for HMPs and LMPs in the future.
When replacing histological biopsy, overdiagnosis of non-cancerous lesions must be minimized. Overdiagnosis of squamous neoplasia by an endoscopist by using ECS will necessitate resection, which carries a certain risk of morbidity and mortality. Therefore, we set the cutoff value for the probability of malignancy at 90%. In some articles that report application of AI for the early detection of cancer, the cutoff value has been set at a relatively low probability so that certain false-positive cases might be included [17, 19].
In our per-patient analysis, the endoscopist applied type classification for ECS diagnosis. Types 1 and 3 corresponded to the non-neoplastic epithelium and ESCC, respectively. Histological biopsy can be avoided in these cases. However, type 2 included various histological situations; therefore, histological biopsy should be performed for treatment planning. In this study, the overall accuracy of the AI exceeded 90%; this was inferior but close to the result obtained by the endoscopist, who was aware of the non-magnified endoscopic view of the lesions. The sensitivity of AI diagnosis for ESCC was 92.9%. Although the number of pictures diagnosed as “malignant” by the AI is relatively small [40.9% (0–88.9)], this result is acceptable in spite of the high cutoff value. In these cases, histological biopsy can be replaced with supporting assurance of AI. However, two cancer cases could not be diagnosed as malignant by the AI but were accurately diagnosed as type 3 by the endoscopist. In these cases, histological biopsy may be replaced without the support of AI because a non-magnified conventional endoscopic view can already clearly indicate malignancy.
On the other hand, the specificity of this study was 89.3%, and 3 non-cancerous cases were diagnosed as malignant by the AI. One case was of grade C GERD and the other two cases were of radiation-related esophagitis. Pathological diagnoses of these lesions were regenerative squamous epithelium, pseudo-malignant erosion, and esophageal ulcer, respectively. Some regenerative squamous epithelia are difficult to distinguish from ESCC even for an experienced pathologist. In addition, pseudo-malignant erosion also includes atypical cells; thus, a pathologist may confuse it with malignancy during diagnosis. ECS can adequately capture the characteristics of these lesions [20]. Furthermore, our test series did not include cases with intraepithelial neoplasia (IN). However, AI has a potential of distinguishing IN from ESCC because most of the IN was not observed in atypical cells at the mucosal surface based on the histological features (total layer of the squamous epithelium was not rearranged by the atypical cells in IN). Among the 3 misdiagnosed non-cancerous lesions by the AI, 2 were diagnosed as type 2 by the endoscopist, for which histological biopsy was recommended for treatment planning. One case of radiation-related esophagitis was diagnosed as type 3 by the endoscopist and as neoplastic lesion by the AI. After irradiation, diagnosis of recurrence or remnant esophageal cancer is sometimes difficult even if histological biopsy has been performed. Histological biopsy must be performed in cases of follow-up after irradiation for esophageal cancer in addition to ECS observation.
In this study, we can point out some limitations. First, the number of test images was small. This may be a reason for the considerably low median percentage of the pictures showing malignancy (40.9%) in per-patient analysis. We expected that deeply learned AI will further improve the outcome of AI diagnosis. In this context, we are planning to obtain a large number of pictures from recorded videos. Second, we used two different ECSs with optical magnification powers of × 400 and × 500, respectively. Differences in the magnification power of these two ECSs may also have affected the results of AI diagnosis. GIF-Y0074, which has an optical magnification power of ×500, is now available in the market [8]. Training images should be limited to the pictures obtained by GIF-Y0074 in the future.
In conclusion, our deep-learning AI is expected to provide strong support to endoscopists in the diagnosis of ESCC accurately based on ECS pictures without referring to histological biopsy. HMPs, rather than LMPs, should be used as a reference for better recognition of nuclear abnormalities.
References
Kumagai Y, Monma K, Kawada K. Magnifying chromoendoscopy of the esophagus: in vivo pathological diagnosis using an endocytoscopy system. Endoscopy. 2004;36:590–4.
Kumagai Y, Kawada K, Yamazaki S, et al. Endocytoscopic observation for esophageal squamous cell carcinoma: can biopsy histology be omitted? Dis Esophagus. 2009;22:505–12.
Kumagai Y, Kawada K, Yamazaki S, et al. Prospective replacement of magnifying endoscopy by a newly developed endocytoscope, the ‘GIF-Y0002’. Dis Esophagus. 2010;23:627–32.
Kumagai Y, Kawada K, Yamazaki S, et al. Current status and limitations of the newly developed endocytoscope GIF-Y0002 with reference to its diagnostic performance for common esophageal lesions. J Dig Dis. 2012;13:393–400.
Kumagai Y, Kawada K, Yamazaki S, et al. Endocytoscopic observation of esophageal squamous cell carcinoma. Dig Endosc. 2010;22:10–6.
Kumagai Y, Kawada K, Higashi M, et al. Endocytoscopic observation of various esophageal lesions at ×600: can nuclear abnormality be recognized? Dis Esophagus. 2015;28:269–75.
Kumagai Y, Takubo K, Kawada K, et al. Endocytoscpic observation of various types of esophagitis. Esophagus. 2016;13:200–7. https://doi.org/10.1007/s10388-015-0517-1.
Kumagai Y, Takubo K, Kawada K, et al. A newly developed continuous zoom-focus endocytoscope. Endoscopy. 2017;49(2):176–80.
Inoue H, Kazawa T, Sato Y, et al. In vivo observation of living cancer cells in the esophagus, stomach, and colon using catheter-type contact endoscope, “Endo-Cytoscopy system”. Gastrointest Endosc Clin N Am. 2004;14:589–94.
Sasajima K, Kudo SE, Inoue H, et al. Real-time in vivo virtual histology of colorectal lesions when using the endocytoscopy system. Gastrointest Endosc. 2006;63(7):1010–7.
Bibault JE, Giraud P, Burgun A. Big Data and machine learning in radiation oncology: state of the art and future prospects. Cancer Lett. 2016;382:110–7.
Esteva A, Kuprel B, Novoa RA, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542:115–8.
Gulshan V, Peng L, Coram M, et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA. 2016;316:2402–10.
Yoshida H, Shimazu T, Kiyuna T, et al. Automated histological classification of whole-slide images of gastric biopsy specimens. Gastric Cancer. 2018;21(2):249–57. https://doi.org/10.1007/s10120-017-0731-8.
Misawa M, Kudo S, Mori Y, et al. Accuracy of computer-aided diagnosis based on narrow-band imaging endocytoscopy for diagnosing colorectal lesions: comparison with experts. Int J Comput Assist Radiol Surg. 2017;12:757–66.
Shichijo S, Nomura S, Aoyama K, et al. Application of convolutional neural networks in the diagnosis of Helicobacter pylori infection based on endoscopic images. EBioMedicine. 2017;25:106–11.
Hirasawa T, Aoyama K, Tanimoto T, et al. Application of artificial intelligence using a convolutional neural network for detecting gastric cancer in endoscopic images. Gastric Cancer. 2018;21(4):653–60. https://doi.org/10.1007/s10120-018-0793-2.
Inoue H, Kazawa T, Sato Y, et al. In vivo observation of living cancer cells in the esophagus, stomach, and colon using catheter-type contact endoscope, “Endo-Cytoscopy system”. Gastrointest Endosc Clin N Am. 2004;14:589–94.
Misawa M, Kudo S, Mori Y, et al. Artificial intelligence-assisted polyp detection for colonoscopy: initial experience. Gastroenterology. 2018;154(8):2027.e3–2029.e3. https://doi.org/10.1053/j.gastro.2018.04.003.
Takubo K. Pathology of the esophagus. 3rd ed. Tokyo: Wiley Publishing Japan; 2017. p. 88–102 (131–134 ).
Acknowledgements
We are grateful to Tomoyuki Kawada for his kind advice and confirmation of statistical analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
T. Tada, Y. Endo and K. Aoyama are employed by AI Medical Service Inc. The other authors declare no conflict of interest.
Human rights statement
All procedures followed were in accordance with the ethical standards of our hospital ethics committee on human experimentation and with the Helsinki Declaration of 1964 and later versions.
Informed consent
Written informed consent for endocytoscopic observation was obtained from all patients. Opt out consent was obtained for AI analysis.
Rights and permissions
About this article
Cite this article
Kumagai, Y., Takubo, K., Kawada, K. et al. Diagnosis using deep-learning artificial intelligence based on the endocytoscopic observation of the esophagus. Esophagus 16, 180–187 (2019). https://doi.org/10.1007/s10388-018-0651-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10388-018-0651-7